Submitted:
21 December 2023
Posted:
25 December 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
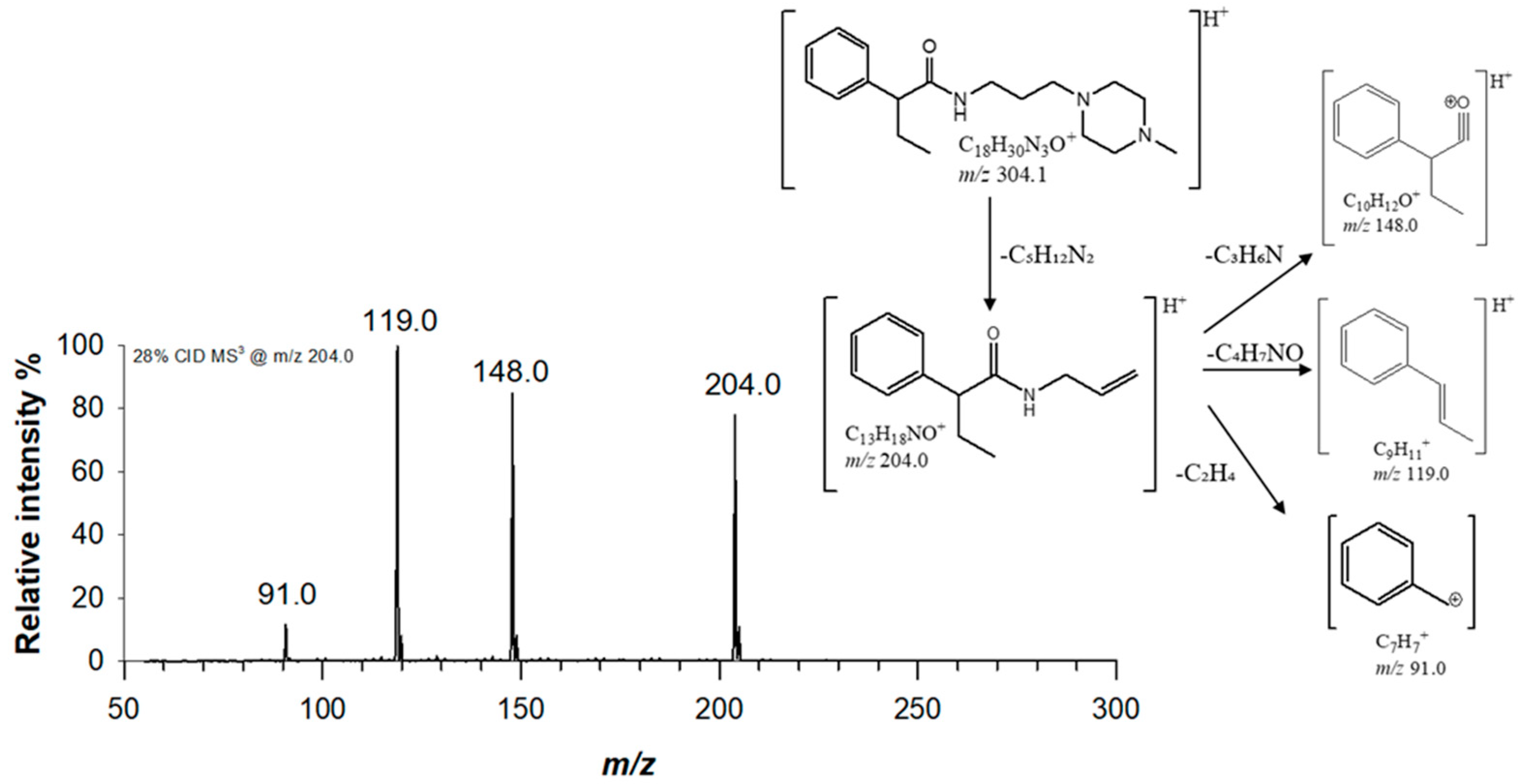
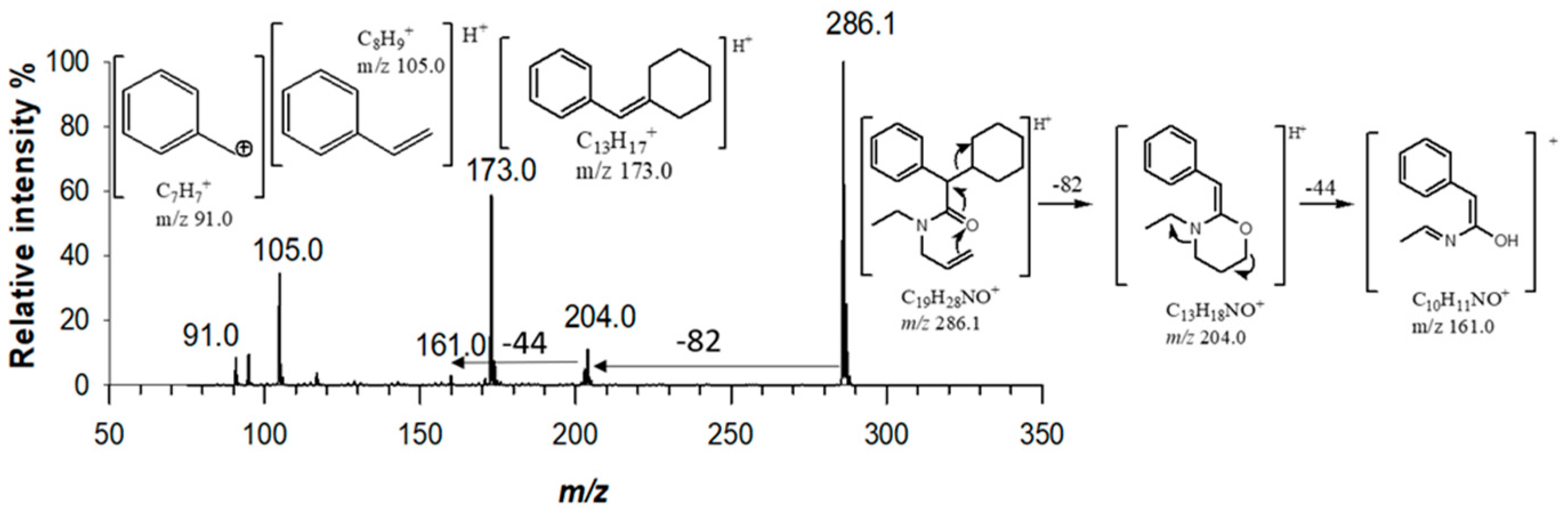
2.1. Mass Spectrometric characterization of Piperazine derivatives
2.2. Mass Spectrometric characterization of Morpholine derivatives
2.3. Antimicrobial activity evaluation
3. Discussion
4. Materials and Methods
4.1. Synthesis of derivative (1)
4.2. Synthesis of derivative (2)
4.3. Synthesis of derivatives (3), (4), (5) and (6)
4.4. Mass spectrometric analysis
4.5. Antimicrobial activity
2.5.1. Bacterial strains and growth conditions
2.5.2. Agar Well Diffusion Assay and Minimum Inhibitory Concentration (MIC)
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Davies, J. Where have all the raises gone? Can. J. Infect. Dis. Med. Microbiol. 2006, 17, 287–290.
- Hutchings, M.; Truman, A.; Wilkinson, B. Antibiotics: past, present and future. Curr. Opin. Microbiol. 2019, 51, 72–80. [CrossRef]
- Gaskins, H.R.; Collier, C.T.; Anderson, D.B. Antibiotics as growth promotants: Mode of action. Anim. Biotechnol. 2002, 13, 29–42. [CrossRef]
- Prescott, J.F. The resistance tsunami, antimicrobial stewardship, and the golden age of microbiology. Vet. Microbiol. 2014, 171, 273–278. [CrossRef]
