Submitted:
01 January 2024
Posted:
03 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Detection Systems: these systems are designed to identify the presence of anomalies in chest X-Rays.
- Classification Systems: these systems are designed to classify anomalies in chest X-Rays based on the type of lung pathology.
- Convolutional Neural Networks (CNNs): CNNs are a class of artificial neural networks designed for image processing. They have been successfully used for recognizing various anomalies in chest X-Rays, including lung nodules, lung infiltrates, and pneumonia.
- Recurrent Neural Networks (RNNs): RNNs are a class of artificial neural networks designed for processing sequence data. RNNs have been successfully used for recognizing anomalies in chest X-Rays that develop over time, such as the progression of lung cancer.
- Generative Adversarial Networks (GANs): GANs are a class of artificial neural networks designed to generate realistic data. GANs have been successfully used for synthesizing chest X-Ray images containing anomalies, improving the training of DL systems.
- Early and Accurate Detection of Lung Pathologies: AI-enhanced image analysis systems can support radiologists in identifying anomalies in chest X-rays, providing technological assistance for a quick and precise diagnosis;
- Personalization of Diagnosis and Treatment: data collected from IoT sensors and DTs can be used to develop personalized diagnoses and treatment plans that are more effective and cost-efficient;
- Improvement of Patients’ Quality of Life: remote monitoring systems enable patients to receive care from home, improving their quality of life and reducing healthcare costs.
- Implementation of a case study that includes a proof-of-concept of a Lung-DT based on a microservices architecture characterized by an artificial intelligence component.
- The proposed Lung-DT architecture is designed to acquire various input signals, including chest X-Ray images (the subject of our study) and blood oxygen saturation (to be integrated later), allowing for a more comprehensive evaluation of lung conditions.
- Extension of the second-level architecture previously proposed in [3]. This extension involves the integration of DTs related to other organs to define the presence of various pathologies. The use of information from different organs enables a more comprehensive detection and classification of pathological conditions.
- Accurate classification of lung pathologies into 5 categories to contribute to a more detailed and specific diagnosis.
- Expansion of the neural network’s training and validation dataset by integrating two public datasets. This integration aims to enhance the predictive capacity of the model, improving its adaptability to a wider range of pathologies. Use of a third dataset completely unknown to the neural network to perform additional testing. This step is aimed at confirming the robustness of the model by evaluating its performance on novel data.
2. Related Work
- More powerful DL architectures: new DL architectures, such as DenseNet, ResNet, Transformers, and Inception, have been developed to effectively capture complex features from X-Ray images.
- Large X-Ray datasets: the availability of large X-Ray datasets, such as the NIH ChestX-ray14 dataset, has allowed the training of more accurate DL models.
- Collective learning: collective learning techniques, combining multiple DL models to improve performance, have been used to achieve the best results in lung disease detection.
- Fast and accurate diagnosis: DL models can be implemented on mobile devices or in hospitals to provide a rapid and accurate diagnosis of lung diseases.
- Early detection of lung cancer: DL models can be used to identify early signs of lung cancer, leading to early treatment and better outcomes for patients.
- Risk classification: DL models can be used to classify patients based on their risk of developing lung diseases, providing useful information for therapeutic decision-making.
- Continuous and personalized patient monitoring: utilize DTs to optimize the real-time collection and processing of data from various medical devices, including wearable sensors, clinical records, and other applications. This data can be used to create a digital model of the patient, predict the risk of diseases, identify anomalies, and optimize therapies.
- Unified patient management platform: use a platform that allows monitoring different organs and pathologies, through the creation of Organ DTs that, working collaboratively, provide the medical world with a tool for a comprehensive view of the patient’s health and optimize treatment planning.
- Optimization in the fusion and sharing of clinical information: consult, through a single frontend, the entirety of patient medical information and create a shared virtual environment where doctors can collaborate on patient care, share data, discuss clinical cases, and plan surgeries.
- Tool to improve procedures and reduce healthcare costs: use DTs in medical scenarios to simulate medical procedures in a safe virtual environment before performing them on a real patient, receive feedback to improve organizational efficiency. These simulations can be a valuable tool to assess the critical points of the entire medical supply chain and optimize the use of medical resources, such as medical devices and staff, to reduce healthcare costs.
