Submitted:
03 September 2023
Posted:
05 September 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
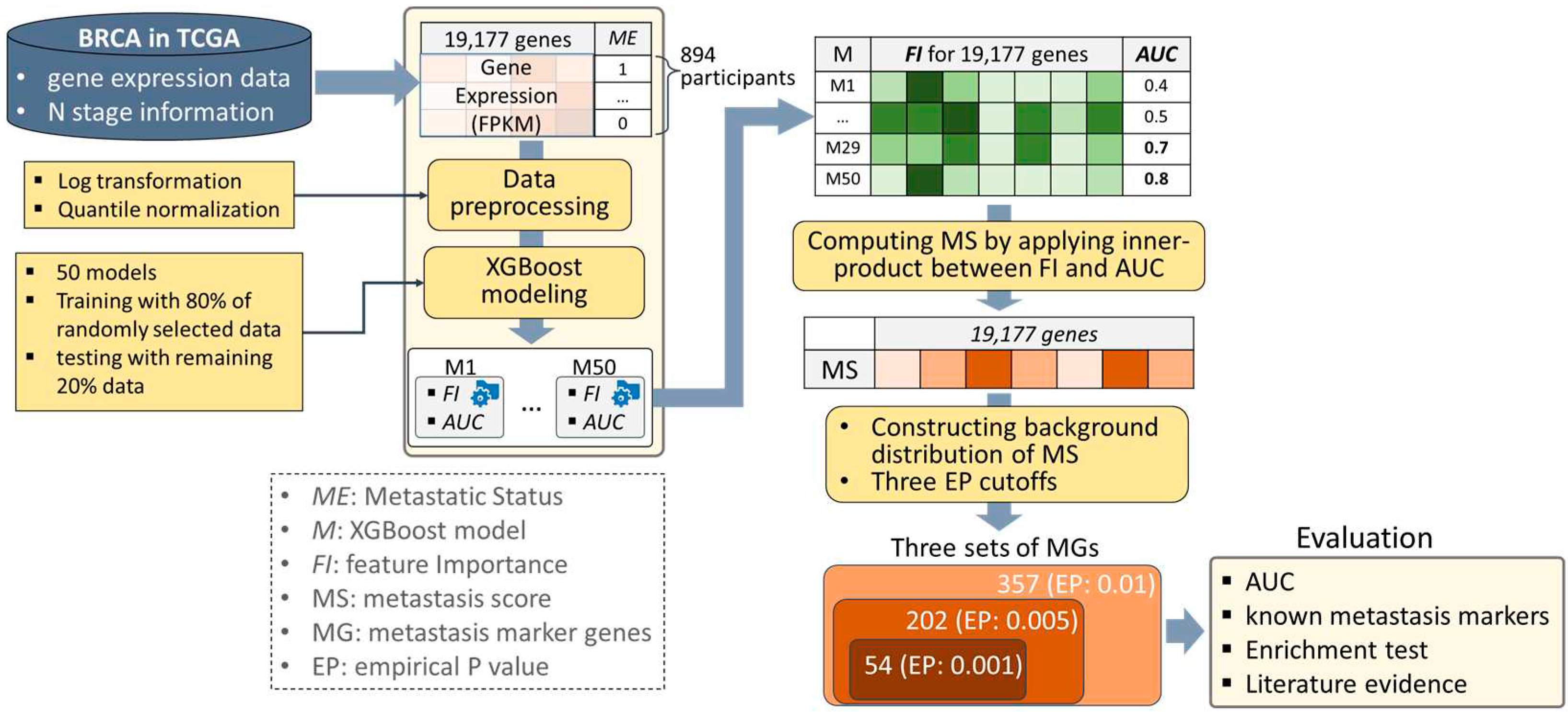
2. Materials and Methods
2.1. Data Preparation
2.2. Data Preprocessing
2.3. XGBoost Modeling
2.4. Characterizing Metastasis Marker Genes
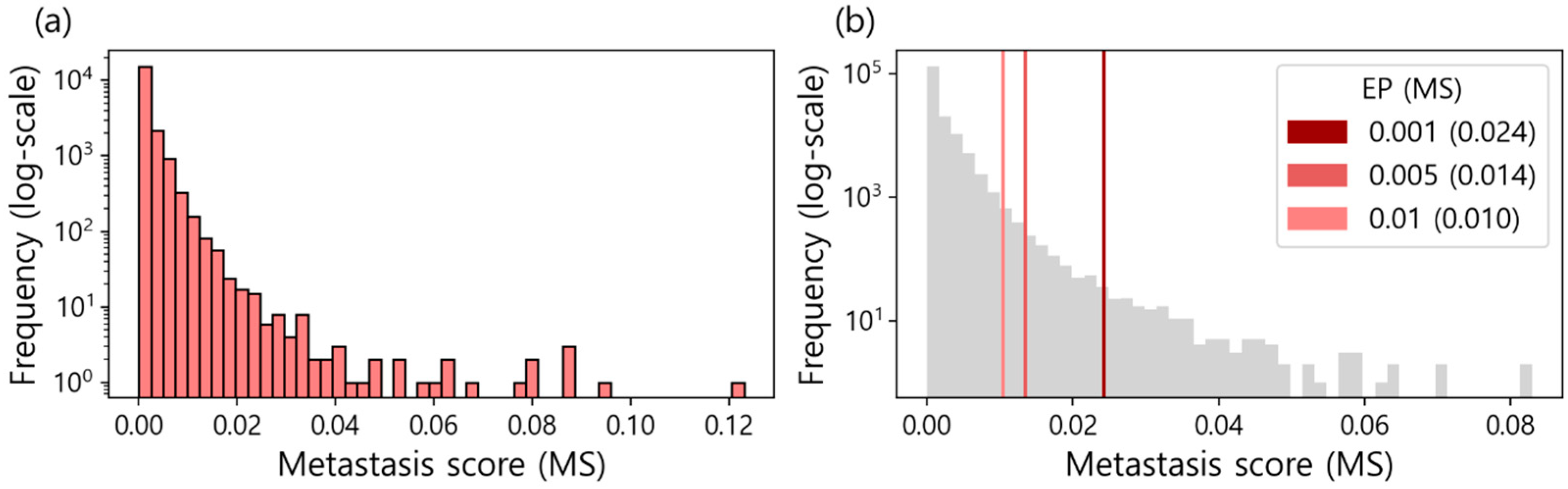
3. Results and Evaluations
3.1. Metastasis Marker Genes
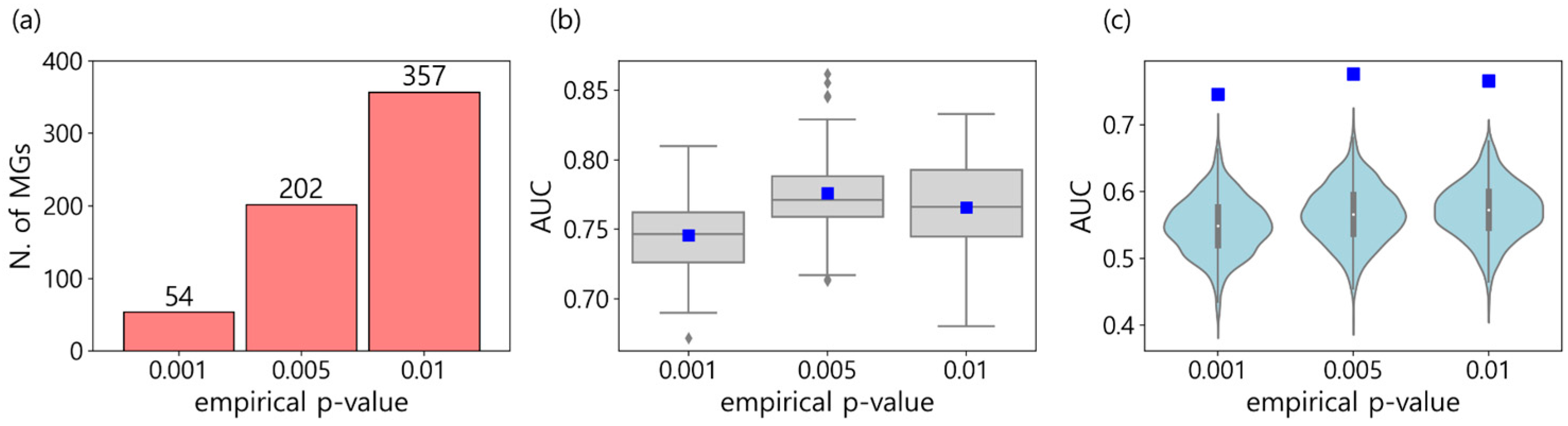
3.2. Comparing with known metastasis markers
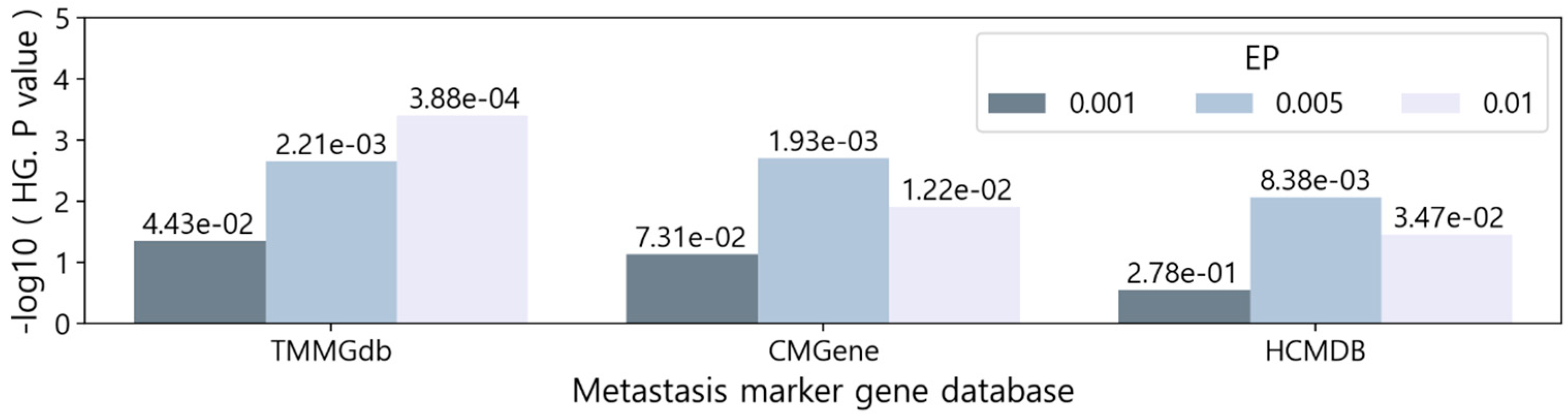
3.3. Enrichment tests on metastasis-related processes
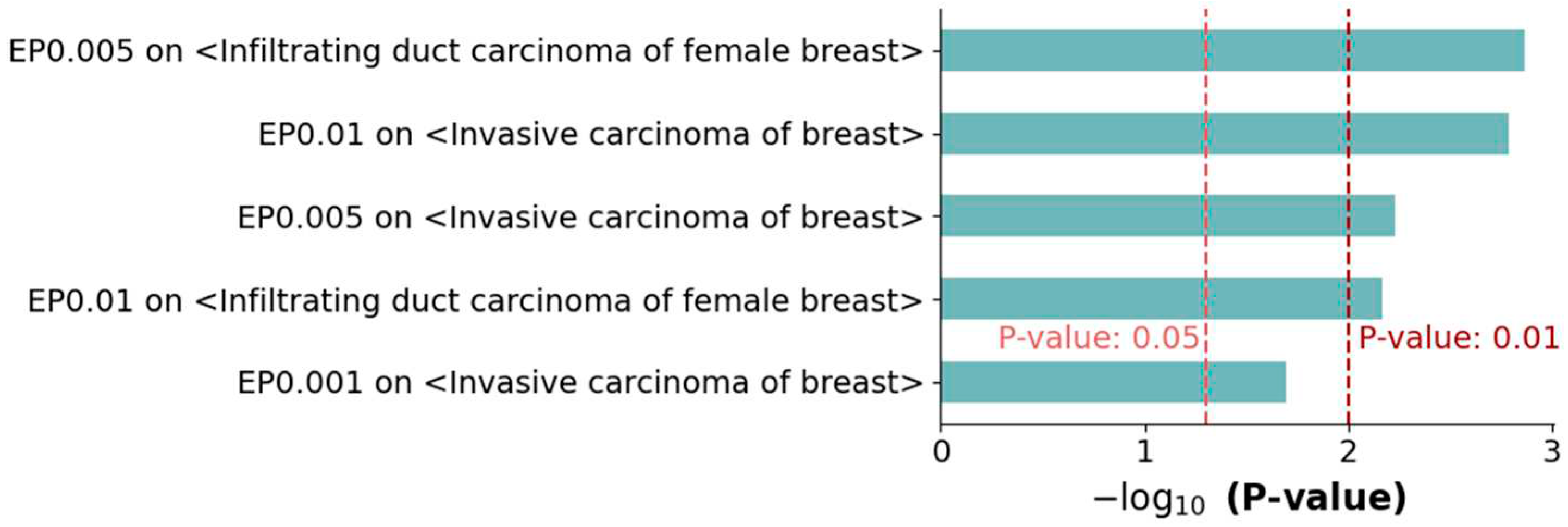
3.4. Literature Evidence
3.4.1. Metastasis marker genes with the highest Metastasis score