- Macgowan, A. Antibiotic resistance. Medicine (Baltimore). 2017, 45, 622–628. [CrossRef]
- Singh, S.B.; Young, K.; Silver, L.L. What is an “ideal” antibiotic? Discovery challenges and path forward. Biochem. Pharmacol. 2017, 133, 63–73. [CrossRef]
- Gettys, K.E.; Ye, Z.; Dai, M. Recent Advances in Piperazine Synthesis. Synth. 2017, 49, 2589–2604. [CrossRef]
- Al-Neaimy, U.I.S.; Saeed, Z.F.; Shehab, S.M. A Review on Analytical Methods for Piperazine Determination. NTU J. Pure Sci. 2022, 1, 1–9. [CrossRef]
- Walker, M.A. Novel tactics for designing water-soluble molecules in drug discovery. Expert Opin. Drug Discov. 2014, 9, 1421–1433. [CrossRef]
- Kourounakis, A.P.; Xanthopoulos, D.; Tzara, A. Morpholine as a privileged structure: A review on the medicinal chemistry and pharmacological activity of morpholine containing bioactive molecules. Med. Res. Rev. 2020, 40, 709–752. [CrossRef]
- Al-Ghorbani, M.; Begum, B.; Zabiulla; Mamatha, V.S.; Khanum, S.A. Piperazine and morpholine: Synthetic preview and pharmaceutical applications. J. Chem. Pharm. Res. 2015, 7, 281–301.
- Kumar Rupak , Srinivasa R Vulichi, K.S. Emphasizing morpholine and its derivatives (MAID): a typical candidates of pharmaceutical importance. Int. J. Chem. Sci. 2016, 14, 1777–1788.
