Submitted:
17 November 2023
Posted:
17 November 2023
You are already at the latest version
Abstract
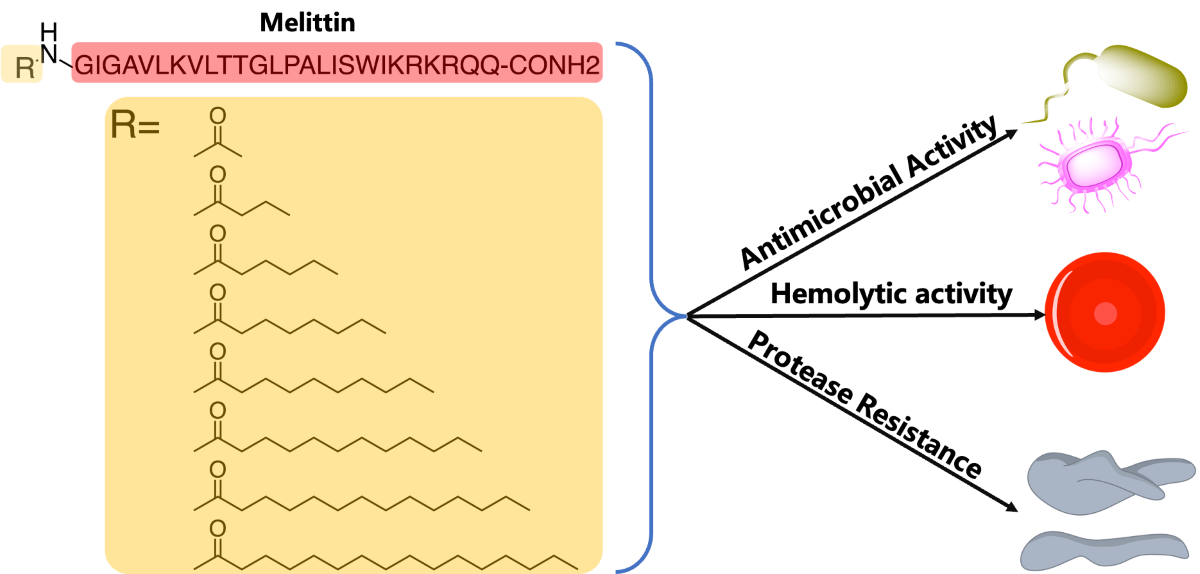
Keywords:
1. Introduction
2. Results and Discussion
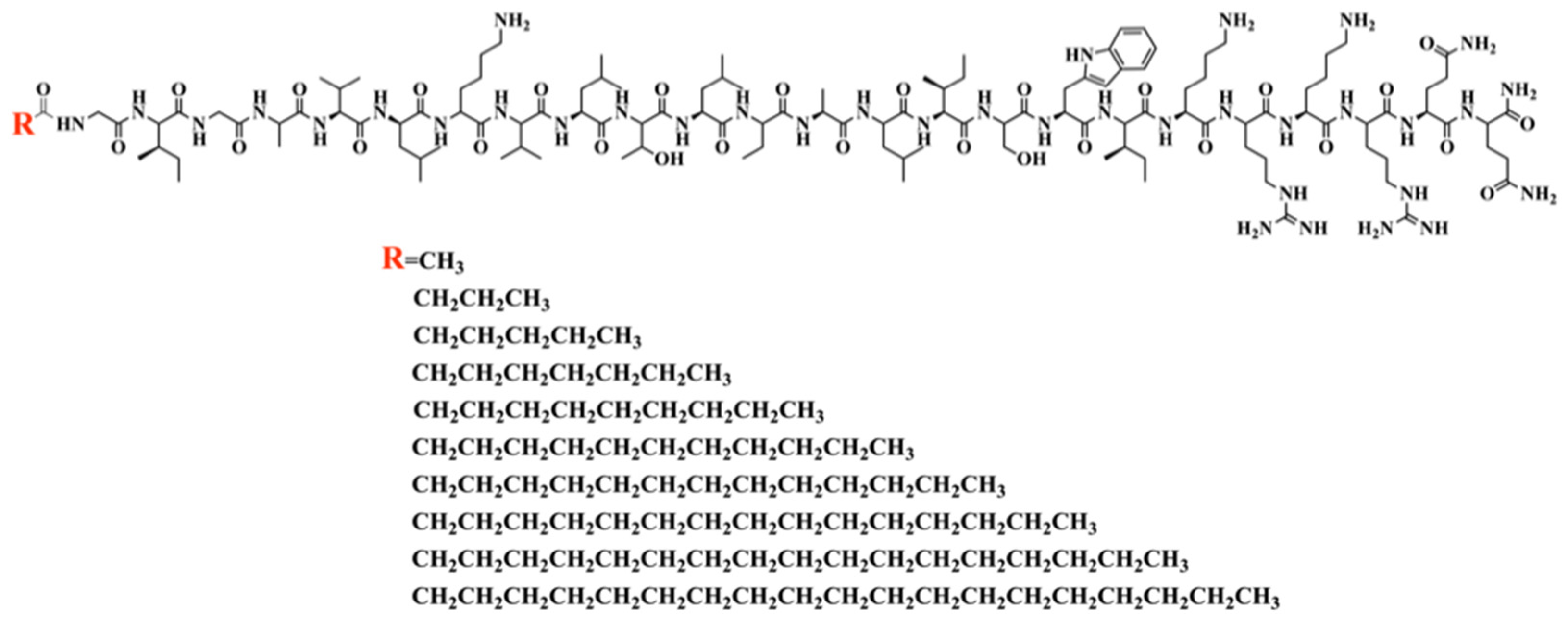
2.1. The Characterization of MDLs
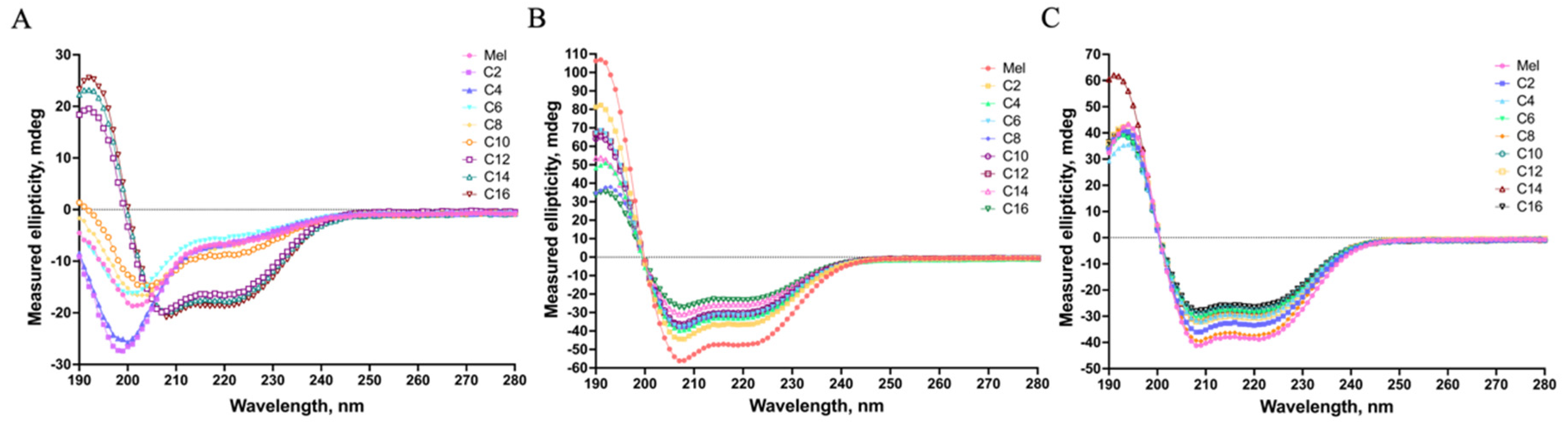
2.2. The Secondary Structure of MDLs
2.3. Antimicrobial Activity in vitro
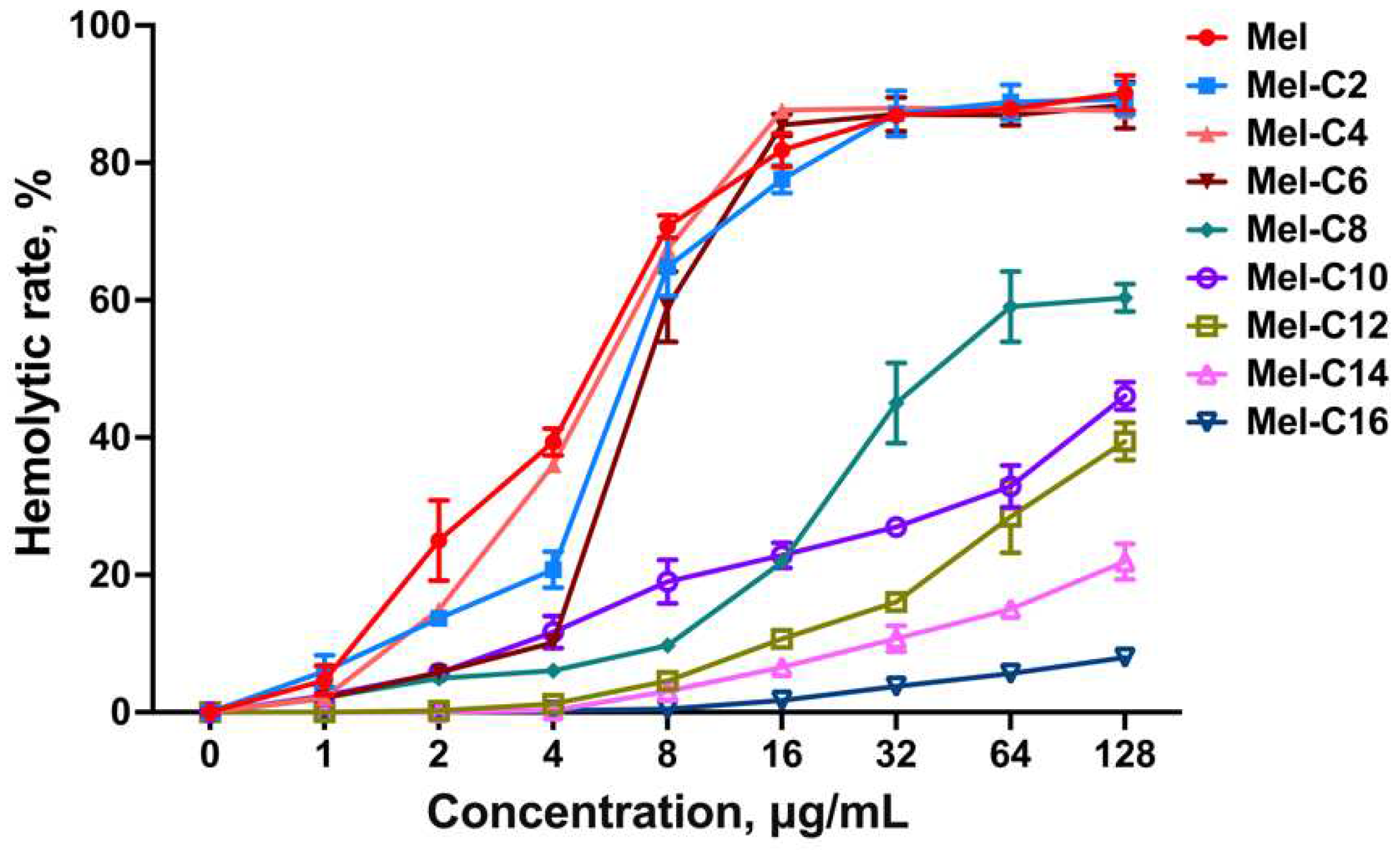
2.5. Hemolytic Assay
2.6. Stability Assay
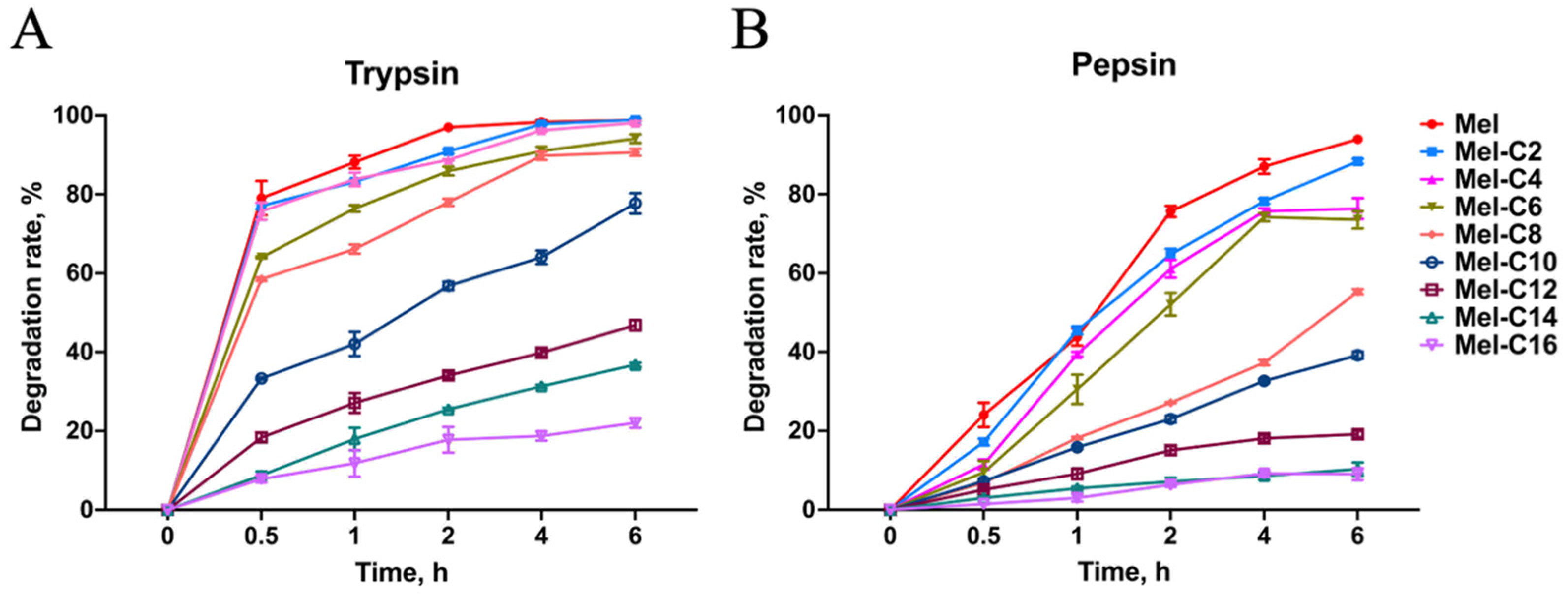
2.6.1. Protease Resistance Assay
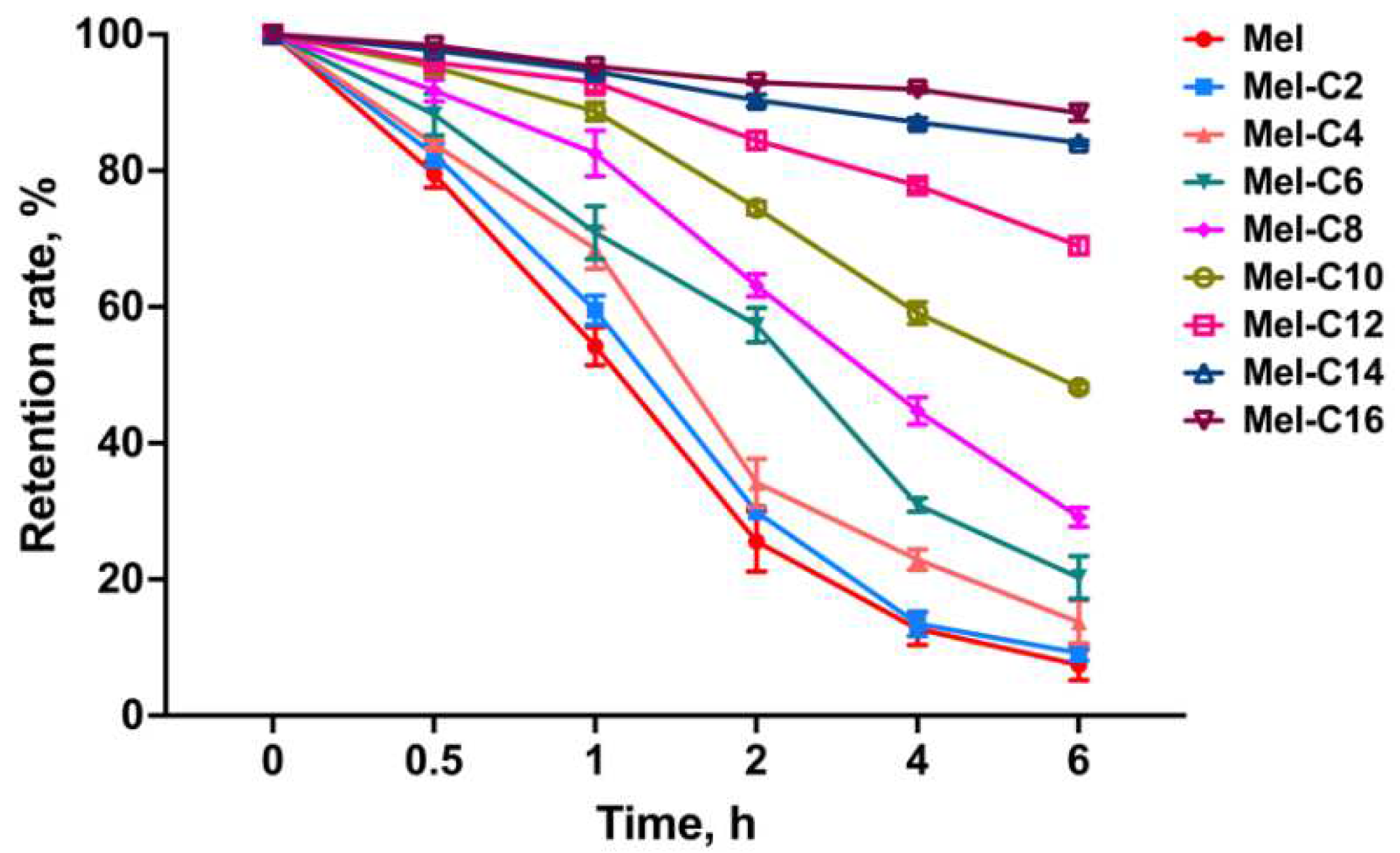
2.6.2. Serum Stability
2.6.3. Salt Sensitivity
2.7. The Antimicrobial Mechanisms of the MDLs
2.7.1. Outer Membrane Permeabilization Assay
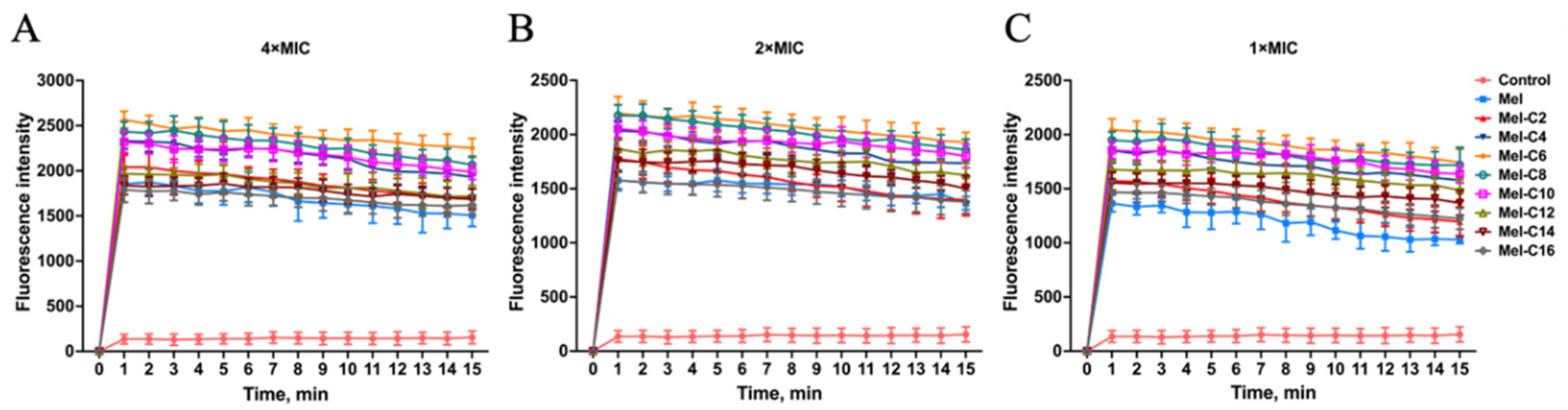
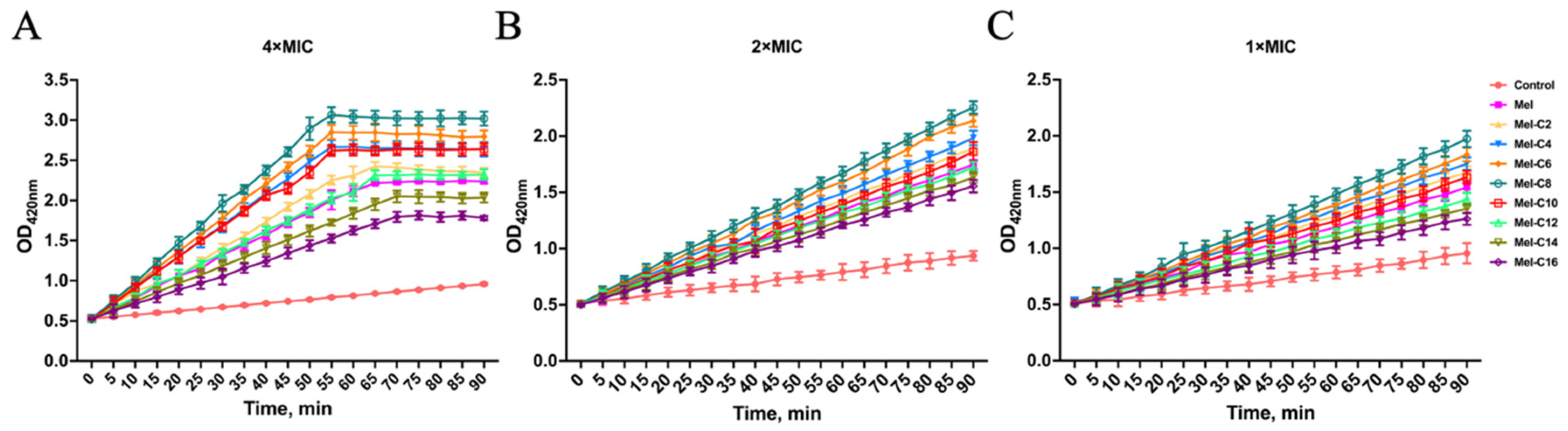
2.7.2. Inner Membrane Permeabilization Assay
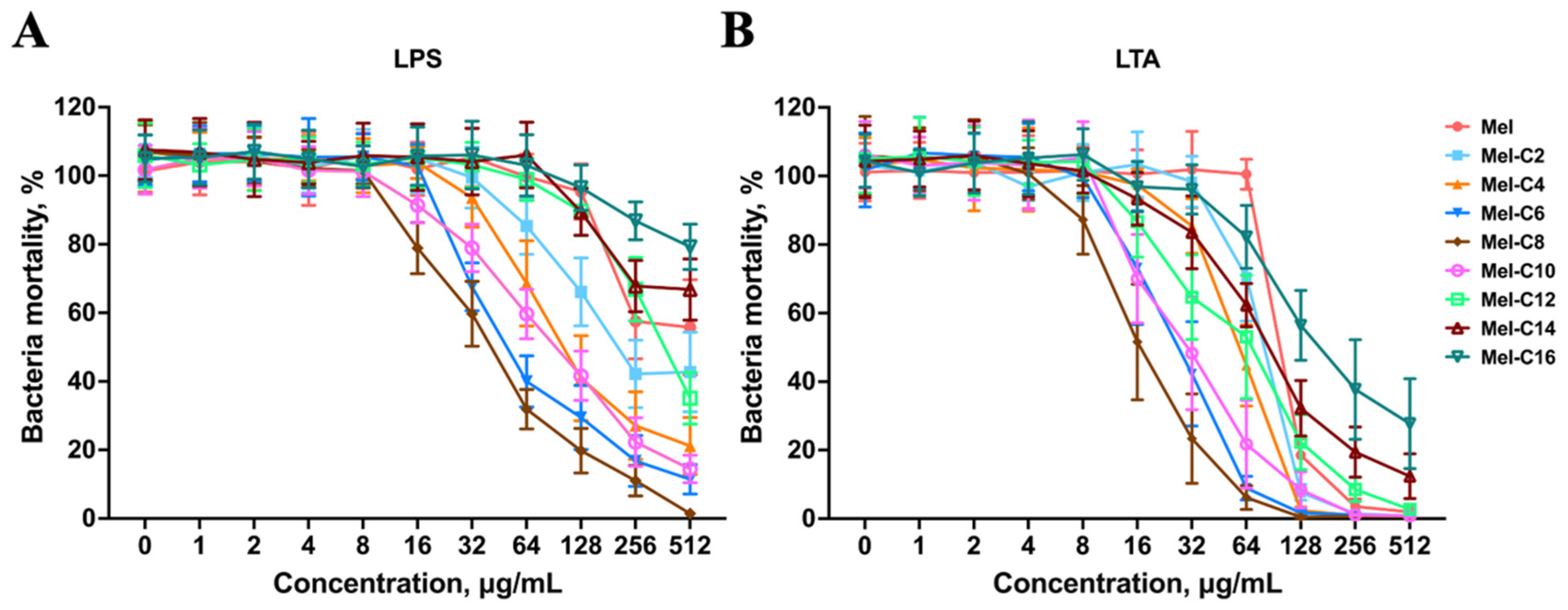
2.7.3. LPS/LTA Competitive Inhibition Assay
3. Materials and Method
3.1. Bacterial Strain
3.2. Peptide Design, Synthesis, and Analysis
3.3. Hydrophobicity Assay
3.4. Circular Dichroism (CD) Measurements
3.5. Minimal Inhibitory Concentration (MIC) Measurements
3.6. Hemolytic Activity Assay
3.7. Stability Assay
3.7.1. Protease Resistance Assay
3.7.2. Serum Stability
3.7.3. Salt Sensitivity
3.8. Antimicrobial Mechanism
3.8.1. Outer Membrane Permeabilization Assay
3.8.2. Inner Membrane Permeabilization Assay
3.8.3. LPS/LTA Competitive Inhibition Assay
4. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Reynolds, D.; Burnham, J. P.; Vazquez Guillamet, C.; McCabe, M.; Yuenger, V.; Betthauser, K.; Micek, S. T.; Kollef, M. H., The threat of multidrug-resistant/extensively drug-resistant Gram-negative respiratory infections: another pandemic Eur. Respir. Rev. 2022, 31, 166. [CrossRef]
- Smith, R. P.; May, H. E.; AbuOun, M.; Stubberfield, E.; Gilson, D.; Chau, K. K.; Crook, D. W.; Shaw, L. P.; Read, D. S.; Stoesser, N.; Vilar, M. J.; Anjum, M. F., A longitudinal study reveals persistence of antimicrobial resistance on livestock farms is not due to antimicrobial usage alone. Front Microbiol 2023, 14, 1070340. [CrossRef]
- Shao, Y.; Wang, Y.; Yuan, Y.; Xie, Y., A systematic review on antibiotics misuse in livestock and aquaculture and regulation implications in China. Sci. Total. Environ. 2021, 798, 149205. [CrossRef]
- Huang, S.; Wang, J.; Sun, H.; Fu, Y.; Wang, Y. Probing Changes in Ca2+-Induced Interaction Forces between Calmodulin and Melittin by Atomic Force Microscopy. Micromachines 2020, 11, 906. [CrossRef]
- Lima, W.G.; de Brito, J.C.M.; Cardoso, V.N.; Fernandes, S.O.A. In-depth characterization of antibacterial activity of melittin against Staphylococcus aureus and use in a model of non-surgical MRSA-infected skin wounds. Eur. J. Pharm. Sci. 2021, 156, 105592. [CrossRef]