3. Lung-DT
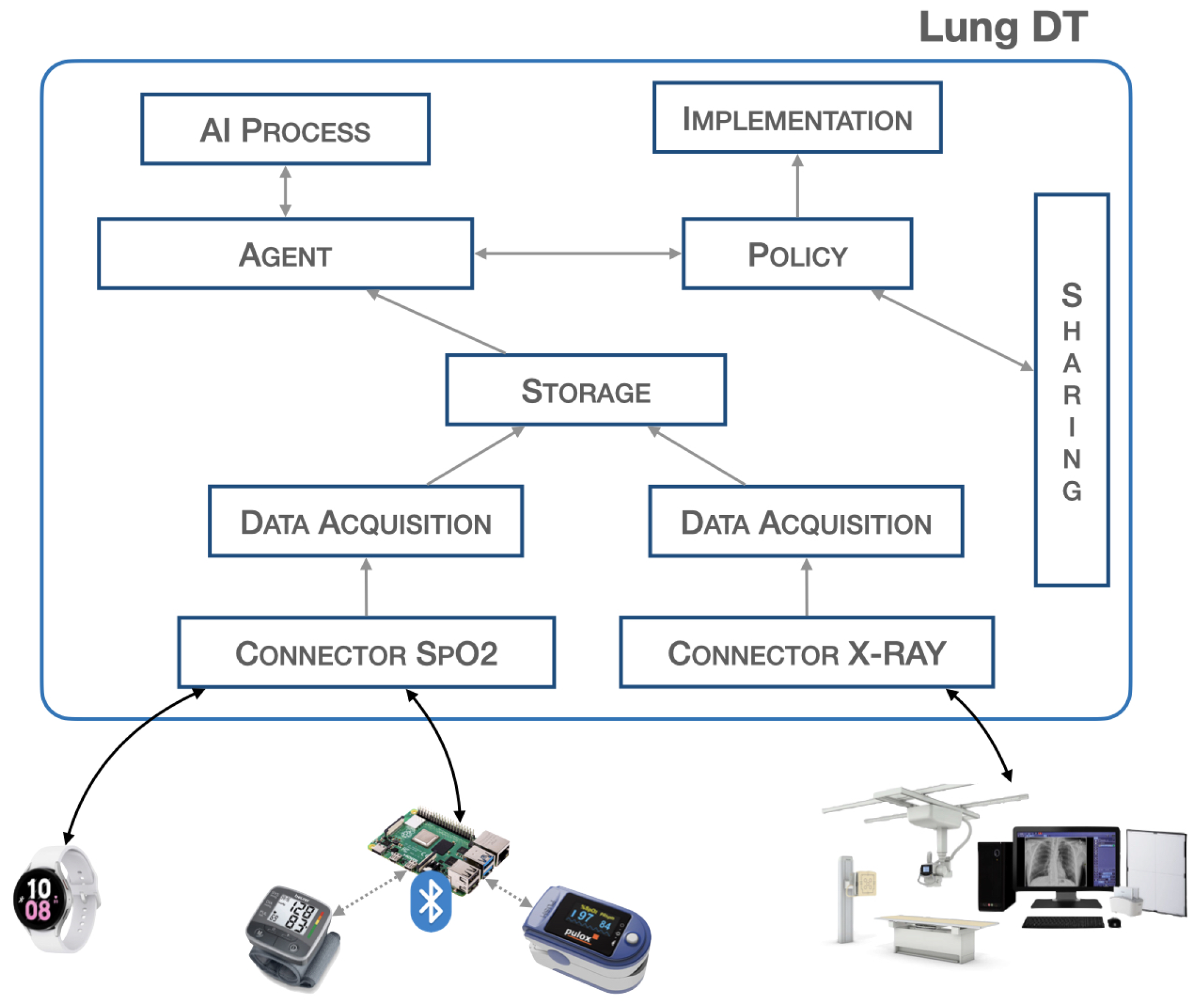
3.1. Lung-DT Architecture
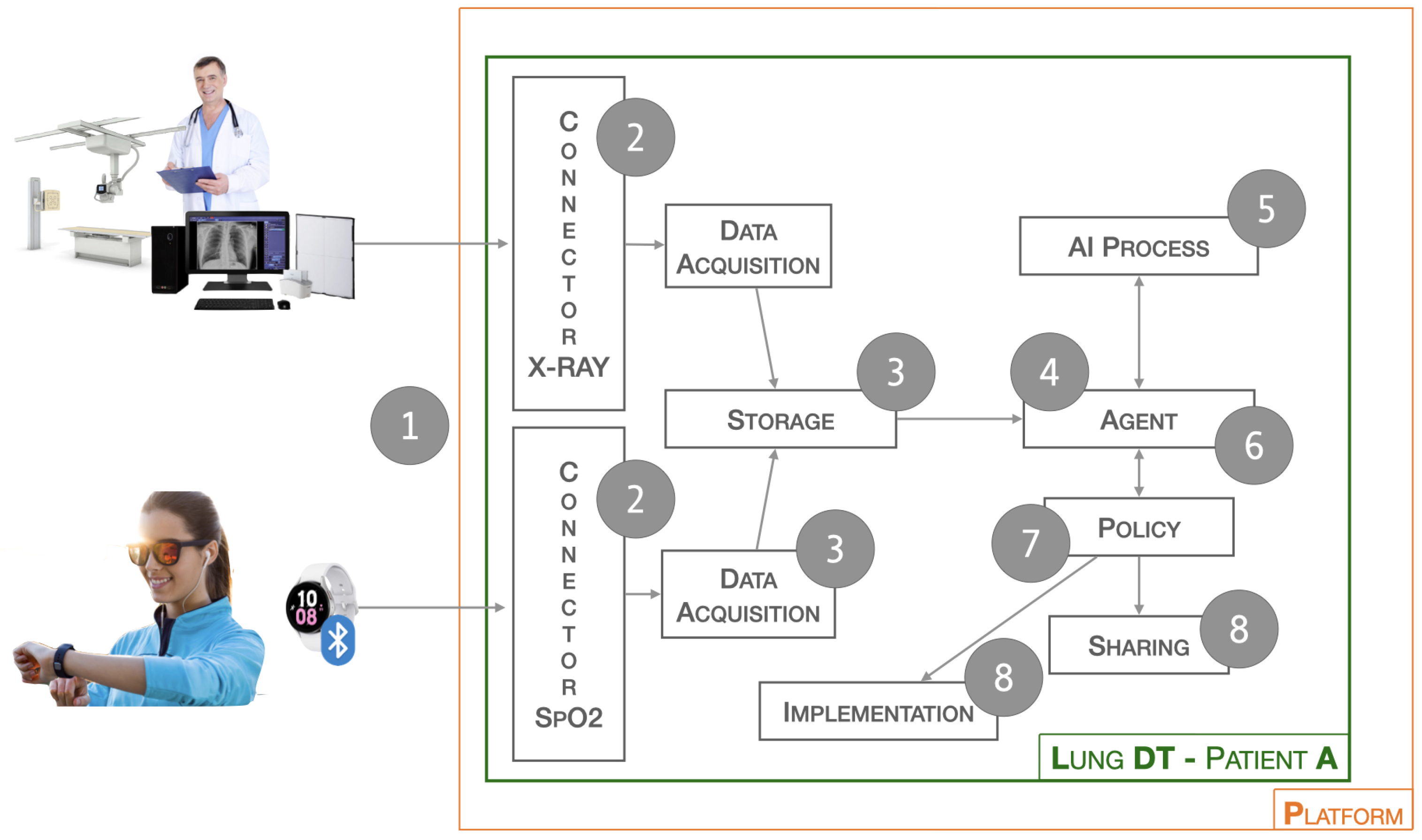
3.2. Lung-DT Workflow
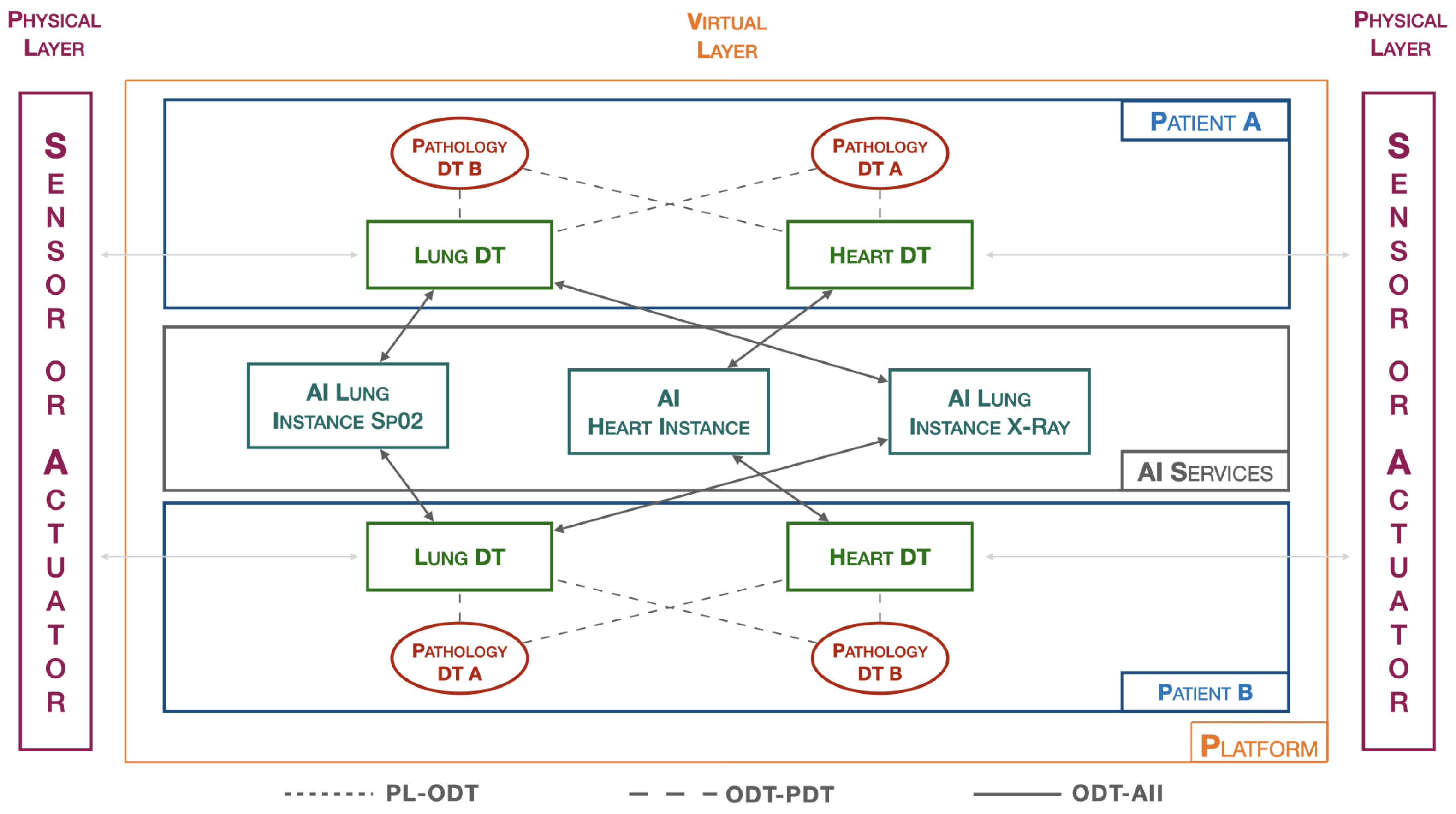
3.3. Healthcare DT platform
4. Testbed setup of Lung-DT
4.1. Datasets
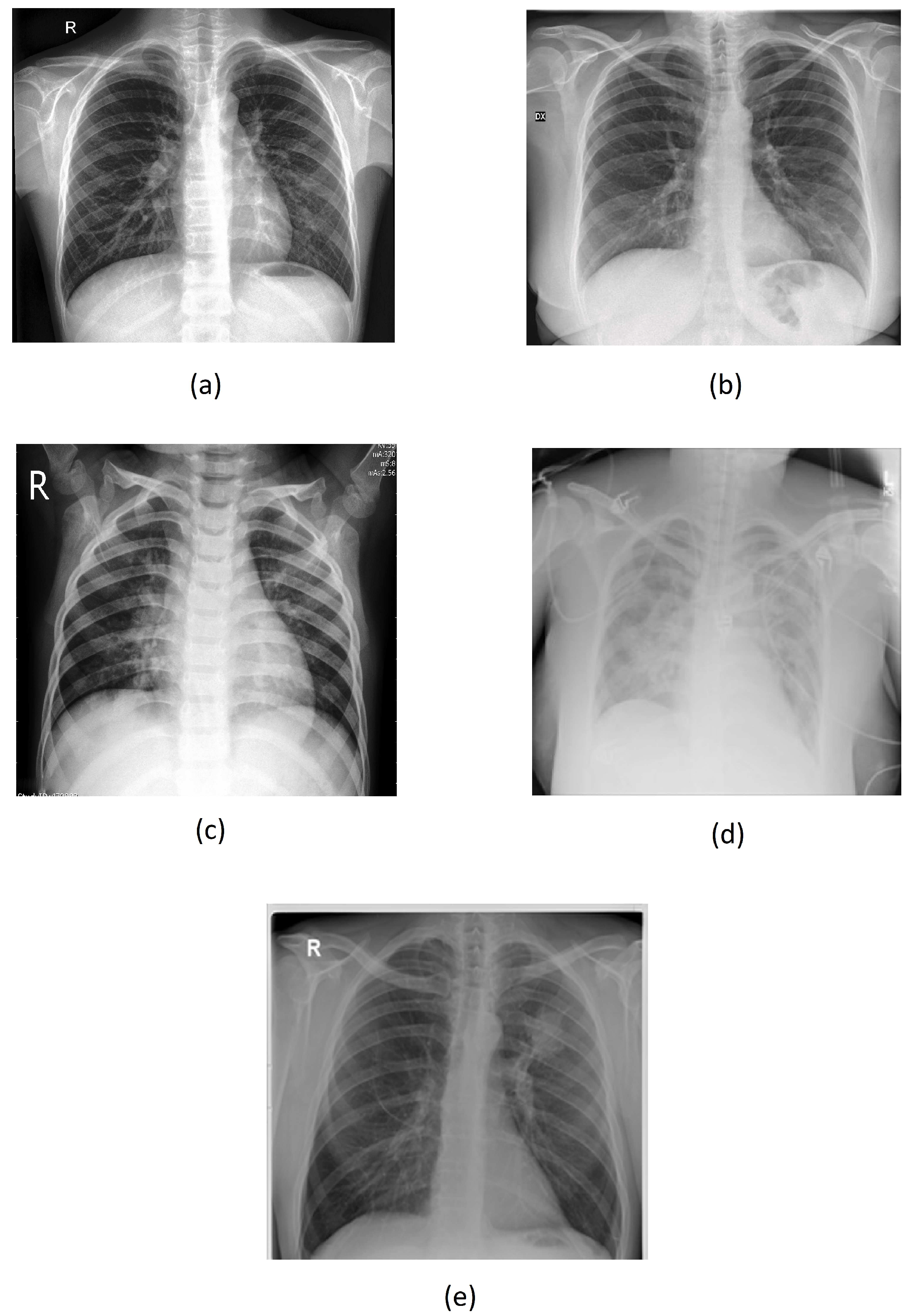
- "Normal": 1,481 images;
- "Covid": 3,997 images;
- "Pneumonia": 5,348 images;
- "Lung Opacity": 4,207 images;
- "Tuberculosis": 1,330 images.
- "Normal": 635 images;
- "Covid": 1,697 images;
- "Pneumonia": 2,291 images;
- "Lung Opacity": 1,803 images;
- "Tuberculosis": 653 images.
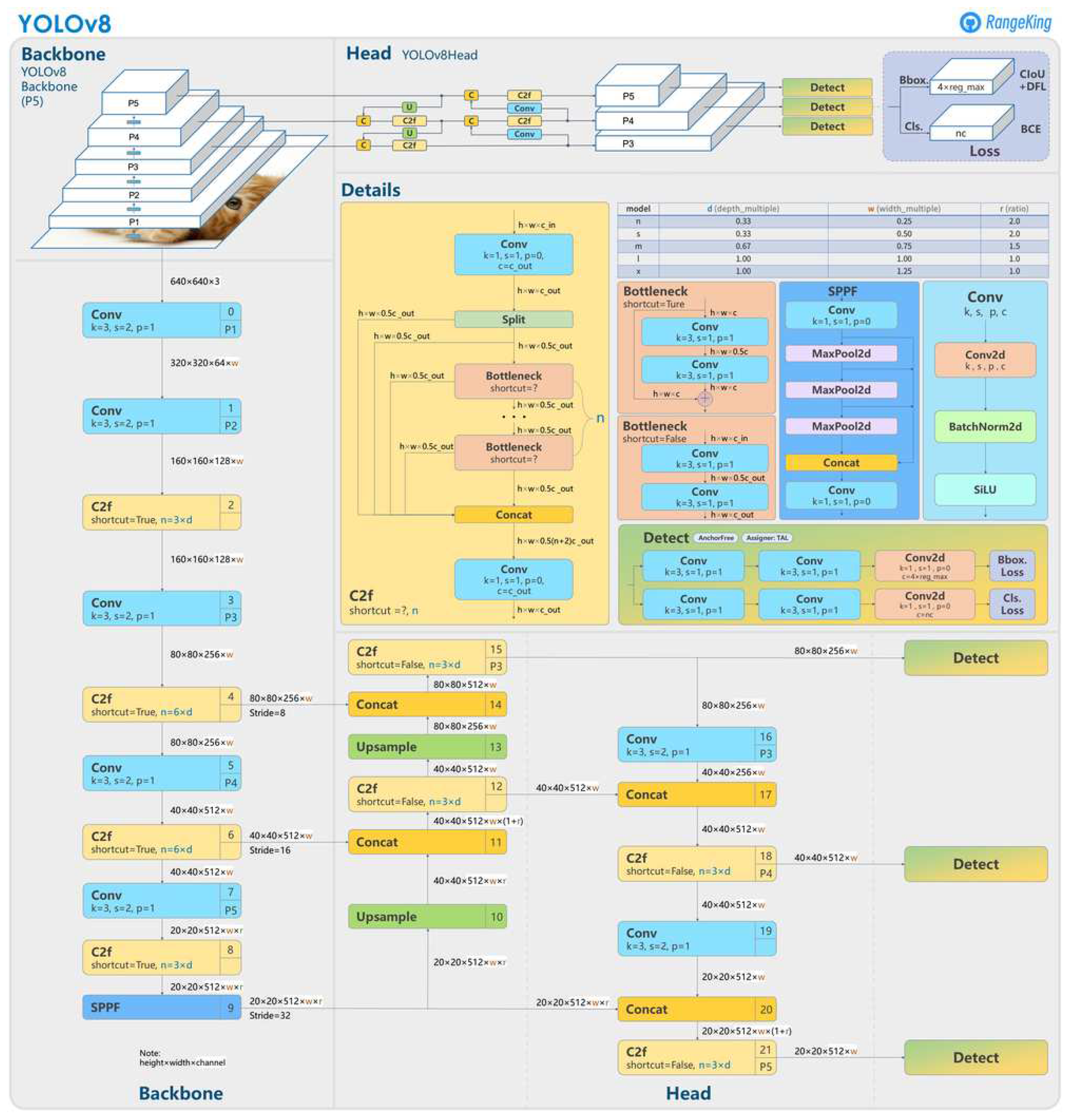
4.2. Convolutional Neural Network: YOLOv8
- High Accuracy: YOLOv8 has demonstrated high accuracy measured through COCO and Roboflow 100 metrics.