- Welz, A.; Koba, M.; Kośliński, P.; Siódmiak, J. Rapid targeted method of detecting abused piperazine designer drugs. J. Clin. Med. 2021, 10. [CrossRef]
- Kőnig, B.; Sztanó, G.; Holczbauer, T.; Soós, T. Syntheses of 2- and 3-Substituted Morpholine Congeners via Ring Opening of 2-Tosyl-1,2-Oxazetidine. J. Org. Chem. 2023, 88, 6182–6191. [CrossRef]
- Niessen, W.M.A.; Rosing, H.; Beijnen, J.H. Interpretation of MS–MS spectra of small-molecule signal transduction inhibitors using accurate-m/z data and m/z-shifts with stable-isotope-labeled analogues and metabolites. Int. J. Mass Spectrom. 2021, 464, 116559. [CrossRef]
- Kim, E.H.; Seo, H.S.; Ki, N.Y.; Park, N.H.; Lee, W.; Do, J.A.; Park, S.; Baek, S.Y.; Moon, B.; Oh, H. Bin; et al. Reliable screening and confirmation of 156 multi-class illegal adulterants in dietary supplements based on extracted common ion chromatograms by ultra-high-performance liquid chromatography-quadrupole/time of flight-mass spectrometry. J. Chromatogr. A 2017, 1491, 43–56. [CrossRef]
- Paneth, A.; Trotsko, N.; Popiołek, Ł.; Grzegorczyk, A.; Krzanowski, T.; Janowska, S.; Malm, A.; Wujec, M. Synthesis and Antibacterial Evaluation of Mannich Bases Derived from 1,2,4-Triazole. Chem. Biodivers. 2019, 16. [CrossRef]
- Turan-Zitouni, G.; Kaplancikli, Z.A.; Yildiz, M.T.; Chevallet, P.; Kaya, D. Synthesis and antimicrobial activity of 4-phenyl/cyclohexyl-5-(1- phenoxyethyl)-3-[N-(2-thiazolyl)acetamido]thio-4H-1,2,4-triazole derivatives. Eur. J. Med. Chem. 2005, 40, 607–613. [CrossRef]
- Patil, M.; Noonikara-Poyil, A.; Joshi, S.D.; Patil, S.A.; Patil, S.A.; Bugarin, A. New urea derivatives as potential antimicrobial agents: Synthesis, biological evaluation, and molecular docking studies. Antibiotics 2019, 8. [CrossRef]
- Kosikowska, U.; Wujec, M.; Trotsko, N.; Płonka, W.; Paneth, P.; Paneth, A. Antibacterial activity of fluorobenzoylthiosemicarbazides and their cyclic analogues with 1,2,4-triazole scaffold. Molecules 2021, 26, 1–18. [CrossRef]
- Wujec, M.; Kosikowska, U.; Siwek, A.; Malm, A. New Derivatives of Thiosemicarbazide and 1,2,4-Triazoline-5-thione with Potential Antimicrobial Activity. Phosphorus, Sulfur Silicon Relat. Elem. 2009, 184, 559–567. [CrossRef]
- Bonomo, M.G.; Cafaro, C.; Russo, D.; Calabrone, L.; Milella, L.; Saturnino, C.; Capasso, A.; Salzano, G. Antimicrobial Activity, Antioxidant Properties and Phytochemical Screening of Aesculus hippocastanum Mother Tincture against Food-borne Bacteria. Lett. Drug Des. Discov. 2018, 17, 48–56. [CrossRef]
- Siwek, A.; Wujec, M.; Dobosz, M.; Jagiełło-Wójtowicz, E.; Chodkowska, A.; Kleinrok, A.; Paneth, P. Synthesis and pharmacological properties of 3-(2-methyl-furan-3-yl)-4-substituted-Δ 2 - 1,2,4-triazoline-5-thiones. Cent. Eur. J. Chem. 2008, 6, 47–53. [CrossRef]
- Kharb, R.; Bansal, K.; Sharma, A.K. A valuable insight into recent advances on antimicrobial activity of piperazine derivatives. Der Pharma Chem. 2012, 4, 2470–2488.
- Lukin, A.; Chudinov, M.; Vedekhina, T.; Rogacheva, E.; Kraeva, L.; Bakulina, O.; Krasavin, M. Exploration of Spirocyclic Derivatives of Ciprofloxacin as Antibacterial Agents. Molecules 2022, 27, 1–20. [CrossRef]
- Hashemian, S.M.R.; Farhadi, T.; Ganjparvar, M. Linezolid: A review of its properties, function, and use in critical care. Drug Des. Devel. Ther. 2018, 12, 1759–1767. [CrossRef]
- De Rosa, M.; Bonomo, M.G.; Vassallo, A.; Palma, G.; Calabrone, L.; Bimonte, S.; Silvestris, N.; Amruthraj, N.J.; Sinicropi, M.S.; Salzano, G.; et al. Linezolid and its derivatives: The promising therapeutic challenge to multidrug-resistant pathogens. Pharmacologyonline 2018, 2, 134–148.
- Bonomo, M.G.; Calabrone, L.; Saturnino, C.; Sinicropi, M.S.; Capasso, A.; Salzano, G. Antibacterial activity of new β-lactam compound. Pharmacologyonline 2019, 3, 185–194.
