Submitted:
22 July 2023
Posted:
25 July 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
ME/CFS Metabolomics Dataset
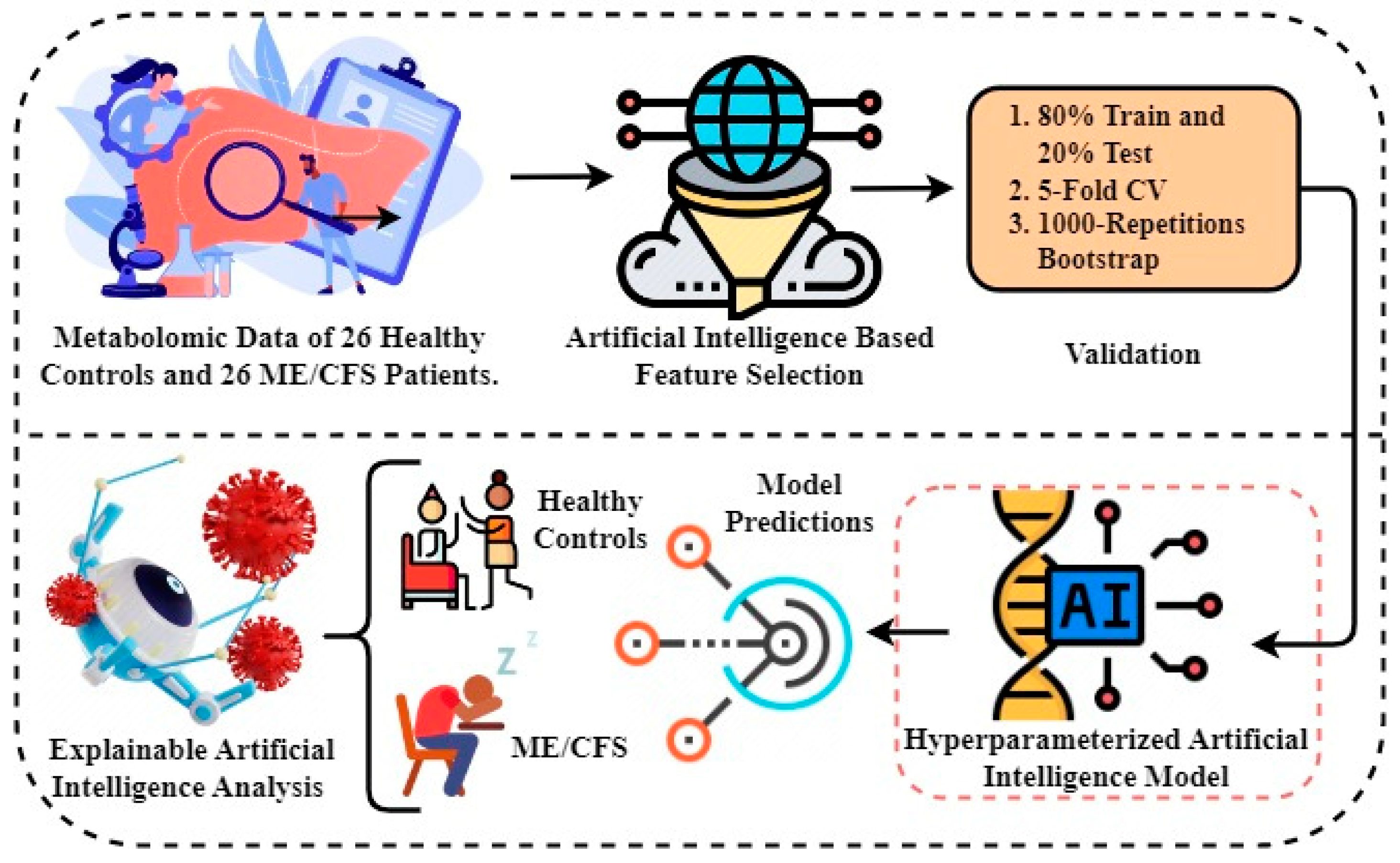
Experimental Setup and Proposed Framework
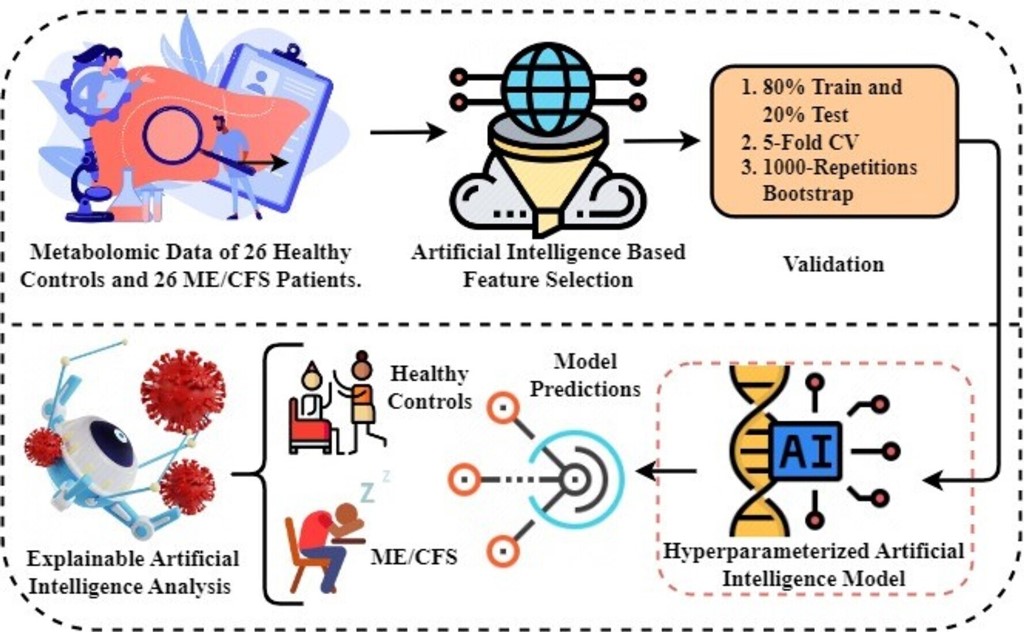
- The first step involves obtaining metabolomics data to be used in experiments. Metabolomics data are based on results from a study of 26 healthy controls and 26 ME/CFS patients aged 22 to 72 years with similar body mass index (BMI).
