Submitted:
11 September 2023
Posted:
13 September 2023
You are already at the latest version
Abstract
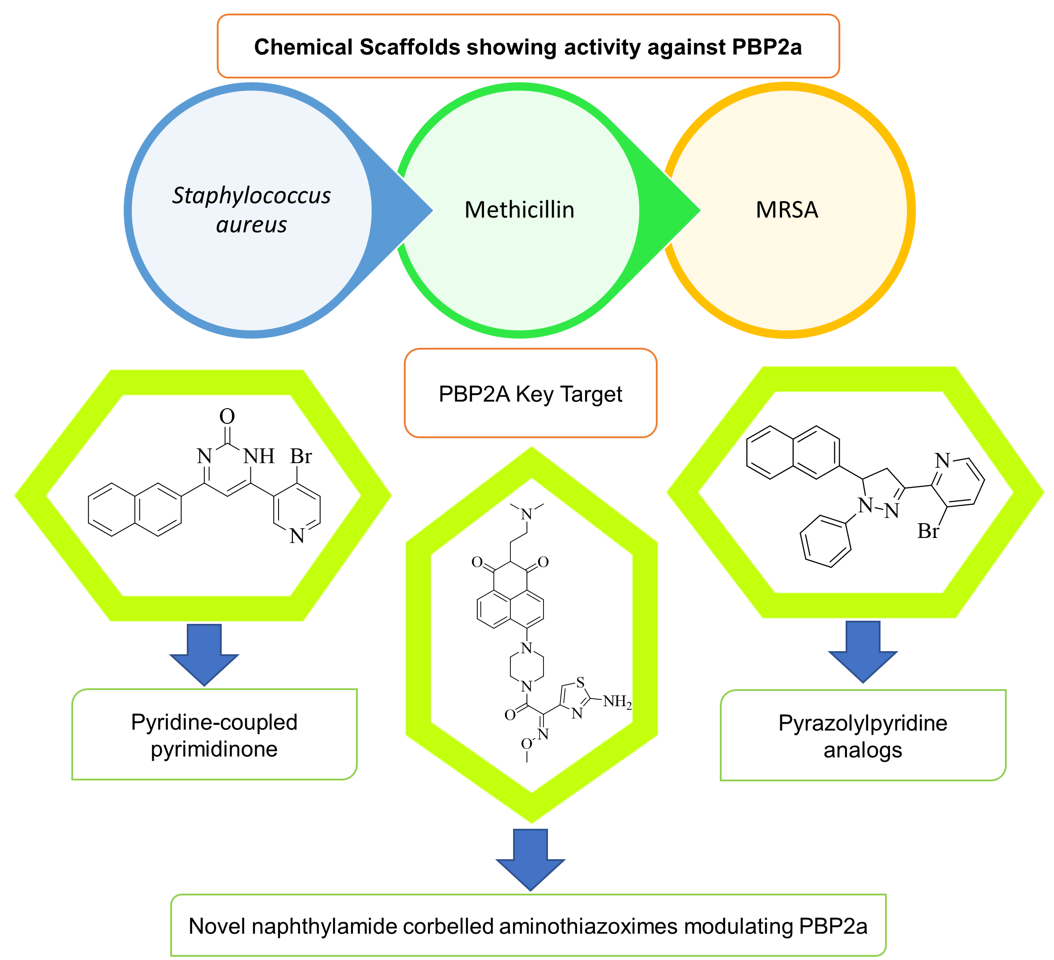
Keywords:
1. Introduction:
1.2. History and Emergence of MRSA:
2. PBPs, Types of PBP, its location and contribution in transpeptidation reaction:
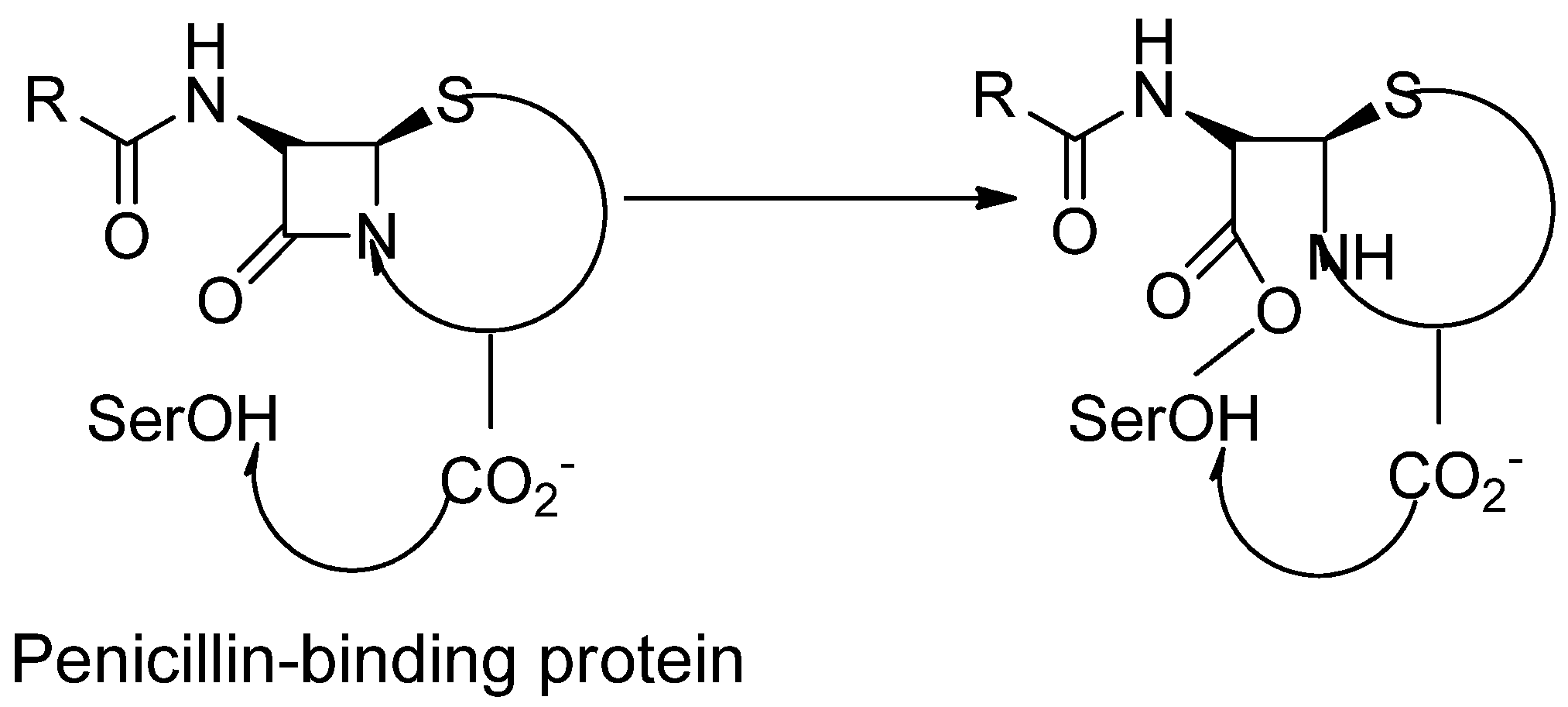
2.1. PBP2a:
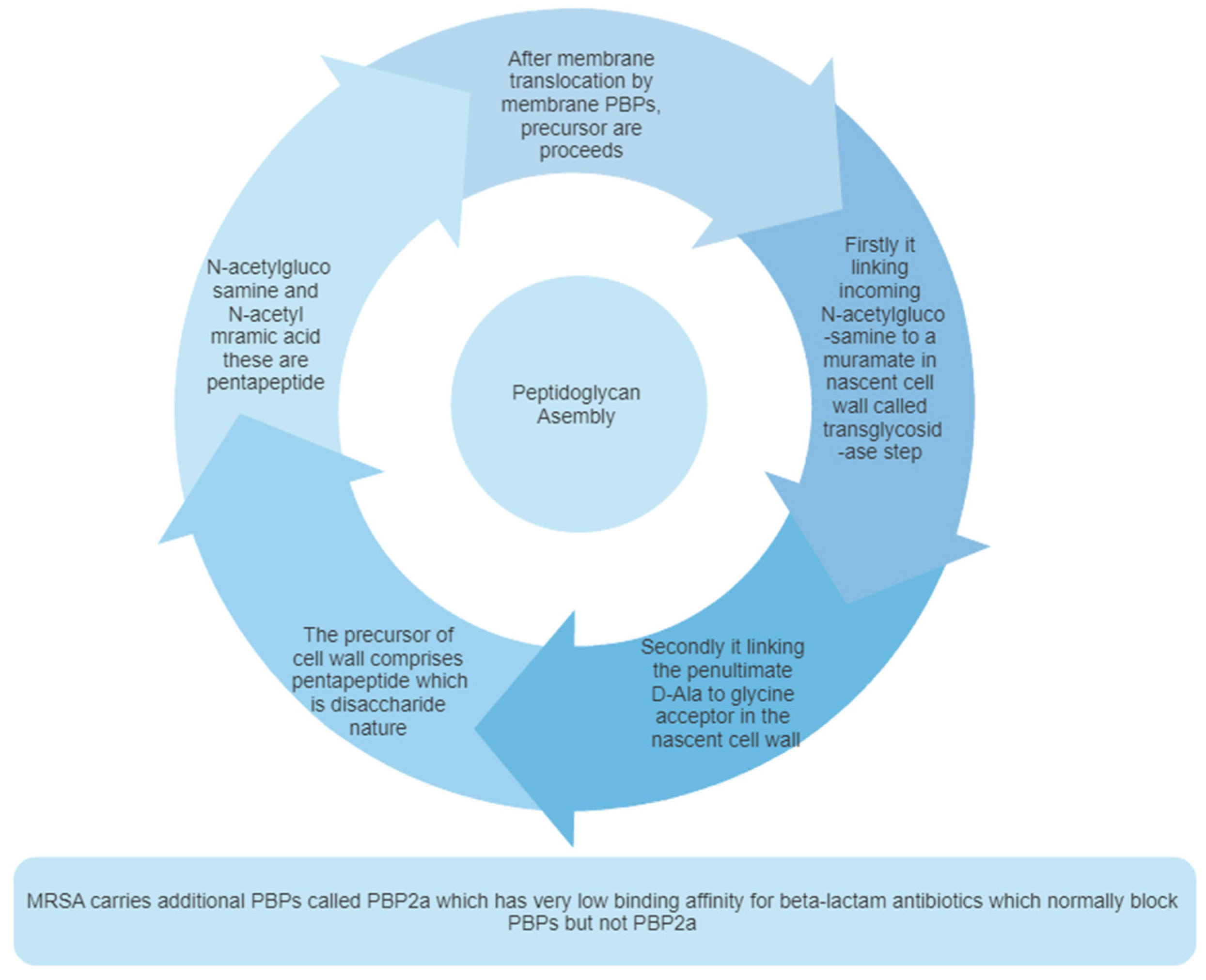
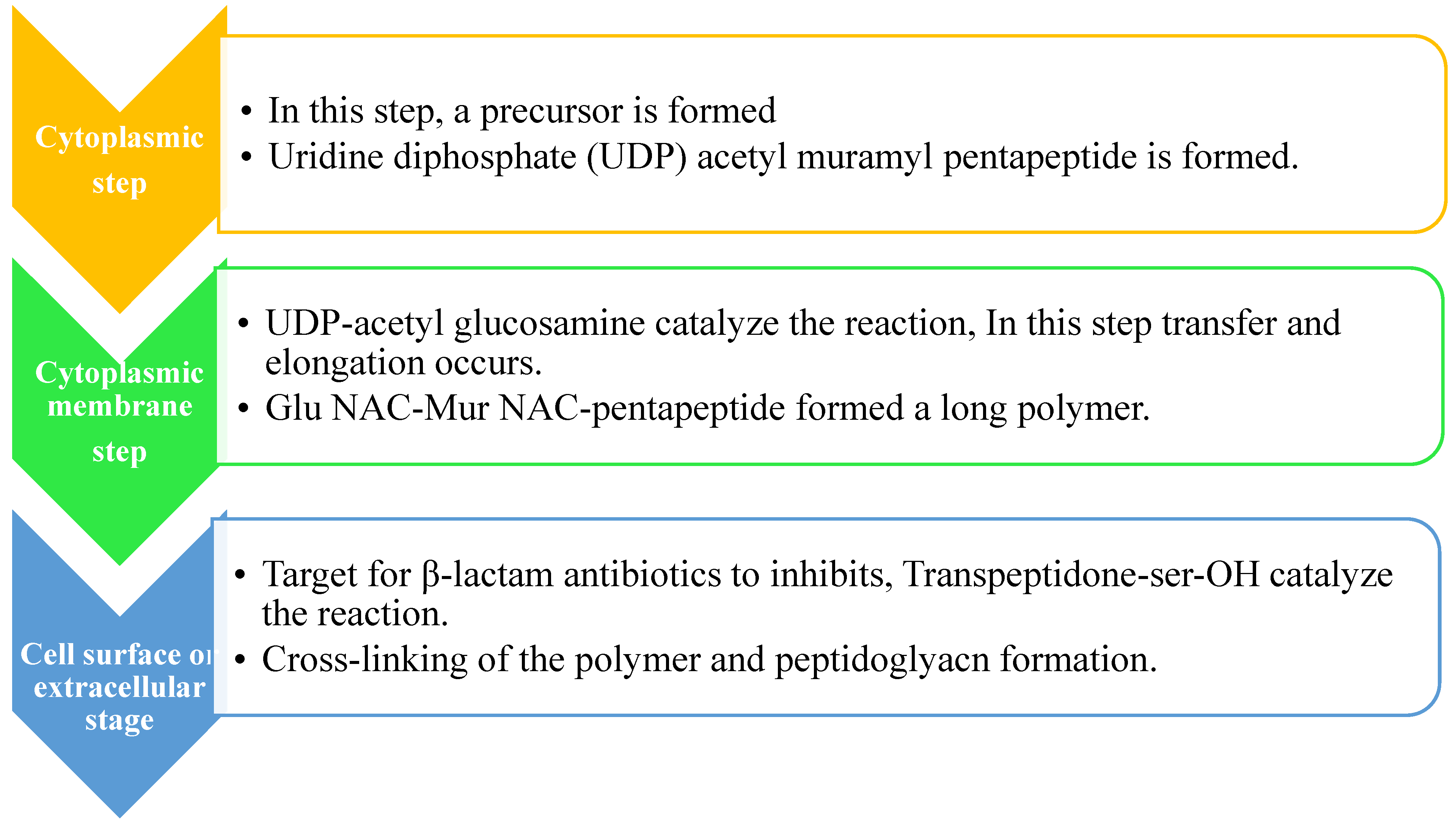
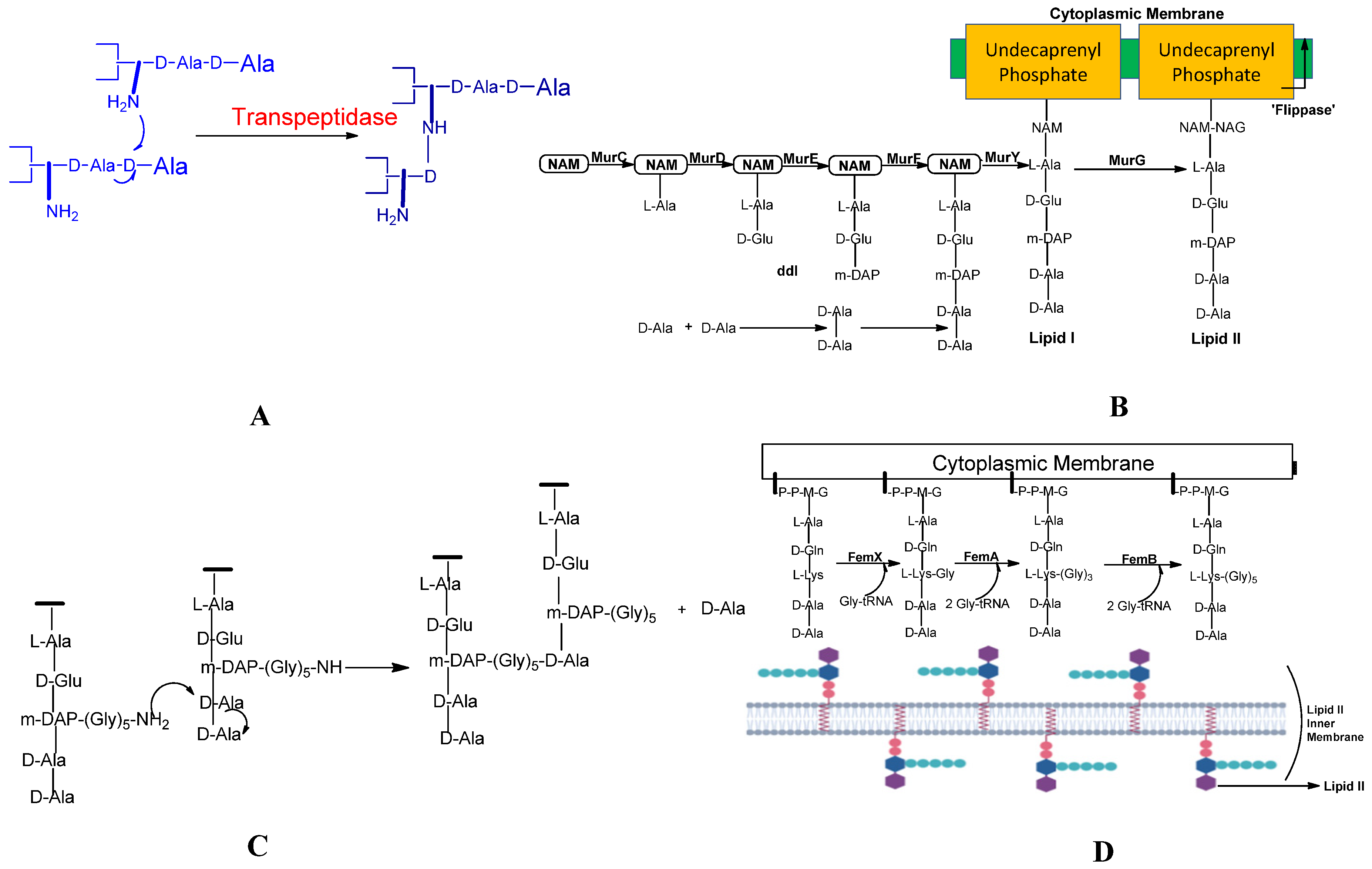
2.2. Peptidoglycan biosynthesis:
| Sr. no | Name of Mur enzymes | Role of enzyme in the biosynthesis of peptidoglycan |
|---|---|---|
| 1 | MurA | The enzyme MurA is responsible for catalysing the intracellular stage of peptidoglycan biosynthesis, which is the first stage in peptidoglycan biosynthesis. MurA enzyme brings such a reaction simply by transferring enol pyruvate. This enol pyruvate is produced by conversion from phosphoenolpyruvate to UDP-N-acetylglucosamine. |
| 2 | MurB | Reduce UDP-GlcNAc-enol pyruvate to UDP-acetylmuramic acid. |
| 3 | The ATP-dependent enzyme, Mur ligases enzymes, Enzymes from MurC to MurF | These enzymes are responsible for linking five amino acid residues. These enzymes relate to the UDP-MurNAc and thus help to form the UDP-MurNAc-pentapeptide. |
| 4 | MraY | Produce lipid I. |
| 5 | MurG | MurG adds GlcNAc moiety to lipid I resulting in the formation of lipid II. |
| 6 | MurJ | To export lipid II to the outer leaflet, MurJ switches between inward and outward conformations. Lipid II reversal is involved by MurJ81. |
2.3. Pentaglycine bridge and transpeptidation reaction in cell wall:
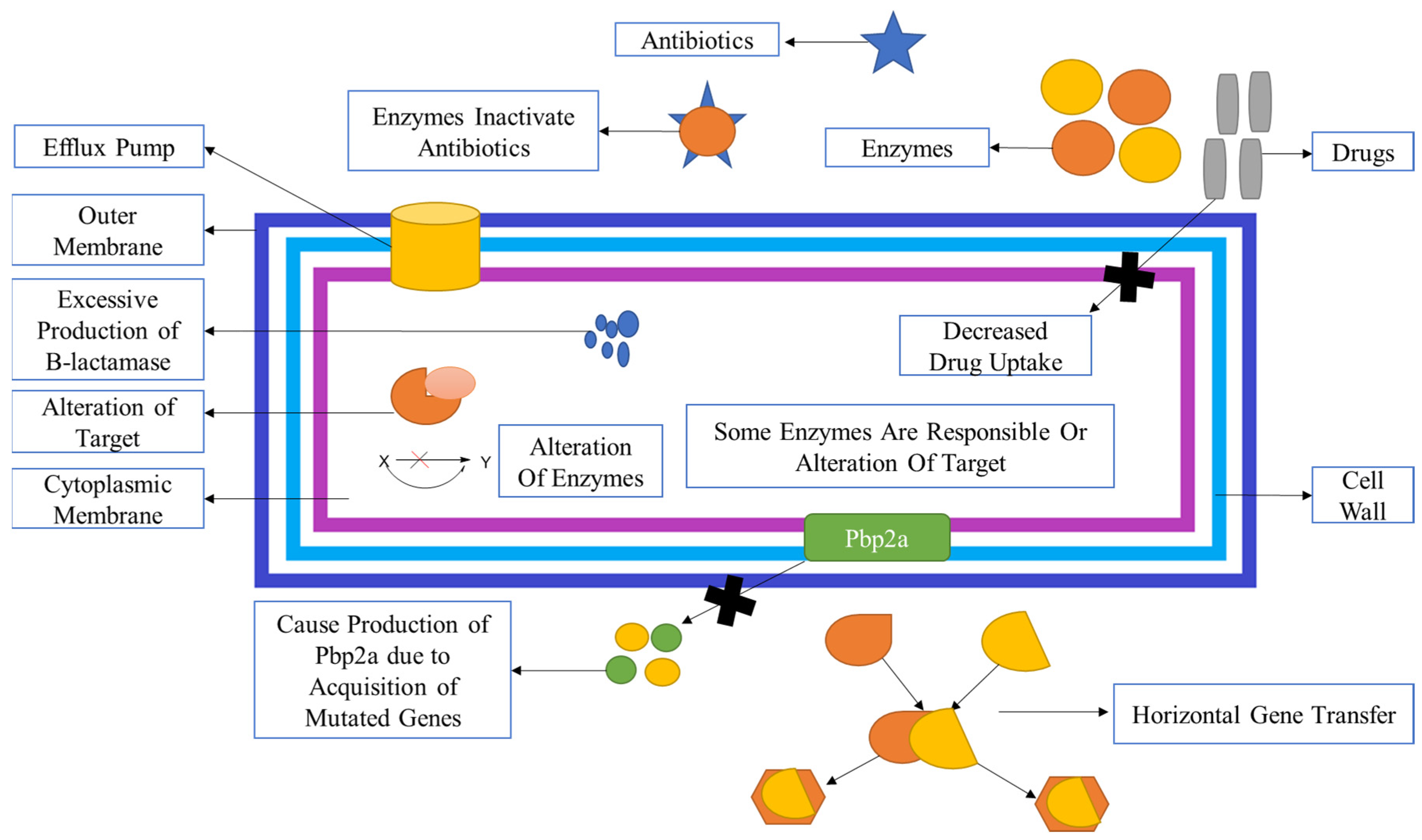
2.4. Mechanism of Methicillin Resistances in S. Aureus:
- 1)
- By formation of PBPs (PBP2a): It is a transpeptidase that mainly causes inhibition resistance by all β-lactam antibiotics and other antibiotics. It is also assisted in maintaining the cross-linking of peptidoglycan. Hence, it allows various MRSA strains for survival, even in the presence of β-lactam antibiotics and other antibiotics.