- Pandey, B.K.; Ahmad, A.; Asthana, N.; Azmi, S.; Srivastava, R.M.; Srivastava, S.; Verma, R.; Vishwakarma, A.L.; Ghosh, J.K. Cell-Selective Lysis by Novel Analogues of Melittin against Human Red Blood Cells and Escherichia coli. Biochemistry 2010, 49, 7920–7929. [CrossRef]
- Wang, H.; Wang, S.; Wang, R.; Wang, X.; Jiang, K.; Xie, C.; Zhan, C.; Wang, H.; Lu, W. Co-delivery of paclitaxel and melittin by glycopeptide-modified lipodisks for synergistic anti-glioma therapy. Nanoscale 2019, 11, 13069–13077. [CrossRef]
- Wang, X.; Yang, X.; Wang, Q.; Meng, D. Unnatural amino acids: promising implications for the development of new antimicrobial peptides. Crit. Rev. Microbiol. 2022, 49, 231–255. [CrossRef]
- Liu, T.; Zhu, N.; Zhong, C.; Zhu, Y.; Gou, S.; Chang, L.; Bao, H.; Liu, H.; Zhang, Y.; Ni, J. Effect of N-methylated and fatty acid conjugation on analogs of antimicrobial peptide Anoplin. Eur. J. Pharm. Sci. 2020, 152, 105453. [CrossRef]
- Lai, Z.; Yuan, X.; Chen, H.; Zhu, Y.; Dong, N.; Shan, A. Strategies employed in the design of antimicrobial peptides with enhanced proteolytic stability. Biotechnol. Adv. 2022, 59, 107962. [CrossRef]
- Grimsey, E.; Collis, D.W.; Mikut, R.; Hilpert, K. The effect of lipidation and glycosylation on short cationic antimicrobial peptides. BBA-Biomembranes 2020, 1862, 183195. [CrossRef]
- Li, R.; He, S.; Yin, K.; Zhang, B.; Yi, Y.; Zhang, M.; Pei, N.; Huang, L. Effects of N-terminal modifications on the stability of antimicrobial peptide SAMP-A4 analogues against protease degradation. J. Pept. Sci. 2021, 27, e3352. [CrossRef]
