Submitted:
06 December 2023
Posted:
14 December 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction:
2. Materials and Methods
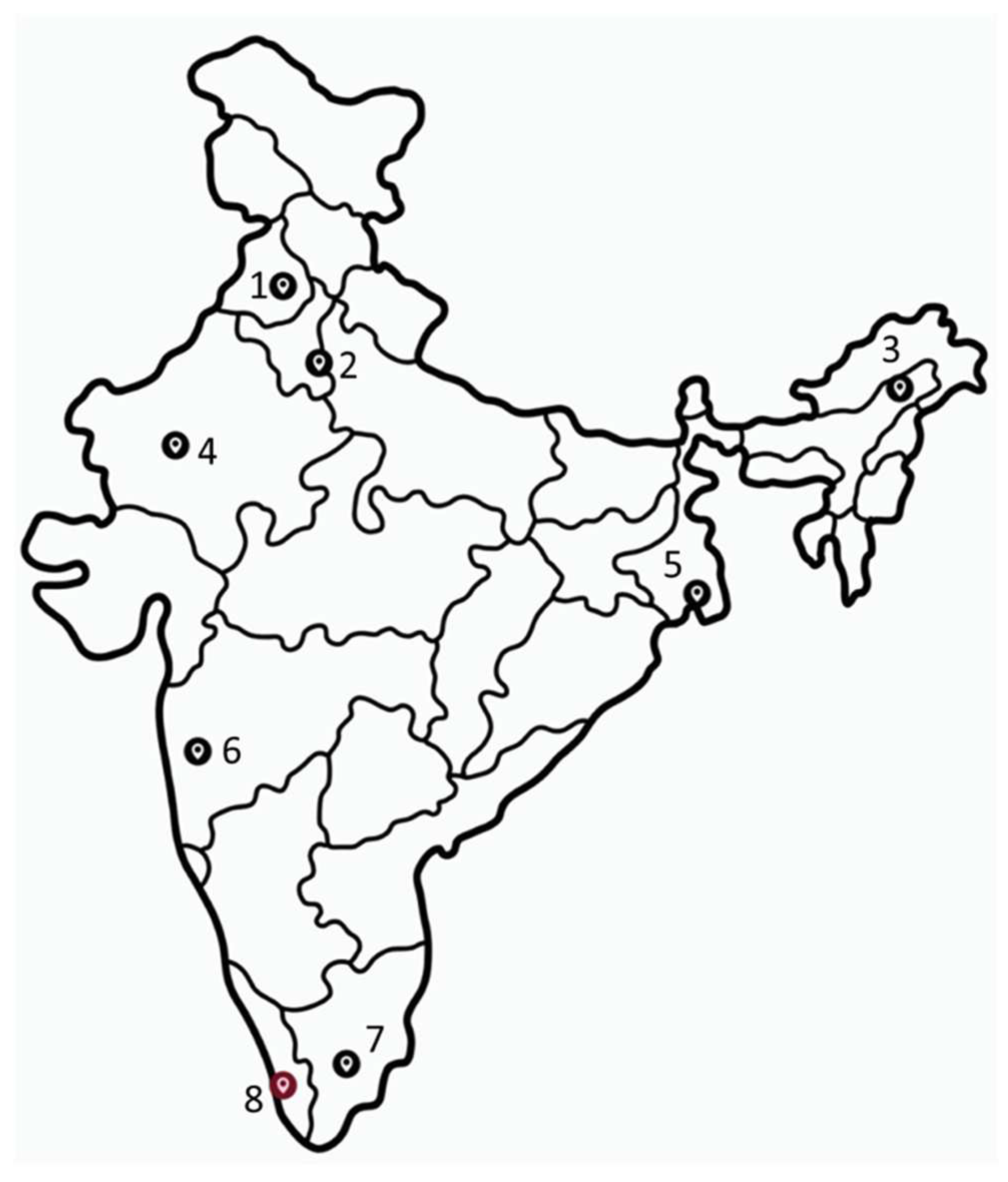
2.1. Collection and storage of wild mosquitoes, larvae and pupae
2.2. Viral RNA extraction, double-stranded complementary DNA (cDNA) preparation, and Sequence-Independent Single Primer Amplification (SISPA)
2.3. Library preparation and sequencing on Oxford Nanopore MinION Mk1C sequencer
2.4. Analysis of raw metagenomics data
2.5. Sanger sequencing to obtain complete Cell fusing agent virus (CFAV) genome sequence
2.5. Phylogenetic analysis of metagenomics derived whole genome sequences of viruses
3. Results:
3.1. Length and count statistics of sequencing data generated on ONT platform
| Sampling location | Average quality score | Average sequence length | Reads analyzed | Reads classified | Reads unclassified | Cumulative reads (Eukaryota, Viruses, Bacteria and Archaea) | Virus reads |
|---|---|---|---|---|---|---|---|
| Dibrugarh | 12.29 | 378 | 2,566,367 | 1,148,796 | 1,409,637 | 1,097,121 | 38,089 |
| Jodhpur | 12.40 | 357 | 1,416,476 | 605,659 | 805,246 | 604,734 | 35,220 |
| Kolkata | 12.29 | 318 | 1,258,839 | 246,022 | 1,008,574 | 214,756 | 52,798 |
| Madurai | 11.91 | 286 | 7,786,762 | 3,749,144 | 3,997,735 | 2,945,046 | 808,370 |
| New Delhi | 12.41 | 280 | 1,728,868 | 761,242 | 960,985 | 595,398 | 140,179 |
