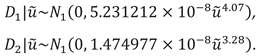Appendix B
B.1.1. Mixed-Effects models
Mixed effects model for unlinked data are described as follows [
11]:
, are the repeated measurements.
, is the subject’s number.
, is the method’s number.
, is a fixed effect and represents the difference in the fixed biases of the methods.
follow independent distributions. This is an interaction term. One interpretation for is the effect of method on subject . These interactions are subject-specific biases of the methods. They are a characteristic of the method-subject combination that remains stable during the measurement period.
follow independent distributions.
follow independent distributions.
, and are mutually independent.
To examine the measures of similarity and agreement we must retrieve the parameters of the assumed model (B1) which produces a bivariate distribution for
. By dropping the subscripts for the sake of simplicity, we have:
Thus, Then the model has a total of 6 unknown parameters .
Linked data are modeled as in model (B1) except for the addition of the term
which represents the random effect of the common time k on the measurements.
Thus, Then the model has a total of 7 unknown parameters .
The distribution of
for unlinked data is the following:
Based on the model parameters the heteroscedastic difference distribution is the following both for the unliked and linked data:
When the errors of models (B1) and (B2) are heteroscedastic,
and
are replaced with
and
. For a given
denoted as
(
for subject
) function
is the variance function and
is a vector of heteroscedasticity parameters, which for
corresponds to homoscedasticity. Variance covariate
is defined in advance (
) and accounts for heteroscedasticity. Choudhary and Nagaraja [
11] set
if method 1 is the reference and
otherwise. For the variance function
, two simple models are introduced. First, the power model, where
. Second, the exponential model,
. The parameters
can be estimated while fitting the model using ML and the “nlme” package [
46]. More details on the choice of the variance function
can be found in [
47]. AIC and BIC can be compared to distinguish between different candidate models for the variance functions.
Models with covariates.
Other factors might affect the agreement between the two methods. Covariates might affect the means of the methods (mean covariates), explaining part of the variability in the measurements. Covariates might also interact with the method or affect the error variance (variance covariates). In any case, the extend of the agreement is affected by the covariates. The mixed-effects models (B1) and (B2) can be extended as follows:
are the mean covariates.
independent and is defined as accounts for possible heteroscedasticity.
independent independent
Choudhary and Nagaraha [
11] describe the detailed methodology for defining mean and variance -specific covariates.
B.1.2. Measurement-Error models
Taffé [
29,
30] and Taffé et al. [
31] proposed various graphical tools to assess bias and precision of measurement methods in method comparison studies by following the measurement error model proposed by [
48] but an alternative estimation procedure based on an empirical Bayes approach. The model is the following:
is the replicate measurement by method on individual , , , denotes the number or repeated measurements per subject.
is a latent trait variable with density representing the true latent trait.
represents measurement errors by method .
The variances of these methods
are heteroscedastic and increase with the level of the true latent trait
which depends on the vectors of unknown parameters
[
29].
The mean value of the latent trait is and the variance is .
It is assumed that the latent variable represents the true unknown but constant value of the trait for individual and therefore .