- Lee, S.K.; Choi, K.H.; Lee, S.J.; Suh, S.W.; Kim, B.M.; Lee, B.J. Peptide deformylase inhibitors with retro-amide scaffold: Synthesis and structure-activity relationships. Bioorganic Med. Chem. Lett. 2010, 20, 4317–4319. [CrossRef]
- Bonomo, M.G.; Giura, T.; Salzano, G.; Longo, P.; Mariconda, A.; Catalano, A.; Iacopetta, D.; Ceramella, J.; Sinicropi, M.S.; Saturnino, C. Bis-Thiourea Quaternary Ammonium Salts as Potential Agents against Bacterial Strains from Food and Environmental Matrices. Antibiotics 2021, 10. [CrossRef]
- Birnie, C.R.; Malamud, D.; Schnaare, R.L. Antimicrobial evaluation of N-Alkyl betaines and N-Alkyl-N,N-dimethylamine oxides with variations in chain length. Antimicrob. Agents Chemother. 2000, 44, 2514–2517. [CrossRef]
- Cui, Y.J.; Rao, X.P.; Shang, S. Bin; Song, Z.Q.; Shen, M.G.; Liu, H. Synthesis, structure analysis and antibacterial activity of N-[5-dehydroabietyl-[1,3,4]thiadiazol-2-yl]-aromatic amide derivatives. J. Saudi Chem. Soc. 2017, 21, S258–S263. [CrossRef]
- Bayrak, H.; Demirbas, A.; Karaoglu, S.A.; Demirbas, N. Synthesis of some new 1,2,4-triazoles, their Mannich and Schiff bases and evaluation of their antimicrobial activities. Eur. J. Med. Chem. 2009, 44, 1057–1066. [CrossRef]
- Saturnino, C.; Grande, F.; Aquaro, S.; Caruso, A.; Iacopetta, D.; Bonomo, M.G.; Longo, P.; Schols, D.; Sinicropi, M.S. Chloro-1,4-dimethyl-9H-carbazole derivatives displaying anti-HIV activity. Molecules 2018, 23. [CrossRef]
- Adamson, J.T.; Hakansson, K. Electrospray Ionization Fourier Transform Ion Cyclotron Resonance Mass Spectrometry for Lectin Analysis. Lectins Anal. Technol. 2007, 343–371. [CrossRef]
- Zimmer, J.S.D.; Monroe, M.E.; Qian, W.J.; Smith, R.D. Advances in proteomics data analysis and display using an accurate mass and time tag approach. Mass Spectrom. Rev. 2006, 25, 450–482. [CrossRef]
- Onzo, A.; Acquavia, M.A.; Pascale, R.; Iannece, P.; Gaeta, C.; Nagornov, K.O.; Tsybin, Y.O.; Bianco, G. Metabolic profiling of Peperoni di Senise PGI bell peppers with ultra-high resolution absorption mode Fourier transform ion cyclotron resonance mass spectrometry. Int. J. Mass Spectrom. 2021, 470, 116722. [CrossRef]
- Onzo, A.; Acquavia, M.A.; Pascale, R.; Iannece, P.; Gaeta, C.; Lelario, F.; Ciriello, R.; Tesoro, C.; Bianco, G.; Di Capua, A. Untargeted metabolomic analysis by ultra-high-resolution mass spectrometry for the profiling of new Italian wine varieties. Anal. Bioanal. Chem. 2022, 7805–7812. [CrossRef]
