Submitted:
21 September 2023
Posted:
25 September 2023
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
- Simplicity: Single-channel ECG processing is simpler and more straightforward, which can make the development and debugging of algorithms easier.
- Data Availability: In some situations, only single-lead ECG data might be available (e.g., Icentia11k DB). Many portable and wearable ECG devices only record a single lead, so algorithms designed for single-lead data can be more broadly applicable.
- Robustness to Noise: Single-lead ECGs might be less susceptible to noise and artifacts that can affect multi-lead recordings. For instance, movement artifacts can affect different leads to different extents, potentially making multi-lead data more challenging to interpret. By analyzing each lead independently we may overcome this.
2. Related Work
- Real world settings are not considered. The electrocardiogram (ECG) data inherently contain various types of noise, including baseline wandering, power line interference, muscle noise and other artifacts related to contact with the electrodes. These noise elements pose significant challenges to the extraction of robust features, consequently affecting the performance of PVC classification in real-world settings. Thus, an algorithm that performs well on a clean, noise-free dataset may not perform as well when deployed in a real-world setting where the noise level is higher or varies unpredictably.
- Testing datasets are not representative: Gender differences in ECG are well documented in the literature. Men and women can have different heart rates, QRS complex durations, QT intervals, and T-wave morphologies, among other characteristics. These differences can affect the performance of PVC detection algorithms if they are not properly accounted for during algorithm development and testing [8,9].
- Training and testing datasets are not separated. A notable limitation of many existing methods lies in their reliance on small or overlapped ECG datasets for training and testing. This practice raises questions about their efficiency and generalizability when applied to a large collection of ECG recordings, an issue that remains largely unaddressed in the literature [10].
2.1. Beat Detection Performances
| Algorithm | Dataset | Se (%) | PPV (%) | F1 |
|---|---|---|---|---|
| Pan and Tompkins [12] | MIT-BIH | 99.76 | 99.56 | 99.66 |
| Christov [13] | MIT-BIH | 99.74 | 99.65 | 99.69 |
| Chiarugi et al [14] | MIT-BIH | 99.76 | 99.81 | 99.78 |
| Chouakri et al [15] | MIT-BIH | 98.68 | 97.24 | 97.95 |
| Elgendi [16] | MIT-BIH | 99.78 | 99.87 | 99.82 |
| BeatLogic [11] | MIT-BIH | 99.60 | 99.78 | 99.69 |
| Liu et al. [17] | MIT-BIH | 99.00 | 99.20 | 99.10 |
| He et al. [18] | MIT-BIH | 99.56 | 99.72 | 99.64 |
| Martinez et al [19] | MIT-BIH VFib excluded | 99.80 | 99.86 | 99.83 |