| Patiala | 12.48 | 300 | 5,966,644 | 3,494,089 | 2,472,555 | 3,414,206 | 188,311 |
| Pune | 12.67 | 401 | 2,415,355 | 774,139 | 344,458 | 612,756 | 276,367 |
| Sampling location | Average quality score | Average sequence length | Reads Analysed | Reads classified | Reads unclassified | Cumulative reads (Eukaryota, Viruses, Bacteria and Archaea) | Total virus reads |
|---|---|---|---|---|---|---|---|
| Pune | 11.27 | 476 | 359,883 | 153,303 | 204,490 | 153,303 | 122,369 |
| Alappuzha | 10.75 | 296 | 514,027 | 185,691 | 306,661 | 185,691 | 61,982 |
3.2. Virus taxa identified through metaviromics
| Location | Patiala (Punjab) | Pune (Maharashtra) | Dibrugarh (Assam) | Madurai (Tamil Nadu) | New Delhi | Jodhpur (Rajasthan) | Kolkata (West Bengal) |
|---|---|---|---|---|---|---|---|
|
Species identified 
|
Phasi Charoen-like phasivirus | Phasi Charoen-like phasivirus | Phasi Charoen-like phasivirus | Phasi Charoen-like phasivirus | Phasi Charoen-like phasivirus | Phasi Charoen-like phasivirus | Phasi Charoen-like phasivirus |
| Choristoneura fumiferana granulovirus | Choristoneura fumiferana granulovirus | Cell fusing agent virus | Choristoneura fumiferana granulovirus | Choristoneura fumiferana granulovirus | Wenzhou sobemo-like virus 4 | Cell fusing agent virus | |
| Cell fusing agent virus | Shamonda orthobunyavirus | Wenzhou sobemo-like virus 4 | Shamonda orthobunyavirus | Cell fusing agent virus | Choristoneura fumiferana granulovirus | Choristoneura fumiferana granulovirus | |