3.4.2. Metastasis marker genes not identified by statistical analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Dillekas, H., M.S. Rogers, and O. Straume, Are 90% of deaths from cancer caused by metastases? Cancer Med, 2019. 8(12): p. 5574–5576. [CrossRef]
- Guan, X., Cancer metastases: Challenges and opportunities. Acta Pharm Sin B, 2015. 5(5): p. 402–18. [CrossRef]
- Albaradei, S., et al., Machine learning and deep learning methods that use omics data for metastasis prediction. Comput. Struct. Biotechnol. J., 2021. 19: p. 5008–5018. [CrossRef]
- Chen, C., et al., Screening and evaluation of the role of immune genes of brain metastasis in lung adenocarcinoma progression based on the TCGA and GEO databases. J Thorac Dis, 2021. 13(8): p. 5016–5034. [CrossRef]
- Kim, G.E., et al., Differentially expressed genes in matched normal, cancer, and lymph node metastases predict clinical outcomes in patients with breast cancer. Appl Immunohistochem Mol Morphol, 2020. 28(2): p. 111–122. [CrossRef]
- Wei, W., et al., Identification of key genes involved in the metastasis of clear cell renal cell carcinoma. Oncol Lett, 2019. 17(5): p. 4321–4328. [CrossRef]
- Metri, R., et al., Identification of a gene signature for discriminating metastatic from primary melanoma using a molecular interaction network approach. Sci Rep, 2017. 7(1): p. 17314. [CrossRef]
- Wei, D., A multigene support vector machine predictor for metastasis of cutaneous melanoma. Mol Med Rep, 2018. 17(2): p. 2907–2914. [CrossRef]
- Burton, M., et al., Gene expression profiles for predicting metastasis in breast cancer: A cross-study comparison of classification methods. Sci. World J., 2012. 2012: p. 380495. [CrossRef]
- Tamar, G. and T. Vasil, THE BURDEN OF BREAST CANCER IN TBILISI IN 2015-2019. European journal of biomedical and life sciences, 2021(4): p. 27-33.
- Tomczak, K., P. Czerwińska, and M. Wiznerowicz, Review the cancer genome atlas (TCGA): An immeasurable source of knowledge. Contemp. Oncol., 2015. 2015(1): p. 68–77. [CrossRef]
- Liu, J., et al., An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell, 2018. 173(2): p. 400–416.e11.