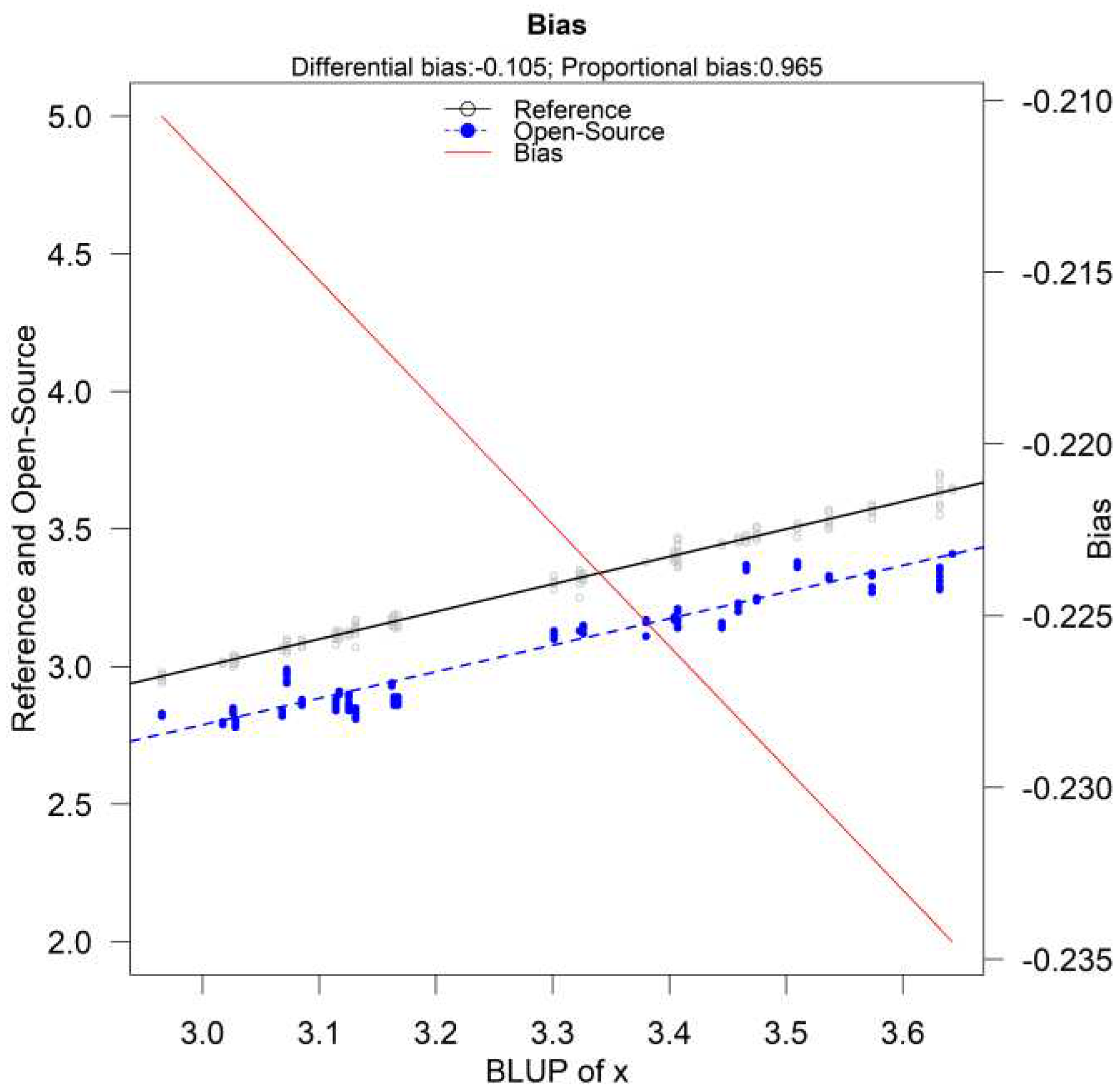
The parameters and are considered fixed and the error produced by them is called systematic. Their values depend on the measurement method. Parameter is defined as the fixed bias (or differential bias) of the method, while is defined as the proportional bias. Fixed bias is the added constant that the measurement method adds to the true value for every measurement. Proportional bias is based on the measured quantity and is the slope of (B5). The true value is multiplied by the proportional bias and is interpreted as the amount of change in the measurement method if the true value changes by 1 unit.
This modification of the classical measurement error model considers that heteroscedasticity depends on the latent trait and not on the observed average, compared to Choudhary’s and Nagaraja’s methodology [
11]). The model is assumed to be linear even though non-linear functions of
can be used and easily interpreted. To visually assess the plausibility of the straight-line model a graphical representation of
versus
provides a good start. Term
is the regression of the absolute values of the residuals
from the linear regression model
on
(the estimate of the latent trait) by ordinary least squares.
B.2. Indices to assess agreement and similarity.
B.2.1. Assessing Agreement.
