Submitted:
27 December 2023
Posted:
27 December 2023
You are already at the latest version
Abstract
Keywords:
Introduction
3. Basis of the decoding tables
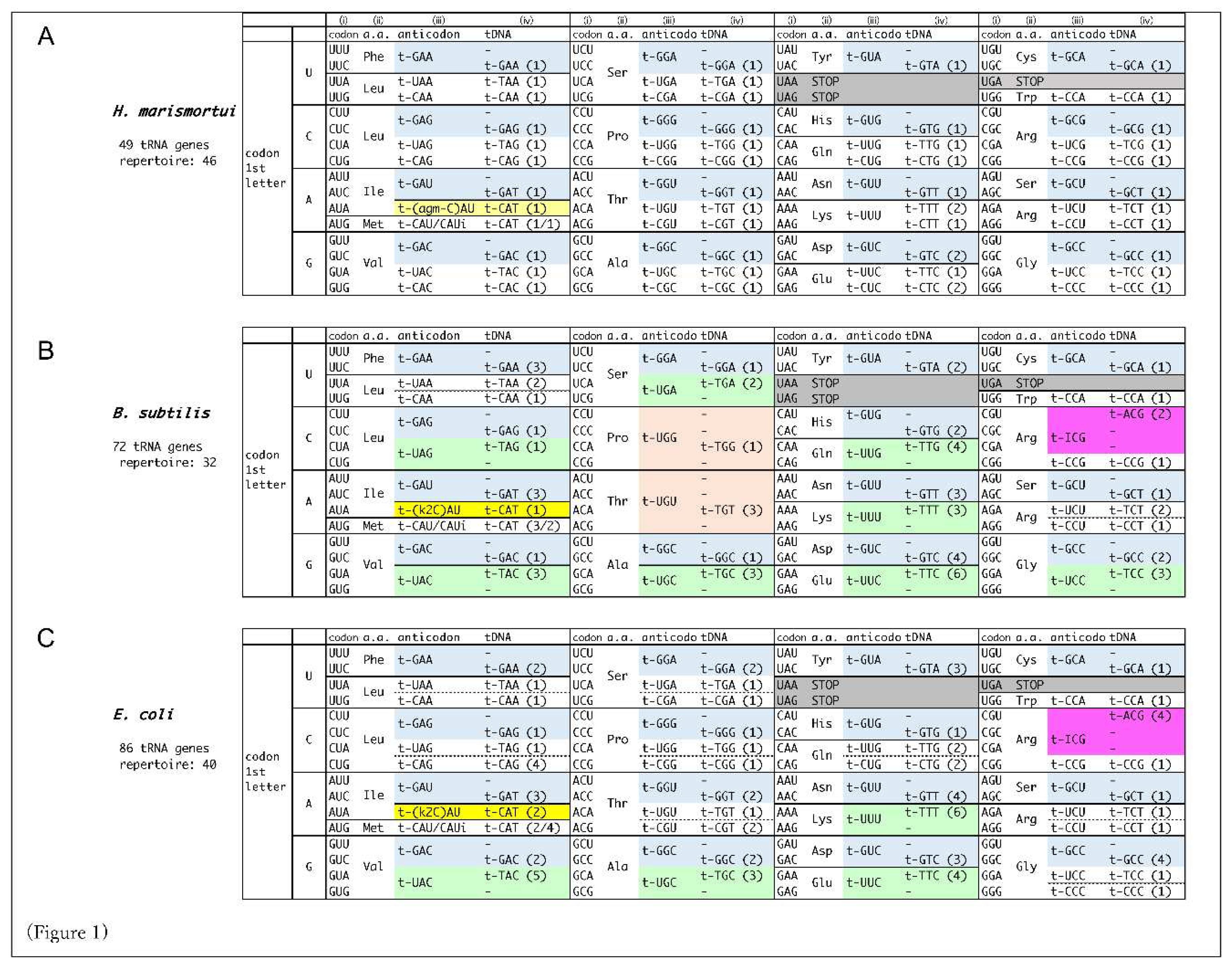
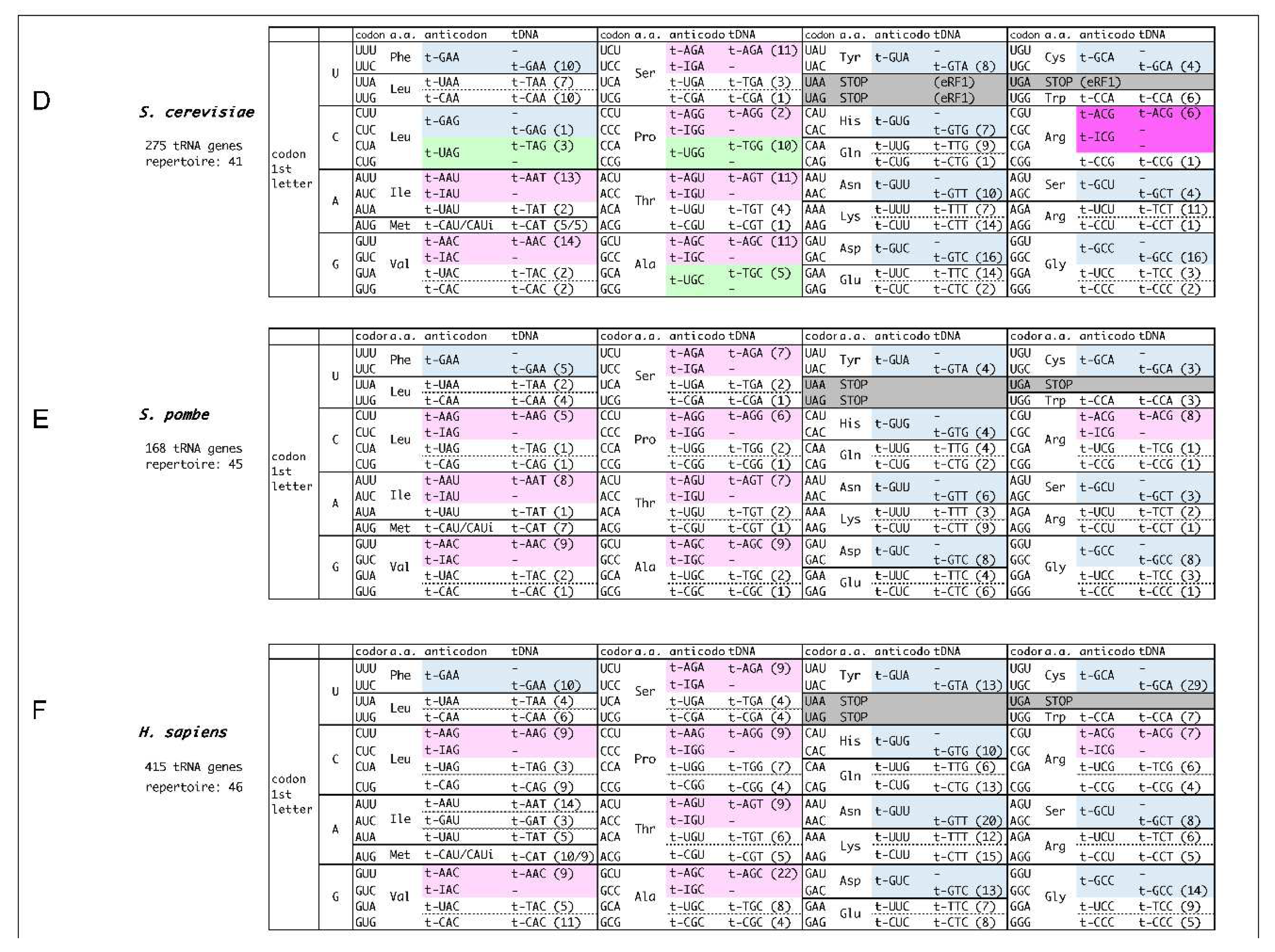
- A set of decoding information comprises (i) codons, (ii) amino acids, (iii) tRNA anticodons, and (iv) tDNA anticodons, described in this order within the boxes. The number of corresponding tRNA genes with the anticodon is shown in the parentheses within (iv). In short, (i) and (iii) represent the codon-anticodon pairing.
- tDNA anticodons (iv) are essentially derived from the tRNA genome database [4], and placed by appropriate codon-anticodon pairings. When a codon (i) forms complete Watson-Crick base pairs with a tDNA anticodon (iv) (either adenine (A)- thymine (T) or guanine (G)- cytosine (C)), the tRNA anticodon (iii) is corresponding RNA anticodon to the tDNA anticodon.
-
When a codon does not make perfect match pairing with any of tDNA anticodons, potential wobble-pairing is introduced into the 1st base of the anticodon (position 34 of tRNA) (iii) from the tDNA anticodon (iv).
- 3-1)
- 3-2)
- The G-U pairing allows the anticodon 1st U to pair with the codon 3rd G, in addition to 3rd A (shown in light green).