- 2)
- Production of β-lactamase: It hydrolyses the amide bond, which is essentially present in the four-membered β-lactam ring and causes inactivation of β-lactam antibiotics penicilloic acid before they reach the PBP target95.
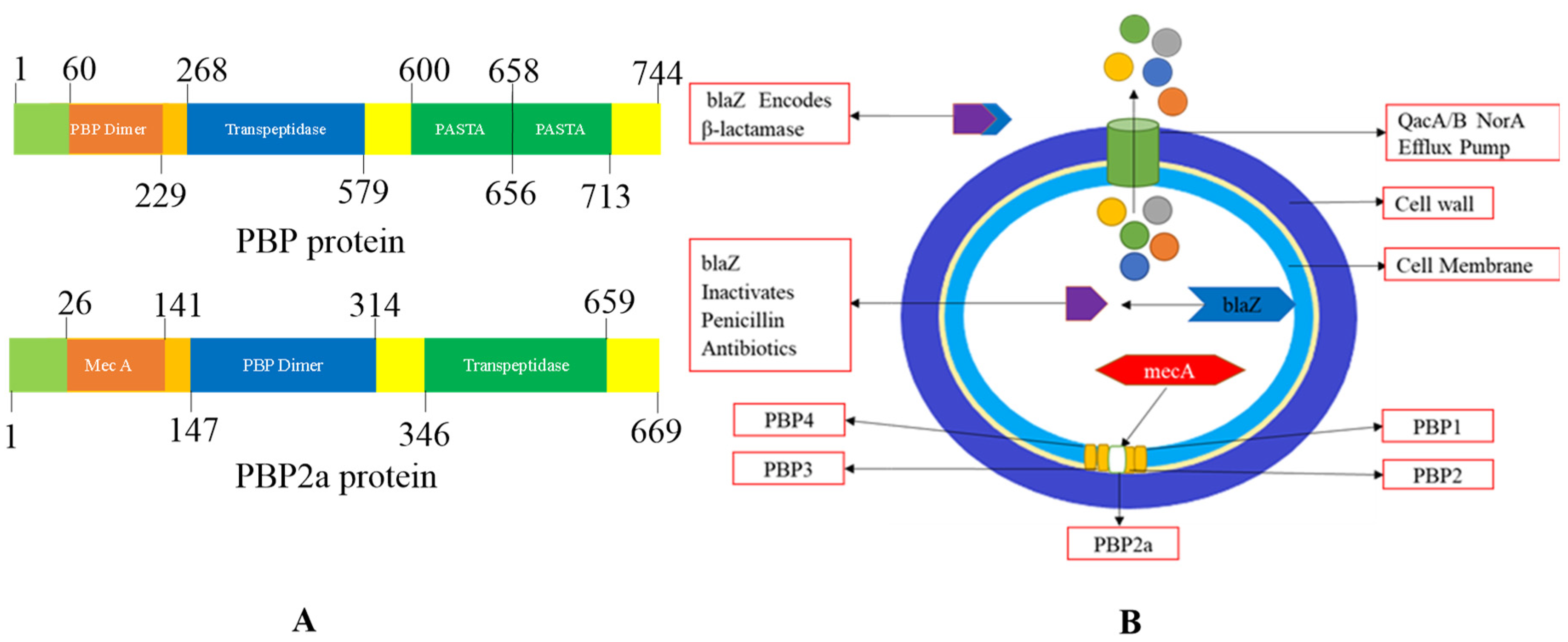
2.5. Staphylococcus Cassette chromosome:
- a)
- The mecA gene is a complex which contains mecA genes coding for a penicillin-binding protein (PBP2) with low affinity for beta-lactam antibiotics.
- b)
- Site-specific recombinase is encoded by the cassette chromosome recombinase (ccr) gene complex112.
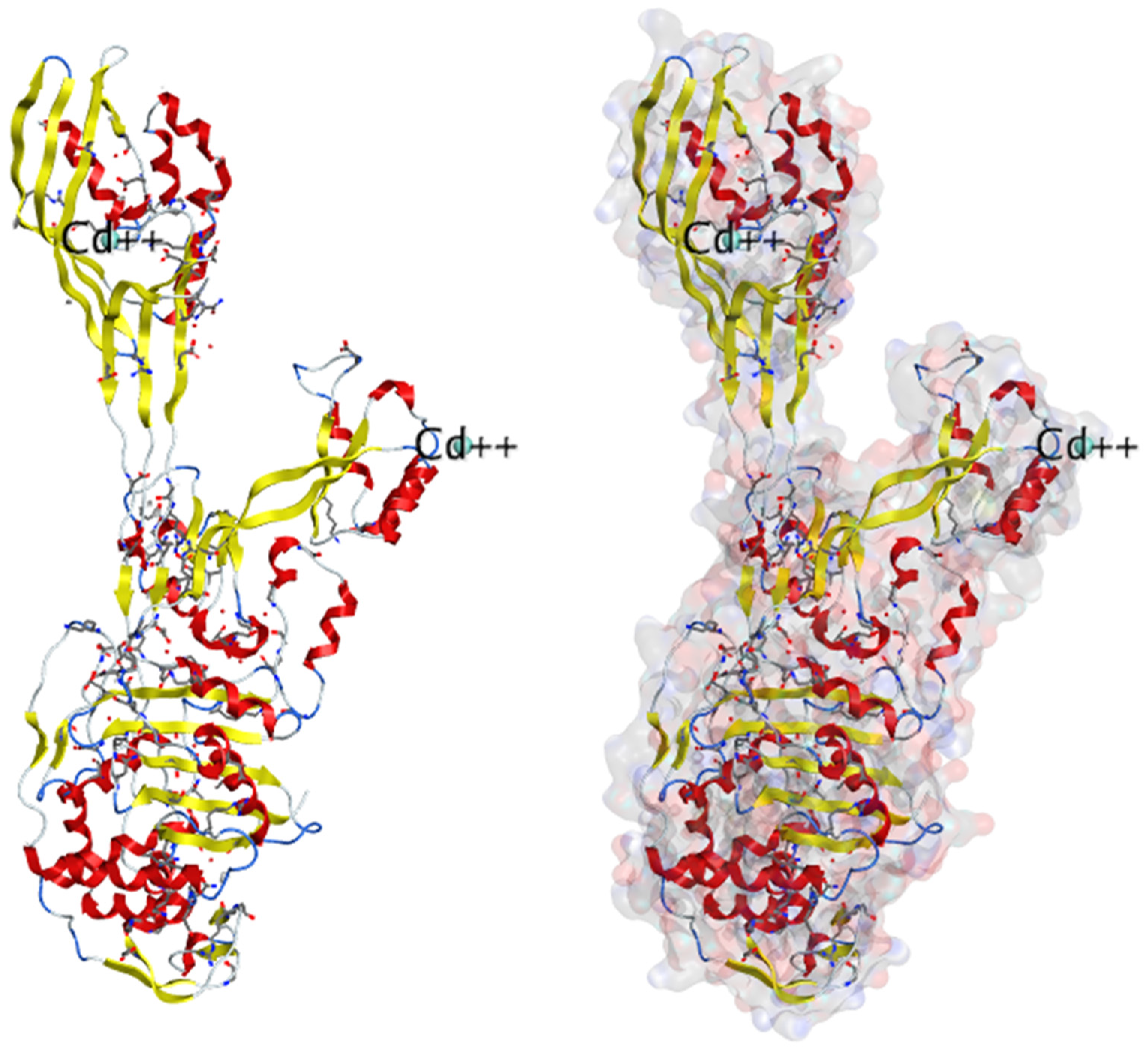
3. Structure basis for PBP2A:
3.2. About Crystal structures for PBP2a:
| Sr. no | Release Year | PDB ID | Resolution | Bound ligand | Species | Ref. |
|---|---|---|---|---|---|---|
| 1 | 2019 | 6Q9N | 2.5 |
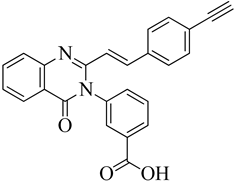 
|
S. aureus subsp. Mu50 | 122 |
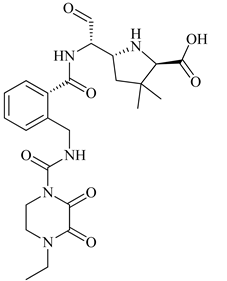
| 2 | 2019 | 6H5O | 2.82 | 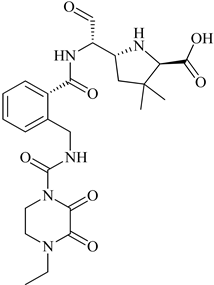 |