- Developer Convenience: the model offers many features for developer convenience, including an easy-to-use command-line interface (CLI) and a well-structured Python package.
- Large Community: YOLO has a large community, and the YOLOv8 community is growing. This means there are many online resources and experts in computer vision who can provide support and guidance.
- Anchor-Free Detection: YOLOv8 adopts an anchor-free model, predicting the object’s center directly rather than the offset from a known anchored box.
- New Convolutions: changes have been made to the model’s structure, including replacing 6x6 convolutions with 3x3 convolutions and modifications to the main building blocks.
- Mosaic Augmentation Closure: YOLOv8 uses mosaic augmentation during online training but disables this technique in the last ten epochs to improve performance.
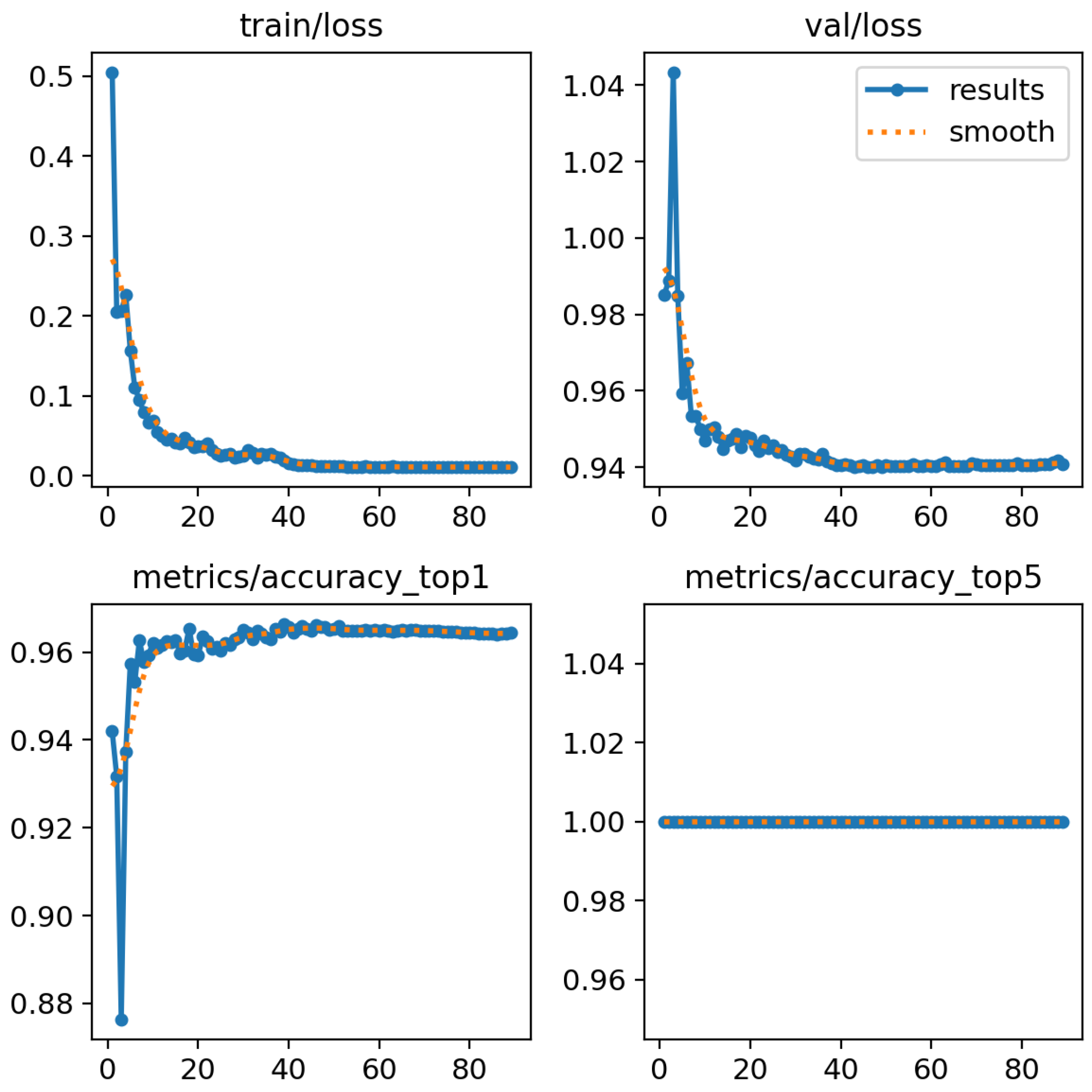
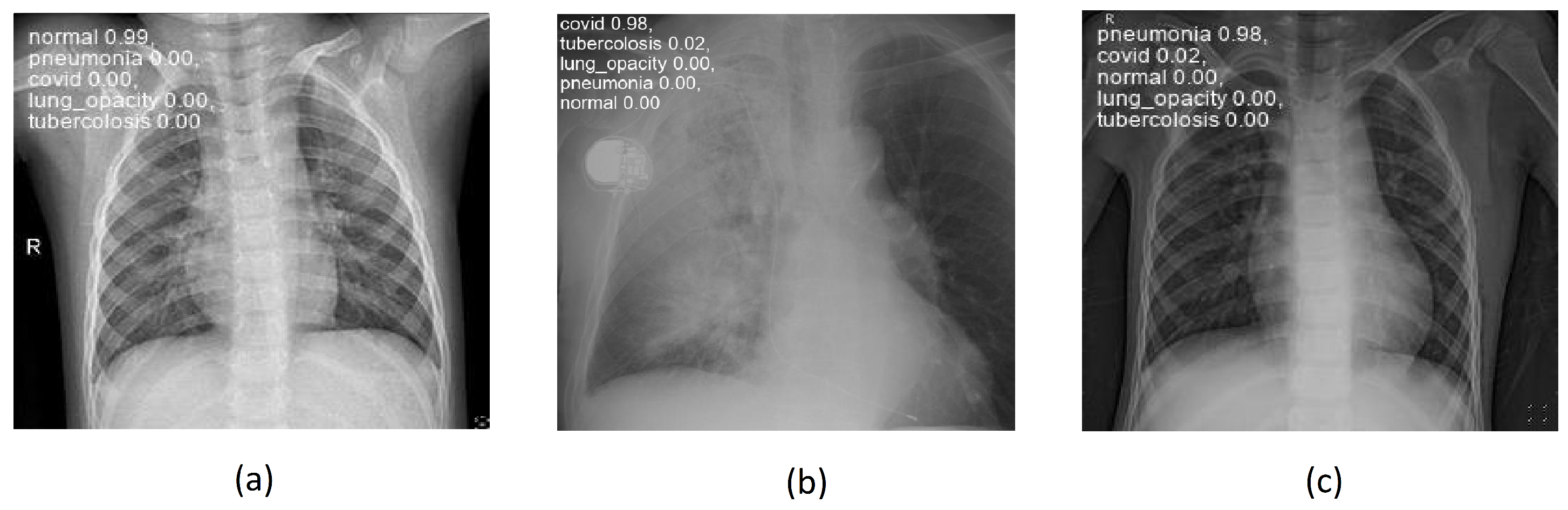
4.3. Results
5. Discussion
6. Conclusion
Author Contributions
Funding
Informed Consent Statement
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| CNN | Convolutional Neural Network |
| DL | Deep Learning |
| DT | Digital Twin |
| FTP | File Tranfer Protocol |
| GAN | Generative Adversarial Network |
| HDT | Heart Digital Twin |
| IoT | Internet of Things |
| MQTT | Message-Queuing Telemetry Transport |
| RNN | Recurrent Neural Network |
| YOLO | You Only Look Once |
References
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. ChestX-Ray8: Hospital-Scale Chest X-Ray Database and Benchmarks on Weakly-Supervised Classification and Localization of Common Thorax Diseases. 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE, 2017. [CrossRef]