| Arzeno et al [20] | MIT-BIH VFib excluded | 99.68 | 99.63 | 99.65 |
| Zidelmal et al [21] | MIT-BIH VFib excluded | 99.64 | 99.82 | 99.73 |
| BeatLogic [11] | MIT-BIH VFib excluded | 99.60 | 99.90 | 99.75 |
2.2. PVC Detection Performances
| Algorithm | Dataset | Se (%) | PPV (%) | F1 |
|---|---|---|---|---|
| de Chazal et al [22] | MIT-BIH 11 | 77.5 | 90.6 | 83.5 |
| Jiang and Kong [23] | MIT-BIH 11 | 94.3 | 95.8 | 95.0 |
| Ince et al [24] | MIT-BIH 11 | 90.3 | 92.2 | 91.2 |
| Kiranyaz et al [2] | MIT-BIH 11 | 95.9 | 96.2 | 96.0 |
| Zhang et al [25] | MIT-BIH 11 | 97.6 | 97.6 | 97.6 |
| BeatLogic [11] | MIT-BIH 11 | 97.9 | 98.9 | 98.4 |
| Liu et al [17] | MIT-BIH (22 Records) | 91.6 | 95.6 | 93.6 |
3. Methods
3.1. Overview
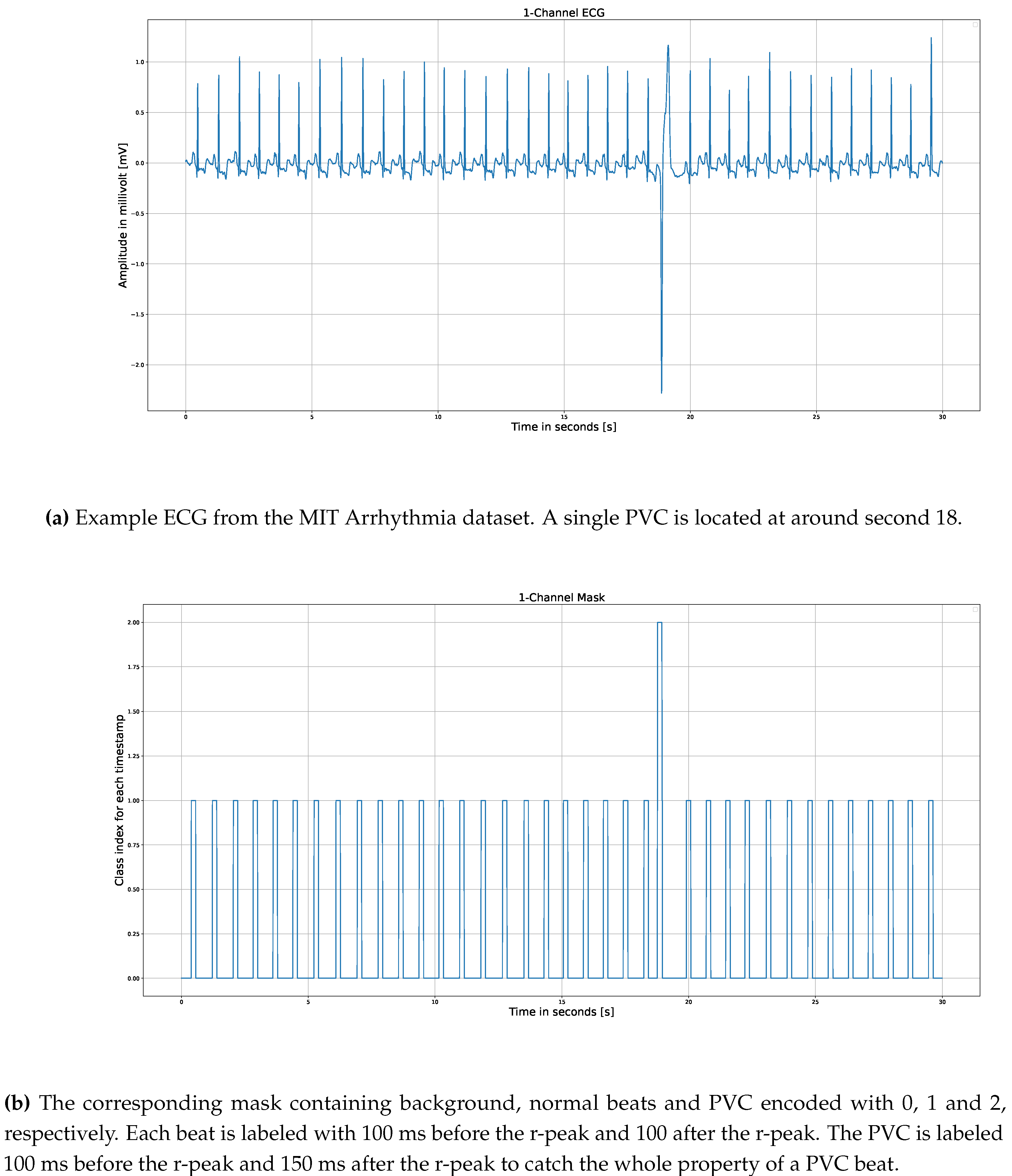
3.2. Data Preparation
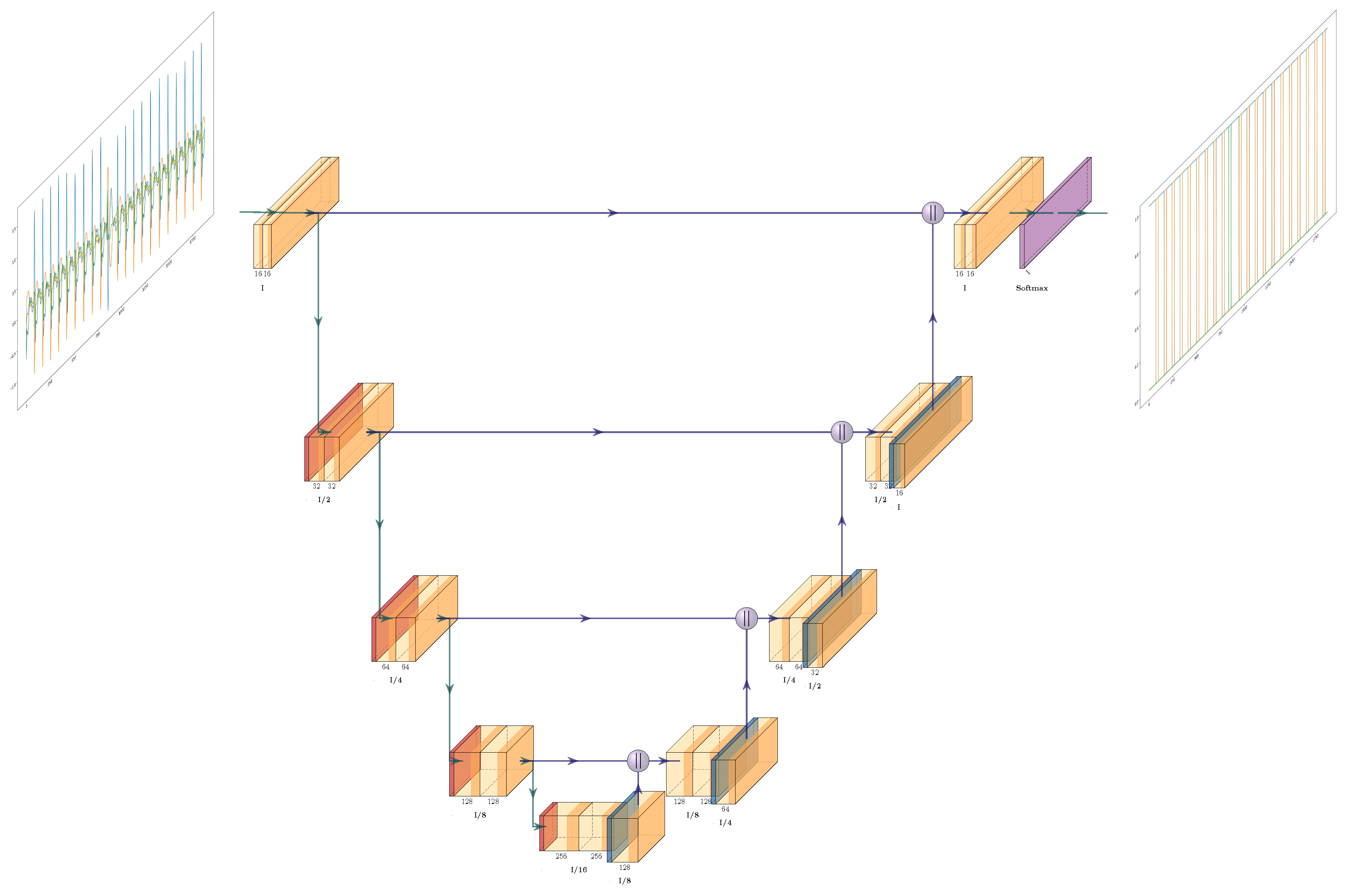
3.3. Model Architecture
3.4. Augmentation
- Scaling the amplitude with a probability of and scaling factor of .
- Offset the amplitude with a probability of and a offset value of .
- Adding gaussian, brown or pink noise with a probability of and a offset value of .
3.5. Post-Processing
- The No Beat Mask: This provides the probability for each time point that it does not lie within any beat.
- The Normal Beat Mask: This provides the probability for each time point that it lies within a normal beat.
- The PVC Mask: This provides the probability for each time point that it lies within a PVC.