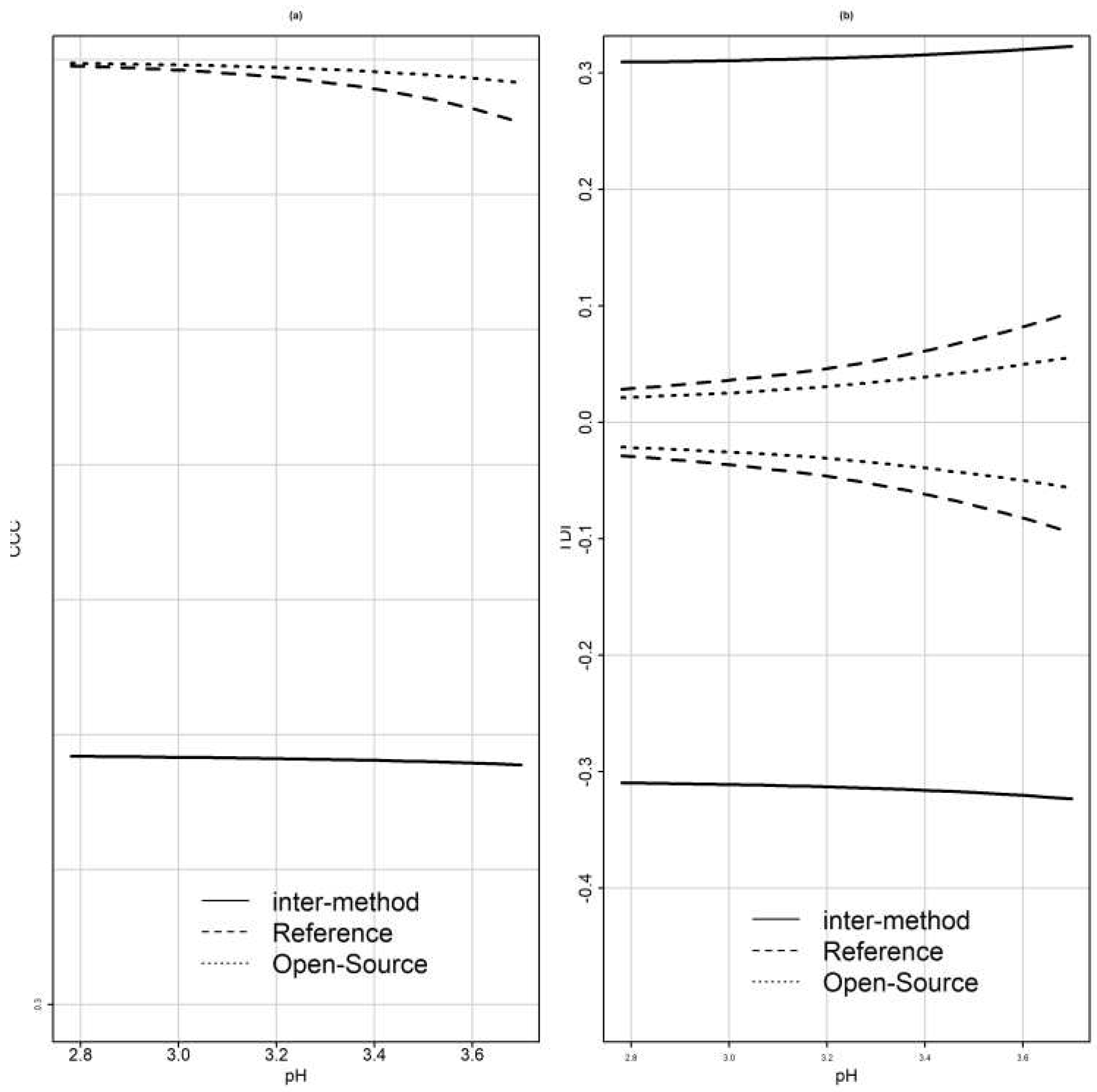
To evaluate the agreement and similarity between the reference and the open-source device, CCC and TDI were calculated using both (B1) and (B2) [
11] and methodologies provided by [
34,
36].