- Kermany, D.S.; Goldbaum, M.; Cai, W.; Valentim, C.C.; Liang, H.; Baxter, S.L.; McKeown, A.; Yang, G.; Wu, X.; Yan, F.; Dong, J.; Prasadha, M.K.; Pei, J.; Ting, M.Y.; Zhu, J.; Li, C.; Hewett, S.; Dong, J.; Ziyar, I.; Shi, A.; Zhang, R.; Zheng, L.; Hou, R.; Shi, W.; Fu, X.; Duan, Y.; Huu, V.A.; Wen, C.; Zhang, E.D.; Zhang, C.L.; Li, O.; Wang, X.; Singer, M.A.; Sun, X.; Xu, J.; Tafreshi, A.; Lewis, M.A.; Xia, H.; Zhang, K. Identifying Medical Diagnoses and Treatable Diseases by Image-Based Deep Learning. Cell 2018, 172, 1122–1131.e9. [Google Scholar] [CrossRef] [PubMed]
- Avanzato, R.; Beritelli, F.; Lombardo, A.; Ricci, C. Heart DT: Monitoring and Preventing Cardiac Pathologies Using AI and IoT Sensors. Future Internet 2023, 15, 223. [Google Scholar] [CrossRef]
- Gazda, M.; Plavka, J.; Gazda, J.; Drotar, P. Self-supervised deep convolutional neural network for chest X-ray classification. IEEE Access 2021, 9, 151972–151982. [Google Scholar] [CrossRef]
- Hussein, F.; Mughaid, A.; AlZu’bi, S.; El-Salhi, S.M.; Abuhaija, B.; Abualigah, L.; Gandomi, A.H. Hybrid clahe-cnn deep neural networks for classifying lung diseases from x-ray acquisitions. Electronics 2022, 11, 3075. [Google Scholar] [CrossRef]
- Avanzato, R.; Beritelli, F. Thorax Disease Classification based on the Convolutional Network SqueezeNet. 12th IEEE International Conference on Intelligent Data Acquisition and Advanced Computing Systems: Technology and Applications, 2023.