| Avianendogenous retrovirus EAV-HP | Lactobacillus virus LP65 | Choristoneura fumiferana granulovirus | Hubei toti-like virus 10 | Shamonda orthobunyavirus | Tobacco mild green mosaic virus | Ilheus virus | |
| Ostreococcus lucimarinus virus 1 | Cyprinid herpesvirus 2 | Hubei mosquito virus 2 | Badu phasivirus | Lactobacillus virus LP65 | Aroa virus | Tobacco mosaic virus | |
| Lactobacillus virus LP65 | Ostreid herpesvirus 1 | Porcine astrovirus 4 | Serratia phage Muldoon | Wenzhou hepe-like virus 1 | Ilheus virus | Aroa virus | |
| Chrysochromulina ericina virus | Bacillus virus G | Astrovirus wild boar/WBAstV-1/2011/HUN | Synechococcus phage S-CAM4 | Drosophila immigrans Nora virus | Culex Flavi-like virus | Shamonda orthobunyavirus | |
| Cotesia congregata bracovirus | Lactobacillus virus LLKu | Ilheus virus | Salinivibrio phage CW02 | Sphingomonas phage PAU | Synechococcus phage S-CAM22 | Shuangao insect virus 7 | |
| Lactobacillus virus LLKu | Chilli ringspot virus | Astrovirus MLB2 | Mycobacterium phage UnionJack | Esparto virus | Chrysochromulina ericina virus | ||
| Badu phasivirus | Aotine betaherpesvirus 1 | Shamonda orthobunyavirus | Jacquemontia yellow vein virus | Trichoplusia ni ascovirus 2c | |||
| Wenzhou sobemo-like virus 4 | Tenacibaculum phage PTm1 | Porcine astrovirus 2 | Esparto virus | Golden Marseillevirus | |||