3.6. Training Data
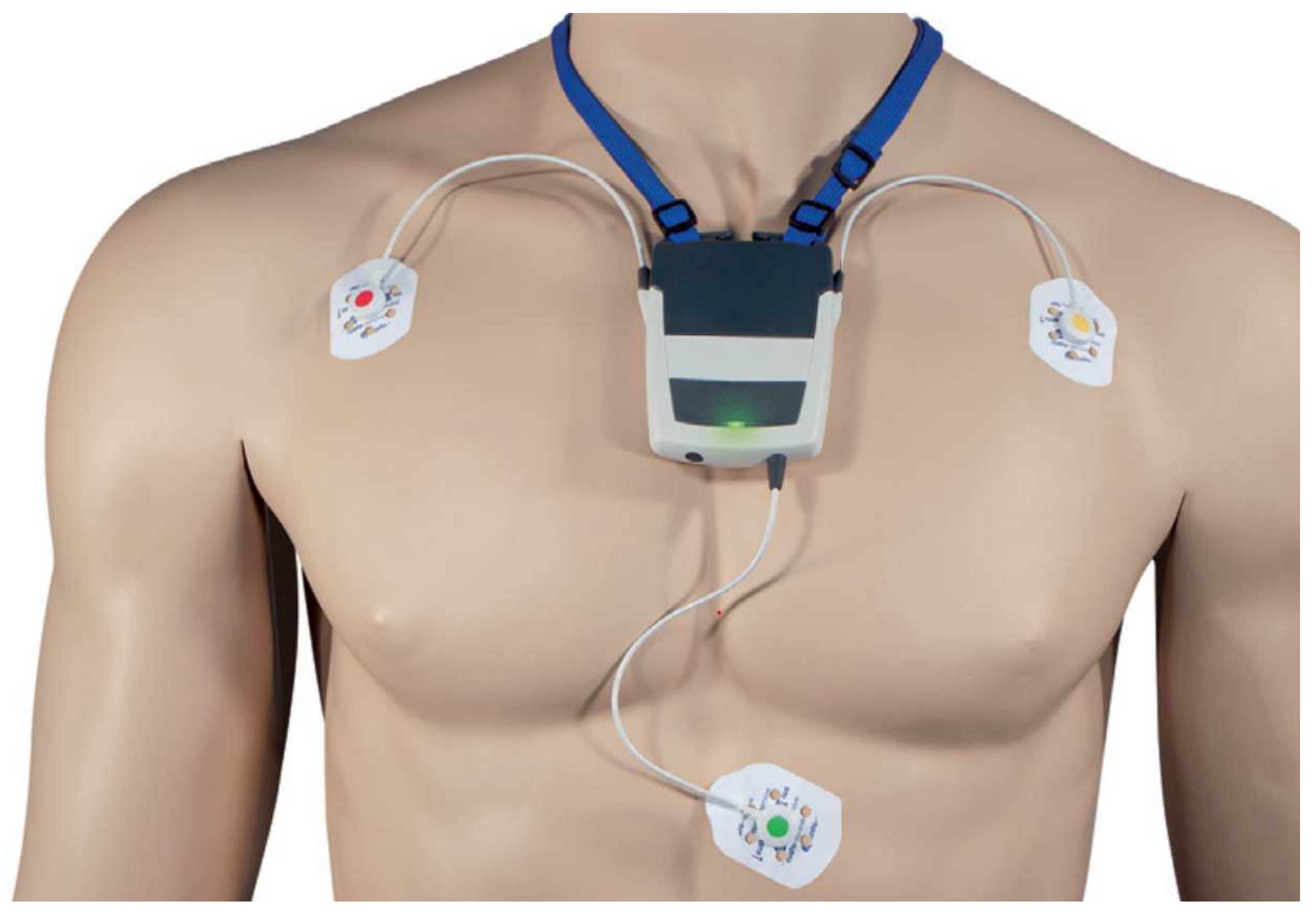
3.6.1. Custo Med Training Dataset
3.6.2. Icentia11k
3.6.3. St. Petersburg INCART 12-lead Arrhythmia Database
3.7. Test data
3.7.1. AHA
3.7.2. NST
3.7.3. MIT
3.7.4. Custo Med Test Dataset
4. Results
4.1. Evaluation Method
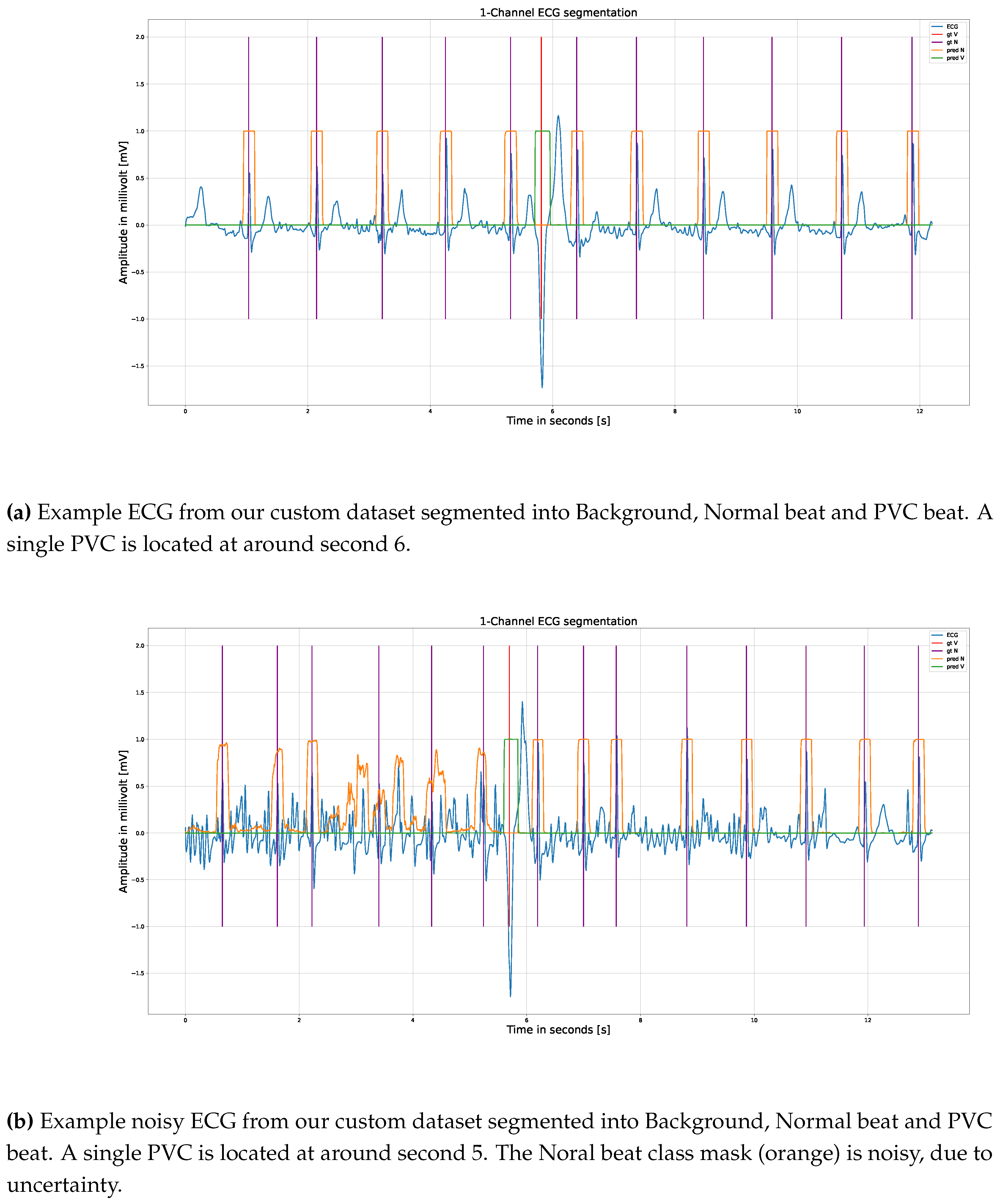
4.2. Model Output
4.3. Productive usage
5. Model Interpretability
6. Discussion
7. Limitations
- Pruning: Our model did not incorporate pruning techniques during the training process. Pruning is a common strategy to reduce the complexity and size of deep learning models, improving computational efficiency and potentially reducing overfitting. Future studies might explore the impacts of various pruning techniques on model performance and efficiency.