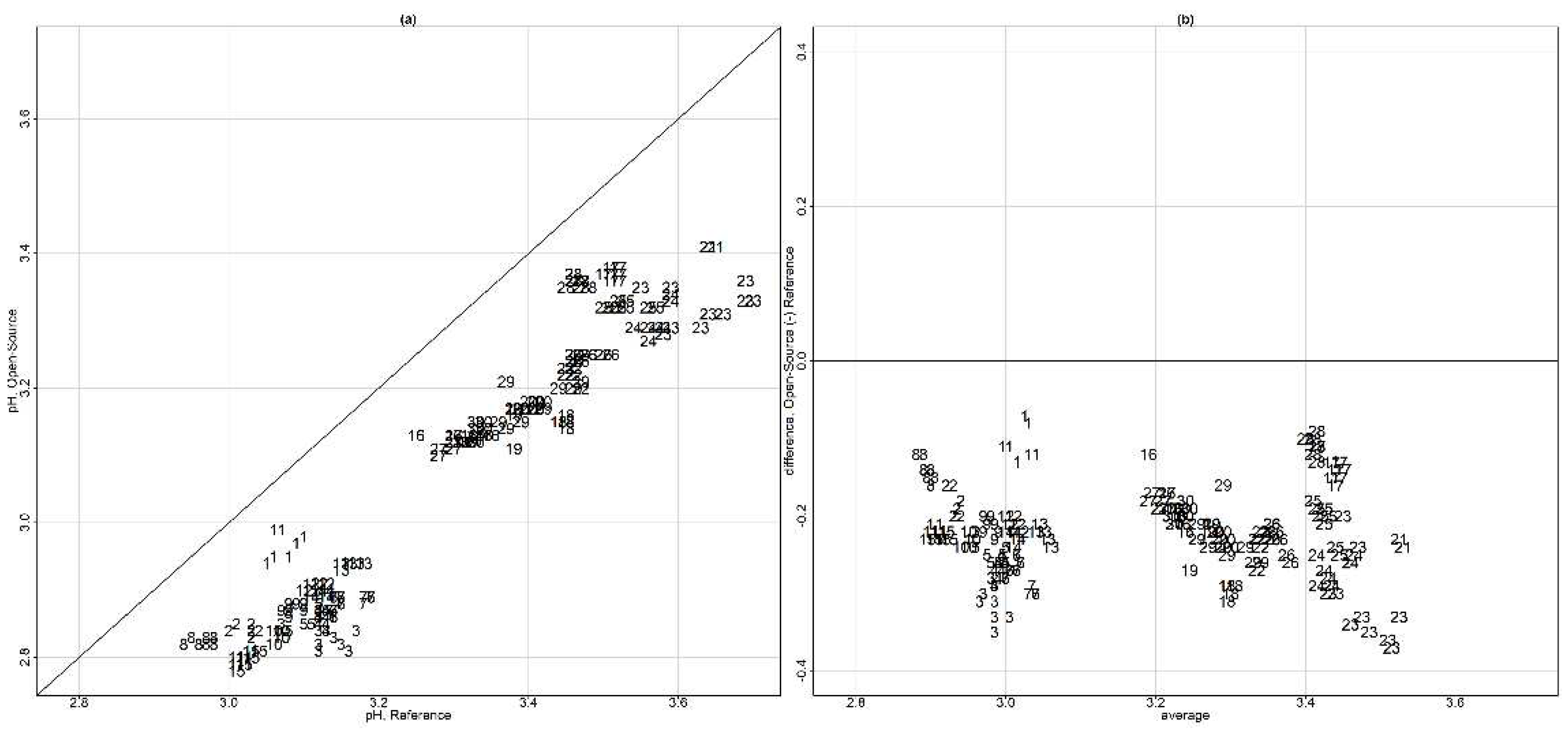
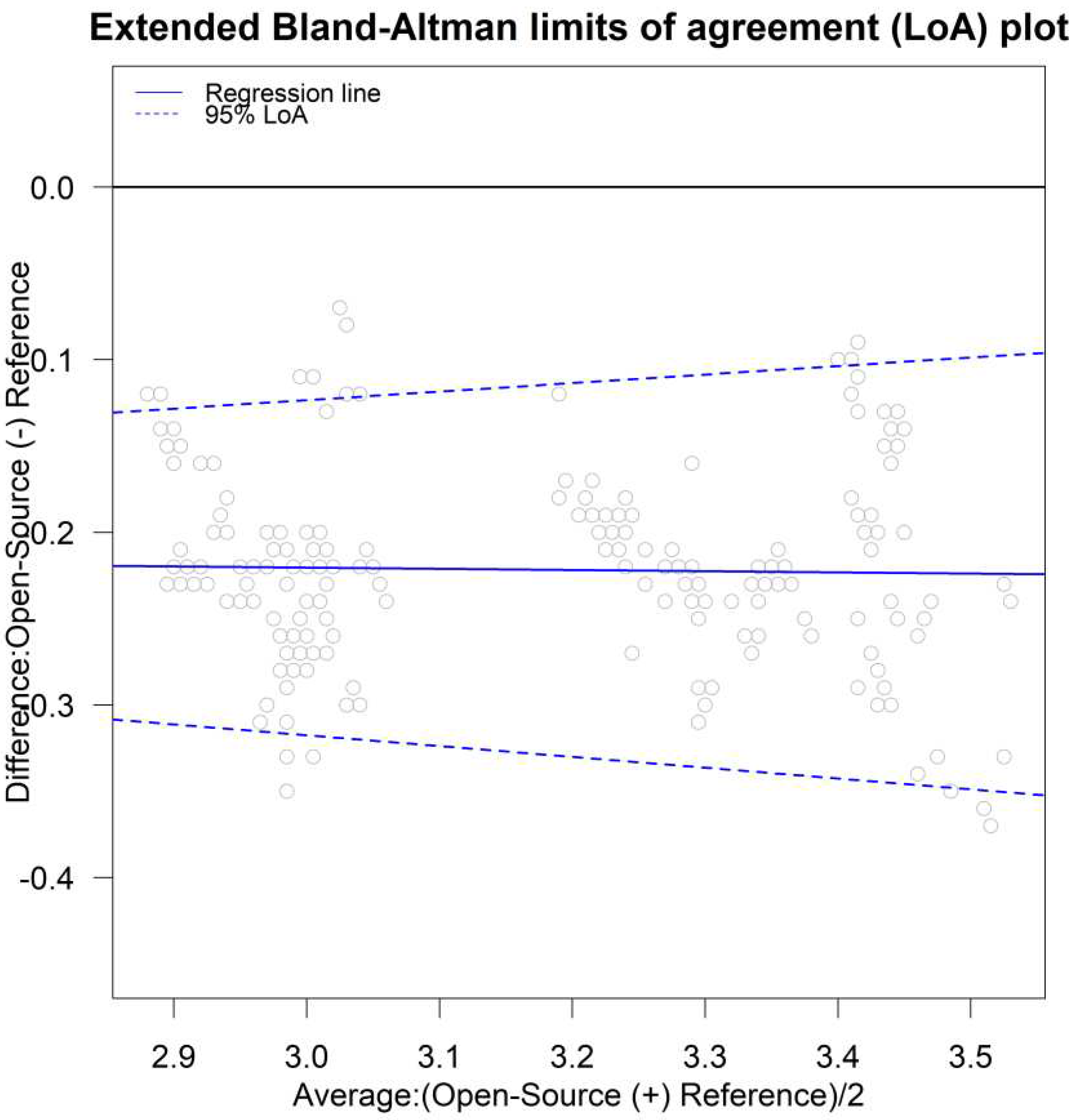
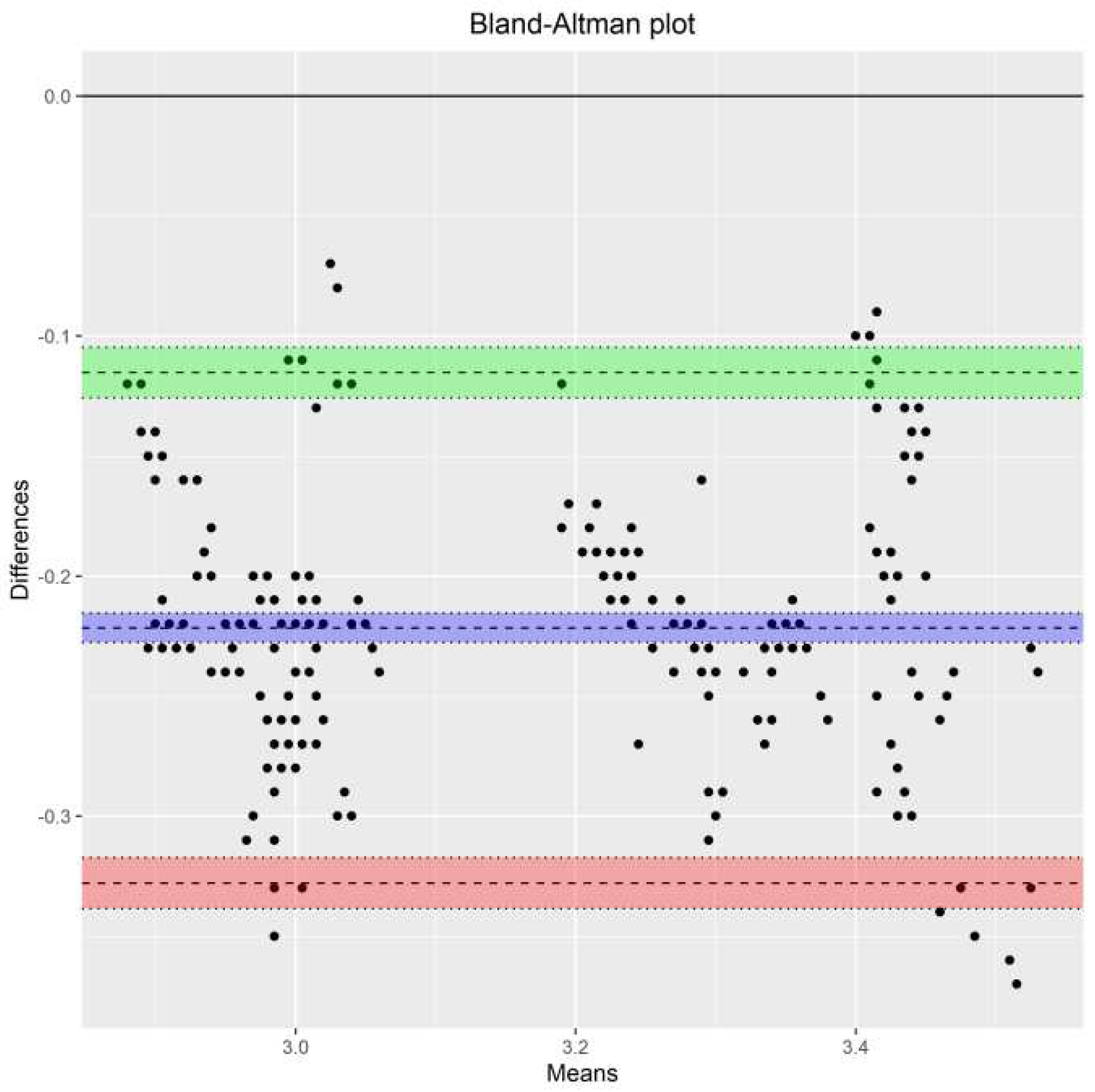
The limits of agreement [
26,
44,
49] for both unlinked and linked data were calculated using the parameters in models (B1) and (B2) via the following formulas provided by [
11]:
For heteroscedastic data and are replaced with and .
For unlinked and linked data, using models (B1, B3) and following the aproach described by [
11] TDI can be calculated using the following formula:

For unlinked data, under model (B1), [
11] proposed the following formula to calculate CCC:
While for linked data under model (B2):
Inference for TDI
Escaramis et al. [
34] proposed tolerance intervals for TDI. The value
is obtained by replacing
and
by their REML estimate counterparts derived from mixed-effects models (available in [
34]) in expression
. For inference, a one-sided tolerance interval is computed that covers the
-percent of the population from D with a stated confidence.