- 3-3)
- In some codon boxes (codons that share 1st and 2nd bases), only one tRNA repertoire with anticodon 1st U is assigned. In these cases, superwobbling, which describes the anticodons with 1st U can pair with any of codon 3rd base, is implemented (shown in orange). Superwobbling has been evidenced in some organisms [16,17].
- 3-4)
- Where the anticodon 1st G is not assigned but 1st A in codon boxes, inosine (I) is implemented to pair with both the codon 3rd C and 3rd U (shown in pale pink). In special cases of tRNAArg (ICG), the anticodon 1st I pairs with the codon 3rd A in addition to 3rd C and U, and the type of pairing is called A-I wobble, as is observed in Escherichia coli and S. cerevisiae (shown in pink) [18]. Intriguingly, the anticodon 1st inosine does not necessarily pair with U, allowing the possibility of the precursor tRNA with anticodon 1st A to be utilized. In this meaning, the extent of usage of the anticodon 1st inosine containing tRNAs (i-tRNAs) is yet to be determined, though i-tRNAs are definitely necessary to decipher C-ending codons in the boxes.
- In prokaryotes and archaea, lysidine(k2) and agmatidine(agm) modifications of the anticodon 1st C of CAU are used to decipher AUA codon, respectively, according to previous reports. These k2-CAU and amg-CAU are deciphered by tRNAIle as isoleucine, distinguishing AUG codon for methionine (shown yellow) [11,12].
4. Comparative wobble pairing usage
5. Around initiation and termination codons
6. Evaluation of the decoding tables
Supplementary Materials
Funding
Acknowledgments
Conflicts of Interest
Abbreviation
References
- Crick:F.H.C. [1966] Codon—anticodon pairing: The wobble hypothesis. Journal of Molecular Biology, 19, 548–555. [CrossRef]
- Holley,R. [1965] Structure of an alanine transfer ribonucleic acid. science, 194, 868–871. [CrossRef]
- Madison,J., Eferett,G. and Kung,H. [1966] Nucleotide sequence of a yeast tyrosine transfer RNA. science, 153, 531–534. [CrossRef]
- Lowe,T.M. and Chan,P.P. [2016] tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Research, 44, W54–W57. [CrossRef]
- Chan,P.P., Lin,B.Y., Mak,A.J. and Lowe,T.M. [2021] classification of transfer RNA genes. 49, 9077–9096. [CrossRef]
- Cherry,J.M., Hong,E.L., Amundsen,C., Balakrishnan,R., Binkley,G., Chan,E.T., Christie,K.R., Costanzo,M.C., Dwight,S.S., Engel,S.R., et al. [2012] Saccharomyces Genome Database: The genomics resource of budding yeast. Nucleic Acids Research, 40, 700–705. [CrossRef]
- Dunin-Horkawicz,S., Czerwoniec,A., Gajda,M.J., Feder,M., Grosjean,H. and Bujnicki,J.M. [2006] MODOMICS: a database of RNA modification pathways. Nucleic acids research, 34, 145–149. [CrossRef]
- Cappannini,A., Ray,A. and Mukherjee,S. [2023] MODOMICS : a database of RNA modi cations and related inf ormation . 2023 updat e.
- Grosjean,H., de Crécy-Lagard,V. and Marck,C. [2010] Deciphering synonymous codons in the three domains of life: Co-evolution with specific tRNA modification enzymes. FEBS Letters, 584, 252–264. [CrossRef]
- Gerber,A.P. and Keller,W. [1999] An adenosine deaminase that generates inosine at the wobble position of tRNAs. Science, 286, 1146–1149. [CrossRef]
- Suzuki,T. and Miyauchi,K. [2010] Discovery and characterization of tRNAIle lysidine synthetase (TilS). FEBS Letters, 584, 272–277. [CrossRef]
- Mandal,D., Köhrer,C., Su,D., Russell,S.P., Krivos,K., Castleberry,C.M., Blum,P., Limbach,P.A., Söll,D. and RajBhandary,U.L. [2010] Agmatidine, a modified cytidine in the anticodon of archaeal tRNA Ile, base pairs with adenosine but not with guanosine. Proceedings of the National Academy of Sciences of the United States of America, 107, 2872–2877. [CrossRef]
- Volume 5 Number 11 November 1978 Nucleic Acids Research [1978] Nucleic Acids Research, 5, 4451–4462. [CrossRef]