- Lin, Z.; Zhu, C.; Xia, H. HRMS studies on the fragmentation pathways of metallapentalyne. Spectrochim. Acta - Part A Mol. Biomol. Spectrosc. 2015, 136, 906–910. [CrossRef]
- Saturnino, C.; Capasso, A.; Lancelot, J.C.; Rault, S.; Buonerba, M.; De Martino, G. Pharmacological studies and synthesis of morpholino alkyl derivatives. Chem. Pharm. Bull. 2002, 50, 1151–1154. [CrossRef]
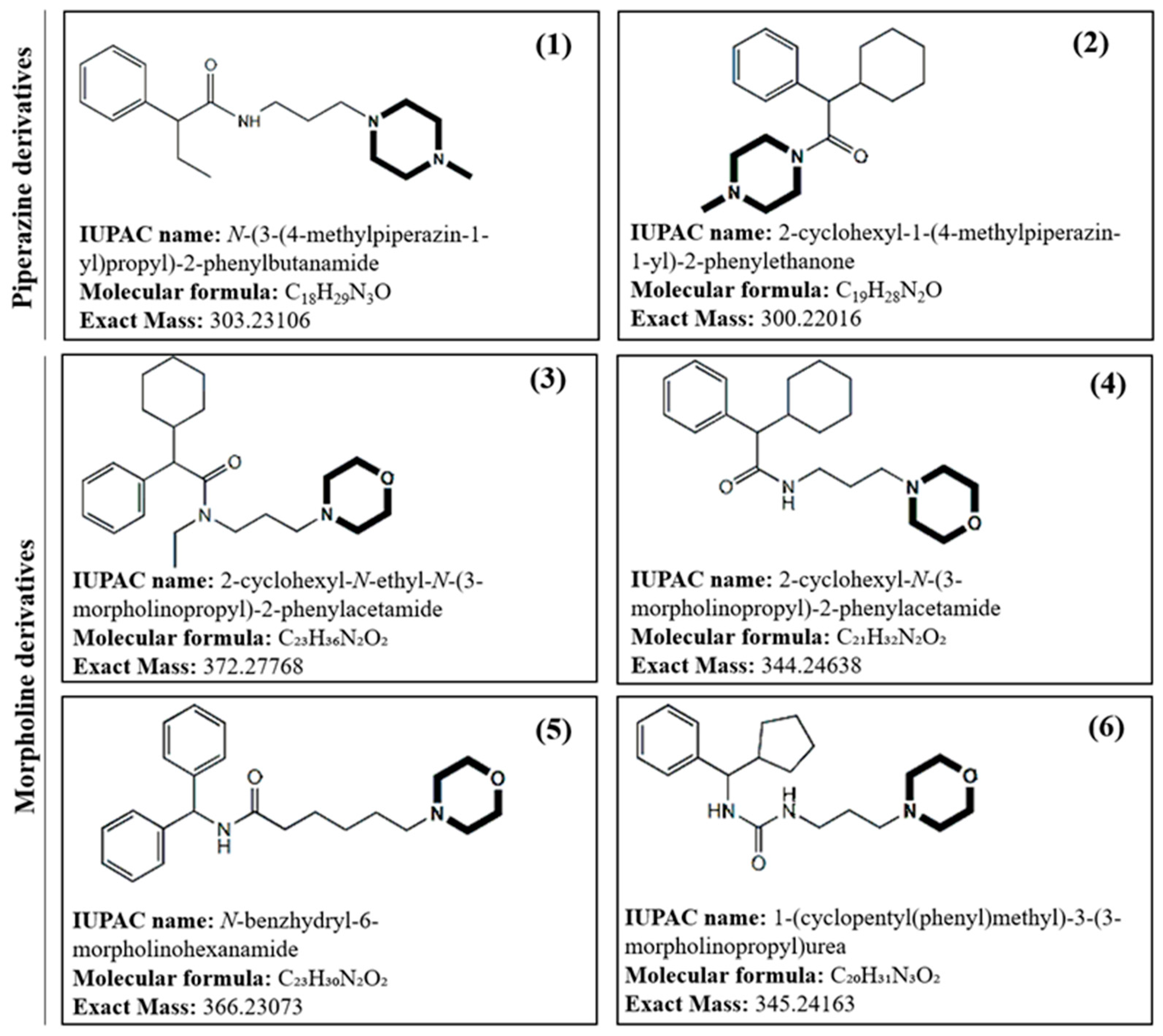
| Compound Molecular Formula Exact mass |
m/za | Relative Intensity (%) | Mass accuracy in RMSb ppm |
ESI-(+)-FT ICR MS spectra |
|---|---|---|---|---|
|
(1) C18H30N3O 303.23106 |
[M+H]+ | 100 | 1.6 | 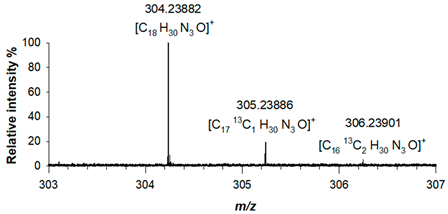 |
| 304.23882 | ||||
| [M+H+1]+ | 19.27 | 1.7 | ||
| 305.23886 | ||||
| [M+H+2]+ | 1.69 | 2.2 | ||
| 306.23901 | ||||
|
(2) C19H28N2O 300.22016 |
[M+H]+ | 100 | 1.3 | 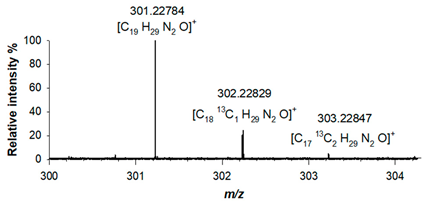 |
| 301.22784 | ||||
| [M+H+1]+ | 20.50 | 2.8 | ||
| 302.22829 | ||||
| [M+H+2]+ | 2.00 | 3.4 | ||
| 303.22847 |
| Compound Molecular Formula Exact mass |
m/za | Relative Intensity (%) | Mass accuracy in RMSb ppm |
ESI-(+)-FT ICR MS spectra |
|---|---|---|---|---|
|
(3) C₂₃H₃₆N₂O₂ 372.27768 |
[M+H]+ | 100 | 2.4 | 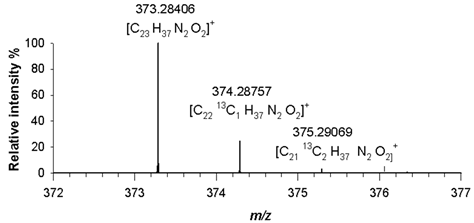 |
| 373.28406 | ||||
| [M+H+1]+ | 24.1 | 2.0 | ||
| 374.28757 | ||||
| [M+H+2]+ | 3.0 | 2.6 | ||
| 375.29069 | ||||
|
(4) C₂₁H₃₂N₂O₂ 344.24638 |
[M+H]+ | 100 | 1.9 | 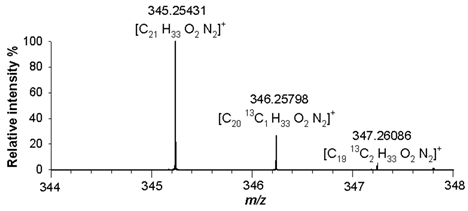 |
| 345.25431 | ||||
| [M+H+1]+ | 22.2 | 2.8 | ||
| 346.25798 | ||||
| [M+H+2]+ | 2.5 | 1.4 | ||
| 347.26086 | ||||
|
(5) C₂₃H₃₀N₂O₂ 366.23073 |
[M+H]+ | 100 | 1.4 | 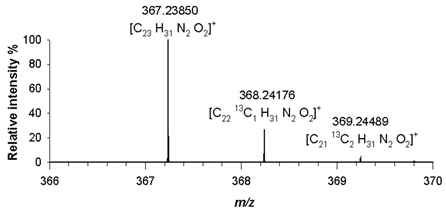 |
| 367.23850 | ||||
| [M+H+1]+ | 24 | 1.1 | ||
| 368.24176 | ||||
| [M+H+2]+ | 3 | 0.5 | ||
| 369.24489 | ||||
|
(6) C₂₀H₃₁N₃O₂ 345.24163 |
[M+H]+ | 100 | 1.8 | 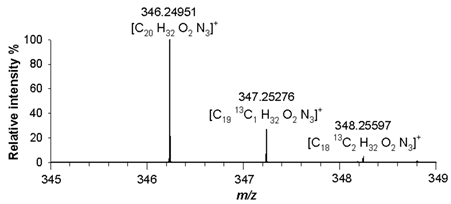 |
| 346.25091 | ||||
| [M+H+1]+ | 21 | 1.4 | ||
| 347.25276 | ||||
| [M+H+2]+ | 2.8 | 1.0 | ||
| 348.25597 |
| Label | Bacterial strains | Compound 3 | Compound 4 | Compound 5 | Compound 6 | ||||
| Inhibition Zone a | MIC (mg/ml) |
Inhibition Zone a | MIC (mg/ml) |
Inhibition Zone a | MIC (mg/ml) |
Inhibition Zone a | MIC (mg/ml) |
||
| 1 | Carnobacterium maltaromaticum | 20 ± 0.98 b | 6.25 ± 0.08 | 23 ± 0.76 | 6.25 ± 0.48 | 27 ± 0.76 | 3.125 ± 0.88 | 20 ± 0.96 | 12.5 ± 0.55 |
| 2 | Carnobacterium divergens | 21 ± 0.11 | 6.25 ± 0.66 | 19 ± 0.98 | 6.25 ± 0.45 | / | / | 18 ± 0.22 | 25 ± 0.72 |