- Absence of Attention Mechanism: The model did not leverage any attention mechanism. Attention models have emerged as powerful tools in deep learning, enabling the model to focus on the most relevant parts of the input for a given task. Incorporating attention mechanisms could improve the model’s performance, especially in tasks where certain parts of the input carry more informative content.
- Lack of Self-supervised Pretraining: Our study did not exploit self-supervised pretraining using multiple data sets. This approach could potentially improve the robustness and generalizability of the model by exposing it to a wider range of data during pretraining.
- Limited Classification: The scope of our model was confined to the detection of normal beats and Premature Ventricular Contractions (PVC). Although this focus has its own merits, the model’s utility could be enhanced by expanding its classification capabilities to detect other types of cardiac events.
- Size of the Test Data sets: Our test data sets were not particularly large. Larger test data sets would provide a more robust estimation of the model’s performance and its ability to generalize to unseen data.
- Single Channel Model: Our model was designed to work with single-channel ECG signals. While this design decision simplifies the model and its input requirements, it might limit the model’s ability to detect cardiac events that are better characterized using multichannel ECG signals. Future research could investigate the benefits of a multi-channel approach.
8. Conclusion
Author Contributions
Funding
Conflicts of Interest
References
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015; Navab, N.; Hornegger, J.; Wells, W.M.; Frangi, A.F., Eds.; Springer International Publishing: Cham, 2015; pp. 234–241.
- Kiranyaz, S.; Ince, T.; Gabbouj, M. Real-time patient-specific ECG classification by 1-D convolutional neural networks. IEEE Transactions on Biomedical Engineering 2015, 63, 664–675. [CrossRef]
- Acharya, U.R.; Fujita, H.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adam, M. Application of deep convolutional neural network for automated detection of myocardial infarction using ECG signals. Information Sciences 2017, 415, 190–198. [CrossRef]
- Hannun, A.Y.; Rajpurkar, P.; Haghpanahi, M.; Tison, G.H.; Bourn, C.; Turakhia, M.P.; Ng, A.Y. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nature medicine 2019, 25, 65–69. [CrossRef]
- Izci, E.; Ozdemir, M.A.; Degirmenci, M.; Akan, A. Cardiac arrhythmia detection from 2d ecg images by using deep learning technique. In Proceedings of the 2019 medical technologies congress (TIPTEKNO). IEEE, 2019, pp. 1–4. [CrossRef]
- Ullah, A.; Anwar, S.M.; Bilal, M.; Mehmood, R.M. Classification of arrhythmia by using deep learning with 2-D ECG spectral image representation. Remote Sensing 2020, 12, 1685. [CrossRef]
- Mazidi, M.H.; Eshghi, M.; Raoufy, M.R. Detection of premature ventricular contraction (PVC) using linear and nonlinear techniques: an experimental study. Cluster Computing 2020, 23, 759–774. [CrossRef]
- Rijnbeek, P.R.; van Herpen, G.; Bots, M.L.; Man, S.C.; Verweij, N.; Hofman, A.; Hillege, H.; Numans, M.E.; Swenne, C.A.; Witteman, J.C.; et al. Normal values of the electrocardiogram for ages 16-90 years. Journal of electrocardiology 2014, 47, 914–921. [CrossRef]
- Linde, C.; Bongiorni, M.G.; Birgersdotter-Green, U.; Curtis, A.B.; Deisenhofer, I.; Furokawa, T.; Gillis, A.M.; Haugaa, K.H.; Lip, G.Y.; Van Gelder, I.; et al. Sex differences in cardiac arrhythmia: a consensus document of the European Heart Rhythm Association, endorsed by the Heart Rhythm Society and Asia Pacific Heart Rhythm Society. Ep Europace 2018, 20, 1565–1565ao. [CrossRef]
- Zhou, F.y.; Jin, L.p.; Dong, J. Premature ventricular contraction detection combining deep neural networks and rules inference. Artificial intelligence in medicine 2017, 79, 42–51. [CrossRef]
- Teplitzky, B.A.; McRoberts, M.; Ghanbari, H. Deep learning for comprehensive ECG annotation. Heart Rhythm 2020, 17, 881–888. Digital Health Special Issue. [CrossRef]
- Pan, J.; Tompkins, W.J. A real-time QRS detection algorithm. IEEE transactions on biomedical engineering 1985, pp. 230–236.