- Colaprico, A., et al., TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic acids research, 2016. 44(8): p. e71-e71. [CrossRef]
- Abawajy, J., A. Darem, and A.A. Alhashmi, Feature subset selection for malware detection in smart IoT platforms. Sensors (Basel), 2021. 21(4): p. 1374. [CrossRef]
- Hughes, G., On the mean accuracy of statistical pattern recognizers. IEEE Trans. Inf. Theory, 1968. 14(1): p. 55–63. [CrossRef]
- Barretina, J., et al., The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature, 2012. 483(7391): p. 603-607. [CrossRef]
- Li, Y., et al., Putative biomarkers for predicting tumor sample purity based on gene expression data. BMC Genom., 2019. 20(1): p. 1021. [CrossRef]
- Pellegrino, E., et al., Machine learning random forest for predicting oncosomatic variant NGS analysis. Sci Rep, 2021. 11(1): p. 21820. [CrossRef]
- Chen, T. and C. Guestrin, Xgboost: A scalable tree boosting system, in Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining. 2016, Association for Computing Machinery: San Francisco, California, USA. p. 785–794.
- Bradley, A.P., The use of the area under the ROC curve in the evaluation of machine learning algorithms. Pattern recognition, 1997. 30(7): p. 1145-1159. [CrossRef]
- Liu, H.-C., et al., TMMGdb-Tumor Metastasis Mechanism-associated Gene Database. Current Bioinformatics, 2023. 18(1): p. 63-75. [CrossRef]
- Liu, Y., et al., CMGene: A literature-based database and knowledge resource for cancer metastasis genes. Journal of Genetics and Genomics, 2017. 44(5): p. 277-279. [CrossRef]
- Zheng, G., et al., HCMDB: The human cancer metastasis database. Nucleic Acids Res, 2018. 46(D1): p. D950–955. [CrossRef]
- Piñero, J., et al., The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res, 2020. 48(D1): p. D845–855. [CrossRef]
- Papatsirou, M., et al., Identification of novel circular RNAs of the human protein arginine methyltransferase 1 (PRMT1) gene, expressed in breast cancer cells. Genes, 2022. 13(7): p. 1133. [CrossRef]
- Vasudevan, S.A., et al., Neuroblastoma-derived secretory protein is a novel secreted factor overexpressed in neuroblastoma. Molecular cancer therapeutics, 2009. 8(8): p. 2478-2489. [CrossRef]
- Keenan, A.B., et al., ChEA3: transcription factor enrichment analysis by orthogonal omics integration. Nucleic acids research, 2019. 47(W1): p. W212-W224. [CrossRef]
- Yong, B.-C., et al., LDOC1 regulates Wnt5a expression and osteosarcoma cell metastasis and is correlated with the survival of osteosarcoma patients. Tumor Biology, 2017. 39(2): p. 1010428317691188. [CrossRef]
- Meyer-Schaller, N., et al., A dual role of Irf1 in maintaining epithelial identity but also enabling EMT and metastasis formation of breast cancer cells. Oncogene, 2020. 39(24): p. 4728-4740. [CrossRef]
- Maubant, S., et al., LRP5 regulates the expression of STK40, a new potential target in triple-negative breast cancers. Oncotarget, 2018. 9(32): p. 22586. [CrossRef]
- Zhang, R., et al., Golgi membrane protein 1 (GOLM1) promotes growth and metastasis of breast cancer cells via regulating matrix metalloproteinase-13 (MMP13). Medical science monitor: international medical journal of experimental and clinical research, 2019. 25: p. 847. [CrossRef]
- Chaudhary, S., et al., MUC16 promotes triple-negative breast cancer lung metastasis by modulating RNA-binding protein ELAVL1/HUR. Breast Cancer Research, 2023. 25(1): p. 1-15. [CrossRef]
- Zhao, Y., et al., A feedback loop comprising EGF/TGFα sustains TFCP2-mediated breast cancer progression. Cancer research, 2020. 80(11): p. 2217-2229. [CrossRef]
- Xu, M.-Y., et al., AZGP1 suppresses epithelial-to-mesenchymal transition and hepatic carcinogenesis by blocking TGFβ1-ERK2 pathways. Cancer letters, 2016. 374(2): p. 241-249. [CrossRef]
- Oughtred, R., et al., The BioGRID interaction database: 2019 update. Nucleic acids research, 2019. 47(D1): p. D529-D541. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