| Euproctis pseudoconspersa nucleopolyhedrovirus | Lactococcus virus KSY1 | Agrotis ipsilon multiple nucleopolyhedrovirus | Cotesia congregata bracovirus | ||||
| Bacillus virus G | Vibrio phage douglas 12A4 | Cyprinid herpesvirus 2 | |||||
| Rhinolophus associated gemykibivirus 2 | Cotesia congregata bracovirus | ||||||
| Pacmanvirus A23 | Acanthamoeba polyphaga mimivirus | ||||||
| Pandoravirus quercus | Amsacta moorei entomopoxvirus | ||||||
| Shamonda orthobunyavirus | Pandoravirus neocaledonia | ||||||
| Emiliania huxleyi virus 86 |
| Location | Alappuzha (Kerala) | Pune (Maharashtra) |
|---|---|---|
| Species identified 
|
Wenzhou sobemo-like virus 4 | Wenzhou sobemo-like virus 4 |
| Choristoneura fumiferana granulovirus | Hubei mosquito virus 2 | |
| Hubei mosquito virus 2 | Choristoneura fumiferana granulovirus | |
| Tobacco mosaic virus | Wenzhou shrimp virus 9 | |
| Shamonda orthobunyavirus | Shamonda orthobunyavirus | |
| Cyprinid herpesvirus 2 | Mythimna unipuncta nucleopolyhedrovirus | |
| Elephant endotheliotropic herpesvirus 4 | ||
| Wilkie partiti-like virus 2 | ||
| Tobacco mild green mosaic virus | ||
| Tomato brown rugose fruit virus | ||
| Tomato mottle mosaic virus | ||
| Hubei picorna-like virus 34 | ||
| Hubei sobemo-like virus 9 | ||
| African swine fever virus | ||
| Pandoravirus quercus |
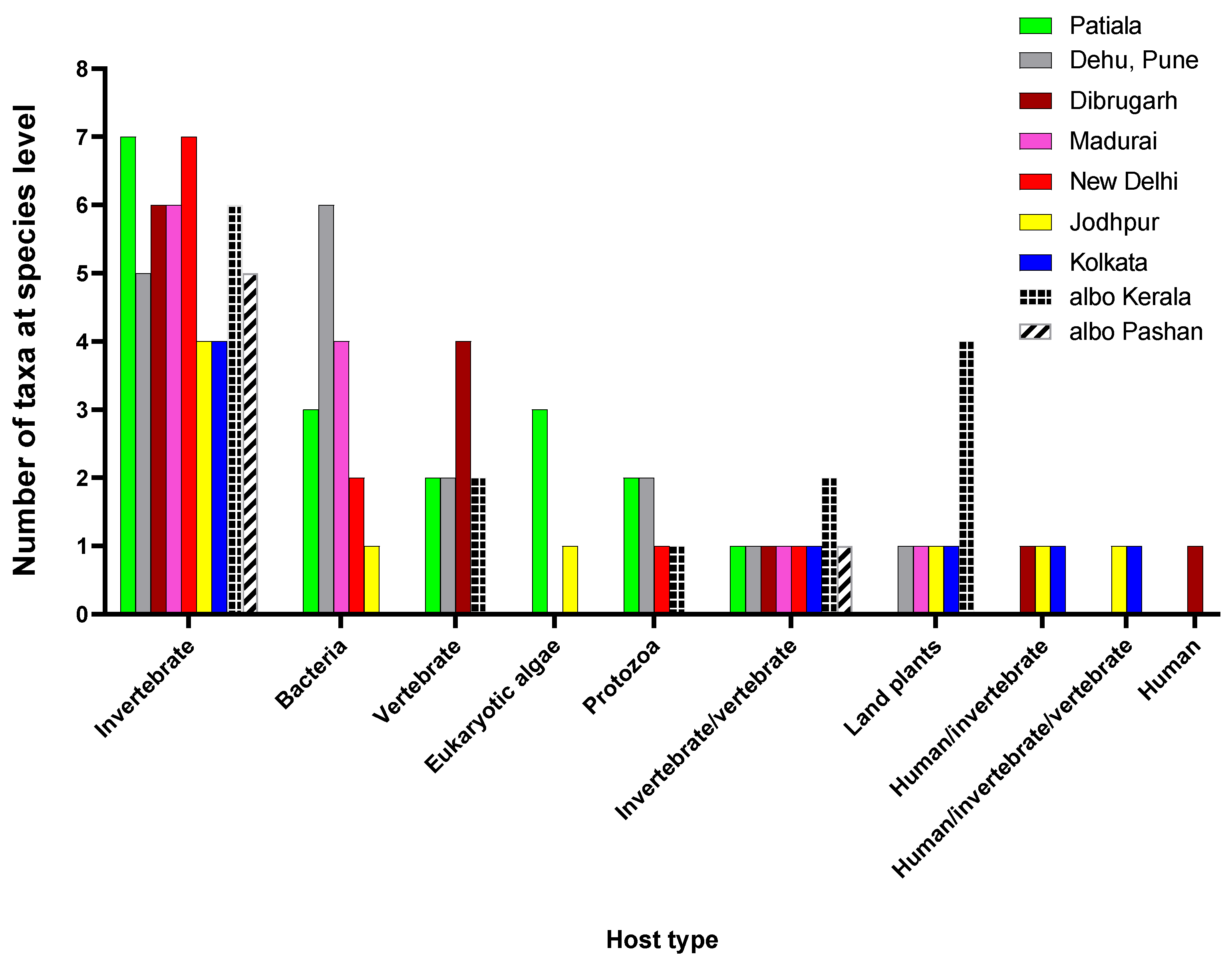
3.3. Viruses identified have a wide range of hosts