- Zhong, C.; Zhu, N.; Zhu, Y.; Liu, T.; Gou, S.; Xie, J.; Yao, J.; Ni, J. Antimicrobial peptides conjugated with fatty acids on the side chain of D-amino acid promises antimicrobial potency against multidrug-resistant bacteria. Eur. J. Pharm. Sci. 2020, 141, 105123. [CrossRef]
- Cui, A.-L.; Hu, X.-X.; Chen, Y.; Jin, J.; Yi, H.; Wang, X.-K.; He, Q.-Y.; You, X.-F.; Li, Z.-R. Design, Synthesis, and Bioactivity of Cyclic Lipopeptide Antibiotics with Varied Polarity, Hydrophobicity, and Positive Charge Distribution. ACS Infect. Dis. 2020, 6, 1796–1806. [CrossRef]
- Balleza, D.; Alessandrini, A.; Beltrán García, M. J., Role of Lipid Composition, Physicochemical Interactions, and Membrane Mechanics in the Molecular Actions of Microbial Cyclic Lipopeptides. J Membrane Biol 2019, 252, 131–157. [CrossRef]
- Neubauer, D.; Jaśkiewicz, M.; Bauer, M.; Gołacki, K.; Kamysz, W. Ultrashort Cationic Lipopeptides–Effect of N-Terminal Amino Acid and Fatty Acid Type on Antimicrobial Activity and Hemolysis. Molecules 2020, 25, 257. [CrossRef]
- Jujjavarapu, S.E.; Dhagat, S. In Silico Discovery of Novel Ligands for Antimicrobial Lipopeptides for Computer-Aided Drug Design. Probiotics Antimicro 2017, 10, 129–141. [CrossRef]
- Chu-Kung, A.F.; Bozzelli, K.N.; Lockwood, N.A.; Haseman, J.R.; Mayo, K.H.; Tirrell, M.V. Promotion of Peptide Antimicrobial Activity by Fatty Acid Conjugation. Bioconjugate Chem. 2004, 15, 530–535. [CrossRef]