Let T be the studentized variable of
. T follows a non-central Student-t distribution with non-centrality parameter
,
, where
is the total possible paired-measurement differences between the two method/devices and the degrees of freedom
, are derived from the residual degrees of freedom. For the case of using individual-device interaction or discarding it, [
34] described different cases for obtaining
. An upper bound for TDI estimate can be constructed by using the following
one-sided tolerance interval, where
is the type
error rate:
It corresponds to the exact one-sided tolerance interval for at least
proportion of the population [
50,
51].
To perform a hypothesis test if the interest is to ensure that at least
-percent of the absolute differences between paired measurements are less than a predefined constant
, Lin’ s form of hypothesis can be followed [
32].
is rejected at level α if:
Choudhary and Nagaraja [
11] use the large-sample theory of ML estimators to compute standard errors, confidence bounds and tolerance intervals. When the sample is not large enough bootstrap confidence intervals are produced. The estimators of the model-based counterparts are obtained from models (B1,B2), depending on the nature of the data (unlinked or linked respectively). To compute simultaneous confidence intervals and bounds, the percentiles of appropriate functions of multivariate normally distributed pivots are needed. Following [
52] the R-package “multcomp” [
53] can be used. Choudhary and Nagaraja [
11] proposed a Bootstrap-t UCB and a modified nearly unbiased estimator (MNUT approximation) for computing the critical value, the p-value and the upper confidence bound (UCB)[
11].
Inference for CCC
Asymptotic distribution of the estimated CCCs can be used for inference if the data are modeled via a large sample size [
54]. Choudhary and Nagaraja [
11] use an asymptotic distribution of the estimated CCCs to produce an upper confidence bound when the sample is large and bootstrap methods when the sample is small. Since the concordance correlation coefficient is related to the intraclass correlation coefficient (ICC), inference methods for ICC can be used for CCC [
54].
B.2.2. Assessing Similarity
Following Choudhary and Nagaraja [
11], to evaluate similarity the marginal distributions of
and
are examined via estimates and two-sided confidence intervals. Their distributions are given by equations (B1, B2), for unlinked and linked data. The fixed bias and the precision ratio are the two measures of similarity that will be evaluated using mixed-effects model. Last, fixed bias, proportional bias and precisions are evaluated under measurement-error models.