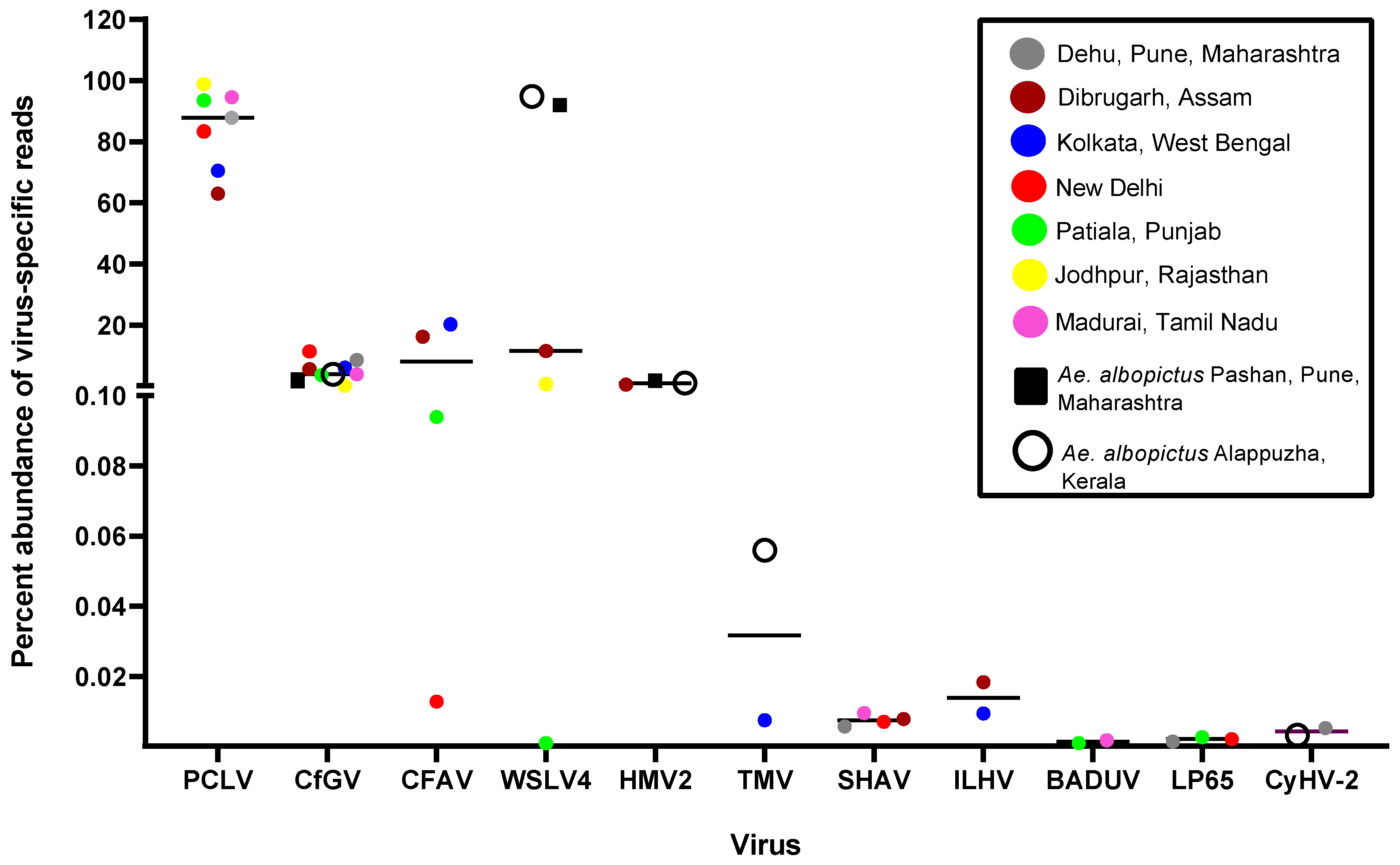
3.4. ISVs dominate the viromes of Ae. aegypti and Ae. albopictus
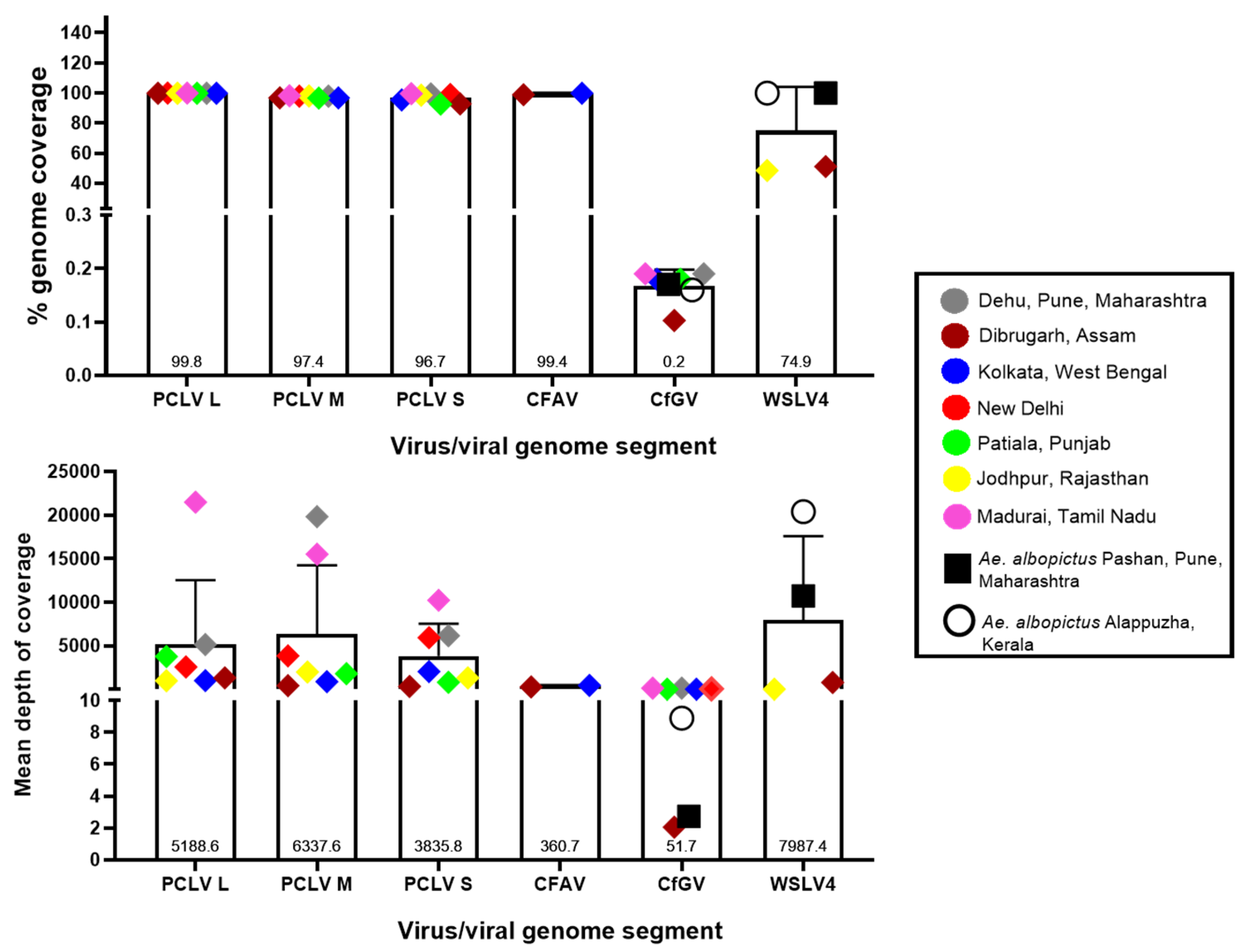
3.5. Read depths and genome coverage
3.6. Aedes albopictus viromes were dominated by unclassified viruses
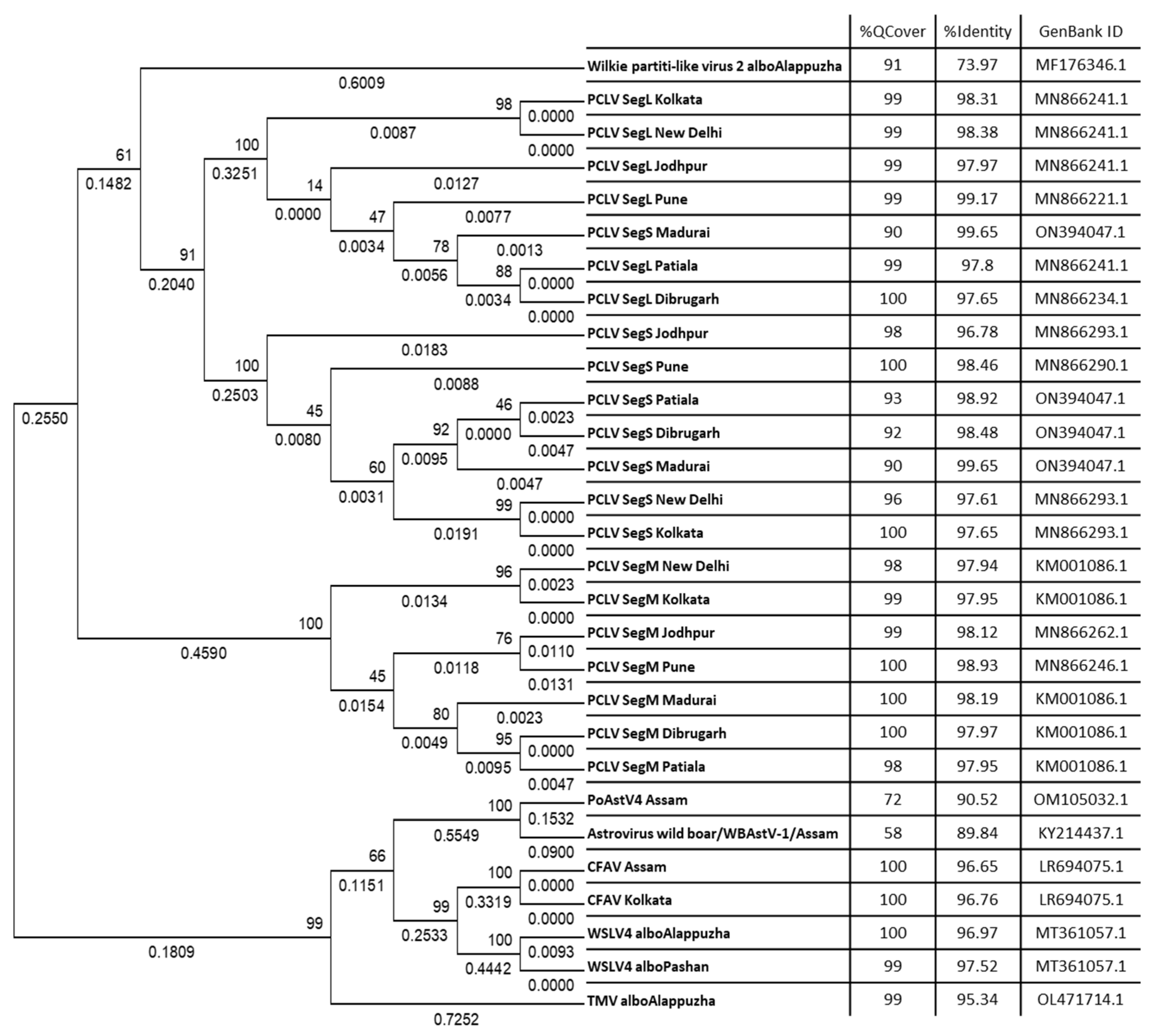
3.7. Phylogenetic analysis of virus sequences derived from Aedes viromes
3.8. PCLV sequences form two distinct clades separated by location and time
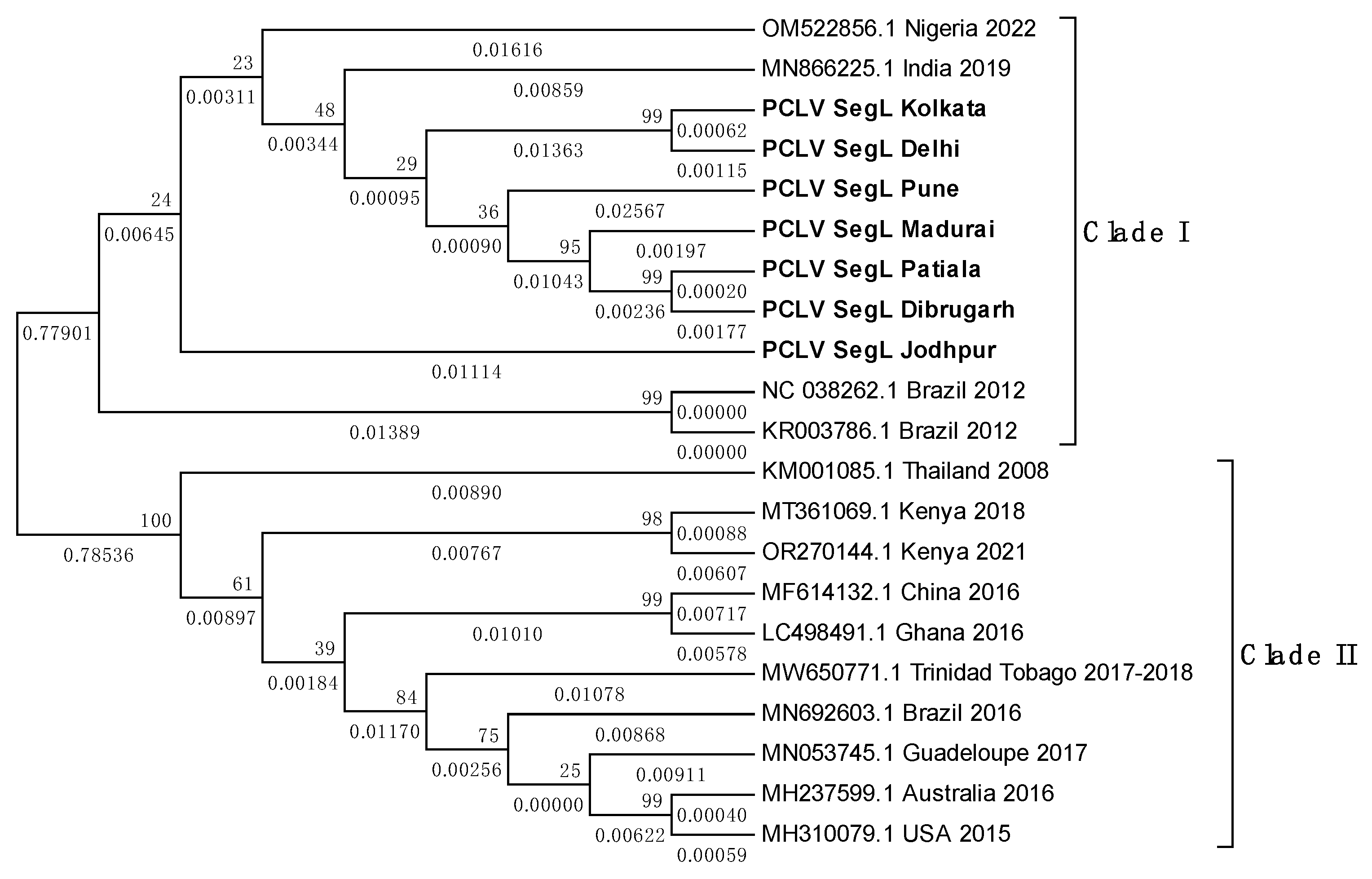
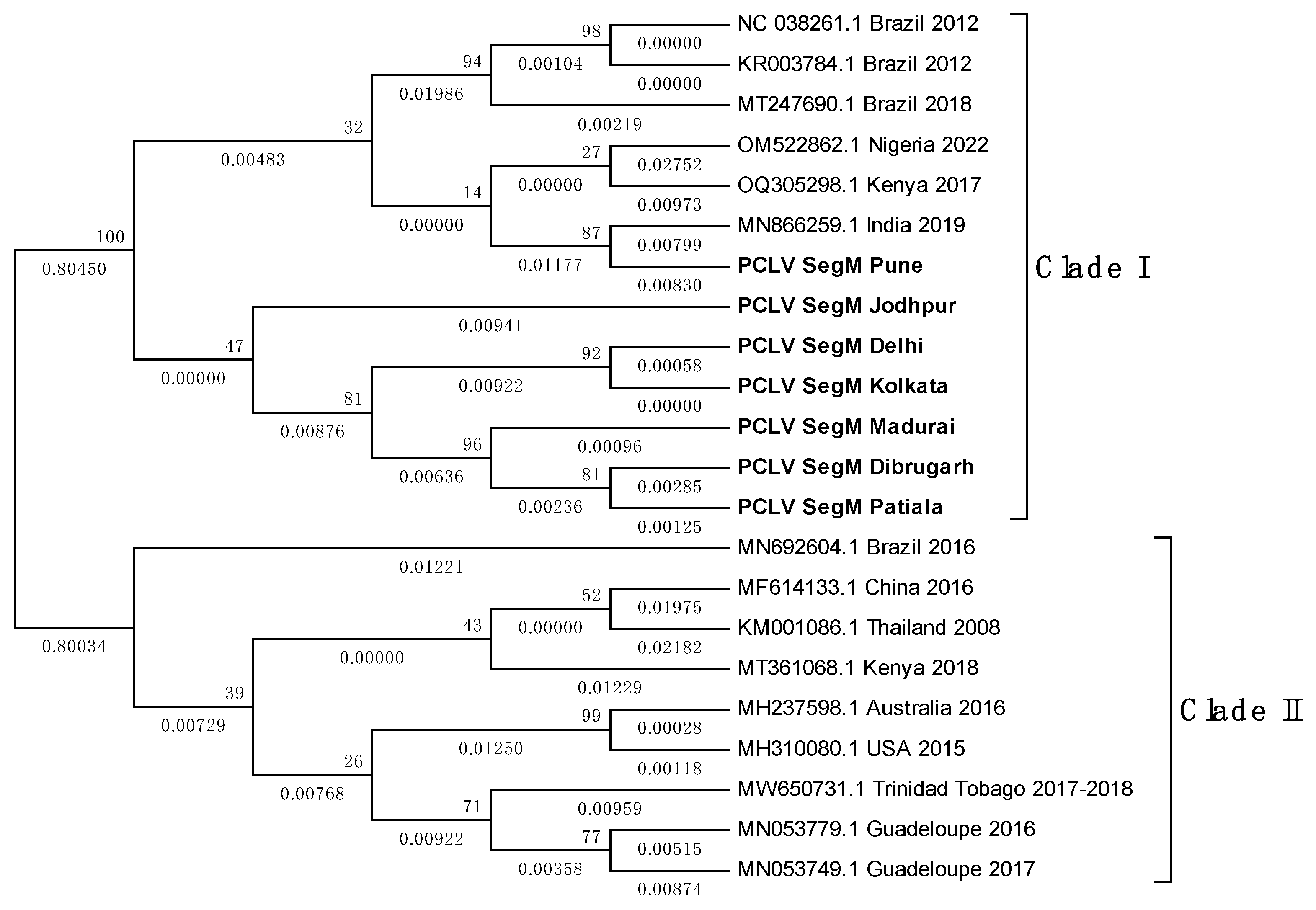
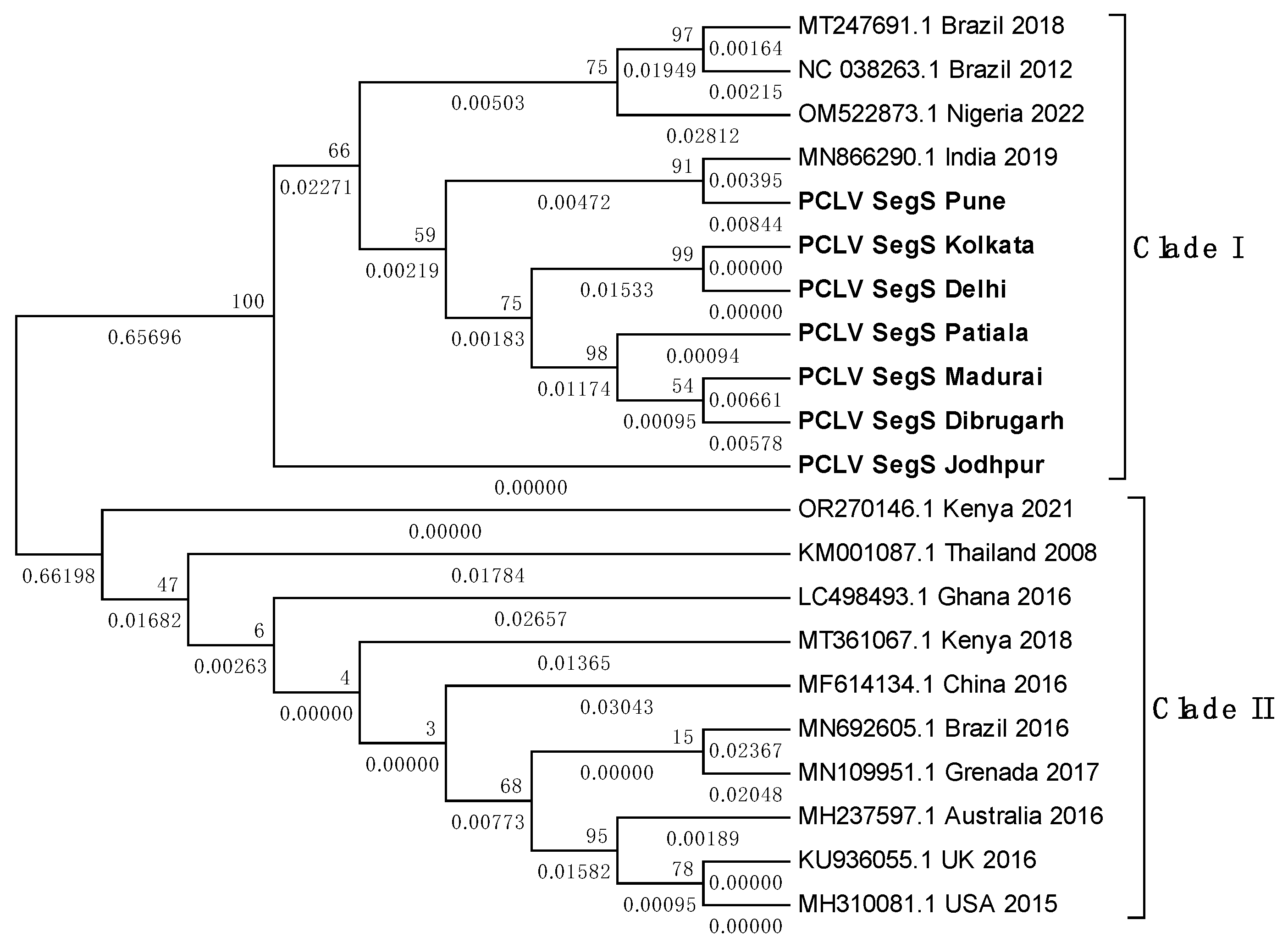
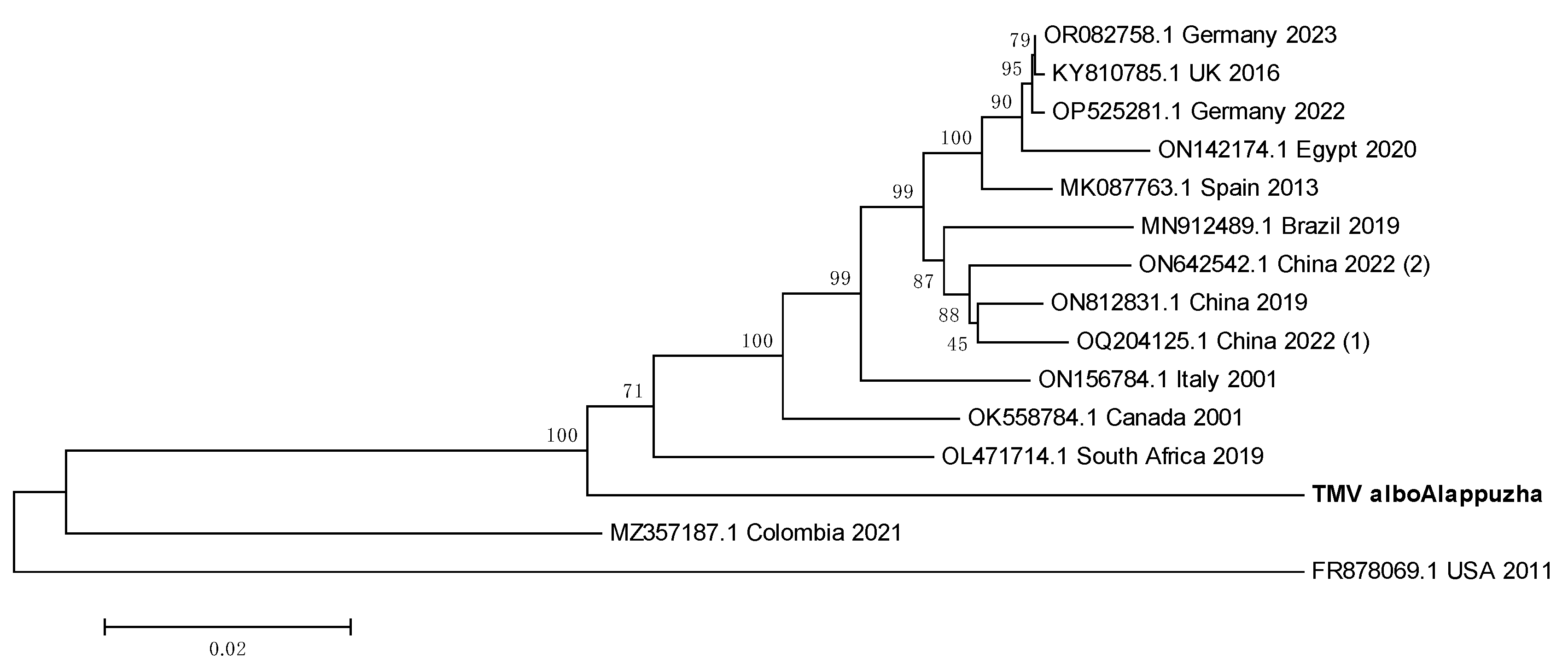
3.9. TMV sequence from Ae. albopictus is closely related to South African TMV sequence
3.10. WBAstV from Belgium is the likely introduction source of the virus into Assam

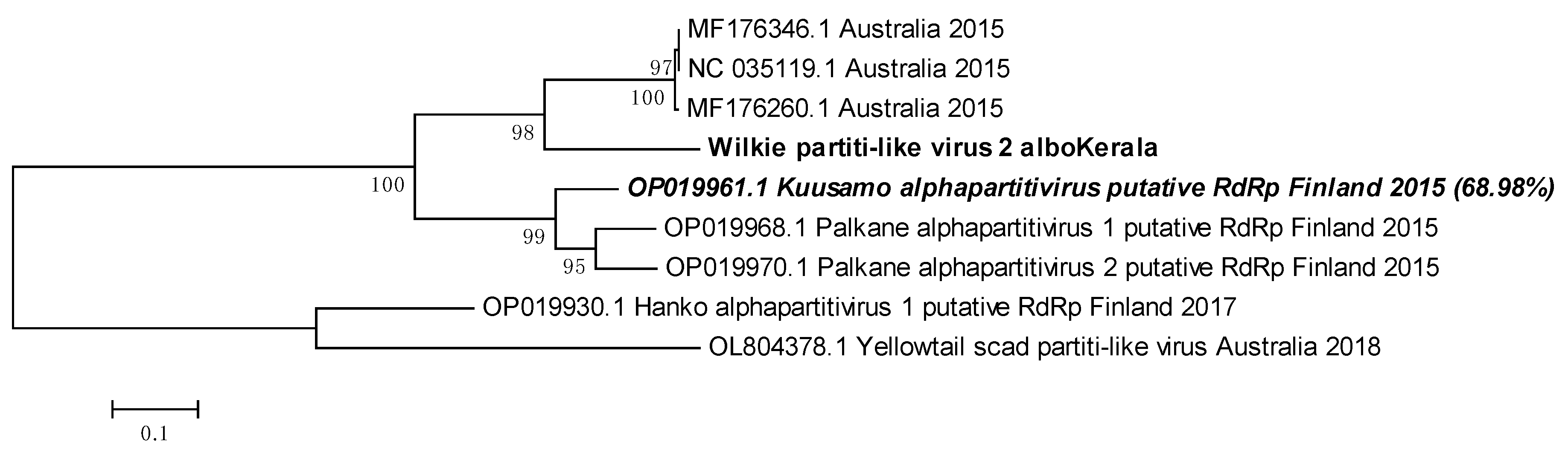
3.11. Wilkie partiti-like virus- 2 is related to the Kuusamo alphapartitivirus
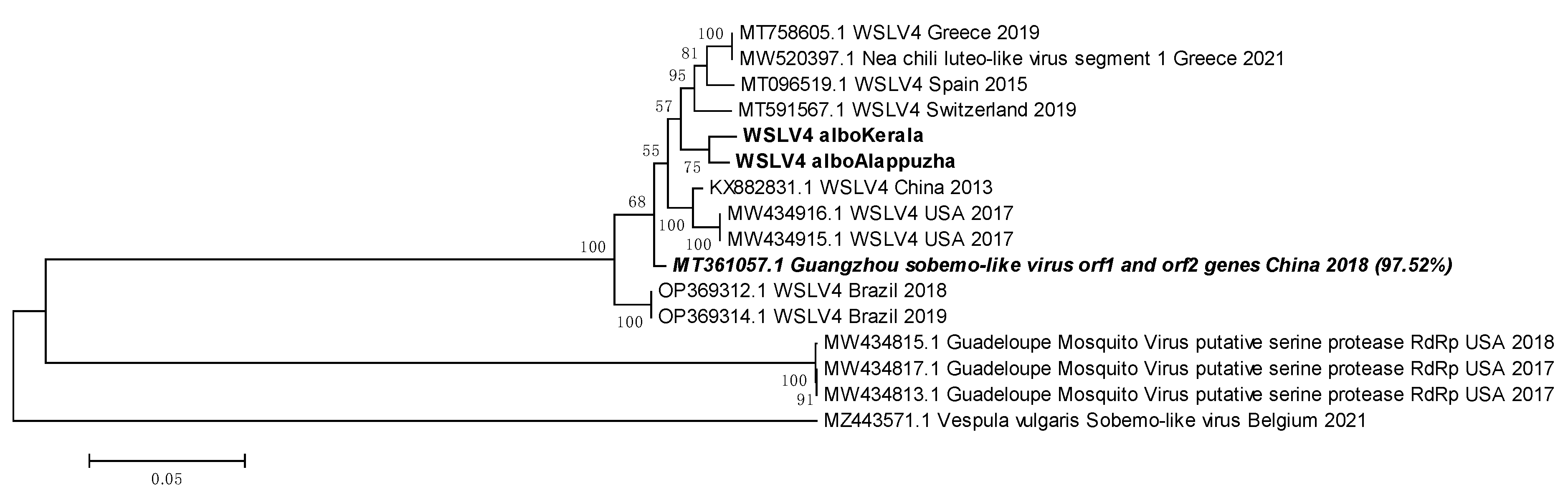
3.12. WSLV4 belongs to either the Solemoviridae family or the Tolivirales order
3.13. Indian CFAV sequences clustered together to form a single clade
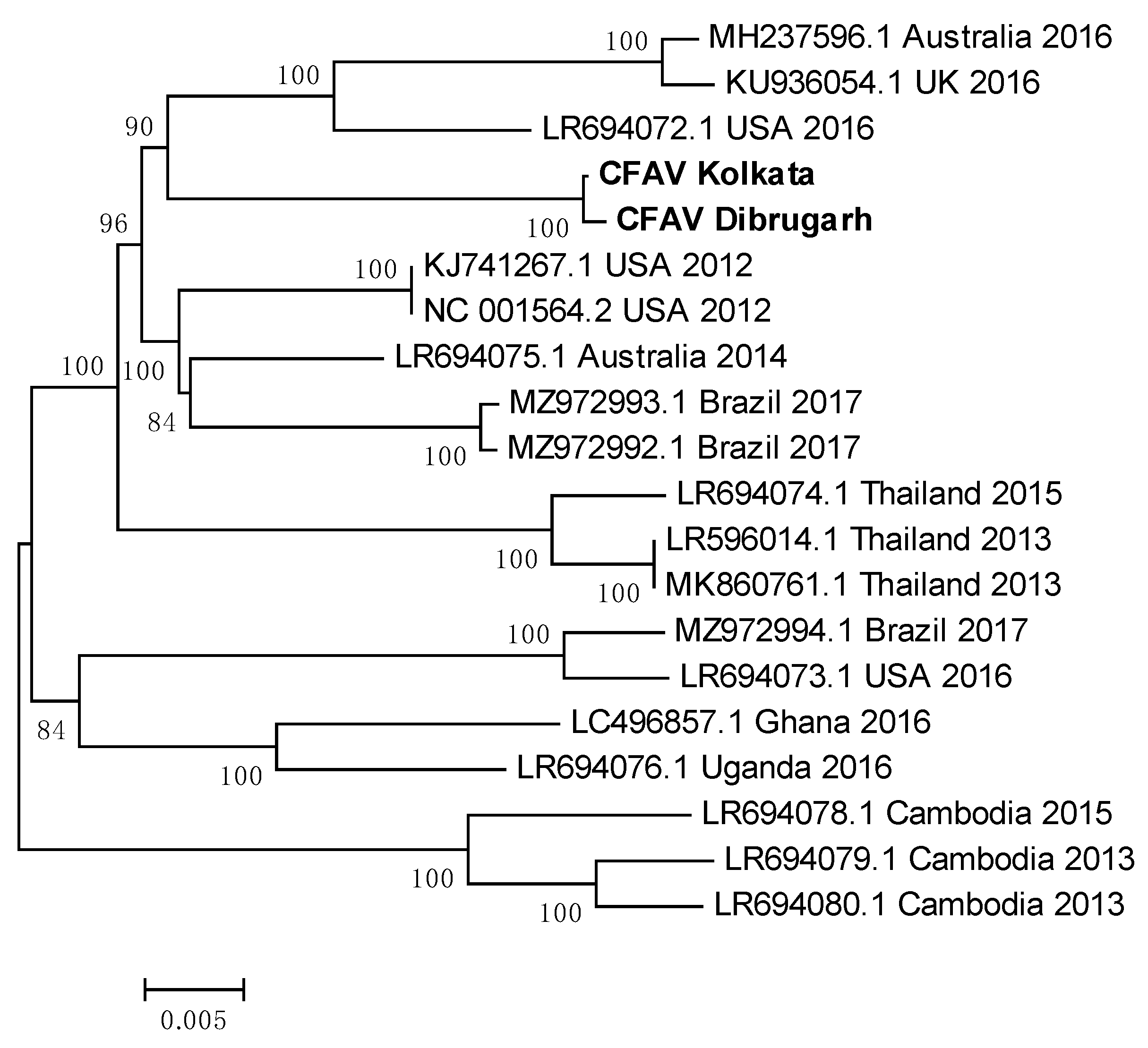
| Sequence/Reference | Country | Year of sample collection/sequence submission | CFAV Kolkata | CFAV Dibrugarh |
|---|---|---|---|---|
| CFAV Kolkata | India | 2022 | 100 | 99.88 |
| CFAV Dibrugarh | 99.88 | 100 | ||
| LR694075.1 | Australia | 2014 | 96.75 | 96.54 |