| 3 | Pseudomonas fragi | 24 ± 0.91 | 6.25 ± 0.78 | 15 ± 0.09 | 12.5 ± 0.56 | / | / | 17 ± 0.59 | 25 ± 0.90 |
| 4 | Hafnia alvei | / | / | 14 ± 0.54 | 50 ± 0.70 | / | / | 13 ± 0.77 | 25 ± 0.87 |
| 5 | Pseudomonas proteamaculans | / | / | 16 ± 0.43 | 25 ± 0.99 | 11 ± 0.53 | 50 ± 0.64 | 14 ± 0.23 | 25 ± 0.46 |
| 6 | Brochothrix thermosphacta | / | / | 8 ± 0.75 | 50 ± 0.69 | 9 ± 0.78 | 50 ± 0.56 | 17 ± 0.59 | 25 ± 0.78 |
| 7 | Escherichia coli | 24 ± 0.28 | 12.5 ± 0.43 | 21 ± 0.38 | 12.5 ± 0.97 | 14 ± 0.90 | 25 ± 0.29 | 17 ± 0.80 | 12.5 ± 0.44 |
| 8 | Enterococcus hirae | 20 ± 0.09 | 3.125 ± 0.61 | 24 ± 0.56 | 6.25 ± 0.76 | / | / | 24 ± 0.82 | 12.5 ± 0.36 |
| 9 | Enterococcus faecium | 20 ± 0.08 | 3.125 ± 0.86 | 19 ± 0.63 | 3.125 ± 0.46 | / | / | 27 ± 0.66 | 6.25 ± 0.93 |
| 10 | Enterococcus faecalis | 19 ± 0.33 | 6.25 ± 0.98 | 15 ± 0.56 | 6.25 ± 0.57 | / | / | 21 ± 0.85 | 6.25 ± 0.76 |
| 11 | Enterococcus casseliflavus | 16 ± 0.77 | 6.25 ± 0.33 | 20 ± 0.86 | 6.25 ± 0.65 | 8 ± 0.33 | 12.5 ± 0.98 | 22 ± 0.47 | 6.25 ± 0.99 |
| 12 | Enteroccus durans | 17 ± 0.45 | 3.125 ± 0.53 | 21 ± 0.90 | 25 ± 0.43 | / | / | 16 ± 0.56 | 25 ± 0.34 |
| 13 | Enterococcus gallinarum | 16 ± 0.68 | 3.125 ± 0.42 | 17 ± 0.86 | 3.125 ± 0.56 | / | / | 15 ± 0.62 | 12.5 ± 0.62 |
| 14 | Listeria innocua | 26 ± 0.71 | 12.5 ± 0.48 | 13 ± 0.33 | 25 ± 0.57 | / | / | 17 ± 0.53 | 25 ± 0.87 |
| 15 | Staphylococcus aureus | 22 ± 0.90 | 6.25 ± 0.90 | 17 ± 0.63 | 12.5 ± 0.88 | / | / | 16 ± 0.56 | 12.5 ± 0.68 |
| 16 | Escherichia coli | 25 ± 0.88 | 6.25 ± 0.79 | 17 ± 0.86 | 12.5 ± 0.89 | / | / | 22 ± 0.58 | 12.5 ± 0.97 |
| 17 | Listeria monocytogenes | 20 ± 0.43 | 25 ± 0.20 | 23 ± 0.56 | 12.5 ± 0.74 | / | / | 20 ± 0.90 | 25 ± 0.47 |
| 18 | Salmonella serovar | 19 ± 0.93 | 6.25 ± 0.99 | 16 ± 0.56 | 25 ± 0.32 | / | / | 18 ± 0.57 | 50 ± 0.85 |
| 19 | Pseudomonas aeruginosa | / | / | 15 ± 0.59 | 6.25 ± 0.71 | / | / | 20 ± 0.83 | 12.5 ± 0.50 |
| 20 | Micrococcus flavus | 15 ± 0.22 | 25 ± 0.68 | 20 ± 0.28 | 12.5 ± 0.86 | / | / | 20 ± 0.77 | 6.25 ± 0.88 |
| 21 | Lysinibacillus fusiformis | / | / | 23 ± 0.94 | 12.5 ± 0.63 | / | / | 2.4 ± 0.89 | 25 ± 0.68 |
| 22 | Bacillus subtilis | 28 ± 0.40 | 3.125 ± 0.49 | 26 ± 0.75 | 6.25 ± 0.90 | 28 ± 0.69 | 3.125 ± 0.96 | 20 ± 0.62 | 12.5 ± 0.33 |
| 23 | Planococcus psychroteleratus | 24 ± 0.87 | 12.5 ± 0.92 | 20 ± 0.38 | 12.5 ± 0.56 | / | / | 21 ± 0. 44 | 12.5 ± 0.87 |