S. aureus Subsp.Mu50 | 123 |
| 3 | 2015 | 4CJN | 1.947 | 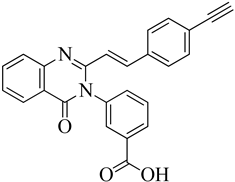 |
S. aureus | 124 |
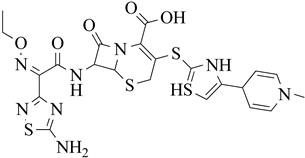
| 4 | 2013 | 3ZG0 | 2.6 |
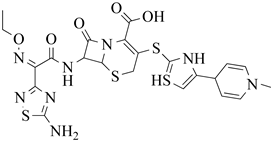 
|
S. aureus Subsp.Mu50 | 125 |
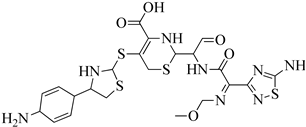
| 5 | 2013 | 3ZFZ | 2.25 |
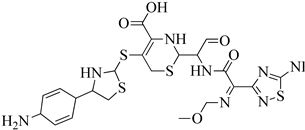 
|
S. aureus subsp. Mu50 | 125 |
| 6 | 2002 | 1MWT | 2.45 | 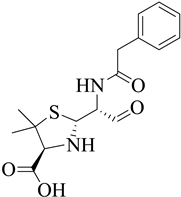 |
S. aureus | 116 |
| 7 | 2002 | 1MWU | 2.60 | 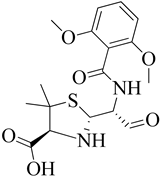 |
S. aureus | 116 |
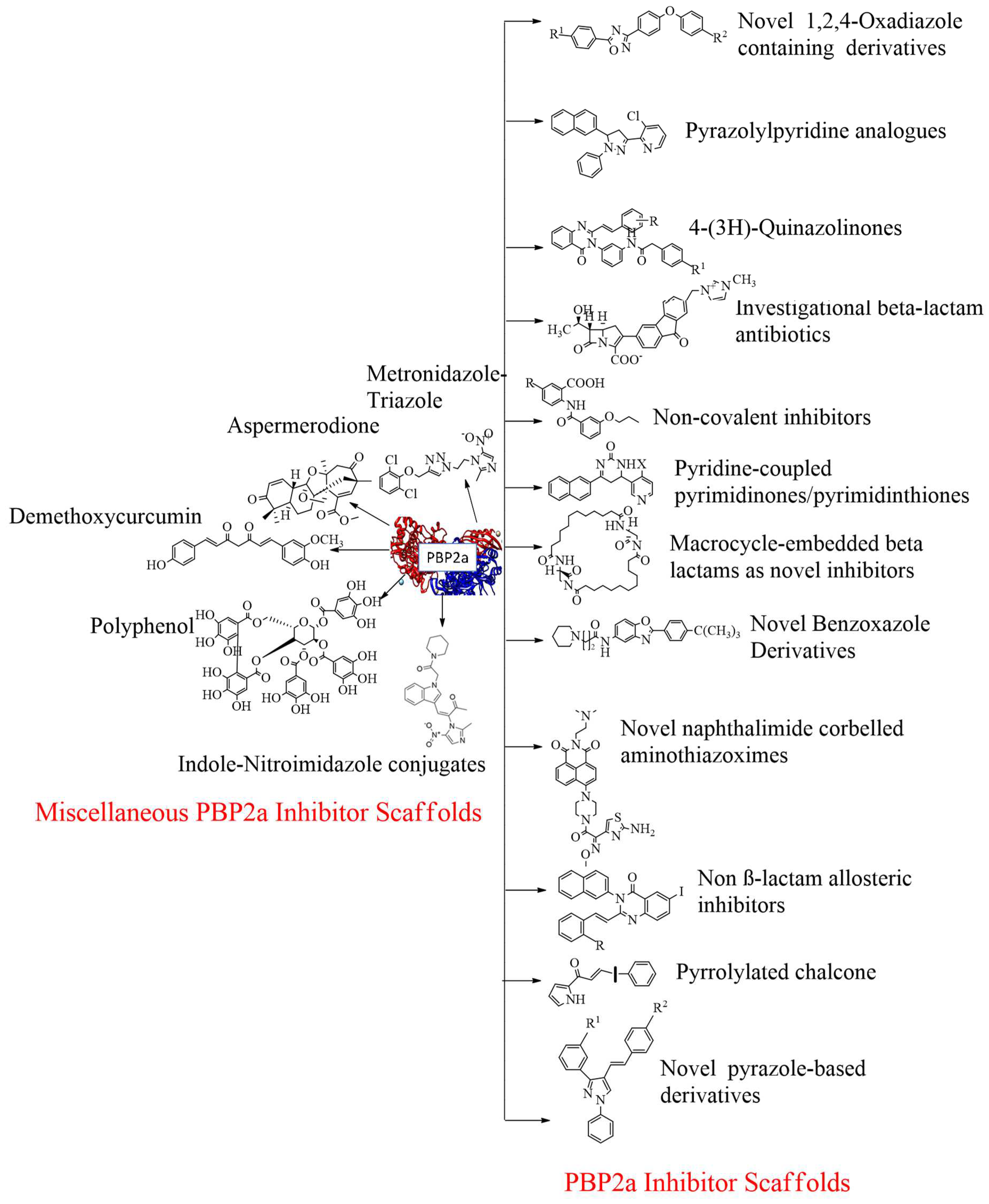
4. Development of PBP2a inhibitors:
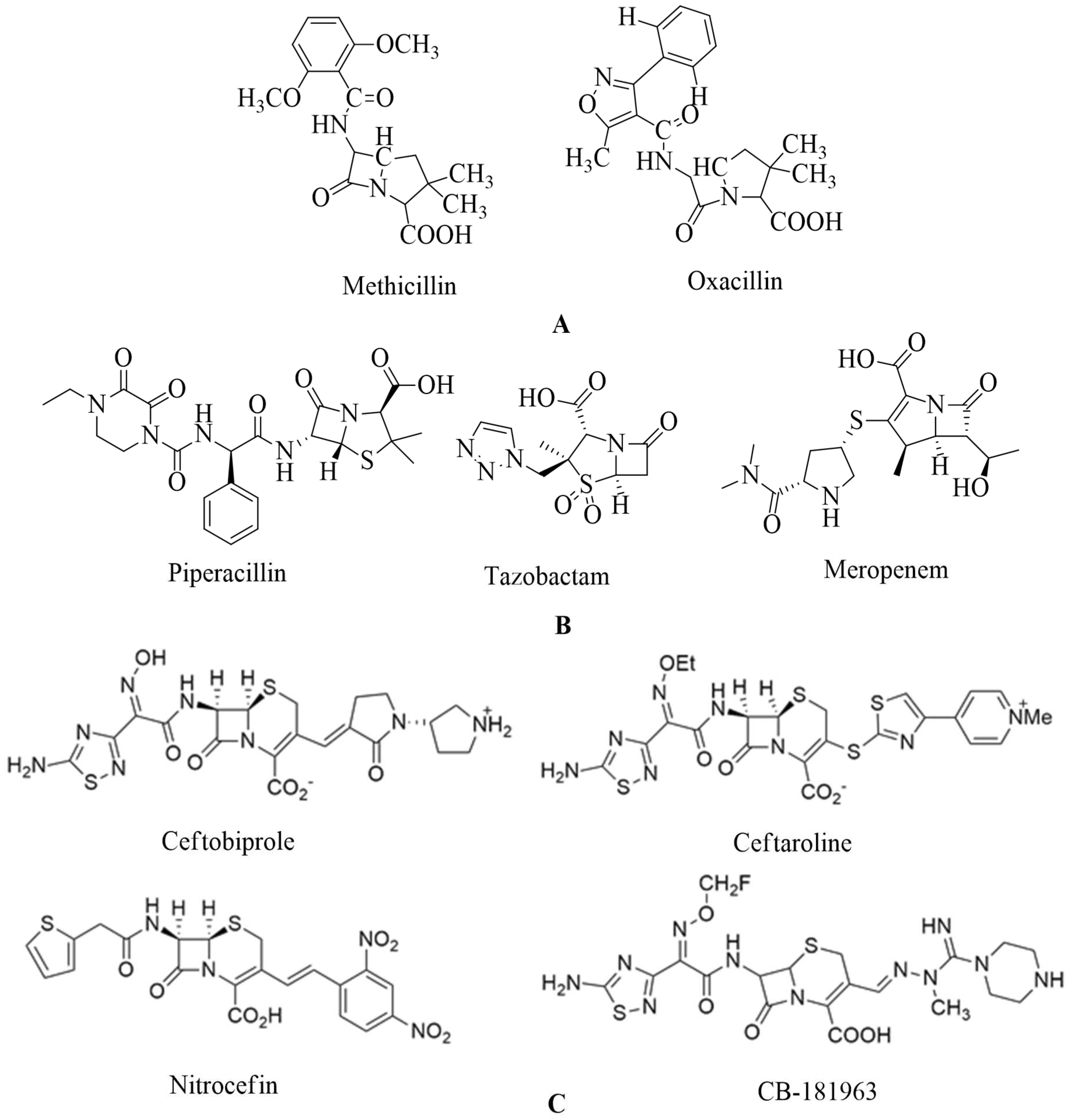
4.1. Penicillin:
4.2. Cephalosporins:
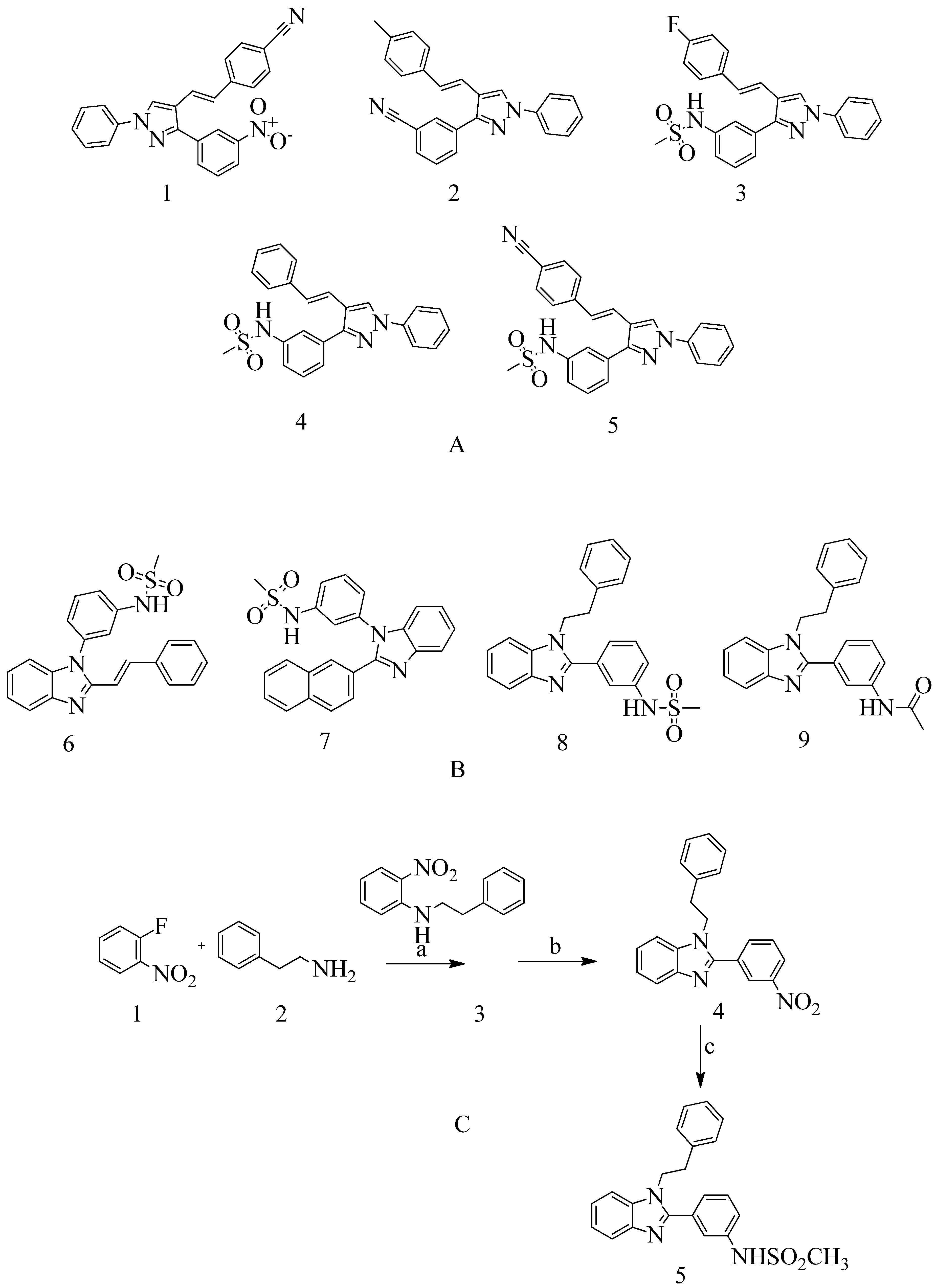
4.3. Novel pyrazole-benzimidazole based derivatives:
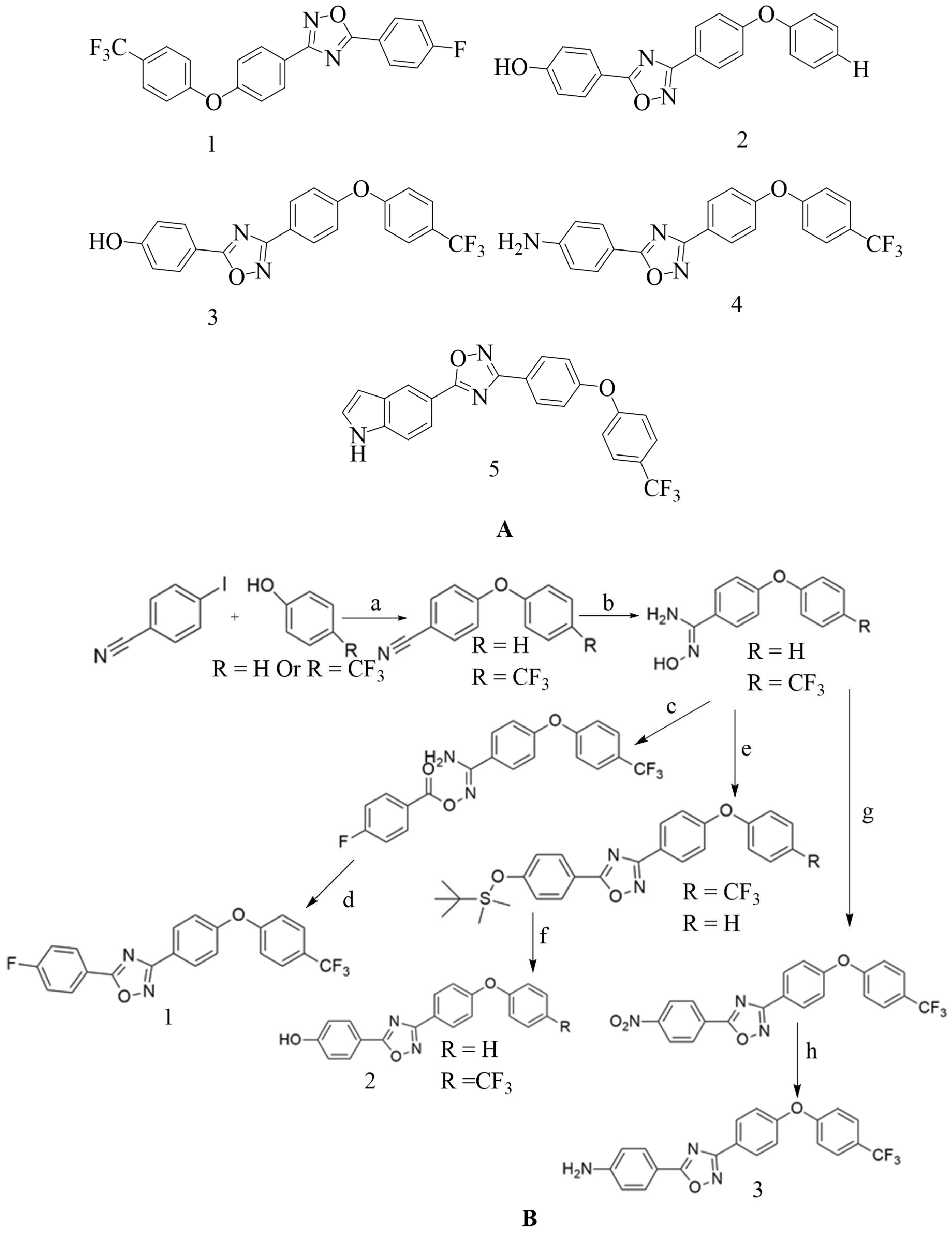
4.4. Novel 1,2,4-oxadiazole-containing derivatives:
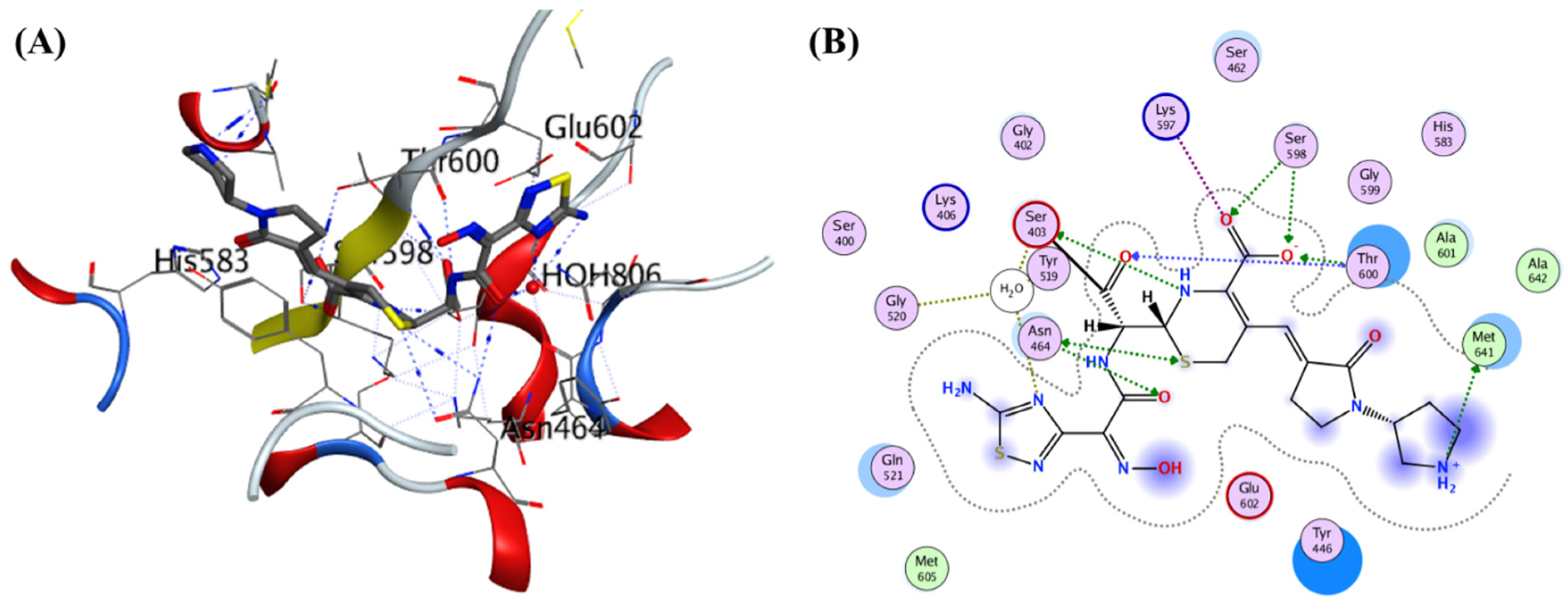
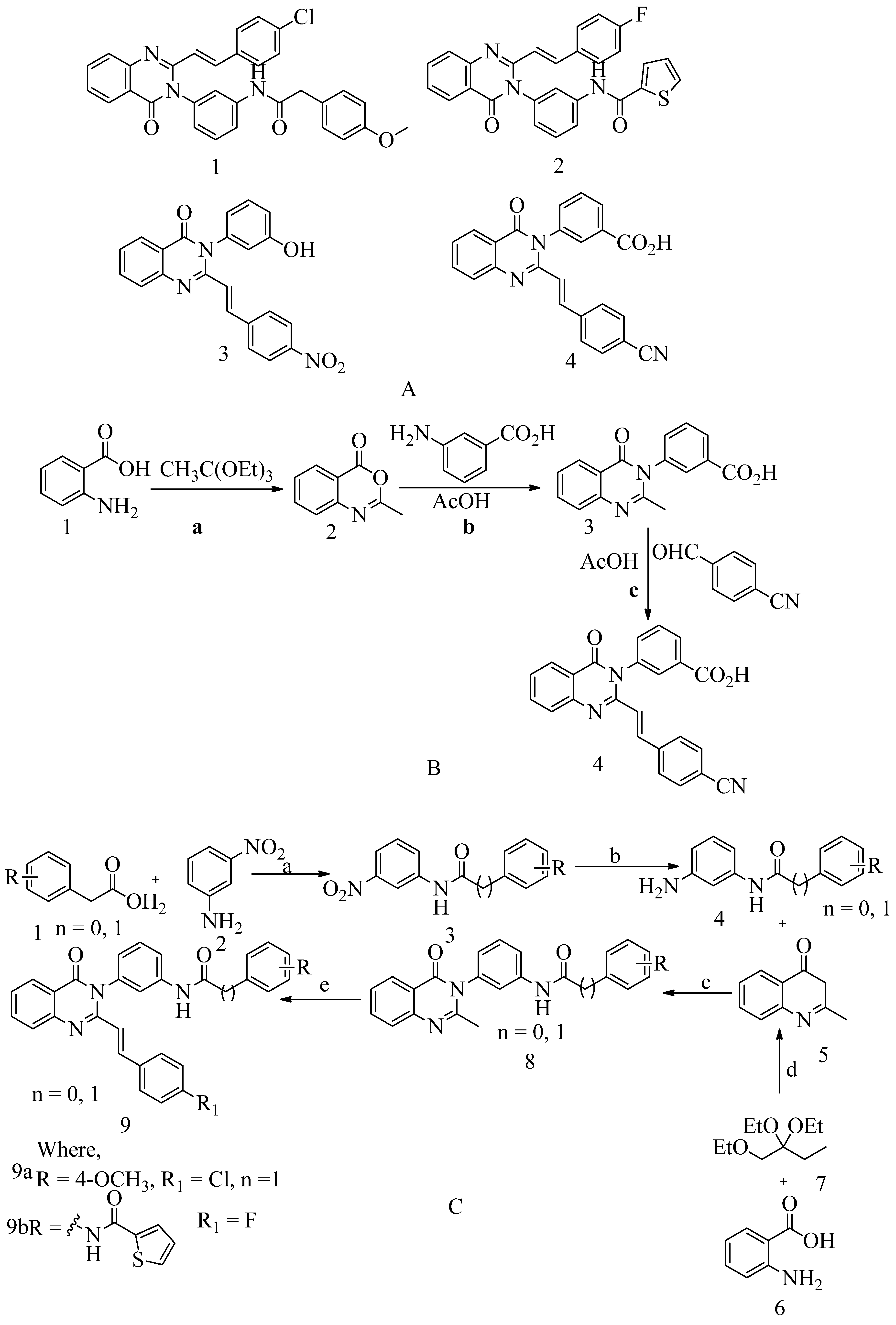