- Christov, I.I. Real time electrocardiogram QRS detection using combined adaptive threshold. Biomedical engineering online 2004, 3, 1–9. [CrossRef]
- Chiarugi, F.; Sakkalis, V.; Emmanouilidou, D.; Krontiris, T.; Varanini, M.; Tollis, I. Adaptive threshold QRS detector with best channel selection based on a noise rating system. In Proceedings of the 2007 Computers in Cardiology. IEEE, 2007, pp. 157–160.
- Chouakri, S.; Bereksi-Reguig, F.; Taleb-Ahmed, A. QRS complex detection based on multi wavelet packet decomposition. Applied Mathematics and Computation 2011, 217, 9508–9525. [CrossRef]
- Elgendi, M. Fast QRS detection with an optimized knowledge-based method: Evaluation on 11 standard ECG databases. PloS one 2013, 8, e73557. [CrossRef]
- Liu, Y.; Li, Q.; He, R.; Wang, K.; Liu, J.; Yuan, Y.; Xia, Y.; Zhang, H. Generalizable beat-by-beat arrhythmia detection by using weakly supervised deep learning. Frontiers in Physiology 2022, 13, 850951. [CrossRef]
- He, R.; Liu, Y.; Wang, K.; Zhao, N.; Yuan, Y.; Li, Q.; Zhang, H. Automatic Detection of QRS Complexes Using Dual Channels Based on U-Net and Bidirectional Long Short-Term Memory. IEEE Journal of Biomedical and Health Informatics 2021, 25, 1052–1061. [CrossRef]
- Martínez, J.P.; Almeida, R.; Olmos, S.; Rocha, A.P.; Laguna, P. A wavelet-based ECG delineator: evaluation on standard databases. IEEE Transactions on biomedical engineering 2004, 51, 570–581. [CrossRef]
- Arzeno, N.M.; Deng, Z.D.; Poon, C.S. Analysis of First-Derivative Based QRS Detection Algorithms. IEEE Transactions on Biomedical Engineering 2008, 55, 478–484. [CrossRef]
- Zidelmal, Z.; Amirou, A.; Adnane, M.; Belouchrani, A. QRS detection based on wavelet coefficients. Computer methods and programs in biomedicine 2012, 107, 490–496.
- De Chazal, P.; O’Dwyer, M.; Reilly, R.B. Automatic classification of heartbeats using ECG morphology and heartbeat interval features. IEEE transactions on biomedical engineering 2004, 51, 1196–1206.
- Jiang, W.; Kong, S.G. Block-based neural networks for personalized ECG signal classification. IEEE Transactions on Neural Networks 2007, 18, 1750–1761.
- Ince, T.; Kiranyaz, S.; Gabbouj, M. A generic and robust system for automated patient-specific classification of ECG signals. IEEE Transactions on Biomedical Engineering 2009, 56, 1415–1426. [CrossRef]
- Zhang, C.; Wang, G.; Zhao, J.; Gao, P.; Lin, J.; Yang, H. Patient-specific ECG classification based on recurrent neural networks and clustering technique. In Proceedings of the 2017 13th IASTED International Conference on Biomedical Engineering (BioMed). IEEE, 2017, pp. 63–67.
- Sudre, C.H.; Li, W.; Vercauteren, T.; Ourselin, S.; Cardoso, M.J. Generalised Dice Overlap as a Deep Learning Loss Function for Highly Unbalanced Segmentations. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer International Publishing, 2017; pp. 240–248. [CrossRef]
- Lin, T.Y.; Goyal, P.; Girshick, R.; He, K.; Dollár, P. Focal Loss for Dense Object Detection. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), 2017, pp. 2999–3007. [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization, 2017, [arXiv:cs.LG/1412.6980].