Fixed bias will be estimated via the model’s counterparts. According to models (B1,B2) the fixed bias is estimated using: for unlinked and linked data.
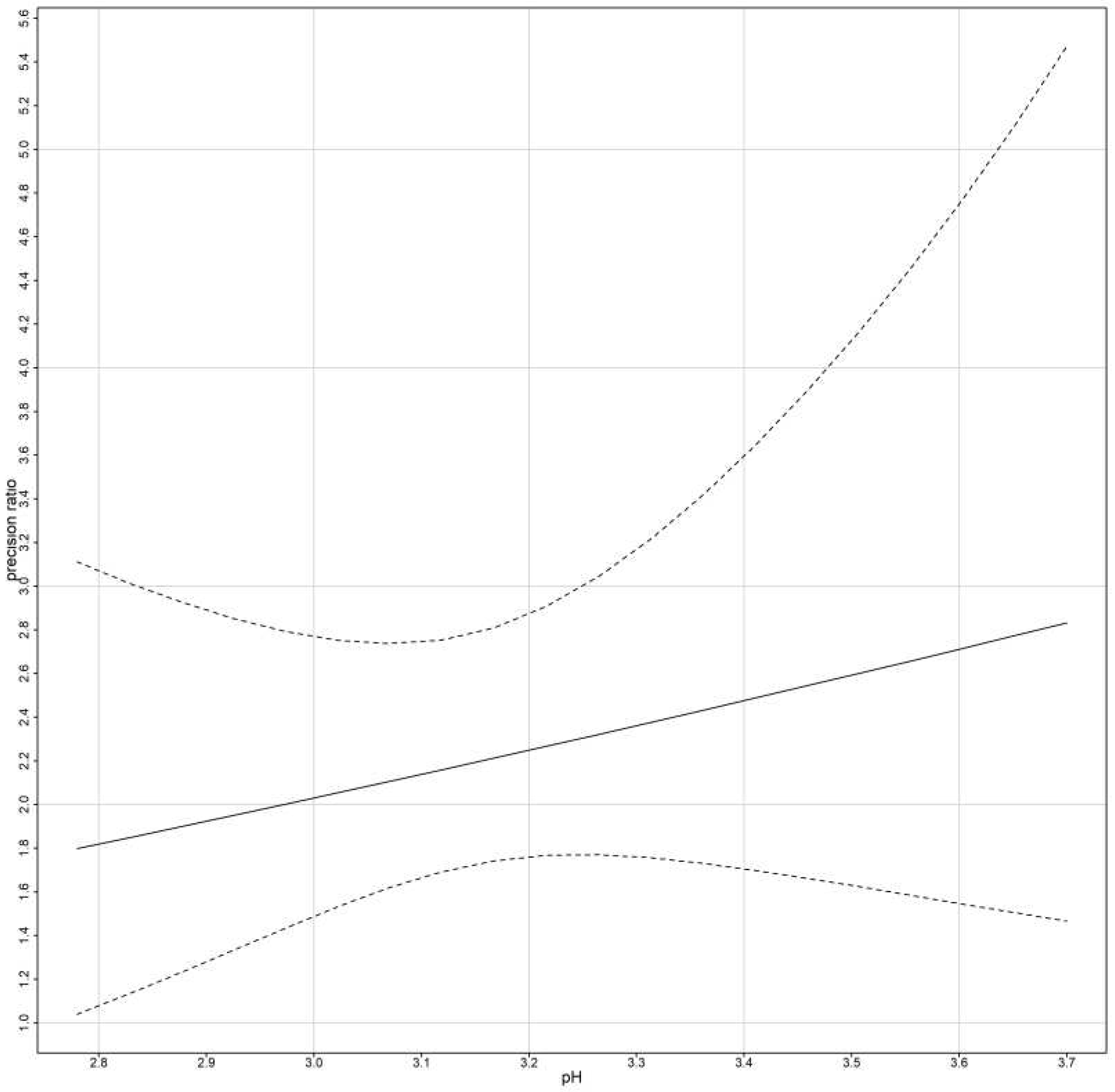
The precision ratio is evaluated in two different cases.
First, for models that ignore subject x method interactions:
Second, for models that include subject x method interactions:
The precision ratios are assumed to be estimated when the errors are homoscedastic. For heteroscedastic data we replace
and
with
and
thus fixed bias remains the same, but the precision ratio is given by:
For inference, the method described by [
48] is used for heteroscedastic data. Specifically, if
is a vector of model’s counterparts then the measure of similarity is a function of
. Denoting the measure of similarity as
, and
a value in the measurement range then
is any measure of similarity in a specific value (the measure is assumed to be scalar). Substituting θ with its corresponding ML estimate,
, in its expression gives its ML estimator
. From delta method [
55], when the sample size is large
, where
can be computed numerically. Thus, approximate
two-sided pointwise confidence interval for
on a grid of values of the measurement range can be computed as
B.3. Assessing Repeatability
Following [
11] and mixed-effect models, for unlinked data, repeated measurements are replications of the same underlying measurement. Instead of using the bivariate distributions
for measurements of the two methods on a randomly selected subject from a population,
is defined as a replication of
, where
denote the two methods/devices. By definition
and
have the same distribution. CCC and TDI are modified and are calculated. By dropping the subscripts for model (B1), for unlinked data:
is induced, similar to (B1) by dropping the subscripts.
where
and
are independent copies of
and
as defined in (B1).
Defining
as the difference in two replications of method j. From (B10) and (B11):
By dropping the subscripts for model (B2) for linked data:
Where , and are independent copies of , and as defined in (B2).
Defining
as the difference in two replications of method j. From (B16) and (B17):
B.4. Recalibration Methods
For measurement-error models, a recalibration procedure is performed by computing:
where
is the estimate of the proportional bias and
is the estimate of the fixed bias and
is the recalibrated value. The method performs well, according to simulations, with a sample size of 100 subjects and 10 to 15 repeated measurements per individual from the reference method and only 1 from the new. It is possible that after the recalibration procedure the novel method turns out to be more precise than the reference. The recalibration procedure can be implemented using
compare_plot() function from “
methodCompare” package [
27].