| 24 | Bacillus amyloliquefaciens | 23 ± 0.24 | 12.5 ± 0.75 | 25 ± 0.84 | 6.25 ± 0.55 | 28 ± 0.66 | 3.125 ± 0.22 | 27 ± 0.70 | 12.5 ± 0.58 |
| 25 | Bacillus anthracis | 29 ± 0.12 | 6.25 ± 0.47 | 22 ± 0.90 | 6.25 ± 0.61 | 21 ± 0.89 | 3.125 ± 0.76 | 24 ± 0.43 | 6.25 ± 0.46 |
| 26 | Bacillus amyloliquefaciens | 31 ± 0.44 | 6.25 ± 0.21 | 25 ± 0.75 | 12.5 ± 0.42 | 25 ± 0.45 | 3.125 ± 0.85 | 25 ± 0.75 | 12.5 ± 0.99 |
| 27 | Pseudomonas orientalis | 29 ± 0.90 | 6.25 ± 0.98 | 26 ± 0.53 | 6.25 ± 0.68 | 29 ± 0.80 | 3.125 ± 0.43 | 24 ± 0.94 | 6.25 ± 0.85 |
| 28 | Bacillus cereus | 23 ± 0.82 | 6.25 ± 0.77 | 23 ± 0.78 | 12.5 ± 0.99 | / | / | 23 ± 065 | 12.5 ± 0.43 |
| 29 | Bacillus cereus | 25 ± 0.90 | 6.25 ± 0.57 | 25 ± 0.39 | 12.5 ± 0.50 | / | / | 30 ± 0.96 | 12.5 ± 0.79 |
| Label | Bacterial strains | Compound 1 | Compound 2 | ||
| Inhibition Zone a | MIC (mg/ml) |
Inhibition Zone a | MIC (mg/ml) |
||
| 1 | Carnobacterium maltaromaticum | / | / | 25 ± 0.77 | 6.25 ± 0.60 |
| 2 | Carnobacterium divergens | / | / | 20 ± 0.34 | 6.25 ± 0.84 |
| 3 | Pseudomonas fragi | / | / | 22 ± 0.84 | 6.25 ± 0.53 |
| 4 | Hafnia alvei | / | / | 19 ± 0.66 | 12.5 ± 0.99 |
| 5 | Pseudomonas proteamaculans | / | / | 19 ± 0.53 | 12.5 ± 0.64 |
| 6 | Brochothrix thermosphacta | / | / | 19 ± 0.78 | 12.5 ± 0.56 |
| 7 | Escherichia coli | / | / | 22 ± 0.90 | 6.25 ± 0.29 |
| 8 | Enterococcus hirae | / | / | 20 ± 0.77 | 12.5 ± 0.85 |
| 9 | Enterococcus faecium | / | / | 20 ± 0.90 | 12.5 ± 0.72 |
| 10 | Enterococcus faecalis | / | / | 21 ± 0.56 | 12.5 ± 0.89 |
| 11 | Enterococcus casseliflavus | / | / | 24 ± 0.33 | 3.125 ± 0.98 |
| 12 | Enteroccus durans | / | / | 21 ± 0.90 | 12.5 ± 0.29 |
| 13 | Enterococcus gallinarum | / | / | 18 ± 0.55 | 12.5 ± 0.80 |
| 14 | Listeria innocua | / | / | 22 ± 0.76 | 12.5 ± 0.68 |
| 15 | Staphylococcus aureus | / | / | 18 ± 0.77 | 12.5 ± 0.34 |
| 16 | Escherichia coli | / | / | 21 ± 0.46 | 12.5 ± 0.88 |
| 17 | Listeria monocytogenes | / | / | 20 ± 0.95 | 12.5 ± 0.58 |
| 18 | Salmonella serovar | / | / | 22 ± 0.35 | 6.25 ± 0.72 |
| 19 | Pseudomonas aeruginosa | / | / | 19 ± 0.59 | 12.5 ± 0.82 |
| 20 | Micrococcus flavus | / | / | 25 ± 0.66 | 6.25 ± 0.77 |
| 21 | Lysinibacillus fusiformis | / | / | 26 ± 0.55 | 6.25 ± 0.32 |
| 22 | Bacillus subtilis | 9 ± 0.67 b | 25 ± 0.84 | 35 ± 0.69 | 3.125 ± 0.96 |
| 23 | Planococcus psychroteleratus | / | / | 32 ± 0.55 | 6.25 ± 0.43 |