| KJ741267.1 | USA | 2012 | 96.7 | 96.51 |
| NC 001564.2 | 96.7 | 96.51 | ||
| MZ972993.1 | Brazil | 2017 | 96.22 | 95.98 |
| MZ972992.1 | 96.22 | 95.98 | ||
| MH237596.1 | Australia | 2016 | 95.56 | 95.34 |
| KU936054.1 | UK | 95.51 | 95.29 | |
| LR694072.1 | USA | 96.31 | 96.11 | |
| LR596014.1 | Thailand | 2013 | 95.41 | 95.22 |
| MK860761.1 | 95.41 | 95.22 | ||
| LR694074.1 | 2015 | 95.35 | 95.16 | |
| LC496857.1 | Ghana | 2016 | 95.28 | 95.04 |
| LR694076.1 | Uganda | 95.48 | 95.25 | |
| MZ972994.1 | Brazil | 2017 | 94.81 | 94.56 |
| LR694073.1 | USA | 2016 | 94.86 | 94.61 |
| LR694079.1 | Cambodia | 2013 | 94.5 | 94.26 |
| LR694080.1 | 94.52 | 94.28 | ||
| LR694078.1 | 2015 | 94.38 | 94.15 |
4. Discussion
Funding
Declarations
References
- Fredericks AC, Russell TA, Wallace LE, Davidson AD, Fernandez-Sesma A, Maringer K. Aedes aegypti (Aag2)-derived clonal mosquito cell lines reveal the effects of pre-existing persistent infection with the insect-specific bunyavirus Phasi Charoen-like virus on arbovirus replication. PLoS Negl Trop Dis. 2019;13: e0007346. [CrossRef]
- Bhatt S, Gething PW, Brady OJ, Messina JP, Farlow AW, Moyes CL, et al. The global distribution and burden of dengue. Nature. 2013;496: 504–507. [CrossRef]
- Romo H, Kenney JL, Blitvich BJ, Brault AC. Restriction of Zika virus infection and transmission in Aedes aegypti mediated by an insect-specific flavivirus. Emerg Microbes Infect. 2018;7: 1–13. [CrossRef]