- Demeshkina,N., Jenner,L., Westhof,E., Yusupov,M. and Yusupova,G. [2013] New structural insights into the decoding mechanism: Translation infidelity via a G·U pair with Watson-Crick geometry. FEBS Letters, 587, 1848–1857. [CrossRef]
- Ananth,P., Goldsmith,G. and Yathindra,N. [2013] An innate twist between Crick’s wobble and Watson-Crick base pairs. Rna, 19, 1038–1053. [CrossRef]
- Rogalski,M., Karcher,D. and Bock,R. [2008] Superwobbling facilitates translation with reduced tRNA sets. Nature Structural and Molecular Biology, 15, 192–198. [CrossRef]
- Lei,L. and Burton,Z.F. [2022] “Superwobbling” and tRNA-34 Wobble and tRNA-37 Anticodon Loop Modifications in Evolution and Devolution of the Genetic Code. Life, 12. [CrossRef]
- Wada,M. and Ito,K. [2023] The CGA codon decoding through tRNAArg(ICG) supply governed by Tad2/Tad3 in Saccharomyces cerevisiae. FEBS Journal, 290, 3480–3489. [CrossRef]
- Lozupone,C.A., Knight,R.D. and Landweber,L.F. [2001] The molecular basis of nuclear genetic code change in ciliates. Current Biology, 11, 65–74. [CrossRef]
- Kim,O.T.P., Sakurai,A., Saito,K., Ito,K., Ikehara,K. and Harumoto,T. [2008] Ciliates use both variant and universal genetic codes: Evidence of omnipotent eRF1s in the class Litostomatea. Gene, 417, 51–58. [CrossRef]
- Novoa,E.M., Pavon-Eternod,M., Pan,T. and Ribas De Pouplana,L. [2012] A role for tRNA modifications in genome structure and codon usage. Cell, 149, 202–213. [CrossRef]
- Rafels-Ybern,À., Torres,A.G., Camacho,N., Herencia-Ropero,A., Frigolé,H.R., Wulff,T.F., Raboteg,M., Bordons,A., Grau-Bove,X., Ruiz-Trillo,I., et al. [2019] The expansion of inosine at the wobble position of tRNAs, and its role in the evolution of proteomes. Molecular Biology and Evolution, 36, 650–662. [CrossRef]
- Ramos-Morales,E., Bayam,E., Del-Pozo-Rodríguez,J., Salinas-Giegé,T., Marek,M., Tilly,P., Wolff,P., Troesch,E., Ennifar,E., Drouard,L., et al. [2021] The structure of the mouse ADAT2/ADAT3 complex reveals the molecular basis for mammalian tRNA wobble adenosine-to-inosine deamination. Nucleic Acids Research, 49, 6529–6548. [CrossRef]
- Wolf,J., Gerber,A.P. and Keller,W. [2002] TadA, an essential tRNA-specific adenosine deaminase from Escherichia coli. EMBO Journal, 21, 3841–3851. [CrossRef]
- Agris,P.F., Eruysal,E.R., Narendran,A., Väre,V.Y.P., Vangaveti,S. and Ranganathan,S. V. [2018] Celebrating wobble decoding: Half a century and still much is new. RNA Biology, 15, 537–553. [CrossRef]
- Ito,K., Uno,M. and Nakamura,Y. [2000] A tripeptide ‘anticodon’ deciphers stop codons in messenger RNA. Nature, 403, 680–684. [CrossRef]
- Wada,M. and Ito,K. [2019] Misdecoding of rare CGA codon by translation termination factors, eRF1/eRF3, suggests novel class of ribosome rescue pathway in S. cerevisiae. FEBS Journal, 286, 788–802. [CrossRef]
- Gomez,M.A.R. and Ibba,M. [2020] Aminoacyl-tRNA synthetases. Rna, 26, 910–936. [CrossRef]
- Gieg,R. and Eriani,G. [2023] The tRNA identity landscape for aminoacylation and beyond. Nucleic Acids Research, 51, 1528–1570. [CrossRef]
- Abelson,J. [1989] tRNA IDENTITY Jennifer Normanl / and John Abelson. Identity.
- Rubio,M.A.T., Ragone,F.L., Gaston,K.W., Ibba,M. and Alfonzo,J.D. [2006] C to U editing stimulates A to I editing in the anticodon loop of a cytoplasmic threonyl tRNA in Trypanosoma brucei. Journal of Biological Chemistry, 281, 115–120. [CrossRef]
- Bertotti,S., Fleming,I., de los Milagros Cámara,M., Cameán,C.C., Carmona,S.J., Agüero,F., Balouz,V., Zahn,A., Di Noia,J.M., Alfonzo,J.D., et al. [2022] Characterization of ADAT2/3 molecules in Trypanosoma cruzi and regulation of mucin gene expression by tRNA editing. Biochemical Journal, 479, 561–580. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