| 24 | Bacillus amyloliquefaciens | / | / | 34 ± 0.66 | 6.25 ± 0.78 |
| 25 | Bacillus anthracis | 17 ± 0.89 | 25 ± 0.79 | 30 ± 0.89 | 6.25 ± 0.94 |
| 26 | Bacillus amyloliquefaciens | 10 ± 0.45 | 12.5 ± 0.82 | 34 ± 0.45 | 6.25 ± 0.65 |
| 27 | Pseudomonas orientalis | 9 ± 0.80 | 25 ± 0.43 | 25 ± 0.80 | 6.25 ± 0.88 |
| 28 | Bacillus cereus | 9 ± 0.66 | 25 ± 0.72 | 24 ± 0.51 | 6.25 ± 0.62 |
| 29 | Bacillus cereus | 9 ± 0.90 | 25 ± 0.27 | 23 ± 0.35 | 6.25 ± 0.70 |
| Label | Bacterial species | Growth conditions | |
|---|---|---|---|
| Temperature | Cultural medium | ||
| 1 | Carnobacterium maltaromaticum | 20°C | Tryptone Soya Yeast Extract Medium |
| 2 | Carnobacterium divergens | 20°C | Tryptone Soya Yeast Extract Medium |
| 3 | Pseudomonas fragi | 20°C | Tryptone Soya Yeast Extract Medium |
| 4 | Hafnia alvei | 30°C | Tryptone Soya Yeast Extract Medium |
| 5 | Pseudomonas proteamaculans | 30°C | Tryptone Soya Yeast Extract Medium |
| 6 | Brochothrix thermosphacta | 20°C | Tryptone Soya Yeast Extract Medium |
| 7 | Escherichia coli | 37°C | Tryptone Soya Yeast Extract Medium |
| 8 | Enterococcus hirae | 37°C | M17 Medium |
| 9 | Enterococcus faecium | 37°C | M17 Medium |
| 10 | Enterococcus faecalis | 37°C | M17 Medium |
| 11 | Enterococcus caseliflavus | 37°C | M17 Medium |
| 12 | Enterococcus durans | 37°C | M17 Medium |
| 13 | Enterococcus gallinarum | 37°C | M17 Medium |
| 14 | Listeria innocua | 30°C | Tryptone Soya Yeast Extract Medium |
| 15 | Staphylococcus aureus | 30°C | Tryptone Soya Yeast Extract Medium |
| 16 | Escherichia coli | 37°C | Tryptone Soya Yeast Extract Medium |
| 17 | Listeria monocytogenes | 30°C | Tryptone Soya Yeast Extract Medium |
| 18 | Salmonella serovar | 30°C | Tryptone Soya Yeast Extract Medium |
| 19 | Pseudomonas aeruginosa | 30°C | Tryptone Soya Yeast Extract Medium |
| 20 | Micrococcus flavus | 37°C | Plate count agar |
| 21 | Lysinibacillus fusiformis | 30°C | Plate count agar |
| 22 | Bacillus subtilis | 30°C | Plate count agar |
| 23 | Planococcus psychrotoleratus | 30°C | Plate count agar |
| 24 | Bacillus amyloliquefaciens | 30°C | Plate count agar |
| 25 | Bacillus anthracis | 30°C | Plate count agar |
| 26 | Bacillus amyloliquefaciens | 30°C | Plate count agar |
| 27 | Pseudomonas orientalis | 30°C | Plate count agar |
| 28 | Bacillus cereus | 30°C | Plate count agar |
| 29 | Bacillus cereus | 30°C | Plate count agar |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