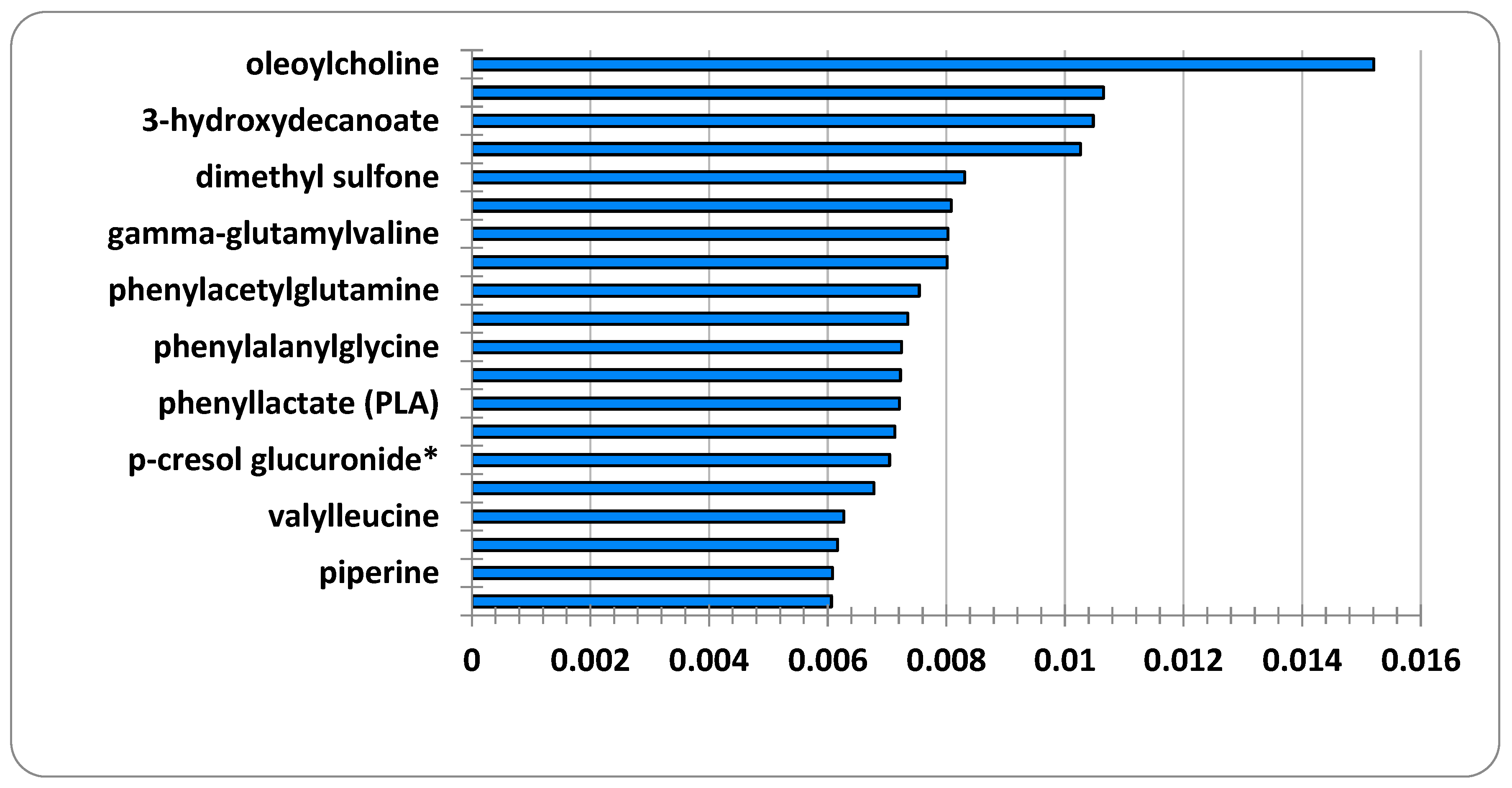
- In the second step, artificial intelligence-based random forest (RF) feature selection is applied to identify biomarker candidate metabolites and to eliminate the high dimensionality problem in omic data. Because the metabolomics data has a large number of feature dimensions, the performance scores of the predicted models may be lower. Therefore, the twenty most important metabolites contributing to improved performance scores in ME/CFS prediction were identified.