- Hermanns K, Zirkel F, Kopp A, Marklewitz M, Rwego IB, Estrada A, et al. Discovery of a novel alphavirus related to Eilat virus. J Gen Virol. 2017;98: 43–49. [CrossRef]
- Datta S, Gopalakrishnan R, Chatterjee S, Veer V. Phylogenetic Characterization of a Novel Insect-Specific Flavivirus Detected in a Culex Pool, Collected from Assam, India. Intervirology. 2015;58: 149–154. [CrossRef]
- Utarini A, Indriani C, Ahmad RA, Tantowijoyo W, Arguni E, Ansari MR, et al. Efficacy of Wolbachia-Infected Mosquito Deployments for the Control of Dengue. N Engl J Med. 2021;384: 2177–2186. [CrossRef]
- Öhlund P, Lundén H, Blomström A-L. Insect-specific virus evolution and potential effects on vector competence. Virus Genes. 2019;55: 127–137. [CrossRef]
- Thomas T, Gilbert J, Meyer F. Metagenomics: a guide from sampling to data analysis. Microb Inform Exp. 2012;2: 3. [CrossRef]
- Santos-Medellin C, Zinke LA, ter Horst AM, Gelardi DL, Parikh SJ, Emerson JB. Viromes outperform total metagenomes in revealing the spatiotemporal patterns of agricultural soil viral communities. ISME J. 2021;15: 1956–1970. [CrossRef]
- Hameed M, Wahaab A, Shan T, Wang X, Khan S, Di D, et al. A Metagenomic Analysis of Mosquito Virome Collected From Different Animal Farms at Yunnan–Myanmar Border of China. Front Microbiol. 2021;11. Available: https://www.frontiersin.org/articles/10.3389/fmicb.2020.591478.