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet. Circulation 2000, 101, e215–e220, http://xxx.lanl.gov/abs/https://www.ahajournals.org/doi/pdf/10.1161/01.CIR.101.23.e215. [CrossRef]
- Tan, S.; Androz, G.; Chamseddine, A.; Fecteau, P.; Courville, A.; Bengio, Y.; Cohen, J.P. Icentia11k: An unsupervised representation learning dataset for arrhythmia subtype discovery. arXiv preprint arXiv:1910.09570 2019.
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the International conference on machine learning. pmlr, 2015, pp. 448–456.
- Zhang, H.; Cisse, M.; Dauphin, Y.N.; Lopez-Paz, D. mixup: Beyond empirical risk minimization. arXiv preprint arXiv:1710.09412 2017.
- Yun, S.; Han, D.; Oh, S.J.; Chun, S.; Choe, J.; Yoo, Y. Cutmix: Regularization strategy to train strong classifiers with localizable features. In Proceedings of the Proceedings of the IEEE/CVF international conference on computer vision, 2019, pp. 6023–6032.
- Moody, G.B.; Muldrow, W.; Mark, R.G. A noise stress test for arrhythmia detectors. Computers in cardiology 1984, 11, 381–384.
- Vijayarangan, S.; R., V.; Murugesan, B.; S.P., P.; Joseph, J.; Sivaprakasam, M. RPnet: A Deep Learning approach for robust R Peak detection in noisy ECG. In Proceedings of the 2020 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), 2020, pp. 345–348. [CrossRef]
- Moody, G.; Mark, R. The impact of the MIT-BIH Arrhythmia Database. IEEE Engineering in Medicine and Biology Magazine 2001, 20, 45–50. [CrossRef]
- Wang, S.; Hou, X.; Zhao, X. Automatic Building Extraction From High-Resolution Aerial Imagery via Fully Convolutional Encoder-Decoder Network With Non-Local Block. IEEE Access 2020, 8, 7313–7322. [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. International Journal of Computer Vision 2019, 128, 336–359. [CrossRef]
| Symbol | Beat Description | Count |
|---|---|---|
| N | Normal | 3,361,174 |
| V | Premature ventricular contraction | 163,592 |
| Symbol | Beat Description | Count |
|---|---|---|
| N | Normal | 2,061,141,216 |
| S | Premature or ectopic supraventricular beat | 19,346,728 |
| V | Premature ventricular contraction | 17,203,041 |
| Q | Undefined: Unclassifiable beat | 676,364,002 |
| Diagnosis | Patients |
|---|---|
| Acute MI | 2 |
| Transient ischemic attack (angina pectoris) | 5 |
| Prior MI | 4 |
| Coronary artery disease with hypertension | 7 |
| Sinus node dysfunction | 1 |
| Supraventricular ectopy | 18 |
| Atrial fibrillation or SVTA | 3 (2 with paroxysmal AF) |
| WPW | 2 |
| AV block | 1 |
| Bundle branch block | 3 |
| Symbol | Beat Description | Count |
|---|---|---|
| N | Normal | 174,260 |
| V | Premature ventricular contraction | 16,296 |
| Symbol | Beat Description | Count |
|---|---|---|
| N | Normal | 100,718 |
| V | Premature ventricular contraction | 7,009 |
| Symbol | Beat Description | Count |
|---|---|---|
| N | Normal | 39,133 |
| V | Premature ventricular contraction | 4,576 |
| DB | ||||||
|---|---|---|---|---|---|---|
| MIT DB | 0.997 | 0.999 | 0.926 | 0.966 | 0.998 | 0.946 |
| MIT 11 DB | 0.999 | 0.999 | 0.976 | 0.991 | 0.999 | 0.986 |
| AHA DB | 0.992 | 0.997 | 0.972 | 0.857 | 0.995 | 0.915 |
| NST DB | 0.954 | 0.893 | 0.936 | 0.881 | 0.924 | 0.909 |
| CST DB | 0.983 | 0.999 | 0.950 | 0.973 | 0.991 | 0.962 |
| CSTStrips DB | 0.993 | 0.997 | 0.960 | 0.932 | 0.995 | 0.946 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