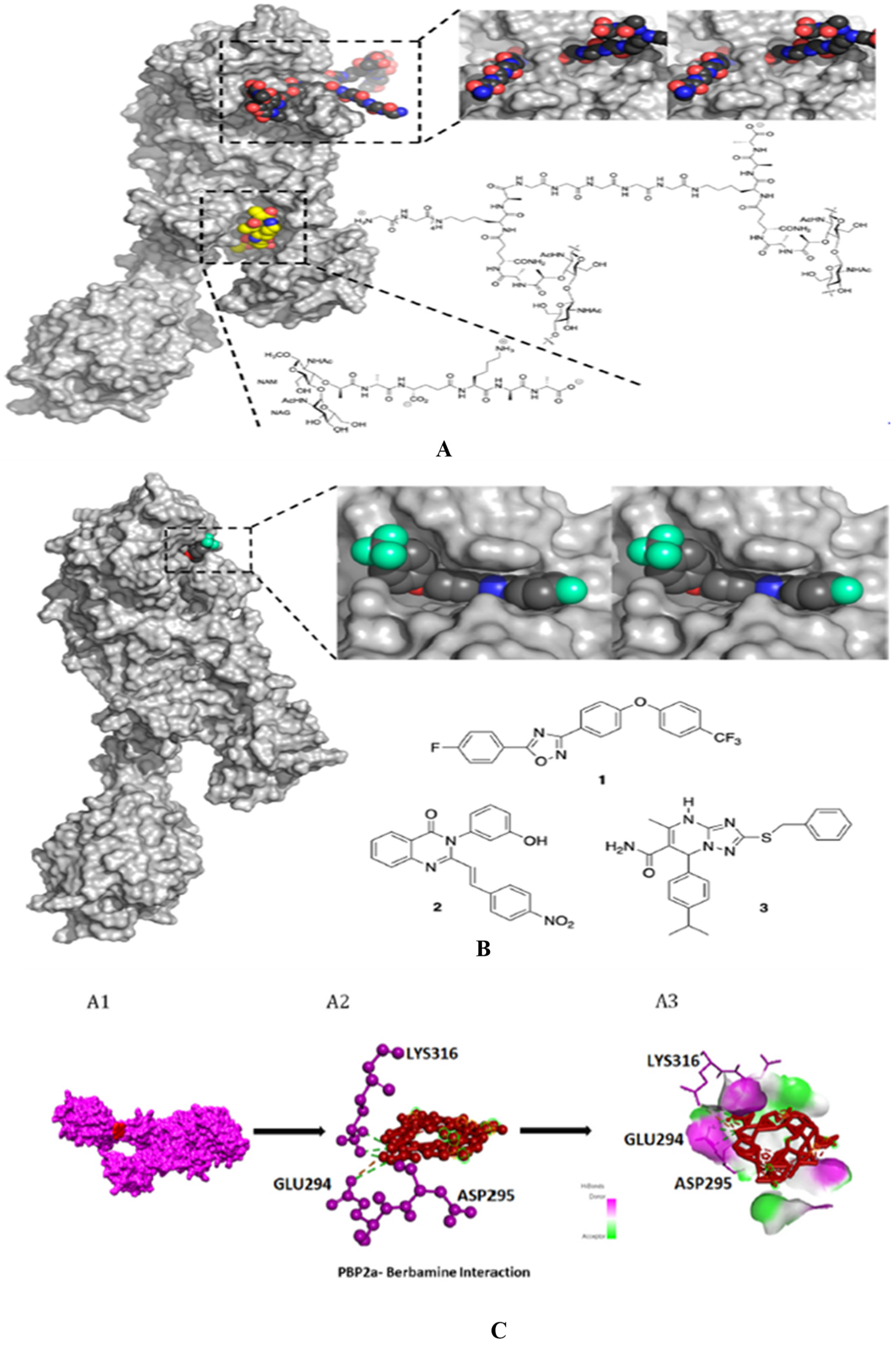
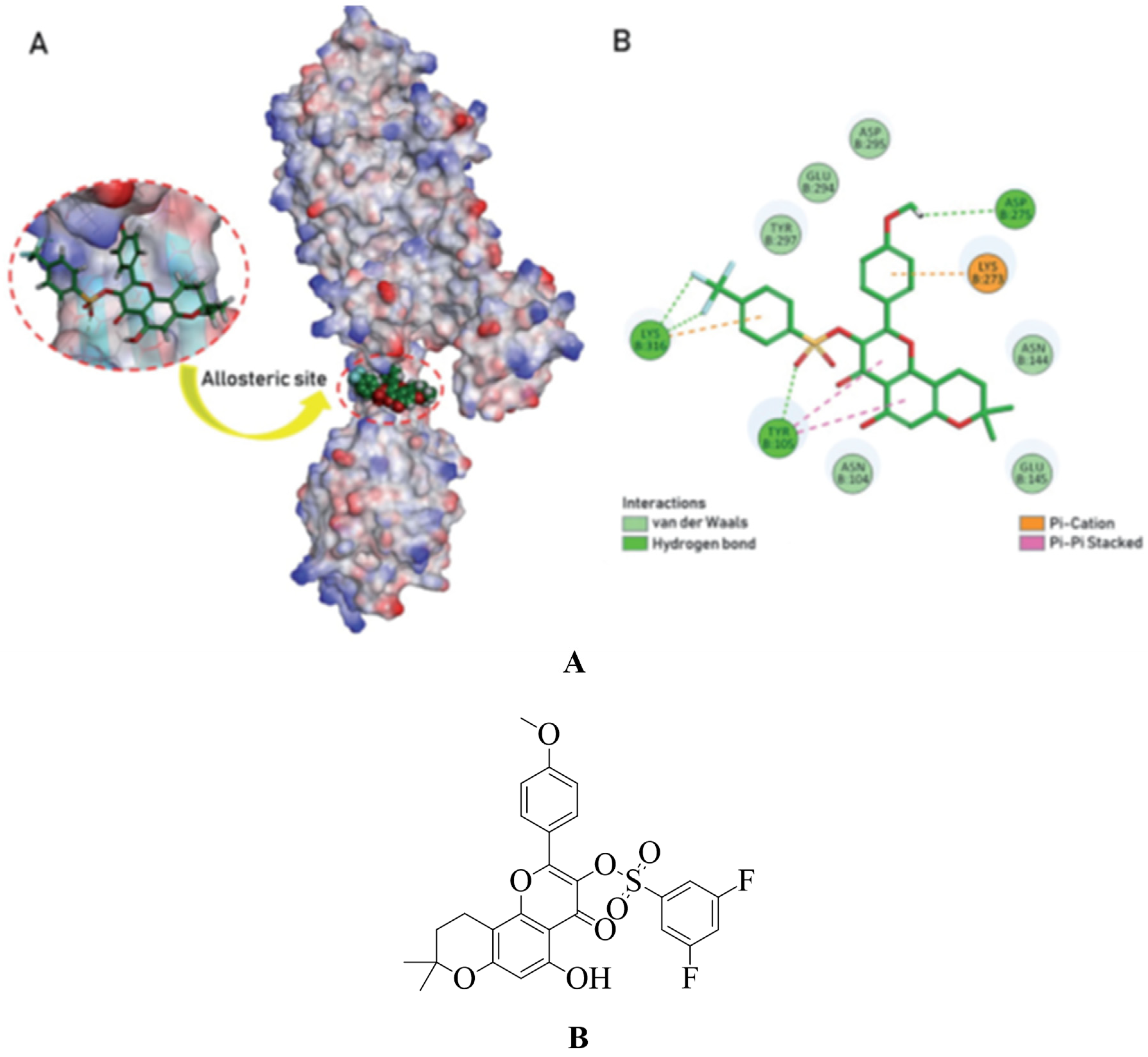
4.5. Non-β-lactam allosteric inhibitors:
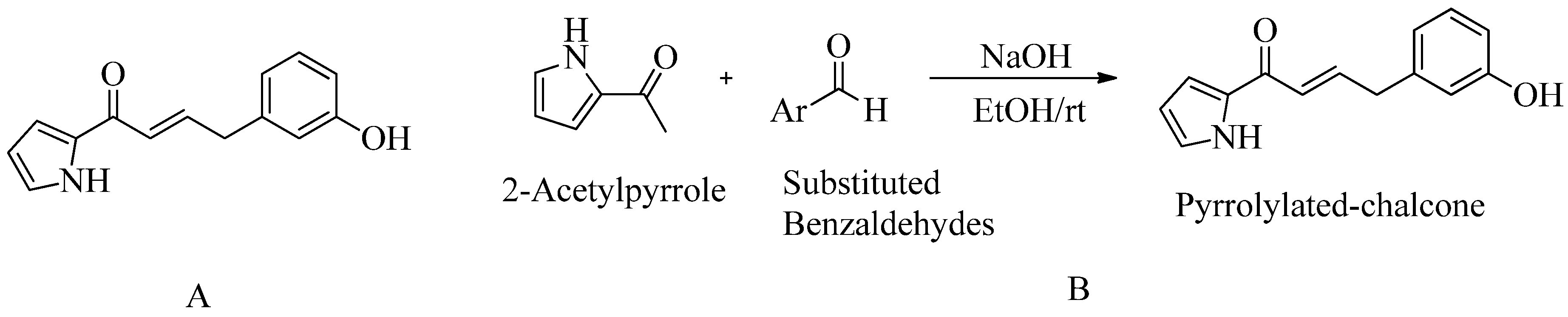
4.7. Pyrrolylated-chalcones:
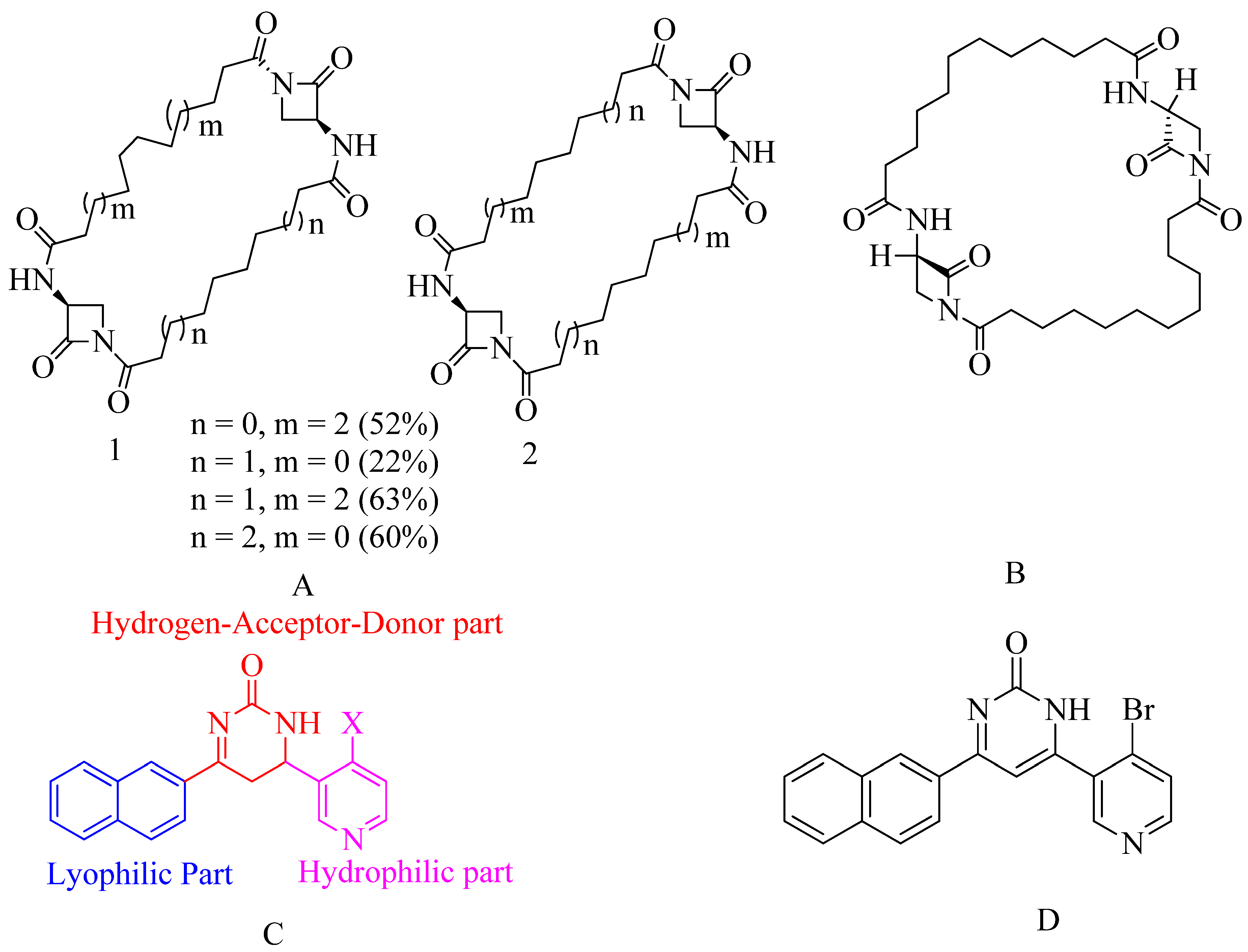
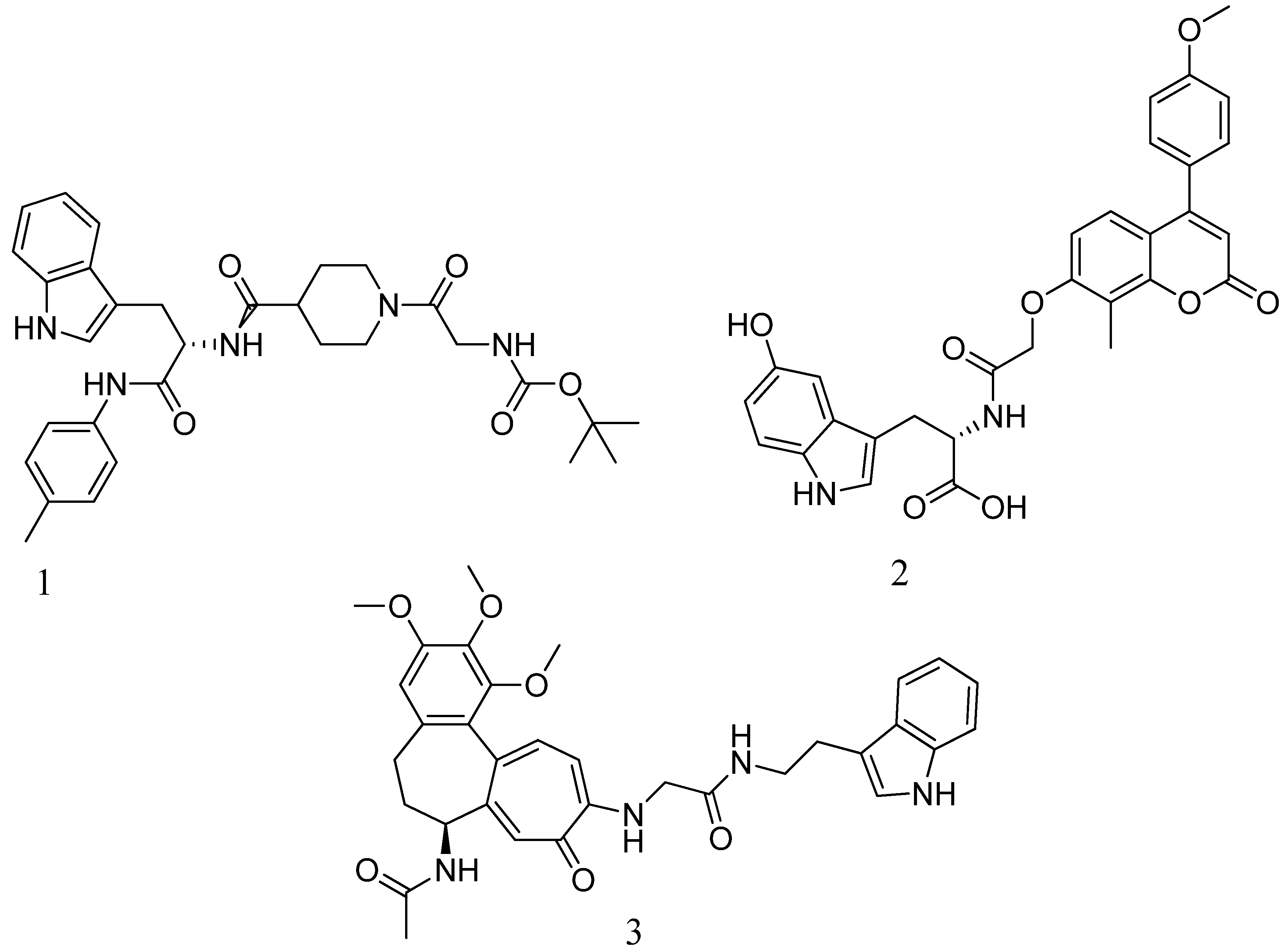
4.8. Bis-2-oxoazetidinyl macrocycle (β-lactams with 1,3-bridges):
4.9. Macrocycle-embedded β-lactams as novel inhibitors:
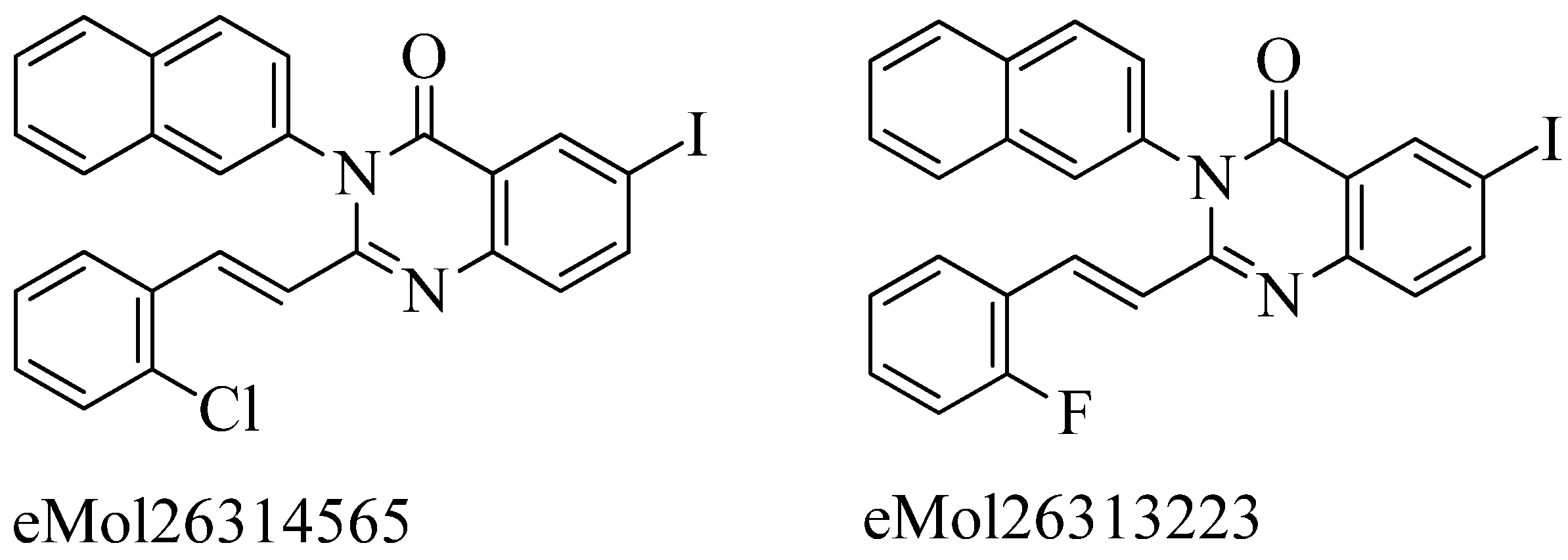
4.10. Pyridine-coupled pyrimidinones/pyrimidinthiones:
4.11. Novel naphthalimide corbelled aminothiazoximes:
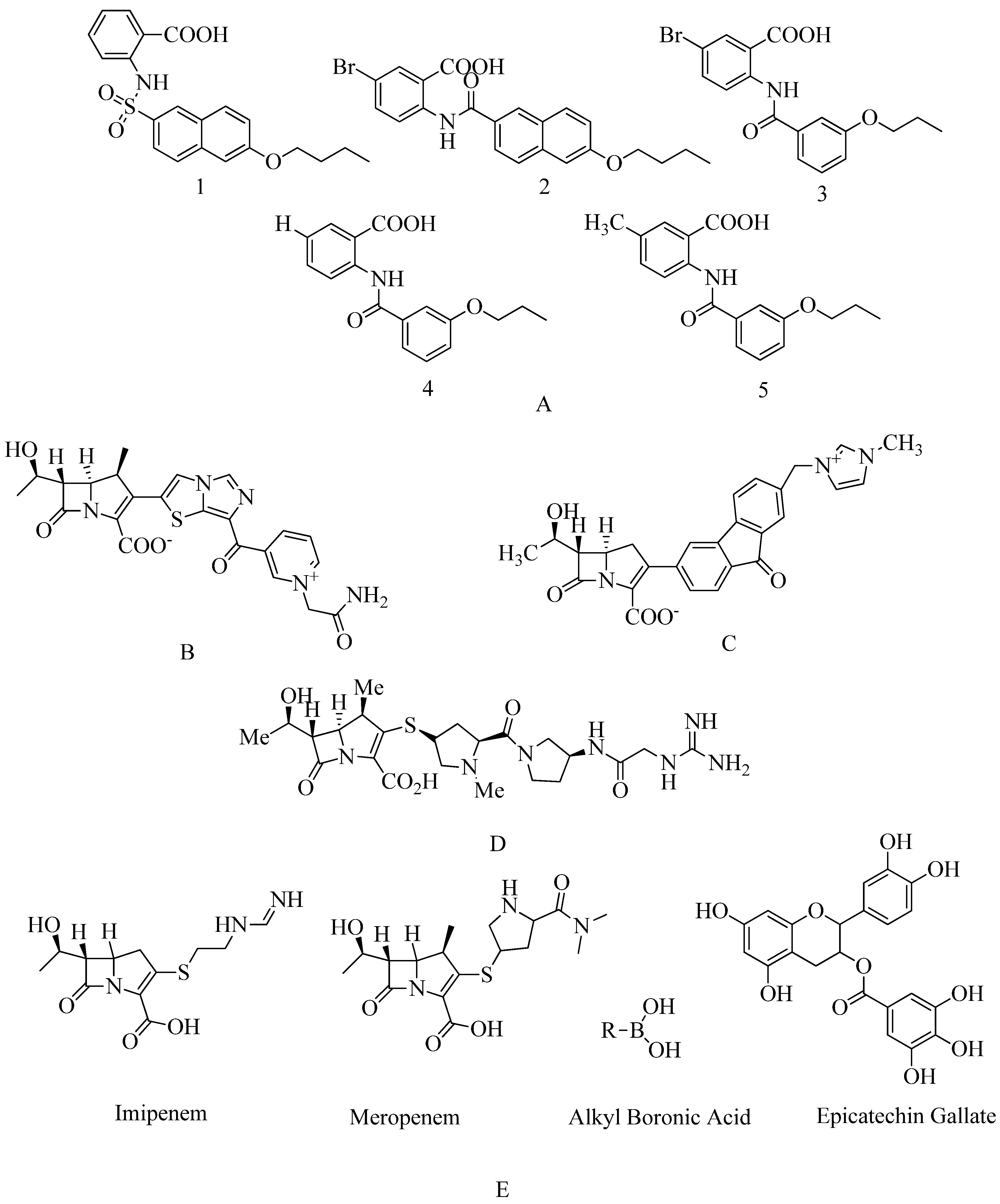
4.12. Non-covalent inhibitors:
4.13. Investigational β-lactam antibiotics:
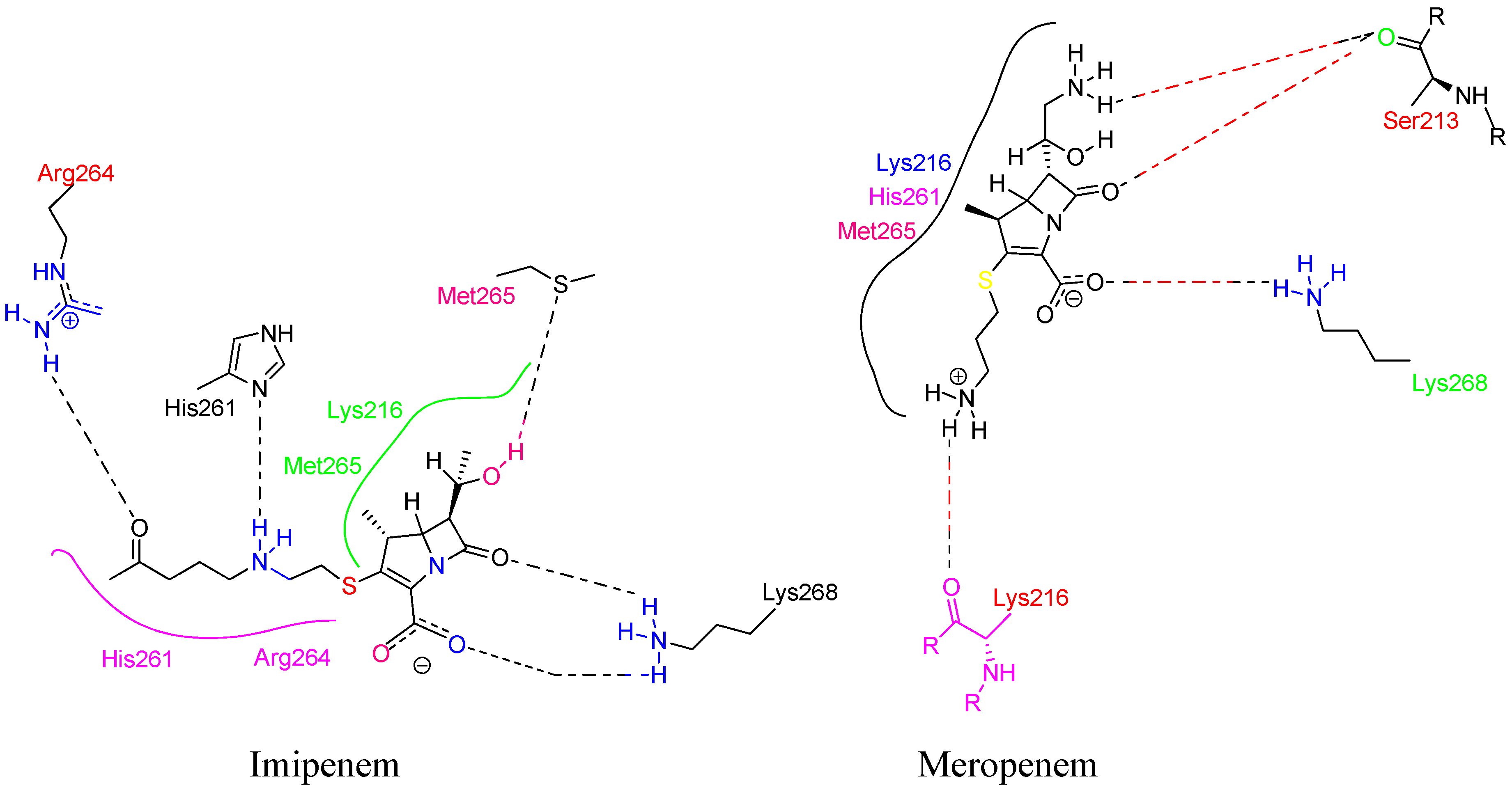
4.14. Carbapenem:
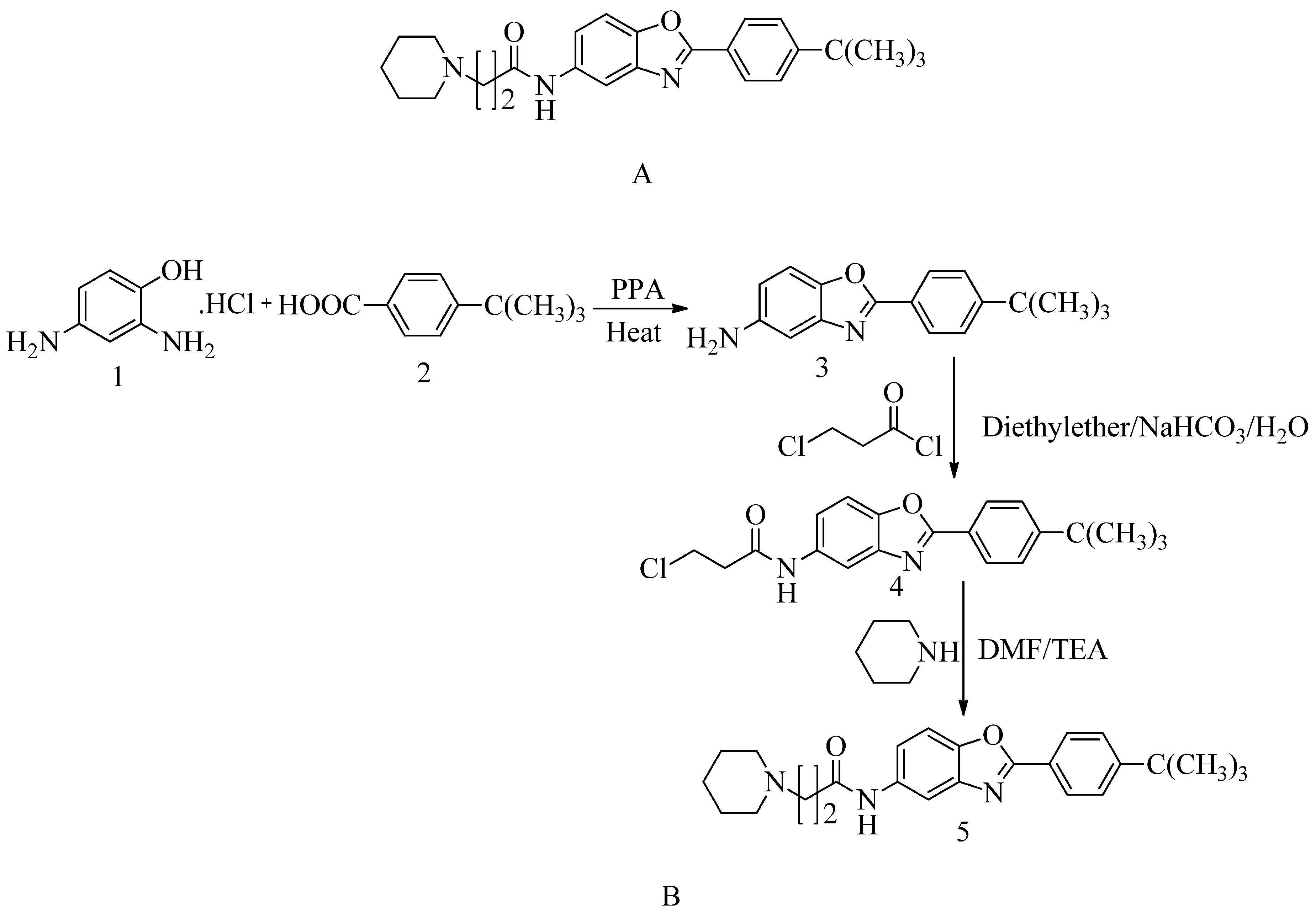
4.15. Novel benzoxazole derivatives:
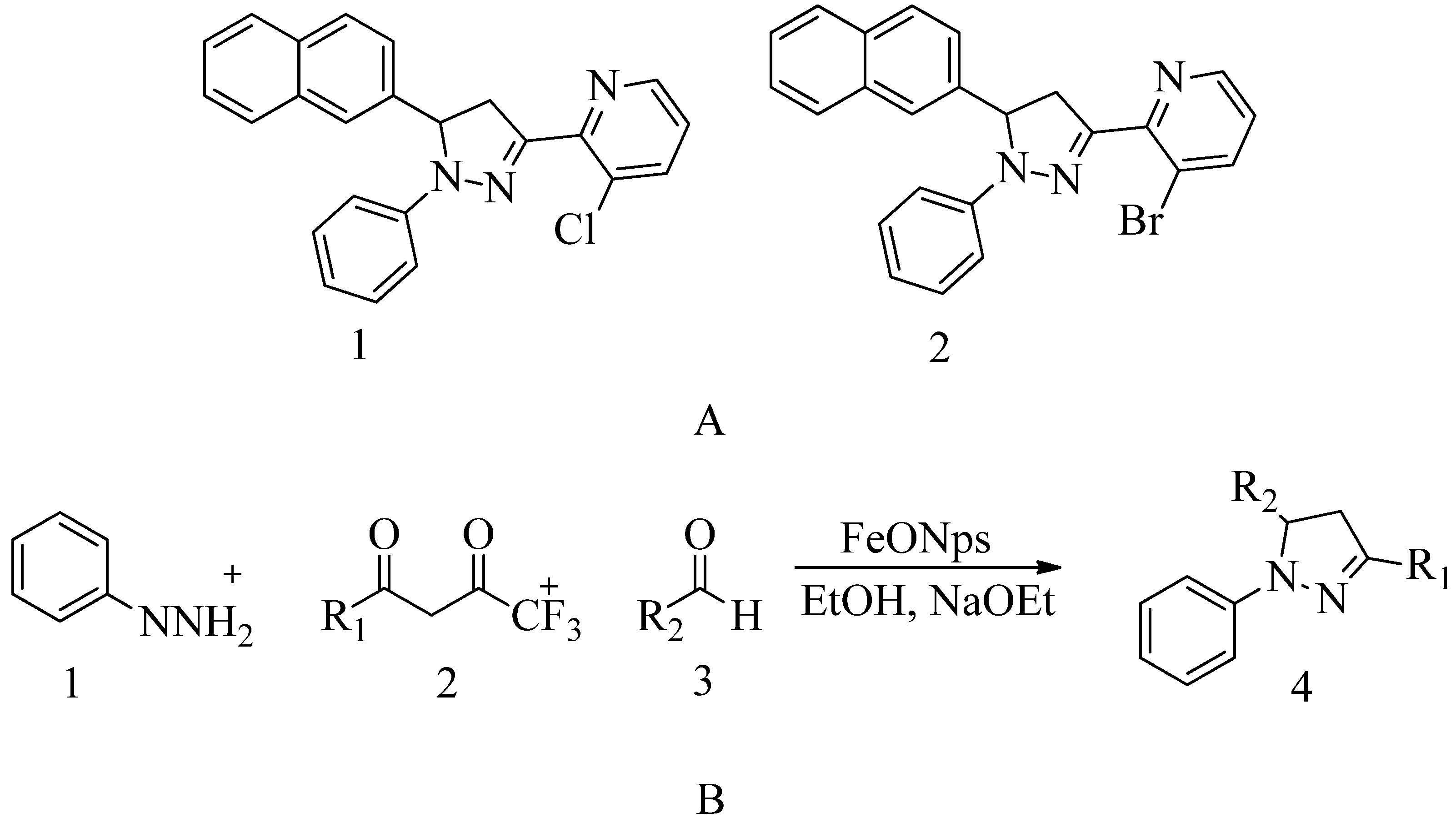
4.16. Pyrazolylpyridine analogues:
4.17. Miscellaneous PBP2a inhibitors:
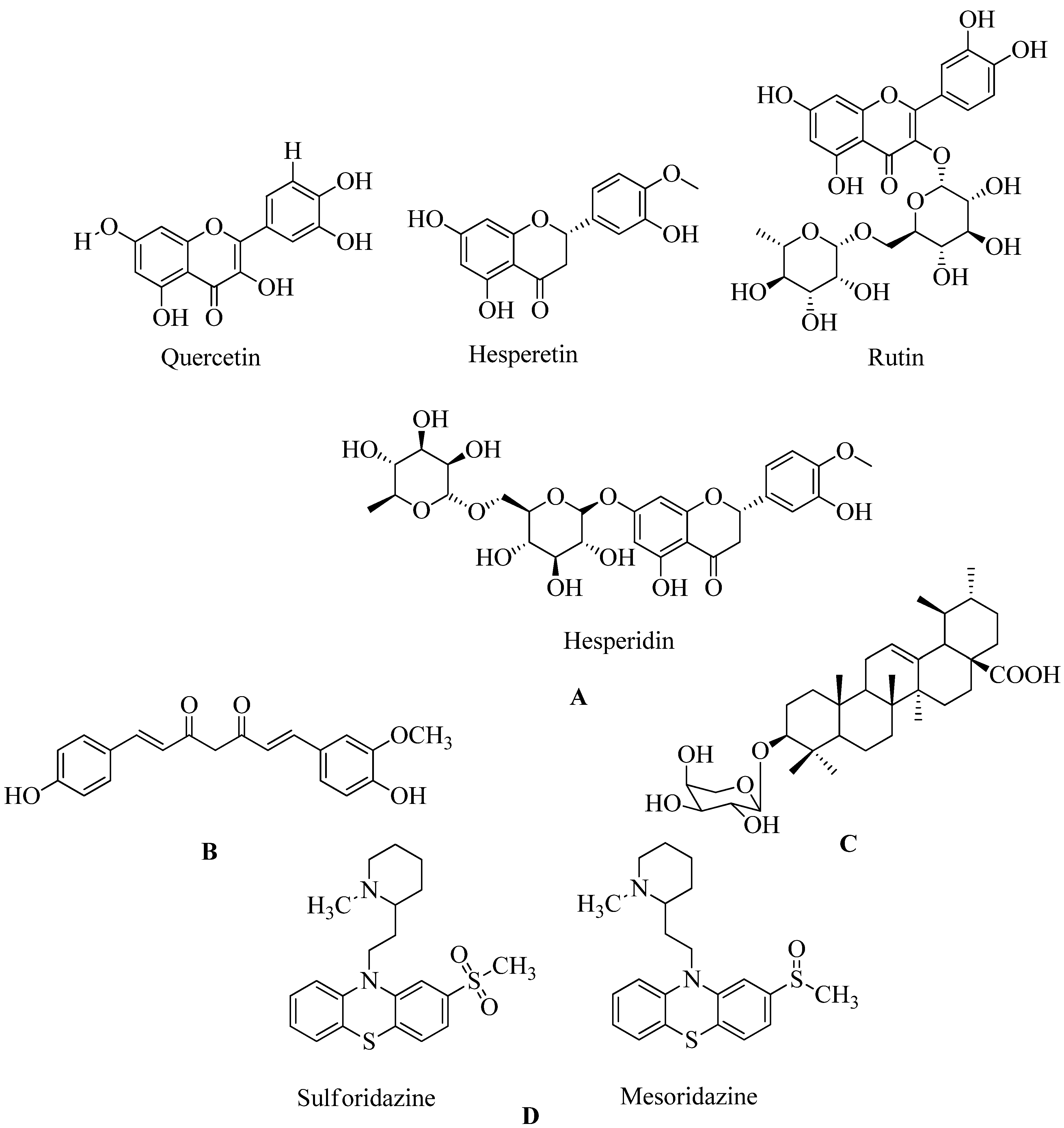
4.17.1. Flavonoids:
4.17.2. Demethoxycurcumin:
4.17.3. Quercetin 3-o-Rutinoside:
4.17.4. Ursolic acid 3-O-α-L-arabinopyranoside with oxacillin:
4.17.5. Thioridazine:
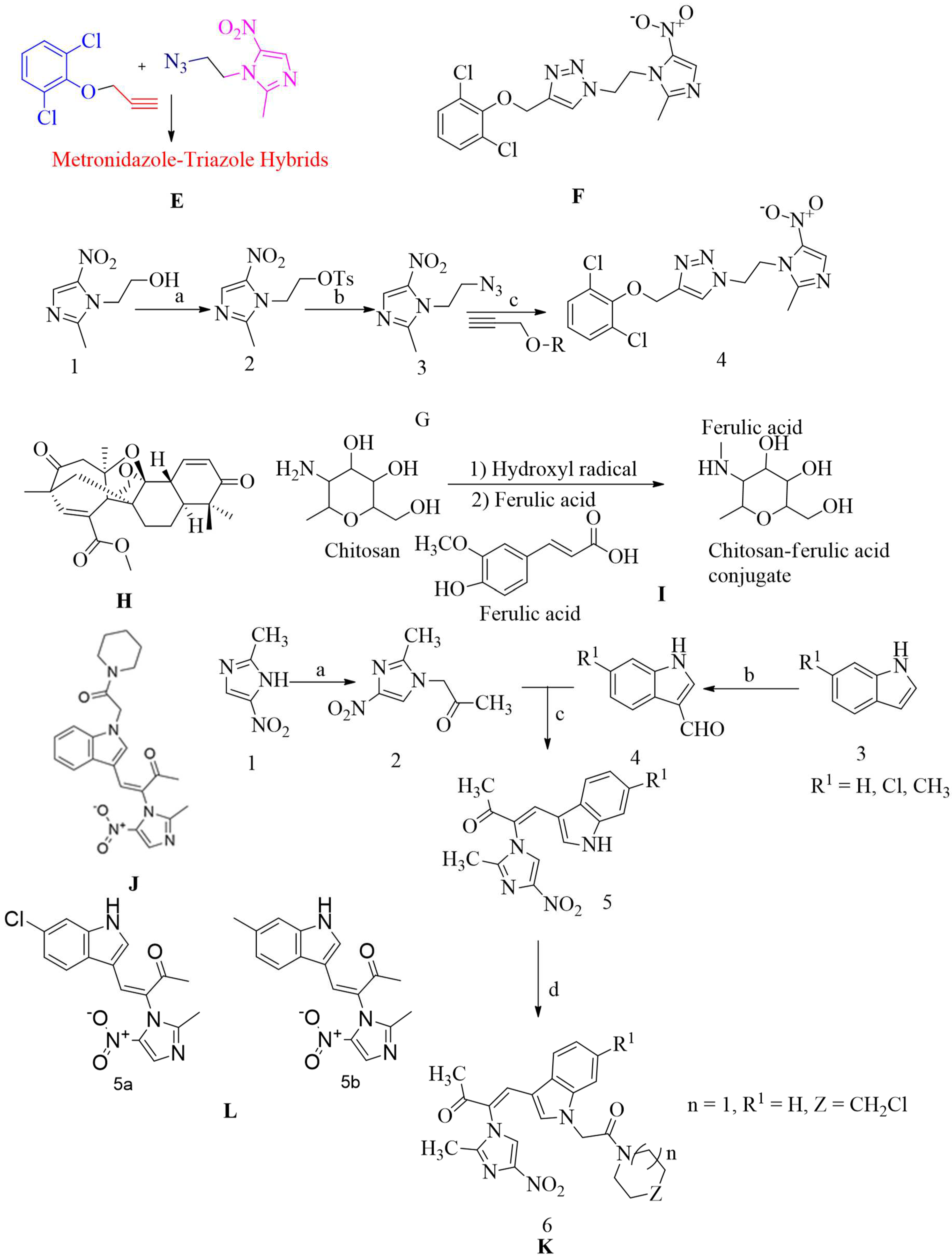
4.17.6. Metronidazole-Triazole Hybrids:
4.17.7. Aspermerodione:
4.17.8. Chitosan-ferulic acid conjugates:
4.17.9. Indole-nitroimidazole conjugates:
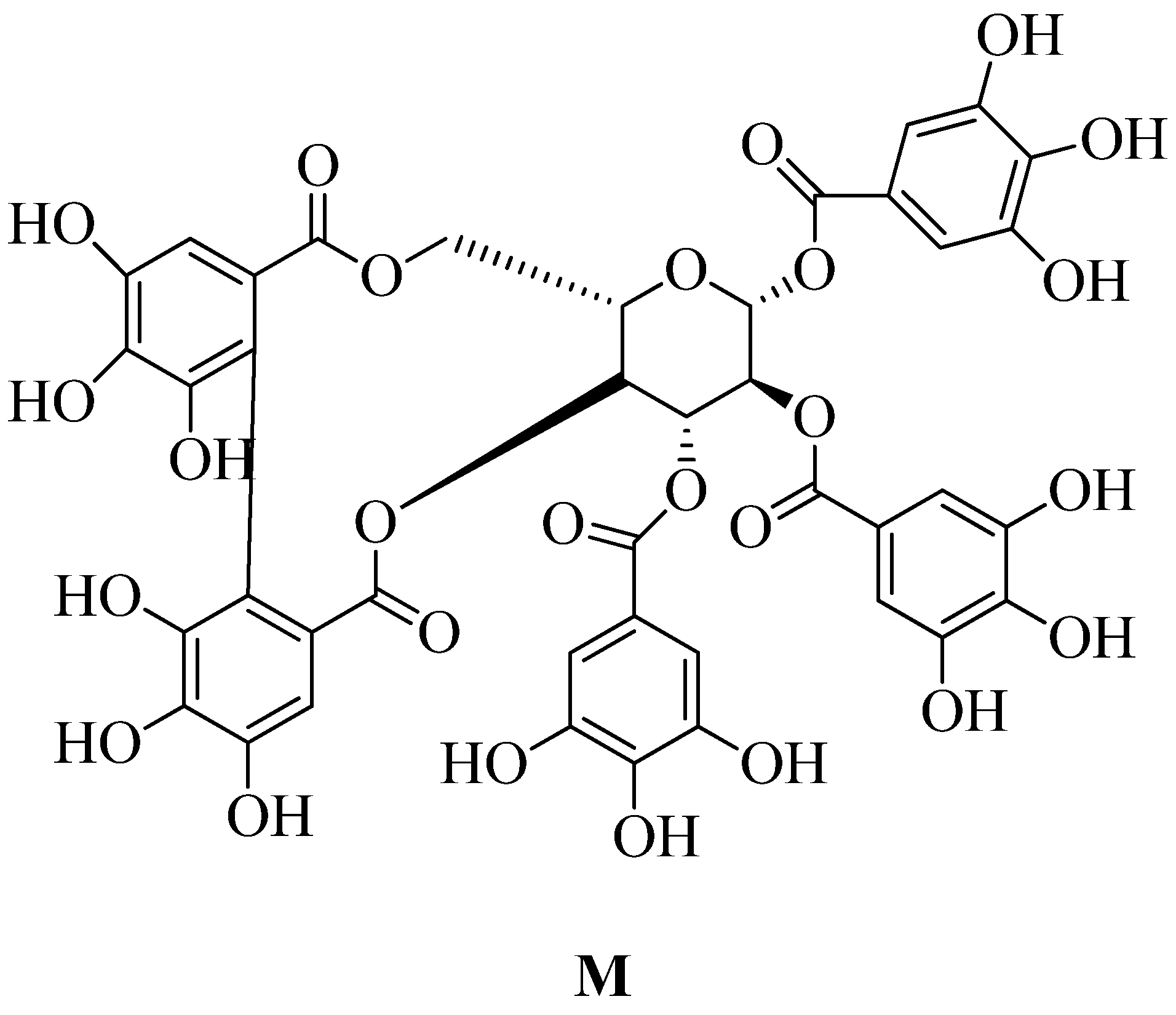
4.17.10. Polyphenol:
5. Screening technologies:
5.1. High throughput screening:
5.2. Computer-aided screening:
5.3. In-silico docking screening:
5.4. Multiple virtual screening techniques:
5.5. X-ray crystallography:
5.6. Miscellaneous method of screening:
5.6.1. Inverse virtual screening:
5.6.2. Reverse virtual screening:
5.6.3. Microtiter plate-based assay:
6. Conclusion:
7. The Outlook:
Author Contributions
Funding resources
Conflicts of Interest
References
- Tansey, E. M. Superbugs And Superdrugs: A History Of Mrsa The Transcript of a Witness Seminar Held by the Wellcome Trust Centre. 2008, 32 (July 2006). 20 July.