- Varela-Santos, S.; Melin, P. A new modular neural network approach with fuzzy response integration for lung disease classification based on multiple objective feature optimization in chest X-ray images. Expert Systems with Applications 2021, 168, 114361. [Google Scholar] [CrossRef]
- Mabrouk, A.; Díaz Redondo, R.P.; Dahou, A.; Abd Elaziz, M.; Kayed, M. Pneumonia detection on chest X-ray images using ensemble of deep convolutional neural networks. Applied Sciences 2022, 12, 6448. [Google Scholar] [CrossRef]
- Shamrat, F.J.M.; Azam, S.; Karim, A.; Islam, R.; Tasnim, Z.; Ghosh, P.; De Boer, F. LungNet22: a fine-tuned model for multiclass classification and prediction of lung disease using X-ray images. Journal of Personalized Medicine 2022, 12, 680. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.; Bu, S. Transfer-learning-based approach for the diagnosis of lung diseases from chest X-ray images. Entropy 2022, 24, 313. [Google Scholar] [CrossRef] [PubMed]
- Alshmrani, G.M.M.; Ni, Q.; Jiang, R.; Pervaiz, H.; Elshennawy, N.M. A deep learning architecture for multi-class lung diseases classification using chest X-ray (CXR) images. Alexandria Engineering Journal 2023, 64, 923–935. [Google Scholar] [CrossRef]
- Bhosale, Y.H.; Patnaik, K.S. PulDi-COVID: Chronic obstructive pulmonary (lung) diseases with COVID-19 classification using ensemble deep convolutional neural network from chest X-ray images to minimize severity and mortality rates. Biomedical Signal Processing and Control 2023, 81, 104445. [Google Scholar] [CrossRef] [PubMed]
- Mezina, A.; Burget, R. Detection of post-COVID-19-related pulmonary diseases in X-ray images using Vision Transformer-based neural network. Biomedical Signal Processing and Control 2024, 87, 105380. [Google Scholar] [CrossRef]
- Karaddi, S.H.; Sharma, L.D. Automated multi-class classification of lung diseases from CXR-images using pre-trained convolutional neural networks. Expert Systems with Applications 2023, 211, 118650. [Google Scholar] [CrossRef]
- Rajagopal, R.; Karthick, R.; Meenalochini, P.; Kalaichelvi, T. Deep Convolutional Spiking Neural Network optimized with Arithmetic optimization algorithm for lung disease detection using chest X-ray images. Biomedical Signal Processing and Control 2023, 79, 104197. [Google Scholar] [CrossRef]
- Yadav, P.; Menon, N.; Ravi, V.; Vishvanathan, S. Lung-GANs: unsupervised representation learning for lung disease classification using chest CT and X-ray images. IEEE Transactions on Engineering Management 2021. [Google Scholar] [CrossRef]
- Sulaiman, A.; Anand, V.; Gupta, S.; Asiri, Y.; Elmagzoub, M.; Reshan, M.S.A.; Shaikh, A. A Convolutional Neural Network Architecture for Segmentation of Lung Diseases Using Chest X-ray Images. Diagnostics 2023, 13, 1651. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, I.; Ahmad, M.; Jeon, G. Integrating digital twins and deep learning for medical image analysis in the era of COVID-19. Virtual Reality & Intelligent Hardware 2022, 4, 292–305. [Google Scholar]
- Tai, Y.; Zhang, L.; Li, Q.; Zhu, C.; Chang, V.; Rodrigues, J.J.; Guizani, M. Digital-Twin-Enabled IoMT System for Surgical Simulation Using rAC-GAN. IEEE Internet of Things Journal 2022, 9, 20918–20931. [Google Scholar] [CrossRef]
- Zhu, L.; Lu, W.; Soleimani, M.; Li, Z.; Zhang, M. Electrical Impedance Tomography Guided by Digital Twins and Deep Learning for Lung Monitoring. IEEE Transactions on Instrumentation and Measurement 2023. [Google Scholar] [CrossRef]
- Xing, L.; Liu, W.; Liu, X.; Li, X. An Enhanced Vision Transformer Model in Digital Twins Powered Internet of Medical Things for Pneumonia Diagnosis. IEEE Journal on Selected Areas in Communications 2023. [Google Scholar] [CrossRef]
- Kaggle, Multiclass Chest X-Ray Disease Dataset. Available online: https://www.kaggle.com/datasets/saifurrahmanshatil/multiclass-chest-xray-disease-dataset (10 December 2023).
- Kaggle, Lungs Disease Dataset (4 types). Available online: https://www.kaggle.com/datasets/omkarmanohardalvi/lungs-disease-dataset-4-types (10 December 2023).