- Zhong, C.; Liu, T.; Gou, S.; He, Y.; Zhu, N.; Zhu, Y.; Wang, L.; Liu, H.; Zhang, Y.; Yao, J., Design and synthesis of new N-terminal fatty acid modified-antimicrobial peptide analogues with potent in vitro biological activity. Eur. J. Med. Chem. 2019, 182, 111636. [CrossRef]
- Storck, P.; Umstätter, F.; Wohlfart, S.; Domhan, C.; Kleist, C.; Werner, J.; Brandenburg, K.; Zimmermann, S.; Haberkorn, U.; Mier, W., Fatty acid conjugation leads to length-dependent antimicrobial activity of a synthetic antibacterial peptide (Pep19-4LF). Antibiotics 2020, 9, 844. [CrossRef]
- Malina, A.; Shai, Y. Conjugation of fatty acids with different lengths modulates the antibacterial and antifungal activity of a cationic biologically inactive peptide. Biochem. J. 2005, 390, 695–702. [CrossRef]
- Liu, H.; Yang, N.; Teng, D.; Mao, R.; Hao, Y.; Ma, X.; Wang, X.; Wang, J. Fatty acid modified-antimicrobial peptide analogues with potent antimicrobial activity and topical therapeutic efficacy against Staphylococcus hyicus. Appl. Microbiol. Biotechnol. 2021, 105, 5845–5859. [CrossRef]
- Chu-Kung, A.F.; Nguyen, R.; Bozzelli, K.N.; Tirrell, M. Chain length dependence of antimicrobial peptide–fatty acid conjugate activity. J. Colloid Interface Sci. 2010, 345, 160–167. [CrossRef]