- Campbell, J.; Singh, A. K.; Santa, J. P.; Jr, M.; Kim, Y.; Swoboda, J. G.; Mylonakis, E.; Wilkinson, B. J.; Walker, S. NIH Public Access. 2012, 6 (1), 106–116. [CrossRef]
- Tong, S. Y. C.; Davis, J. S.; Eichenberger, E.; Holland, T. L.; Fowler, V. G. Staphylococcus Aureus Infections : Epidemiology , Pathophysiology , Clinical Manifestations , and Management. 2015, 623. [CrossRef]
- Guo, Y.; Song, G.; Sun, M.; Wang, J.; Wang, Y.; Wang, Y. Prevalence and Therapies of Antibiotic-Resistance in Staphylococcus Aureus. 2020, 10 (March), 1–11. [CrossRef]
- I, J. F. D. C.; I, Í. M. D. A.; Borges, K.; Rocha, F. Oxacillin Magnetically Targeted for the Treatment Of. 32 (1), 46–55.
- Lowy, F. D.; Lowy, F. D. Antimicrobial Resistance : The Example of Staphylococcus Aureus Find the Latest Version : Antimicrobial Resistance : The Example of Staphylococcus Aureus. 2003, 111 (9), 1265–1273. [CrossRef]
- Aureus, T. The Chromosome, as Well as the Extrachromosomal El- Ements. 6 These Genes Are Transferred between Staphy- Lococcal Strains, Species, or Other Gram-Positive Bacte- Rial Species through the Extrachromosomal Elements. 7. 1998.
- Foster, T. J.; Foster, T. J. The Staphylococcus Aureus “ Superbug ” The Staphylococcus Aureus “ Superbug .” 2004, 114 (12), 1693–1696.
- Saraiva, F. B.; de Araújo, A. C. C.; de Araújo, A. É. V.; Senna, J. P. M. Monoclonal Antibody Anti-PBP2a Protects Mice against MRSA (Methicillin-Resistant Staphylococcus Aureus) Infections. PLoS ONE2019, 14 (11), 1–13. [CrossRef]
- Deng, J. Z. Methicillin/per-6-(4-Methoxylbenzyl)-Amino-6-Deoxy-β-Cyclodextrin 1:1 Complex and Its Potentiation in Vitro against Methicillin-Resistant Staphylococcus Aureus. Journal of Antibiotics2013, 66 (9), 517–521. [CrossRef]
- Vernozy-rozand, C.; Mazuy, C.; Lasne, Y.; Fiedler, F.; Etienne, J. Staphylococcus Fleurettii Sp . Nov ., Isolated from Goat ’ s Milk Cheeses. 2000, 1521–1527.
- Berndsen, R.; Cunningham, T.; Kaelin, L.; Callender, M.; Boldog, W. D.; Viering, B.; King, A.; Labban, N.; Pollock, J. A.; Miller, H. B.; Blackledge, M. S. Identi Fi Cation and Evaluation of Brominated Carbazoles as a Novel Antibiotic Adjuvant Sca Ff Old in MRSA. 2022. [CrossRef]
- Hao, H.; Dai, M.; Wang, Y.; Huang, L.; Yuan, Z. Key Genetic Elements and Regulation Systems in Methicillin-Resistant Staphylococcus Aureus. Future Microbiology2012, 7 (11), 1315–1329. [CrossRef]
- Barber, M.; Rozwadowska-dowzenko, M. Patients 2. 1948, 641–644.
- Sciences, V. Review Article. 2015.
- Lyont, B. R.; Skurray, R. O. N. Antimicrobial Resistance of Staphylococcus Aureus : Genetic Basis. 1987, 51 (1), 88–134.
- Kumosani, T. Nosocomial Infections in Saudi Arabia Caused by Methicillin Resistance Staphylococcus Aureus (MRSA). Clinical Microbiology: Open Access2014, 03 (03). [CrossRef]
- Z; vizdi, AmraHuki, M. STAPHYLOCOCCUS AUREUS ( MRSA ) AS A CAUSE OF NOSOCOMIAL WOUND. 10 (1), 32–37.
- Grundmann, H.; Aires-de-sousa, M.; Boyce, J.; Tiemersma, E. Emergence and Resurgence of Meticillin-Resistant Staphylococcus Aureus as a Public-Health Threat. 2006, 874–885. [CrossRef]
- Liscano, Y.; Amú, A.; González, A.; Oñate-Garzón, J.; Salamanca, C. H. In Silico Characterization of the Interaction between the Pbp2a “Decoy” Protein of Resistant Staphylococcus Aureus and the Monomeric Units of Eudragit e-100 and Poly(Maleic Acid-Alt-Octadecene) Polymers. Polymers (Basel)2021, 13 (14). [CrossRef]
- Jr, R. C. M. MRSA: The First Half Century. 2012, No. October 2011, 4–11. 20 October. [CrossRef]
- Oehler, R. L. MRSA: Historical Perspective Richard L. Oehler, MD. 2000, 1961 (Table 1).
- Andrade, M. M.; Luiz, W. B.; Oliveira, S.; Amorim, J. H. The History of Methicillin-Resistant Staphylococcus Aureus in Brazil. 2020, 2020.
- Haamann, F.; Dulon, M.; Nienhaus, A. MRSA as an Occupational Disease : A Case Series. 2011, 259–266. [CrossRef]
- Diep, B. A.; Carleton, H. A.; Chang, R. F.; Sensabaugh, G. F. Roles of 34 Virulence Genes in the Evolution of Hospital- and Community-Associated Strains of Methicillin-Resistant Staphylococcus Aureus. 2006, 94110, 1495–1503.
- Enright, M. C.; Robinson, D. A.; Randle, G.; Feil, E. J.; Grundmann, H.; Spratt, B. G. The Evolutionary History of Methicillin-Resistant Staphylococcus Aureus ( MRSA ). 2002, 99 (11).
- Stefani, S.; Varaldo, P. E. Epidemiology of Methicillin-Resistant Staphylococci in Europe. 2003.
- Janout, V. CURRENT KNOWLEDGE OF METHICILLIN-RESISTANT STAPHYLOCOCCUS AUREUS AND COMMUNITY-ASSOCIATED METHICILLIN-RESISTANT STAPHYLOCOCCUS AUREUS Ivanka Matouskova , Vladimir Janout. 2008, 152 (2), 191–202.
- Majed, H.; Johnston, T.; Kelso, C.; Monachino, E.; Jergic, S.; Dixon, N. E.; Mylonakis, E.; Kelso, M. J. Structure-Activity Relationships of Pyrazole-4-Carbodithioates as Antibacterials against Methicillin−Resistant Staphylococcus Aureus. Bioorganic & Medicinal Chemistry Letters2018. [CrossRef]
- Since January 2020 Elsevier Has Created a COVID-19 Resource Centre with Free Information in English and Mandarin on the Novel Coronavirus COVID- Research That Is Available on the COVID-19 Resource Centre - Including This for Unrestricted Research Re-Use and Analyses in Any Form or by Any Means with Acknowledgement of the Original Source . These Permissions Are Meticillin-Resistant Staphylococcus Aureus. 2020, No. January.
- Ippolito, G.; Leone, S.; Lauria, F. N.; Nicastri, E.; Wenzel, R. P. International Journal of Infectious Diseases Methicillin-Resistant Staphylococcus Aureus : The Superbug. 2010, 4, 7–11. [CrossRef]
- Pantosti, A.; Venditti, M. What Is MRSA? 2009, 34 (5), 1190–1196. [CrossRef]
- Wong, S. C. Y.; Tse, H.; Chen, J. H. K. Multidrug-Resistant Staphylococcus Aureus , Colistin-Resistant Enterobacteriaceae Carrying the Mcr-1 Gene among Patients in Hong Kong. 2016, 22 (9), 2015–2016.
- Mehta, Y.; Hegde, A.; Pande, R.; Zirpe, K. G.; Gupta, V.; Ahdal, J.; Qamra, A.; Motlekar, S. Methicillin-Resistant Staphylococcus Aureus in Intensive Care Unit Setting of India : A Review of Clinical Burden , Patterns of Prevalence , Preventive Measures , and Future Strategies. 2020, 0–7.
- Correspondence; 2016; Vol. 132, pp 371–372. [CrossRef]
- Ld, I. M.; David, M. G.; Esposito, S.; Garau, J.; Lina, G.; Mazzei, T.; Peters, G. International Journal of Antimicrobial Agents New Insights into Meticillin-Resistant Staphylococcus Aureus ( MRSA ) Pathogenesis , Treatment and Resistance ଝ. 2012, 39, 96–104. [CrossRef]
- Baba, K.; Ishihara, K.; Ozawa, M.; Tamura, Y.; Asai, T. International Journal of Antimicrobial Agents Isolation of Meticillin-Resistant Staphylococcus Aureus ( MRSA ) from Swine in Japan. International Journal of Antimicrobial Agents2010, 36 (4), 352–354. [CrossRef]
- Outbreak, H. W.; Toleman, M. S.; Reuter, S.; Coll, F.; Harrison, E. M.; Peacock, S. J. Local Persistence of Novel MRSA Lineage after Community-Acquired Clostridium Difficile Infection,. 2016, 22 (9), 2011–2012.
- Huang, Y.; Chen, C. International Journal of Antimicrobial Agents Community-Associated Meticillin-Resistant Staphylococcus Aureus in Children In. International Journal of Antimicrobial Agents2011, 38 (1), 2–8. [CrossRef]
- Peacock, S. J.; Paterson, G. K. Mechanisms of Methicillin Resistance in Staphylococcus Aureus. Annual Review of Biochemistry2015, 84, 577–601. [CrossRef]
- Id, C. B.; Schwab, F.; Id, R. V. Infection Control Measures in Nosocomial MRSA Outbreaks — Results of a Systematic Analysis. 2021, 1–10. [CrossRef]
- Kumar, P. A Review on Quinoline Derivatives as Anti-Methicillin Resistant Staphylococcus Aureus (MRSA) Agents. BMC Chemistry2020, 14 (1), 1–14. [CrossRef]
- Jiang, W.; Li, B.; Zheng, X.; Liu, X.; Cen, Y.; Li, J.; Pan, X.; Cao, H.; Zheng, J.; Zhou, H. International Immunopharmacology Artesunate in Combination with Oxacillin Protect Sepsis Model Mice Challenged with Lethal Live Methicillin-Resistant Staphylococcus Aureus ( MRSA ) via Its Inhibition on Proin Fl Ammatory Cytokines Release and Enhancement on Antibacterial Activity of Oxacillin. International Immunopharmacology2011, 11 (8), 1065–1073. [CrossRef]
- Kong, Z.; Zhao, P.; Liu, H.; Yu, X.; Qin, Y.; Su, Z. Whole-Genome Sequencing for the Investigation of a Hospital Outbreak of MRSA in China. 2016, 1–12. [CrossRef]
- Stryjewski, M. E.; Corey, G. R. Methicillin-Resistant Staphylococcus Aureus : An Evolving Pathogen. 2014, 58 (Suppl 1), 10–19. [CrossRef]
- Mrsa, C. S. Methicillin-Resistant Staphylococcus Aureus Infections. 2008, 16 (5).
- Sahare, P.; Moon, A. Penicillin Binding Proteins: An Insight Into Novel Antibacterial Drug Target. 2014, 5 (8), 13–23.
- Goffin, C.; Ghuysen, J. Biochemistry and Comparative Genomics of SxxK Superfamily Acyltransferases Offer a Clue to the Mycobacterial Paradox : Presence of Penicillin-Susceptible Target Proteins versus Lack of Efficiency of Penicillin as Therapeutic Agent. 2002, 66 (4), 702–738. [CrossRef]
- Gordon, E.; Mouz, N.; Due, E.; Dideberg, O.; Horowitz, J. The Crystal Structure of the Penicillin-Binding Protein 2x from Streptococcus Pneumoniae and Its Acyl-Enzyme Form : Implication in Drug Resistance. 2000, 477–485. [CrossRef]
- Frere_Joris_1985_CRC_299.Pdf.
- Mouz, N.; Gordon, E.; Hopkins, J.; Dideberg, O. Crystal Structure of PBP2x from a Highly Penicillin-Resistant Streptococcus Pneumoniae Clinical Isolate. 2001, 276 (48), 45106–45112. [CrossRef]
- Sauvage, E.; Terrak, M.; Ayala, J. A.; Charlier, P. The Penicillin-Binding Proteins : Structure and Role in Peptidoglycan Biosynthesis. 2008, 32, 234–258. [CrossRef]
- EZervosen, A.; Sauvage, E.; Frère, J.; Charlier, P.; Luxen, A. Development of New Drugs for an Old Target — The Penicillin Binding Proteins. 2012, 12478–12505. [CrossRef]
- Green, D. W. The Bacterial Cell Wall as a Source of Antibacterial Targets. Expert Opinion on Therapeutic Targets2002, 6 (1), 1–20. [CrossRef]
- Resistance, B. MINIREVIEW Penicillin-Binding Proteins and Bacterial Resistance to , 3-Lactams. 1993, 37 (10), 2045–2053.
- Scheffers, D.; Pinho, M. G. Bacterial Cell Wall Synthesis : New Insights from Localization Studies. 2005, 69 (4), 585–607. [CrossRef]
- Anderson, J. S.; Matsuhashi, M.; Haskin, M. A.; Strominger, J. L. Biosythesis of the Peptidoglycan of Bacterial Cell Walls. II. Phospholipid Carriers in the Reaction Sequence. Journal of Biological Chemistry1967, 242 (13), 3180–3190. [CrossRef]
- Id, P. D. A. R.; Buss, J.; Id, S. I. S.; Id, G. R. S.; Srisuknimit, V.; Id, M. S.; Cho, H.; Sjodt, M.; Id, A. C. K.; Id, E. C. G.; Walker, S.; Kahne, D. E.; Id, T. G. B. A Central Role for PBP2 in the Activation of Peptidoglycan Polymerization by the Bacterial Cell Elongation Machinery. 2018, 1–25.
- Contreras-martel, C.; Vernet, T. Penicillin-Binding Proteins and b -Lactam Resistance. 2008. [CrossRef]
- Sauvage, E.; Kerff, F.; Terrak, M.; Ayala, J. A.; Charlier, P. The Penicillin-Binding Proteins: Structure and Role in Peptidoglycan Biosynthesis. FEMS Microbiology Reviews2008, 32 (2), 234–258. [CrossRef]
- Dave, K.; Palzkill, T.; Pratt, R. F. Neutral β-Lactams Inactivate High Molecular Mass Penicillin-Binding Proteins of Class B1, Including PBP2a of MRSA. ACS Medicinal Chemistry Letters2014, 5 (2), 154–157. [CrossRef]
- Pinho, M. G.; Errington, J. Recruitment of Penicillin-Binding Protein PBP2 to the Division Site of Staphylococcus Aureus Is Dependent on Its Transpeptidation Substrates. 2005, 55, 799–807. [CrossRef]
- hang Wei Wu; De Lencastre, H.; Tomasz, SA. Recruitment of the MecA Gene Homologue of Staphylococcus Sciuri into a Resistance Determinant and Expression of the Resistant Phenotype in Staphylococcus Aureus. Journal of Bacteriology2001, 183 (8), 2417–2424. [CrossRef]
- De Jonge, B. L. M.; Chang, Y. S.; Gage, D.; Tomasz, A. Peptidoglycan Composition of a Highly Methicillin-Resistant Staphylococcus Aureus Strain. The Role of Penicillin Binding Protein 2A. Journal of Biological Chemistry1992, 267 (16), 11248–11254. [CrossRef]
- Oliveira, D. C.; Wu, S. W. Genetic Organization of the Downstream Region of the MecA Element in Methicillin-Resistant Staphylococcus Aureus Isolates Carrying Different Polymorphisms of This Region. 2000, 44 (7), 1906–1910. [CrossRef]
- Lencastre, H. De; Jonge, B. L. M. De; Matthews, P. R.; Tomasz, A. Review Molecular Aspects of Methicillin Resistance in Staphylococcus Aureus. 1994, 7–24.
- Murakami, K. Nucleotide Sequence of the Structural Gene for the Penicillin-Binding Protein 2 of Staphylococcus Aureus and the Presence of a Homologous Gene in Other Staphylococci. 1994, 117, 131–136. [CrossRef]
- Filipe, R. R.; Pinho, M. G.; Tomasz, A. Complementation of the Essential Peptidoglycan Transpeptidase Function of Penicillin-Binding Protein 2 ( PBP2 ) by the Drug Resistance Protein PBP2A in Staphylococcus Aureus. 2001, 183 (22), 6525–6531. [CrossRef]
- Fuda, C.; Hesek, D.; Lee, M.; Morio, K. I.; Nowak, T.; Mobashery, S. Activation for Catalysis of Penicillin-Binding Protein 2a from Methicillin-Resistant Staphylococcus Aureus by Bacterial Cell Wall. J Am Chem Soc2005, 127 (7), 2056–2057. [CrossRef]
- Hakenbeck, R.; Coyette, J. Resistant Penicillin-Binding Proteins. 1998, 54, 332–340. [CrossRef]
- Fuda, C.; Suvorov, M.; Vakulenko, S. B.; Mobashery, S. The Basis for Resistance to β-Lactam Antibiotics by Penicillin-Binding Protein 2a of Methicillin-Resistant Staphylococcus Aureus. Journal of Biological Chemistry2004, 279 (39), 40802–40806. [CrossRef]
- Stapleton, P. D.; Taylor, P. W. Europe PMC Funders Group Methicillin Resistance in Staphylococcus Aureus : Methicillin Resistance. 2007, 85 (Pt 1), 1–14.
- Srisuknimit, V.; Qiao, Y.; Schaefer, K.; Kahne, D.; Walker, S. Peptidoglycan Cross-Linking Preferences of Staphylococcus Aureus Penicillin-Binding Proteins Have Implications for Treating MRSA Infections. J Am Chem Soc2017, 139 (29), 9791–9794. [CrossRef]
- Contreras-martel, C.; Martins, A.; Ecobichon, C.; Trindade, D. M.; Matteï, P.; Hicham, S.; Hardouin, P.; Ghachi, M. El; Boneca, I. G.; Dessen, A. Molecular Architecture of the PBP2–MreC Core Bacterial Cell Wall Synthesis Complex. Nature Communications 1–10. [CrossRef]
- Schleifer, K. H.; Kandler, O. Peptidoglycan Types of Bacterial Cell Walls and Their Taxonomic Implications. Bacteriol Rev1972, 36 (4), 407–477. [CrossRef]
- Contreras-martel, C.; Amoroso, A.; Woon, E. C. Y.; Zervosen, A.; Inglis, S.; Martins, A.; Verlaine, O.; Rydzik, A. M.; Job, V. Structure-Guided Design of Cell Wall Biosynthesis Inhibitors That Overcome β -Lactam Resistance in Staphylococcus Aureus (MRSA). 2011, 943–951. [CrossRef]
- Tahir99-VRG & Vip.Persianss.Ir.