- Shi C, Beller L, Deboutte W, Yinda KC, Delang L, Vega-Rúa A, et al. Stable distinct core eukaryotic viromes in different mosquito species from Guadeloupe, using single mosquito viral metagenomics. Microbiome. 2019;7: 121. [CrossRef]
- Xiao P, Han J, Zhang Y, Li C, Guo X, Wen S, et al. Metagenomic Analysis of Flaviviridae in Mosquito Viromes Isolated From Yunnan Province in China Reveals Genes From Dengue and Zika Viruses. Front Cell Infect Microbiol. 2018;8. Available: https://www.frontiersin.org/articles/10.3389/fcimb.2018.00359.
- Lole K, Ramdasi A, Patil S, Thakar S, Nath A, Ghuge O, et al. Abundance of Phasi-Charoen-like virus in Aedes aegypti mosquito populations in different states of India. PLOS ONE. 2022;17: e0277276. [CrossRef]
- Chrzastek K, Lee D, Smith D, Sharma P, Suarez DL, Pantin-Jackwood M, et al. Use of Sequence-Independent, Single-Primer-Amplification (SISPA) for rapid detection, identification, and characterization of avian RNA viruses. Virology. 2017;509: 159–166. [CrossRef]
- Schoch CL, Ciufo S, Domrachev M, Hotton CL, Kannan S, Khovanskaya R, et al. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database J Biol Databases Curation. 2020;2020: baaa062. [CrossRef]
- Mihara T, Nishimura Y, Shimizu Y, Nishiyama H, Yoshikawa G, Uehara H, et al. Linking Virus Genomes with Host Taxonomy. Viruses. 2016;8: 66. [CrossRef]
- Vilsker M, Moosa Y, Nooij S, Fonseca V, Ghysens Y, Dumon K, et al. Genome Detective: an automated system for virus identification from high-throughput sequencing data. Bioinformatics. 2019;35: 871–873. [CrossRef]
- Madeira F, Pearce M, Tivey ARN, Basutkar P, Lee J, Edbali O, et al. Search and sequence analysis tools services from EMBL-EBI in 2022. Nucleic Acids Res. 2022;50: W276–W279. [CrossRef]
- Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25: 3389–3402. [CrossRef]
- Zakrzewski M, Rašić G, Darbro J, Krause L, Poo YS, Filipović I, et al. Mapping the virome in wild-caught Aedes aegypti from Cairns and Bangkok. Sci Rep. 2018;8: 4690. [CrossRef]
- Leinonen R, Sugawara H, Shumway M, on behalf of the International Nucleotide Sequence Database Collaboration. The Sequence Read Archive. Nucleic Acids Res. 2011;39: D19–D21. [CrossRef]
- Wragg D, Mason AS, Yu L, Kuo R, Lawal RA, Desta TT, et al. Genome-wide analysis reveals the extent of EAV-HP integration in domestic chicken. BMC Genomics. 2015;16: 784. [CrossRef]
- Bai L, Wang L, Vega-Rodríguez J, Wang G, Wang S. A Gut Symbiotic Bacterium Serratia marcescens Renders Mosquito Resistance to Plasmodium Infection Through Activation of Mosquito Immune Responses. Front Microbiol. 2019;10: 1580. [CrossRef]
- Wu P, Sun P, Nie K, Zhu Y, Shi M, Xiao C, et al. A Gut Commensal Bacterium Promotes Mosquito Permissiveness to Arboviruses. Cell Host Microbe. 2019;25: 101-112.e5. [CrossRef]
- Hejazi A, Falkiner FR. Serratia marcescens. J Med Microbiol. 1997;46: 903–912. [CrossRef]
- Khanna A, Khanna M, Aggarwal A. Serratia marcescens- A Rare Opportunistic Nosocomial Pathogen and Measures to Limit its Spread in Hospitalized Patients. J Clin Diagn Res JCDR. 2013;7: 243–246. [CrossRef]
- Rohit A, Suresh Kumar D, Dhinakaran I, Joy J, Vijay Kumar D, Kumar Ballamoole K, et al. Whole-genome-based analysis reveals multiclone Serratia marcescens outbreaks in a non-Neonatal Intensive Care Unit setting in a tertiary care hospital in India. J Med Microbiol. 2019;68: 616–621. [CrossRef]
- Accoti A, Damiani C, Nunzi E, Cappelli A, Iacomelli G, Monacchia G, et al. Anopheline mosquito saliva contains bacteria that are transferred to a mammalian host through blood feeding. Front Microbiol. 2023;14: 1157613. [CrossRef]
- Esnault G, Earley B, Cormican P, Waters SM, Lemon K, Cosby SL, et al. Assessment of Rapid MinION Nanopore DNA Virus Meta-Genomics Using Calves Experimentally Infected with Bovine Herpes Virus-1. Viruses. 2022;14: 1859. [CrossRef]
- Greninger AL, Naccache SN, Federman S, Yu G, Mbala P, Bres V, et al. Rapid metagenomic identification of viral pathogens in clinical samples by real-time nanopore sequencing analysis. Genome Med. 2015;7: 99. [CrossRef]
- Jia X, Hu L, Wu M, Ling Y, Wang W, Lu H, et al. A streamlined clinical metagenomic sequencing protocol for rapid pathogen identification. Sci Rep. 2021;11: 4405. [CrossRef]
- Regnault B, Bigot T, Ma L, Pérot P, Temmam S, Eloit M. Deep Impact of Random Amplification and Library Construction Methods on Viral Metagenomics Results. Viruses. 2021;13: 253. [CrossRef]
- Phenuiviridae ~ ViralZone. [cited 17 Sep 2023]. Available: https://viralzone.expasy.org/7101.
- Wommack KE, Bhavsar J, Ravel J. Metagenomics: Read Length Matters. Appl Environ Microbiol. 2008;74: 1453–1463. [CrossRef]
- Otake S, Dee SA, Rossow KD, Moon RD, Pijoan C. Mechanical transmission of porcine reproductive and respiratory syndrome virus by mosquitoes, Aedes vexans (Meigen). Can J Vet Res. 2002;66: 191–195.
- Yang X, Hou L, Ye J, He Q, Cao S. Detection of Porcine Circovirus Type 2 (PCV2) in Mosquitoes from Pig Farms by PCR.
- Garcia LAT, Viancelli A, Rigotto C, Pilotto MR, Esteves PA, Kunz A, et al. Surveillance of human and swine adenovirus, human norovirus and swine circovirus in water samples in Santa Catarina, Brazil. J Water Health. 2012;10: 445–452. [CrossRef]
- Powell JR, Gloria-Soria A, Kotsakiozi P. Recent History of Aedes aegypti: Vector Genomics and Epidemiology Records. Bioscience. 2018;68: 854–860. [CrossRef]
- Rose NH, Badolo A, Sylla M, Akorli J, Otoo S, Gloria-Soria A, et al. Dating the origin and spread of specialization on human hosts in Aedes aegypti mosquitoes. Perry GH, Lou N, editors. eLife. 2023;12: e83524. [CrossRef]
- Palatini U, Alfano N, Carballar RL, Chen X-G, Delatte H, Bonizzoni M. Virome and nrEVEome diversity of Aedes albopictus mosquitoes from La Reunion Island and China. Virol J. 2022;19: 190. [CrossRef]


| Primer name | Sequence | Reference |
|---|---|---|
| Random anchored reverse primer RT5 | CATCACATAGGCGTCCGCTGNNNNNN | [12] |
| Anchored forward primer P5 | CATCACATAGGCGTCCGCTG | |
| Random anchored reverse primer RT12 | GGTGGGCGTGTGAAATCGACNNNNNN | |
| Anchored forward primer P12 | GGTGGGCGTGTGAAATCGAC | |
| Random anchored reverse primer IDT-K-8N | GACCATCTAGCGACCTCCACNNNNNNNN | [14] |
| Anchored forward primer IDT-K | GACCATCTAGCGACCTCCAC | |
| CFAV-Capsid-32F | CAGTTTGGGTCACGCTTA | This study |
| CFAV-Capsid-536R | ATGTCAATCACCACGCAT |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