- Koh, J.-J.; Lin, H.; Caroline, V.; Chew, Y.S.; Pang, L.M.; Aung, T.T.; Li, J.; Lakshminarayanan, R.; Tan, D.T.H.; Verma, C.; et al. N-Lipidated Peptide Dimers: Effective Antibacterial Agents against Gram-Negative Pathogens through Lipopolysaccharide Permeabilization. J. Med. Chem. 2015, 58, 6533–6548. [CrossRef]
- Watala, C.; Gwoździński, K. Melittin-induced alterations in dynamic properties of human red blood cell membranes. Chem. Interactions 1992, 82, 135–149. [CrossRef]
- Rounds, T.; Straus, S.K. Lipidation of Antimicrobial Peptides as a Design Strategy for Future Alternatives to Antibiotics. Int. J. Mol. Sci. 2020, 21, 9692. [CrossRef]
- Gao, J.; Xie, C.; Zhang, M.; Wei, X.; Yan, Z.; Ren, Y.; Ying, M.; Lu, W. RGD-modified lipid disks as drug carriers for tumor targeted drug delivery. Nanoscale 2016, 8, 7209–7216. [CrossRef]
- Lai, Z.; Tan, P.; Zhu, Y.; Shao, C.; Shan, A.; Li, L. Highly Stabilized α-Helical Coiled Coils Kill Gram-Negative Bacteria by Multicomplementary Mechanisms under Acidic Condition. ACS Appl. Mater. Interfaces 2019, 11, 22113–22128. [CrossRef]
- Tam, J.P.; Lu, Y.-A.; Yang, J.-L. Correlations of Cationic Charges with Salt Sensitivity and Microbial Specificity of Cystine-stabilized β-Strand Antimicrobial Peptides. J. Biol. Chem. 2002, 277, 50450–50456. [CrossRef]
- Wu, G.; Ding, J.; Li, H.; Li, L.; Zhao, R.; Fan, X.; Shen, Z., Effects of cations and pH on antimicrobial activity of thanatin and s-thanatin against Escherichia coli ATCC25922 and B. subtilis ATCC 21332. Curr Microbiol 2008, 57, 552–557.