- Hrast, M.; Rožman, K.; Jukič, M.; Patin, D.; Gobec, S.; Sova, M. Synthesis and Structure–Activity Relationship Study of Novel Quinazolinone-Based Inhibitors of MurA. Bioorganic and Medicinal Chemistry Letters2017, 27 (15), 3529–3533. [CrossRef]
- Nikolaidis, I.; Dessen, A. Resistance to Antibiotics Targeted to the Bacterial Cell Wall. 2014, 23, 243–259. [CrossRef]
- Lewis, R. J. Structural Basis for the Coordination of Cell Division with the Synthesis of the Bacterial Cell Envelope. 2019, No. August, 2042–2054. [CrossRef]
- Caveney, N. A.; Li, F. K.; Strynadka, N. C. Enzyme Structures of the Bacterial Peptidoglycan and Wall Teichoic Acid Biogenesis Pathways. Current Opinion in Structural Biology2018, 53, 45–58. [CrossRef]
- Macheboeuf, P.; Contreras-martel, C.; Job, V.; Dideberg, O. Penicillin Binding Proteins : Key Players in Bacterial Cell Cycle and Drug Resistance Processes. 2006, 30, 673–691. [CrossRef]
- Gautam, A.; Vyas, R.; Tewari, R. Peptidoglycan Biosynthesis Machinery : A Rich Source of Drug Targets. 2011, 31 (April 2010), 295–336. 20 April. [CrossRef]
- Naclerio, G. A.; Sintim, H. O. Multiple Ways to Kill Bacteria via Inhibiting Novel Cell Wall or Membrane Targets. 2020.
- Kovac, A. Cytoplasmic Steps of Peptidoglycan Biosynthesis. 2008, 32, 168–207. [CrossRef]
- Rogers, H. J.; Perkins, H. R.; Ward, J. B. Biosynthesis of Peptidoglycan. Microbial Cell Walls and Membranes1980, 239–297. [CrossRef]
- Monteiro, J. M.; Covas, G.; Rausch, D.; Filipe, S. R.; Schneider, T.; Sahl, H. G.; Pinho, M. G. The Pentaglycine Bridges of Staphylococcus Aureus Peptidoglycan Are Essential for Cell Integrity. Scientific Reports2019, 9 (1), 1–10. [CrossRef]
- Goldman, R. C.; Gange, D. Inhibition of Transglycosylation Peptidoglycan Synthesis Involved in Bacterial. 2000, No. 609, 801–820.
- Gondokesumo, M. E.; Kurniawan, I. M. Molecular Docking Study of Sappan Wood Extract to Inhibit PBP2A Enzyme on Methicillin-Resistant Staphylococcus Aureus (MRSA). Journal of Basic and Clinical Physiology and Pharmacology2019, 30 (6), 1–9. [CrossRef]
- Young, K. D. Peptidoglycan. 2001, 1–11.
- Walsh, C. T. Enzymes in the D-Alanine Branch of Bacterial Cell Wall Peptidoglycan Assembly. Journal of Biological Chemistry1989, 264 (5), 2393–2396. [CrossRef]
- Berger-Bächi, B.; Tschierske, M. Role of Fem Factors in Methicillin Resistance. Drug Resistance Updates1998, 1 (5), 325–335. [CrossRef]
- Kourtis, A. P.; Hatfield, K.; Baggs, J.; Mu, Y.; See, I.; Epson, E.; Nadle, J.; Kainer, M. A.; Dumyati, G. Vital Signs : Epidemiology and Recent Trends in Methicillin-Resistant and in Methicillin-Susceptible Staphylococcus Aureus Bloodstream Infections — United States. 2020.
- Cedraro, N.; Cannalire, R.; Astolfi, A.; Mangiaterra, G.; Felicetti, T.; Vaiasicca, S.; Cernicchi, G.; Massari, S.; Biavasco, F.; Sabatini, S. From Quinoline to Quinazoline-Based S . Aureus NorA Efflux Pump Inhibitors by Coupling a Focused Scaffold Hopping Approach and a Pharmacophore Search. 2021, 3044–3059. [CrossRef]
- Lade, H.; Kim, J. S. Bacterial Targets of Antibiotics in Methicillin-Resistant Staphylococcus Aureus. Antibiotics2021, 10 (4). [CrossRef]
- Kaczor, A.; Witek, K.; Podlewska, S.; Czekajewska, J.; Lubelska, A.; Zesławska, E.; Nitek, W.; Latacz, G.; Alibert, S.; Pagès, J. M.; Karczewska, E.; Kieć-Kononowicz, K.; Handzlik, J. 5-Arylideneimidazolones with Amine at Position 3 as Potential Antibiotic Adjuvants against Multidrug Resistant Bacteria. Molecules2019, 24 (3). [CrossRef]
- Jousselin, A.; Manzano, C.; Biette, A.; Reed, P.; Pinho, M. G.; Rosato, A. E.; Kelley, W. L.; Renzoni, A. The Staphylococcus Aureus Chaperone PrsA Is a New Auxiliary Factor of Oxacillin Resistance Affecting Penicillin-Binding Protein 2A. Antimicrobial Agents and Chemotherapy2016, 60 (3), 1656–1666. [CrossRef]
- Murphy, J. T.; Walshe, R.; Devocelle, M. A Computational Model of Antibiotic-Resistance Mechanisms in Methicillin-Resistant Staphylococcus Aureus (MRSA). Journal of Theoretical Biology2008, 254 (2), 284–293. [CrossRef]
- Mohamed, S. B.; Adlan, T. A.; Khalafalla, N. A.; Abdalla, N. I.; Ali, Z. S. A.; Munir KA, A.; Hassan, M. M.; Elnour, M. A. B. Proteomics and Docking Study Targeting Penicillin-Binding Protein and Penicillin-Binding Protein2a of Methicillin-Resistant Staphylococcus Aureus Strain SO-1977 Isolated from Sudan. Evolutionary Bioinformatics2019, 15. 2019; 15. [CrossRef]
- Babic, M.; Hujer, A. M.; Bonomo, R. A.; Stokes, L.; Medical, A. What ’ s New in Antibiotic Resistance ? Focus on Beta-Lactamases. 2006, 9, 142–156. [CrossRef]
- Wielders, C. L. C.; Fluit, A. C.; Brisse, S.; Verhoef, J.; Schmitz, F. J. MecA Gene Is Widely Disseminated in Staphylococcus Aureus Population. 2002, 40 (11), 3970–3975. [CrossRef]
- Tsubakishita, S.; Kuwahara-arai, K.; Sasaki, T.; Hiramatsu, K. Origin and Molecular Evolution of the Determinant of Methicillin Resistance in Staphylococci. 2010, 54 (10), 4352–4359. [CrossRef]
- Webster, J. A.; Bannerman, T. L.; Hubner, R. J.; Ballard, D. N.; Cole, E. M.; Bruce, J. L.; Fiedler, F.; Schubert, K.; Kloos, W. E. Identification of the Staphylococcus Sciuri Species Group with EcoRI Fragments Containing RRNA Sequences and Description of Staphylococcus Vitulus Sp . Nov . 1994, 460 (July), 454–460.
- Schleifer, K. H.; Geyer, U.; Kilpper-balz, R.; Devriese, L. A. Elevation of Staphylo Coccus Sciuri Subsp . Lentus ( Kloos et al .) to Species Status : Staphylococcus Lentus ( Kloos et al .) Comb . Nov . 1983, 387, 382–387. [CrossRef]
- Ouchenane, Z.; Agabou, A.; Smati, F.; Rolain, J. M.; Raoult, D. Staphylococcal Cassette Chromosome Mec Characterization of Methicillin-Resistant Staphylococcus Aureus Strains Isolated at the Military Hospital of Constantine/Algeria. Pathologie Biologie2013, 61 (6), 280–281. [CrossRef]
- Chromosome, S. C. Crossm Evolutionary Origin of the Staphylococcal Cassette Chromosome. 2017, 61 (6), 1–16.
- Katayama, Y.; Ito, T.; Hiramatsu, K. A New Class of Genetic Element , Staphylococcus Cassette Chromosome Mec , Encodes Methicillin Resistance in Staphylococcus Aureus. 2000, 44 (6), 1549–1555.
- Liu, J.; Chen, D.; Peters, B. M.; Li, L.; Li, B.; Xu, Z.; Shirliff, M. E. Staphylococcal Chromosomal Cassettes Mec (SCCmec): A Mobile Genetic Element in Methicillin-Resistant Staphylococcus Aureus. Microbial Pathogenesis2016, 101, 56–67. [CrossRef]
- Ito, T.; Ma, X. X.; Takeuchi, F.; Okuma, K.; Yuzawa, H.; Hiramatsu, K. Novel Type V Staphylococcal Cassette Chromosome Mec Driven by a Novel Cassette Chromosome Recombinase , CcrC. 2004, 48 (7), 2637–2651. [CrossRef]
- Ender, M.; Berger-Bächi, B.; McCallum, N. Variability in SCCmecN1 Spreading among Injection Drug Users in Zurich, Switzerland. BMC Microbiology2007, 7, 1–10. [CrossRef]
- hongtrakool, P.; Ito, T.; Ma, X. X.; Trakulsomboon, S.; Tiensasitorn, C.; Jamklang, M.; Chavalit, T.; Song, J.; Chongtrakool, P.; Ito, T.; Ma, X. X.; Kondo, Y.; Trakulsomboon, S. Staphylococcal Cassette Chromosome Mec ( SCC Mec ) Typing of Methicillin-Resistant Staphylococcus Aureus Strains Isolated in 11 Asian Countries : A Proposal for a New Nomenclature for SCC Mec Elements Staphylococcal Cassette Chromosome Mec ( SCC Mec ) Typing of Methicillin-Resistant Staphylococcus Aureus Strains Isolated in 11 Asian Countries : A Proposal for a New Nomenclature for SCC Mec Elements. 2006. [CrossRef]
- Tsubakishita, S.; Kuwahara-arai, K.; Baba, T.; Hiramatsu, K. Staphylococcal Cassette Chromosome Mec -Like Element in Macrococcus Caseolyticus. 2010, 54 (4), 1469–1475. [CrossRef]
- Uehara, Y. Current Status of Staphylococcal Cassette. 2022, 1–12. [CrossRef]
- Berger-Bächi, B.; Rohrer, S. Factors Influencing Methicillin Resistance in Staphylococci. Archives of Microbiology2002, 178 (3), 165–171. [CrossRef]
- Fishovitz, J.; Hermoso, J. A.; Chang, M.; Mobashery, S. Penicillin-Binding Protein 2a of Methicillin-Resistant Staphylococcus Aureus. IUBMB Life2014, 66 (8), 572–577. [CrossRef]
- Lim, D.; Strynadka, N. C. J. Structural Basis for the β-Lactam Resistance of PBP2a from Methicillin-Resistant Staphylococcus Aureus. Nature Structural Biology2002, 9 (11), 870–876. [CrossRef]
- Kumar, K. M.; Anitha, P.; Sivasakthi, V.; Bag, S.; Lavanya, P.; Anbarasu, A.; Ramaiah, S. In Silico Study on Penicillin Derivatives and Cephalosporins for Upper Respiratory Tract Bacterial Pathogens. 2014, 241–251. [CrossRef]
- Chiang, Y. C.; Wong, M. T. Y.; Essex, J. W. Molecular Dynamics Simulations of Antibiotic Ceftaroline at the Allosteric Site of Penicillin-Binding Protein 2a (PBP2a). Israel Journal of Chemistry2020, 60 (7), 754–763. [CrossRef]
- Lovering, A. L.; Gretes, M. C.; Safadi, S. S.; Danel, F.; Castro, L. De; Page, M. G. P.; Strynadka, N. C. J. Structural Insights into the Anti-Methicillin-Resistant Staphylococcus Aureus ( MRSA ) Activity of Ceftobiprole *. 2012, 287 (38), 32096–32102. [CrossRef]
- Hermoso, J. A.; Chang, M.; Mobashery, S. HHS Public Access. 2018, 139 (5), 2102–2110. [CrossRef]
- My, N. H.; Hirao, H.; Van, D. U.; Morokuma, K. Computational Studies of Bacterial Resistance to β-Lactam Antibiotics: Mechanism of Covalent Inhibition of the Penicillin-Binding Protein 2a (PBP2a). Journal of Chemical Information and Modeling2011, 51 (12), 3226–3234. [CrossRef]
- Janardhanan, J.; Bouley, R.; Peng, Z.; Batuecas-mordillo, M.; Meisel, J. E.; Ding, D.; Schroeder, V. A.; Wolter, W. R.; Mahasenan, K. V; Hermoso, J. A.; Mobashery, S.; Chang, M. Crossm The Quinazolinone Allosteric Inhibitor of PBP 2a Synergizes. 2019, 63 (5), 1–12.
- Janardhanan, J.; Bouley, R.; Peng, Z.; Batuecas-mordillo, M.; Meisel, J. E.; Ding, D.; Schroeder, V. A.; Wolter, W. R.; Mahasenan, K. V; Hermoso, J. A.; Mobashery, S.; Chang, M. Crossm The Quinazolinone Allosteric Inhibitor of PBP 2a Synergizes. 2019, 63 (5), 1–12.
- Bouley, R.; Kumarasiri, M.; Peng, Z.; Otero, L. H.; Song, W.; Mark, A.; Schroeder, V. A.; Wolter, W. R.; Lastochkin, E.; Antunes, N. T.; Pi, H.; Vakulenko, S.; Hermoso, J. A.; Chang, M.; Mobashery, S. Discovery of Antibiotic ( E ) -3- ( 3-Carboxyphenyl ) -2- ( 4- Cyanostyryl ) Quinazolin-4 ( 3 H ) -One. 2015, 4. [CrossRef]
- Otero, L. H.; Rojas-Altuve, A.; Llarrull, L. I.; Carrasco-López, C.; Kumarasiri, M.; Lastochkin, E.; Fishovitz, J.; Dawley, M.; Hesek, D.; Lee, M.; Johnson, J. W.; Fisher, J. F.; Chang, M.; Mobashery, S.; Hermoso, J. A. How Allosteric Control of Staphylococcus Aureus Penicillin Binding Protein 2a Enables Methicillin Resistance and Physiological Function. Proc Natl Acad Sci U S A2013, 110 (42), 16808–16813. [CrossRef]
- Lv, N.; Kong, Q.; Zhang, H.; Li, J. Discovery of Novel Staphylococcus Aureus Penicillin Binding Protein 2a Inhibitors by Multistep Virtual Screening and Biological Evaluation. Bioorganic and Medicinal Chemistry Letters2021, 41 (April), 2–9. [CrossRef]
- Lavanya, P.; Ramaiah, S.; Anbarasu, A. A Molecular Docking and Dynamics Study to Screen Potent Anti-Staphylococcal Compounds Against Ceftaroline Resistant MRSA. Journal of Cellular Biochemistry2016, 117 (2), 542–548. [CrossRef]
- Matys, A.; Podlewska, S.; Witek, K.; Witek, J.; Bojarski, A. J.; Schabikowski, J.; Otrębska-Machaj, E.; Latacz, G.; Szymańska, E.; Kieć-Kononowicz, K.; Molnar, J.; Amaral, L.; Handzlik, J. Imidazolidine-4-One Derivatives in the Search for Novel Chemosensitizers of Staphylococcus Aureus MRSA: Synthesis, Biological Evaluation and Molecular Modeling Studies. European Journal of Medicinal Chemistry2015, 101, 313–325. [CrossRef]
- Eom, S.; Kang, S.; Lee, D.; Myeong, J.; Lee, J.; Kim, H.; Kim, K.; Je, J.; Jung, W.; Kim, Y. Synergistic Antibacterial Effect and Antibacterial Action Mode of Chitosan - Ferulic Acid Conjugate against Methicillin-Resistant Staphylococcus Aureus. 2016, 26 (0), 784–789. [CrossRef]
- Kumar, M.; Kumar, D.; Raj, V. Studies on Imidazole and Its Derivatives with Particular Emphasis on Their Chemical/Biological Applications as Bioactive Molecules/Intermediated to Bioactive Molecule. Current Synthetic and Systems Biology2017, 05 (01), 1–10. [CrossRef]
- Bush, K. Synergistic MRSA Combinations. Nature Publishing Group2015, 11 (11), 832–833. [CrossRef]
- Gonzales, P. R.; Pesesky, M. W.; Bouley, R.; Ballard, A.; Biddy, B. A.; Suckow, M. A.; Wolter, W. R.; Schroeder, V. A.; Burnham, C. D.; Mobashery, S.; Chang, M.; Dantas, G. Combinations Suppress Resistance in MRSA. 2015, No. september. [CrossRef]
- Stryjewski, M. E.; Jones, R. N.; Corey, G. R. SC. Diagnostic Microbiology and Infectious Disease2014. [CrossRef]
- Pii, I. PT SC. Integrative Medicine Research2018. [CrossRef]
- Saravolatz, L. D.; Stein, G. E.; Johnson, L. B. Ceftaroline: A Novel Cephalosporin with Activity against Methicillin-Resistant Staphylococcus Aureus. Clinical Infectious Diseases2011, 52 (9), 1156–1163. [CrossRef]
- Soriano, A. Ceftaroline. Revista Espanola de Quimioterapia2021, 34 (7), 29–31. [CrossRef]
- Otero, L. H.; Rojas-Altuve, A.; Llarrull, L. I.; Carrasco-López, C.; Kumarasiri, M.; Lastochkin, E.; Fishovitz, J.; Dawley, M.; Hesek, D.; Lee, M.; Johnson, J. W.; Fisher, J. F.; Chang, M.; Mobashery, S.; Hermoso, J. A. How Allosteric Control of Staphylococcus Aureus Penicillin Binding Protein 2a Enables Methicillin Resistance and Physiological Function. Proc Natl Acad Sci U S A2013, 110 (42), 16808–16813. [CrossRef]
- Hernandez, P. O.; Lema, S.; Tyring, S. K.; Mendoza, N. Ceftaroline in Complicated Skin and Skin-Structure Infections. Infection and Drug Resistance2012, 5 (1), 23–35. [CrossRef]
- Schaumburg, F.; Peters, G.; Alabi, A.; Becker, K.; Idelevich, E. A. Missense Mutations of PBP2a Are Associated with Reduced Susceptibility to Ceftaroline and Ceftobiprole in African MRSA. 2016, No. October 2015, 41–44. [CrossRef]
- Livermore, D. M. Can β-Lactams Be Re-Engineered to Beat MRSA? Clinical Microbiology and Infection2006, 12 (SUPPL. 2), 11–16. (SUPPL. 2). [CrossRef]
- Vidaillac, C.; Rybak, M. J. Ceftobiprole: First Cephalosporin with Activity against Methicillin-Resistant Staphylococcus Aureus. Pharmacotherapy2009, 29 (5), 511–525. [CrossRef]
- Nigo, M.; Lorena Diaz, Lina P. Carvajal, Truc T. Tran, R. R.; Panesso, D.; Garavito, J. D.; Miller, W. R.; Wanger, A.; George Weinstock; Munita, J. M.; Arias, C. A.; (Commentator), H. F. C. Ceftaroline-Resistant, Daptomycin- Tolerant, and Heterogeneous Vancomycin-Intermediate Methicillin- Resistant Staphylococcus Aureus Causing Infective Endocarditis. Antimicrobial Agents and Chemotherapy2017, 61 (3), 1–7.
- Miller, K.; Storey, C.; Stubbings, W. J.; Hoyle, A. M.; Hobbs, J. K.; Chopra, I. Antistaphylococcal Activity of the Novel Cephalosporin. 2005, No. February, 579–582. [CrossRef]
- Yadav, G.; Ganguly, S. Structure Activity Relationship (SAR) Study of Benzimidazole Scaffold for Different Biological Activities: A Mini-Review. European Journal of Medicinal Chemistry2015, 97 (1), 419–443. [CrossRef]
- Malasala, S.; Ahmad, N.; Akunuri, R.; Shukla, M. Jo Ur Na l P Re Of. European Journal of Medicinal Chemistry2020, 112996. [CrossRef]
- Tunçbilek, M.; Kiper, T.; Altanlar, N. Synthesis and in Vitro Antimicrobial Activity of Some Novel Substituted Benzimidazole Derivatives Having Potent Activity against MRSA. European Journal of Medicinal Chemistry2009, 44 (3), 1024–1033. [CrossRef]
- Shalaby, M.-A.; Dokla, E.; Serya, R.; Abouzid, K. Identification of Novel Pyrazole and Benzimidazole Based Derivatives as PBP2a Inhibitors: Design, Synthesis, and Biological Evaluation. Archives of Pharmaceutical Sciences Ain Shams University2019, 3 (2), 228–245. [CrossRef]
- O’Daniel, P. I.; Peng, Z.; Pi, H.; Testero, S. A.; Ding, D.; Spink, E.; Leemans, E.; Boudreau, M. A.; Yamaguchi, T.; Schroeder, V. A.; Wolter, W. R.; Llarrull, L. I.; Song, W.; Lastochkin, E.; Kumarasiri, M.; Antunes, N. T.; Espahbodi, M.; Lichtenwalter, K.; Suckow, M. A.; Vakulenko, S.; Mobashery, S.; Chang, M. Discovery of a New Class of Non-β-Lactam Inhibitors of Penicillin-Binding Proteins with Gram-Positive Antibacterial Activity. J Am Chem Soc2014, 136 (9), 3664–3672. [CrossRef]
- Buommino, E.; Marino, S. De; Sciarretta, M.; Auria, M. V. D.; Festa, C. With Oxacillin against Methicillin-Resistant Staphylococcus Aureus. 2021, 1–16.