- Kaggle, Multi Classe Chest X-Ray DATASET(VERSION 2). Available online: https://www.kaggle.com/datasets/sourov509/multi-classe-chest-x-ray-datasetversion-2 (10 December 2023).
- YOLOv8, Roboflow,. Available online: https://blog.roboflow.com/whats-new-in-yolov8/ (15 December 2023).
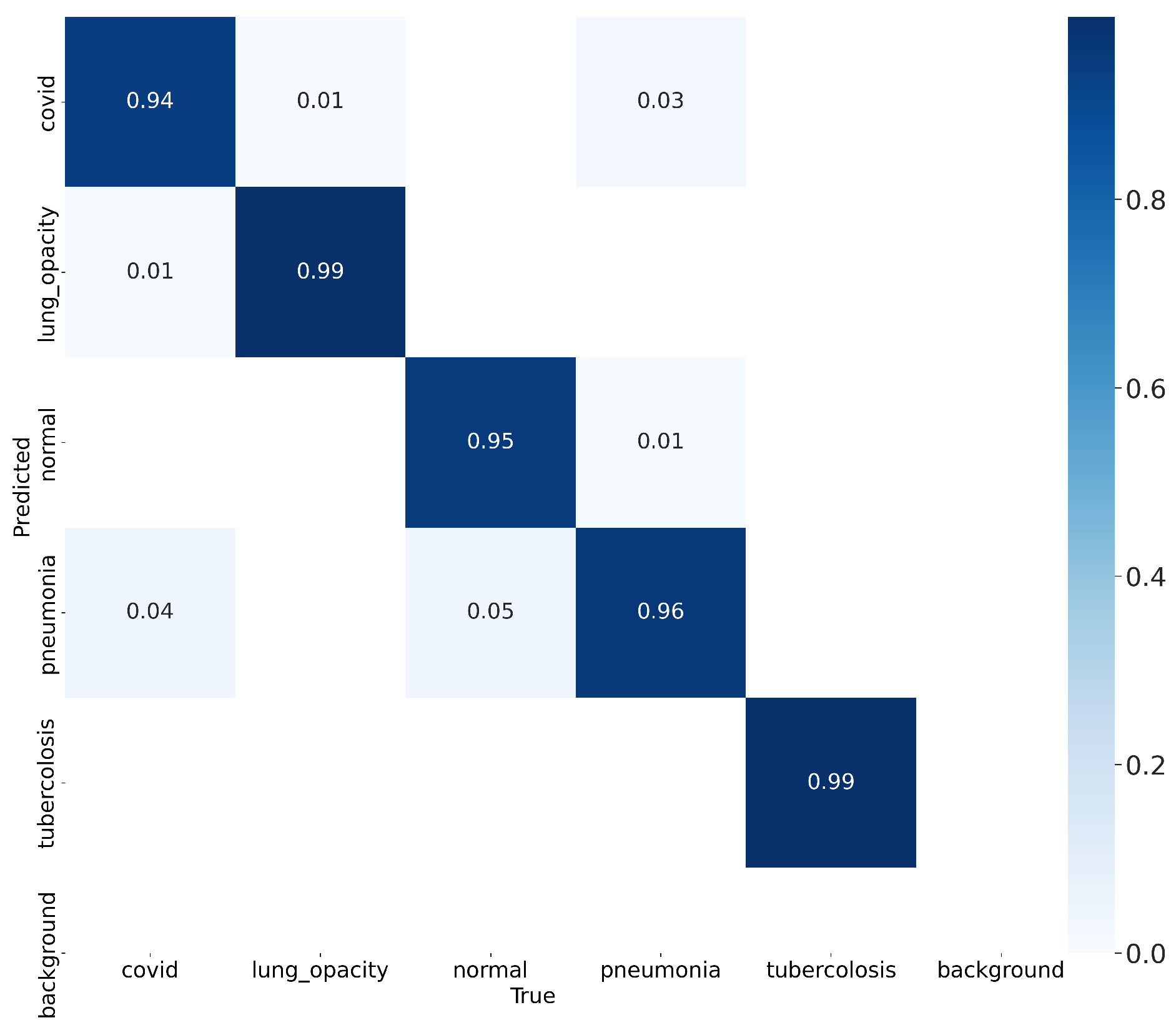
| Class Test | Average | Standard Deviation |
|---|---|---|
| Normal | 0.99 | ± 0.01 |
| Covid | 0.99 | ± 0.05 |
| Pneumonia | 0.96 | ± 0.08 |
| Ref. | Technology Used | Classes | Systems’ Accuracy [%] |
|---|---|---|---|
| [5] | Adaptive histogram equalization (CLAHE), (SVM), VGG19, and CNN networks | COVID - LUNG OPACITY - NORMAL - VIRAL PNEUMONIA | 91 |
| [6] | CNN - SqueezeNet | NORMAL - PNEUMONIA | 94 |
| [8] | CNN, DenseNet169, MobileNetV2, Vision Transformer | NORMAL - PNEUMONIA | 93 |
| [7] | Multi-objective genetic algorithm, neural networks with fuzzy logic | NORMAL - PNEUMONIA | 95 - 99 |
| [18] | RCNN, DT | COVID | 94 |
| [21] | Enhanced Vision Transformer Model, DT | NORMAL - COVID - PNEUMONIA | 88 |
| Our method | CNN - YOLOv8, IoT, DT | NORMAL - COVID - PNEUMONIA - LUNG OPACITY - TUBERCULOSIS | 96.6 - 98 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