- Choi, J.-H.; Lee, T.-K.; Byun, J.-W.; Lee, Y.-S. Preparation of a core–shell type MBHA resin and its application for solid-phase peptide synthesis. Tetrahedron Lett. 2009, 50, 4272–4275. [CrossRef]
- Kowalska-Krochmal, B.; Dudek-Wicher, R. The Minimum Inhibitory Concentration of Antibiotics: Methods, Interpretation, Clinical Relevance. Pathogens 2021, 10, 165. [CrossRef]
- Zhao, X.; Wang, X.; Shukla, R.; Kumar, R.; Weingarth, M.; Breukink, E.; Kuipers, O.P. Brevibacillin 2V, a Novel Antimicrobial Lipopeptide With an Exceptionally Low Hemolytic Activity. Front. Microbiol. 2021, 12, 693725. [CrossRef]
- Yu, H.; Shang, L.; Yang, G.; Dai, Z.; Zeng, X.; Qiao, S. Biosynthetic Microcin J25 Exerts Strong Antibacterial, Anti-Inflammatory Activities, Low Cytotoxicity Without Increasing Drug-Resistance to Bacteria Target. Front. Immunol. 2022, 13, 811378. [CrossRef]
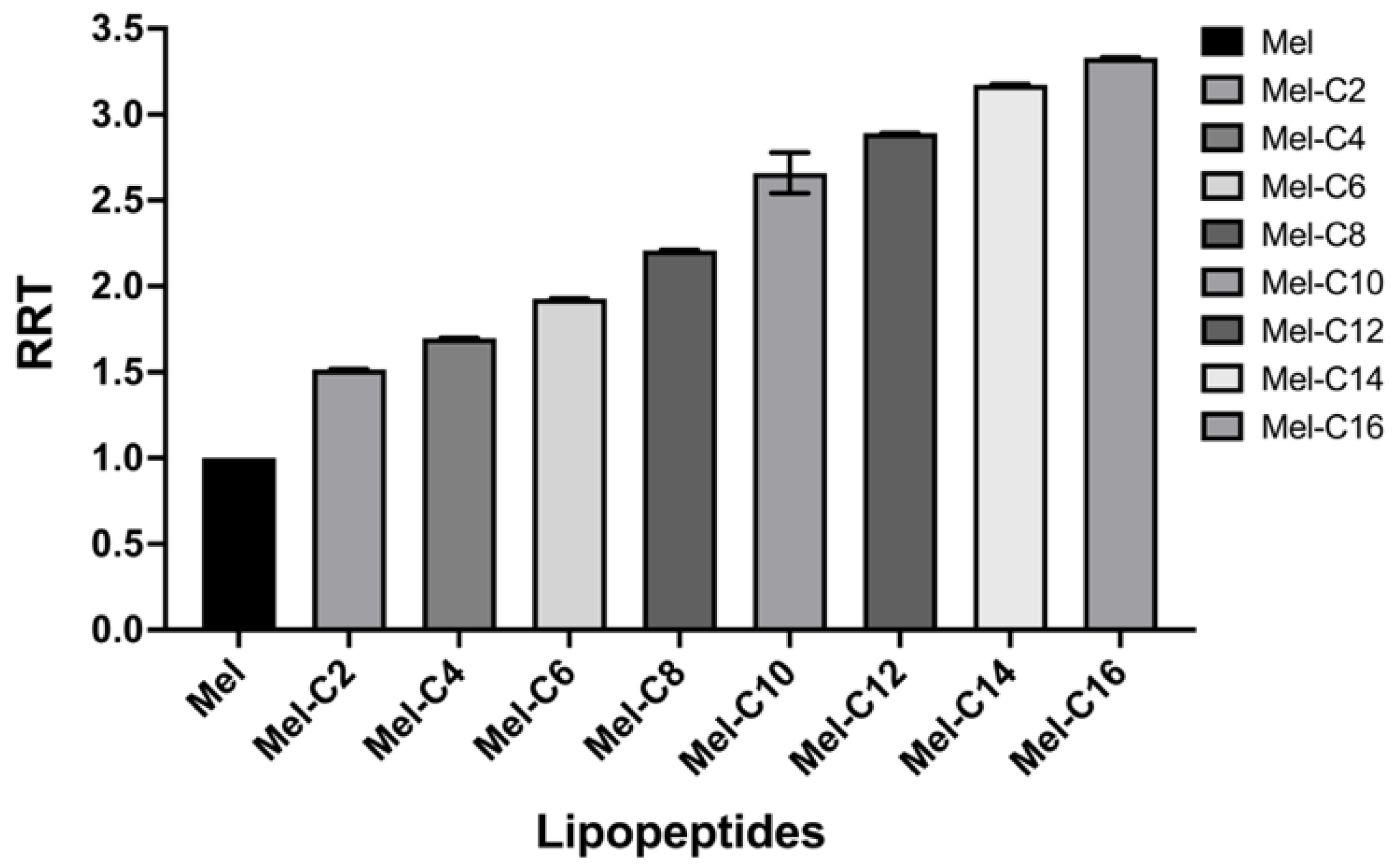
| Peptide | Sequence | Theoretical MW | Measured MWa | Net charge | tR, minb |
|---|---|---|---|---|---|
| Mel | GIGAVLKVLTTGLPALISWIKRKRQQ | 2846.46 | 2846.15 | +6 | 8.076±0.014 |
| C2-Mel | C2-GIGAVLKVLTTGLPALISWIKRKRQQ | 2889.50 | 2889.45 | +6 | 12.247±0.021 |
| C4-Mel | C4-GIGAVLKVLTTGLPALISWIKRKRQQ | 2917.56 | 2917.65 | +6 | 13.704±0.005 |
| C6-Mel | C6-GIGAVLKVLTTGLPALISWIKRKRQQ | 2945.61 | 2945.70 | +6 | 15.567±0.006 |
| C8-Mel | C8-GIGAVLKVLTTGLPALISWIKRKRQQ | 2973.66 | 2973.80 | +6 | 17.845±0.013 |
| C10-Mel | C10-GIGAVLKVLTTGLPALISWIKRKRQQ | 3001.71 | 3001.60 | +6 | 20.541±0.029 |
| C12-Mel | C12-GIGAVLKVLTTGLPALISWIKRKRQQ | 3029.77 | 3029.70 | +6 | 23.344±0.004 |
| C14-Mel | C14-GIGAVLKVLTTGLPALISWIKRKRQQ | 3057.82 | 3057.6 | +6 | 25.621±0.002 |
| C16-Mel | C16-GIGAVLKVLTTGLPALISWIKRKRQQ | 3085.87 | 3085.7 | +6 | 26.892±0.005 |
| Peptide | PBS buffer | 50% TFE | 25 mM SDS | |||
|---|---|---|---|---|---|---|
| α-helix, % | β-fold, % | α-helix, % | β-fold, % | α-helix, % | β-fold, % | |
| Mel | 6.8 | 38.5 | 34.5 | 5.5 | 21.5 | 17.9 |