- Verma, S. K.; Verma, R.; Kumar, K. S. S.; Banjare, L.; Shaik, A. B.; Bhandare, R. R.; Rakesh, K. P.; Rangappa, K. S. A Key Review on Oxadiazole Analogs as Potential Methicillin-Resistant Staphylococcus Aureus (MRSA) Activity: Structure-Activity Relationship Studies. European Journal of Medicinal Chemistry2021, 219, 113442. [CrossRef]
- Shalaby, M. A. W.; Dokla, E. M. E.; Serya, R. A. T.; Abouzid, K. A. M. Penicillin Binding Protein 2a: An Overview and a Medicinal Chemistry Perspective. European Journal of Medicinal Chemistry2020, 199, 112312. [CrossRef]
- Ibrahim, M. A. A.; Abdeljawaad, K. A. A.; Abdelrahman, A. H. M.; Alzahrani, O. R.; Alshabrmi, F. M.; Khalaf, E.; Moustafa, M. F.; Alrumaihi, F.; Allemailem, K. S.; Soliman, M. E. S.; Paré, P. W.; Hegazy, M. E. F.; Atia, M. A. M. Non-β-Lactam Allosteric Inhibitors Target Methicillin-Resistant Staphylococcus Aureus: An in Silico Drug Discovery Study. Antibiotics2021, 10 (8). [CrossRef]
- Qureshi, S. I.; Chaudhari, H. K. Design, Synthesis, in-Silico Studies and Biological Screening of Quinazolinone Analogues as Potential Antibacterial Agents against MRSA. Bioorganic and Medicinal Chemistry2019, 27 (12), 2676–2688. [CrossRef]
- Soc, J. A. C. HHS Public Access. 2015, 137 (5), 1738–1741. [CrossRef]
- Gunasekharan, M.; Choi, T. I.; Rukayadi, Y.; Latif, M. A. M.; Karunakaran, T.; Faudzi, S. M. M.; Kim, C. H. Preliminary Insight of Pyrrolylated-Chalcones as New Anti-Methicillin-Resistant Staphylococcus Aureus (Anti-Mrsa) Agents. Molecules2021, 26 (17). [CrossRef]
- Chem, M.; Marchand-brynaert, J. MedChemComm. 2012, No. 3, 344–351. [CrossRef]
- Nanjundaswamy, S.; Bindhu, S.; Renganathan, R. R. A.; Nagashree, S.; Mallu, P.; Rai, V. R.; Bindhu, S.; Renganathan, R. R. A.; Nagashree, S. Design , Synthesis of Pyridine Coupled Pyrimidinone / Pyrimidinthione as Anti-MRSA Agent : Validation by Molecular Docking and Dynamics Simulation. Journal of Biomolecular Structure and Dynamics2021, 0 (0), 1–12. [CrossRef]
- Zhang, P.; Gopala, L.; Zhang, S.; Cai, G. European Journal of Medicinal Chemistry An Unanticipated Discovery towards Novel Naphthalimide Corbelled Aminothiazoximes as Potential Anti-MRSA Agents and Allosteric Modulators for PBP2a. European Journal of Medicinal Chemistry2022, 229, 114050. [CrossRef]
- Shalaby, M. A. W.; Dokla, E. M. E.; Serya, R. A. T.; Abouzid, K. A. M. Penicillin Binding Protein 2a: An Overview and a Medicinal Chemistry Perspective. European Journal of Medicinal Chemistry2020, 199, 112312. [CrossRef]
- Kurazono, M.; Ida, T.; Yamada, K.; Hirai, Y.; Maruyama, T.; Shitara, E.; Yonezawa, M. In Vitro Activities of ME1036 ( CP5609 ), a Novel Parenteral Carbapenem , against Methicillin-Resistant Staphylococci. 2004, 48 (8), 2831–2837. [CrossRef]
- Toney, J. H.; Hammond, G. G.; Leiting, B.; Pryor, K. A. D.; Wu, J. K.; Cuca, G. C.; Pompliano, D. L. Soluble Penicillin-Binding Protein 2a: β-Lactam Binding and Inhibition by Non-β-Lactams Using a 96-Well Format. Analytical Biochemistry1998, 255 (1), 113–119. [CrossRef]
- Manuscript, A. NIH Public Access. 2013, 8 (9), 1095–1116. [CrossRef]
- Hao, H.; Cheng, G.; Dai, M.; Wu, Q.; Yuan, Z. Inhibitors Targeting on Cell Wall Biosynthesis Pathway of MRSA. Molecular BioSystems2012, 8 (11), 2828–2838. [CrossRef]
- Shiota, S.; Shimizu, M.; Sugiyama, J.; Morita, Y.; Mizushima, T.; Tsuchiya, T. Mechanisms of Action of Corilagin and Tellimagrandin I That Remarkably Potentiate the Activity of  -Lactams against Methicillin-Resistant Staphylococcus Aureus. 2004, 48 (1), 67–73. [CrossRef]
- Erol, M.; Celik, I.; Temiz-Arpaci, O.; Kaynak-Onurdag, F.; Okten, S. Design, Synthesis, Molecular Docking, Density Functional Theory and Antimicrobial Studies of Some Novel Benzoxazole Derivatives as Structural Bioisosteres of Nucleotides. Journal of Biomolecular Structure and Dynamics2021, 39 (9), 3080–3091. [CrossRef]
- Karthik, C. S.; Ananda, A. P.; Nagashree, S.; Mallu, P.; Rai, V. R. Design , Synthesis , and in-Silico Studies of Pyrazolylpyridine Analogues : A Futuristic Antibacterial Contender against Coagulase Positive. Journal of Molecular Structure2022, 1255, 132400. [CrossRef]
- Sangeetha, M.; Saranya, T. S.; Sathianarayanan, S.; Hima Vyshnavi, A. M.; Krishnan Namboori, P. K. Design and Development of Potential Flavonoid Moiety for Pbp2a Inhibition for Mrsa Therapy-A Computational Technique. Biomedical and Pharmacology Journal2020, 13 (2), 687–692. 2020; 13, 687–692. [CrossRef]
- Alhadrami, H. A.; Hamed, A. A.; Hassan, H. M.; Belbahri, L.; Rateb, M. E.; Sayed, A. M. Flavonoids as Potential Anti-MRSA Agents through Modulation of PBP2A: A Computational and Experimental Study. Antibiotics2020, 9 (9), 1–16. [CrossRef]
- Li, Q. Q.; Kang, O. H.; Kwon, D. Y. Study on Demethoxycurcumin as a Promising Approach to Reverse Methicillin-Resistance of Staphylococcus Aureus. International Journal of Molecular Sciences2021, 22 (7). [CrossRef]
- Neto, D. A.; Ramos, A.; Max, W.; Silva, J.; Silva, E.; Coelho, B.; Odorico, M.; Moraes, D. Microbial Pathogenesis Anti-MRSA Activity of Curcumin in Planktonic Cells and Biofilms and Determination of Possible Action Mechanisms. 2021, 155 (March). [CrossRef]
- Rani, N.; Saravanan, V.; Lakshmi, P. T. V. Allosteric Site Mediated Active Site Inhibition of PBP2a Using Quercetin 3-O-Rutinoside and Its Combination. 2015, 1102 (September). [CrossRef]
- Rani, N.; Vijayakumar, S.; Thanga Velan, L. P.; Arunachalam, A. Quercetin 3-O-Rutinoside Mediated Inhibition of PBP2a: Computational and Experimental Evidence to Its Anti-MRSA Activity. Molecular BioSystems2014, 10 (12), 3229–3237. [CrossRef]
- Zhou, T.; Li, Z. H. I.; Kang, O. K. H. W. A.; Mun, S. U. H.; Seo, Y. U. N. S. O. O.; Kong, R. Antimicrobial Activity and Synergism of Ursolic Acid 3-O- α -L-Arabinopyranoside with Oxacillin against Methicillin-Resistant Staphylococcus Aureus. 2017, 1285–1293. [CrossRef]
- Tozar, T.; Costa, S. S.; Udrea, A. M.; Nastasa, V.; Couto, I.; Viveiros, M.; Pascu, M. L.; Romanitan, M. O. Anti - Staphylococcal Activity and Mode of Action of Thioridazine Photoproducts. Scientific Reports2020, 1–12. [CrossRef]
- Negi, B.; Kumar, D.; Kumbukgolla, W.; Jayaweera, S.; Ponnan, P.; Singh, R.; Agarwal, S.; Rawat, D. S. Anti-Methicillin Resistant Staphylococcus Aureus Activity, Synergism with Oxacillin and Molecular Docking Studies of Metronidazole-Triazole Hybrids. European Journal of Medicinal Chemistry2016, 115, 426e–4437. [CrossRef]
- Qiao, Y.; Zhang, X.; He, Y.; Sun, W.; Feng, W.; Liu, J.; Hu, Z.; Xu, Q.; Zhu, H.; Zhang, J.; Luo, Z.; Wang, J.; Xue, Y.; Zhang, Y. Aspermerodione, a Novel Fungal Metabolite with an Unusual 2,6-Dioxabicyclo[2.2.1]Heptane Skeleton, as an Inhibitor of Penicillin-Binding Protein 2a. Scientific Reports2018, 8 (1), 2–12. [CrossRef]
- Li, Z. Z.; Tangadanchu, V. K. R.; Battini, N.; Bheemanaboina, R. R. Y.; Zang, Z. L.; Zhang, S. L.; Zhou, C. H. Indole-Nitroimidazole Conjugates as Efficient Manipulators to Decrease the Genes Expression of Methicillin-Resistant Staphylococcus Aureus. European Journal of Medicinal Chemistry2019, 179, 723–735. [CrossRef]
- Chang, Y. W.; Huang, W. C.; Lin, C. Y.; Wang, W. H.; Hung, L. C.; Chen, Y. H. Tellimagrandin Ii, a Type of Plant Polyphenol Extracted from Trapa Bispinosa Inhibits Antibiotic Resistance of Drug-Resistant Staphylococcus Aureus. International Journal of Molecular Sciences2019, 20 (22), 1–17. [CrossRef]
- Chowdhury, A.; Paul, P.; Bhattacharjee, A.; Talukdar, A. Das; Choudhury, M. D. Virtual High Throughput Screening of Carbapenem Derivatives as New Generation Carbapenemase and Penicillin Binding Protein Inhibitors : A Hunt to Save Drug of Last Resort. 2015, 18–23. [CrossRef]
- Lian, X.; Xia, Z.; Li, X.; Karpov, P.; Jin, H.; Tetko, I. V; Xia, J.; Wu, S. Bioorganic Chemistry Anti-MRSA Drug Discovery by Ligand-Based Virtual Screening and Biological Evaluation. Bioorganic Chemistry2021, 114 (2), 105042. 2021; 114. [CrossRef]
- Skariyachan, S.; Krishnan, R. S.; Siddapa, S. B.; Salian, C.; Bora, P.; Sebastian, D. Computer Aided Screening and Evaluation of Herbal Therapeutics against MRSA Infections. Bioinformation2011, 7 (5), 222–233. [CrossRef]
- Chang, M.; Mahasenan, K. V; Hermoso, J. A.; Mobashery, S. Unconventional Antibacterials and Adjuvants. 2021. [CrossRef]
- Abedi, F.; Balmeh, N.; Sanjari, S.; Plantago-, A. Informatics in Medicine Unlocked In-Silico Investigation of Antibacterial Herbal Compounds in Order to Find New Antibiotic against Staphylococcus Aureus and Its Resistant Subtypes ☆. Informatics in Medicine Unlocked2022, 28, 100843. [CrossRef]
- Davies, D. R. Chapter 23 Screening Ligands by X-Ray Crystallography. 2014, 1140, 315–323. [CrossRef]
- Fishovitz, J.; Rojas-altuve, A.; Otero, L. H.; Dawley, M.; Carrasco-lo, C.; Chang, M.; Hermoso, J. A.; Mobashery, S. Of Resistance to Antibiotics. 2014, 1–4.
- Articles, O. Synthesis and Antibacterial Activity of Novel Icariin Derivatives. 2019, 74 (ii), 73–78. [CrossRef]
- Mart, A.; Pascual, M.; Sheth, C. C.; Veses, V. Icariin in Combination with Amoxycillin-Clavulanate and Ampicillin , but Not Vancomycin , Increases Antibiotic Sensitivity and Growth Inhibition against Methicillin-Resistant Staphylococcus Aureus. 2022.
- Marchand-brynaert, J.; Urbach, A.; Dive, G.; Tinant, B. European Journal of Medicinal Chemistry Large Ring 1 , 3-Bridged 2-Azetidinones : Experimental and Theoretical Studies. 2009, 44, 2071–2080. [CrossRef]
- Cheung, G. Y. C.; Bae, J. S.; Otto, M. Pathogenicity and Virulence of Staphylococcus Aureus ABSTRACT. Virulence2021, 12 (1), 547–569. [CrossRef]
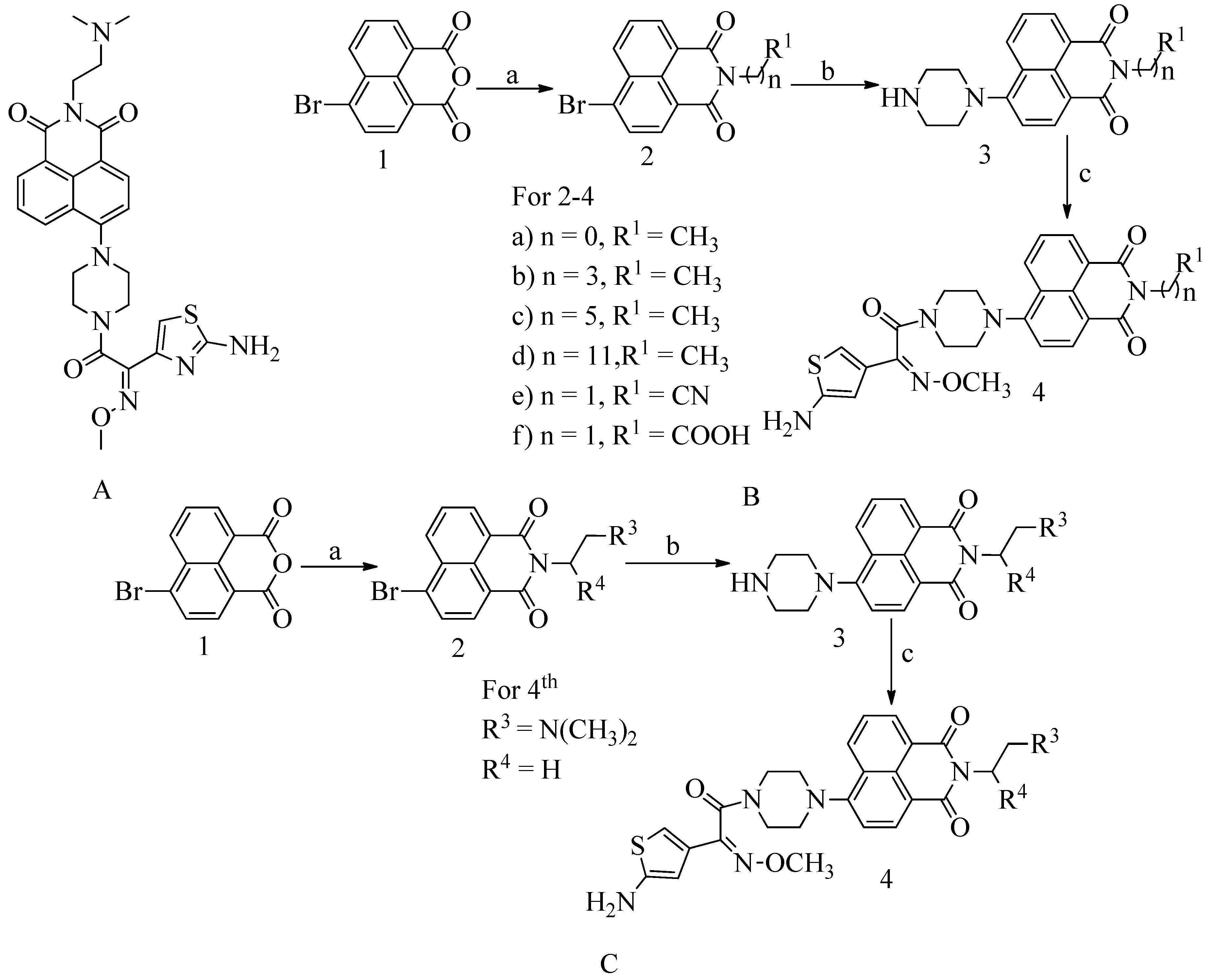
| Sr. No | Name of genes | Role in MRSA |
|---|---|---|
| s1 | Mec (mecA, mecR1-mecIs) | The close relationship between two regulatory elements, one is mec and the second one is bla. BlaR2 or a close relative may be participating in mecA activation here too. |
| 2 | Chromosomal gene | FEM means factors essential for methicillin resistance. Its inactivation reduces methicillin-resistant and the aux (auxiliary) factors usually present in staphylococcal aureus. The majority of elements are engaged in cell wall biosynthesis and some turnover of bacteria. |
| 3 | The PBP2a operon in normal S. aureus contributes to resistance | The mecA is responsible for the production of PBP2a, which helps in the biosynthesis of bacteria's cell walls. However, its role in the resistance of S. aureus is still unclear. |
| 4 | Fmt | Reduce methicillin resistance. |
| 5 | femX, femAB | When reducing the length of the glycine side chain occurs, it is because of the inactivation of the femAB factor. This inactivation and reduced length lead to impairment in the growth of peptidoglycan and also the turnover of cell-wall being reduced. All these reactions are responsible for lowering peptidoglycan cross-linking and hypersusceptibility to all β-lactam antibiotics and other antibiotics, which leads to the decrease in methicillin resistance. |
| 6 |
glnRA (femC) (Mutated gene) |
The precursors of muropeptide use an amino acid which is a non-amidated D-glutamate amino acid, and it takes part in the stem peptide of the transpeptidation reactions less readily, which leads to a reduction of methicillin resistance |
| 7 | glmM (femD, femR) | When the rate of precursor formation for peptidoglycan is reduced, it reduces methicillin resistance. It increases teicoplanin susceptibility and thus decreases methicillin resistance in MRSA. |
| 8 | Lytic enzymes (Hydrolysis Enzyme) |
Murein, which is present in Staphylococcus aureus, causes hydrolysis. It is then needed for the growth of peptidoglycan, which is a precious cell wall content. This play one of the leading roles in peptidoglycan growth. |
| 9 | murE (femF) | In the formation of peptidoglycan precursors, MurE necessitates the presence of this factor. Because the abnormal precursors of peptidoglycan are present, which causes a decline in methicillin resistance, it is also possible that there may be abnormal shortening precursors involved in the peptidoglycan biosynthesis. It is still unclear. |
| 10 | Llm gene | The llm gene encodes lipophilic membrane proteins, affecting methicillin resistance levels and necessarily causing bacterial cell lysis rate reduction. Its functions, however, are still unclear. |
| 11 | Global regulators: sar, agr, sigB | The global regulators like sar and agr control cell density-dependent synthesis of cell wall factors that are cell wall-associated and extracellular virulence factors. In heterogeneous MRSA, this global regulator appears to have just a slight effect on methicillin resistance in MRSA. |
| 12 | ctaA | Reduced the resistance to methicillin in MRSA |
| 13 | Blaz | The blaZ gene has been mutated in Staphylococcus aureus, and it is responsible for the breakdown of the β-lactam ring of mainly penicillin antibiotics. |
| 14 | blaI | blaI is a repressor protein that is responsible for preventing the transcription of mutated genes like blaZ or mecA. |
| 15 | blaR1 | blaR1 is a transmembrane protein that signals blaZor mecA transcription. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