| C2-MEL | 6 | 32.8 | 26.5 | 17.8 | 20.9 | 19.9 |
| C4-MEL | 6.1 | 33.7 | 20.7 | 25.4 | 19.3 | 23.8 |
| C6-MEL | 6.5 | 39.3 | 22.4 | 27 | 18.4 | 24.7 |
| C8-MEL | 7 | 40 | 18.8 | 25.5 | 18.2 | 25.4 |
| C10-MEL | 7.5 | 41.1 | 18.4 | 22.1 | 17.1 | 28.8 |
| C12-MEL | 11.5 | 31.3 | 19 | 21.5 | 15.9 | 29.5 |
| C14-MEL | 12 | 31 | 16 | 25.2 | 15.5 | 31.7 |
| C16-MEL | 12.5 | 30.4 | 14.3 | 26.8 | 15.2 | 33 |
| Peptide | MIC | ||||||
|---|---|---|---|---|---|---|---|
| Gram-positive | Gram-negative | ||||||
| S.aureus ATCC 43300 | L. monocytogenes CVCC 3764 | E. faecalis ATCC 29212 | B. cereus CVCC 4101 | E. coli ATCC 25922 | S dysenteriae CGMCC 1.1869 | multidrug-resistant strain E. coli | |
| Mel | 8 | 32 | 16 | 16 | 32 | 32 | 64 |
| C2-Mel | 4 | 32 | 4 | 8 | 16 | 8 | 64 |
| C4-Mel | 2 | 8 | 2 | 8 | 8 | 8 | 32 |
| C6-Mel | 1 | 4 | 2 | 4 | 4 | 8 | 8 |
| C8-Mel | 0.5 | 4 | 1 | 2 | 2 | 4 | 4 |
| C10-Mel | 2 | 16 | 8 | 8 | 8 | 32 | 8 |
| C12-Mel | 8 | 32 | 16 | 64 | 32 | 64 | 32 |
| C14-Mel | 32 | 128 | 32 | >256 | 128 | 128 | 128 |
| C16-Mel | 64 | >256 | 64 | >256 | >256 | 128 | >256 |
| Bacterial strain | Peptide | Control | Salt | |||
|---|---|---|---|---|---|---|
| NaCl | KCl | MgCl2 | FeCl3 | |||
| E. coliATCC 25922 | Mel | 32 | 32 | 32 | 64 | 64 |
| Mel-C2 | 16 | 16 | 16 | 32 | 32 | |
| Mel-C4 | 8 | 8 | 16 | 32 | 32 | |
| Mel-C6 | 4 | 4 | 8 | 16 | 16 | |
| Mel-C8 | 2 | 2 | 8 | 8 | 8 | |
| Mel-C10 | 8 | 8 | 16 | 32 | 32 | |
| Mel-C12 | 16 | 32 | 32 | 128 | 128 | |
| Mel-C14 | 32 | 128 | 64 | 256 | 256 | |
| Mel-C16 | 128 | >256 | 256 | >256 | >256 | |
| S.aureus ATCC 43300 | Mel | 8 | 8 | 8 | 32 | 32 |
| Mel-C2 | 4 | 4 | 4 | 16 | 16 | |
| Mel-C4 | 2 | 2 | 4 | 16 | 16 | |
| Mel-C6 | 1 | 1 | 2 | 8 | 8 | |
| Mel-C8 | 0.5 | 0.5 | 1 | 8 | 8 | |
| Mel-C10 | 2 | 2 | 8 | 32 | 32 | |
| Mel-C12 | 8 | 16 | 32 | 64 | 64 | |
| Mel-C14 | 32 | 64 | 64 | 256 | 256 | |
| Mel-C16 | 64 | 128 | 256 | >256 | >256 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