- In the third step, 80%-20% split, 5-fold cross-validation (CV), and 1000 replicates Bootstrap approaches were used to validate the prediction models to be generated using the selected biomarker candidate metabolites, and the results were compared.
- In the fourth step, Bayesian hyper-parameter optimization was used to determine the optimal parameters.
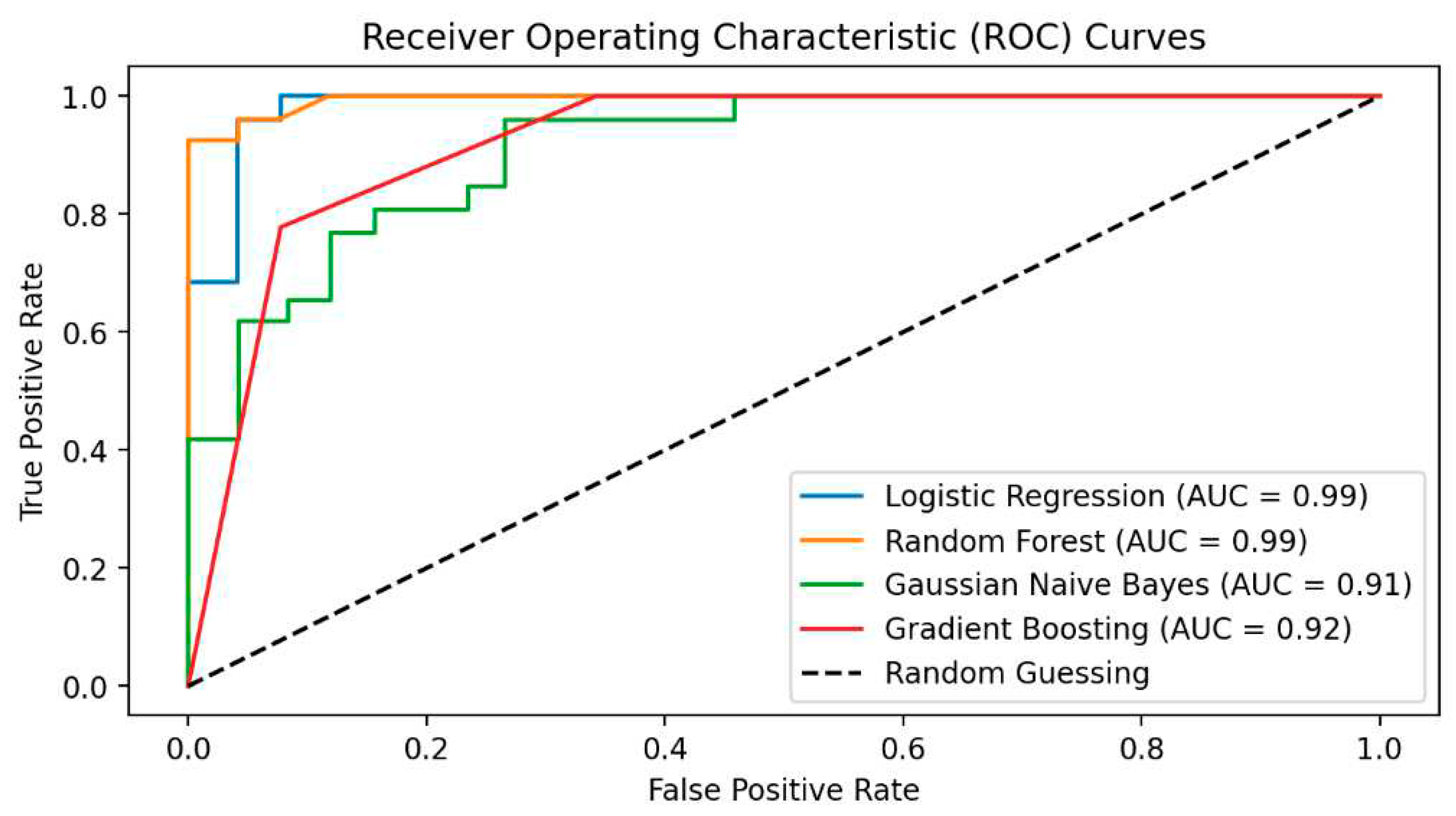
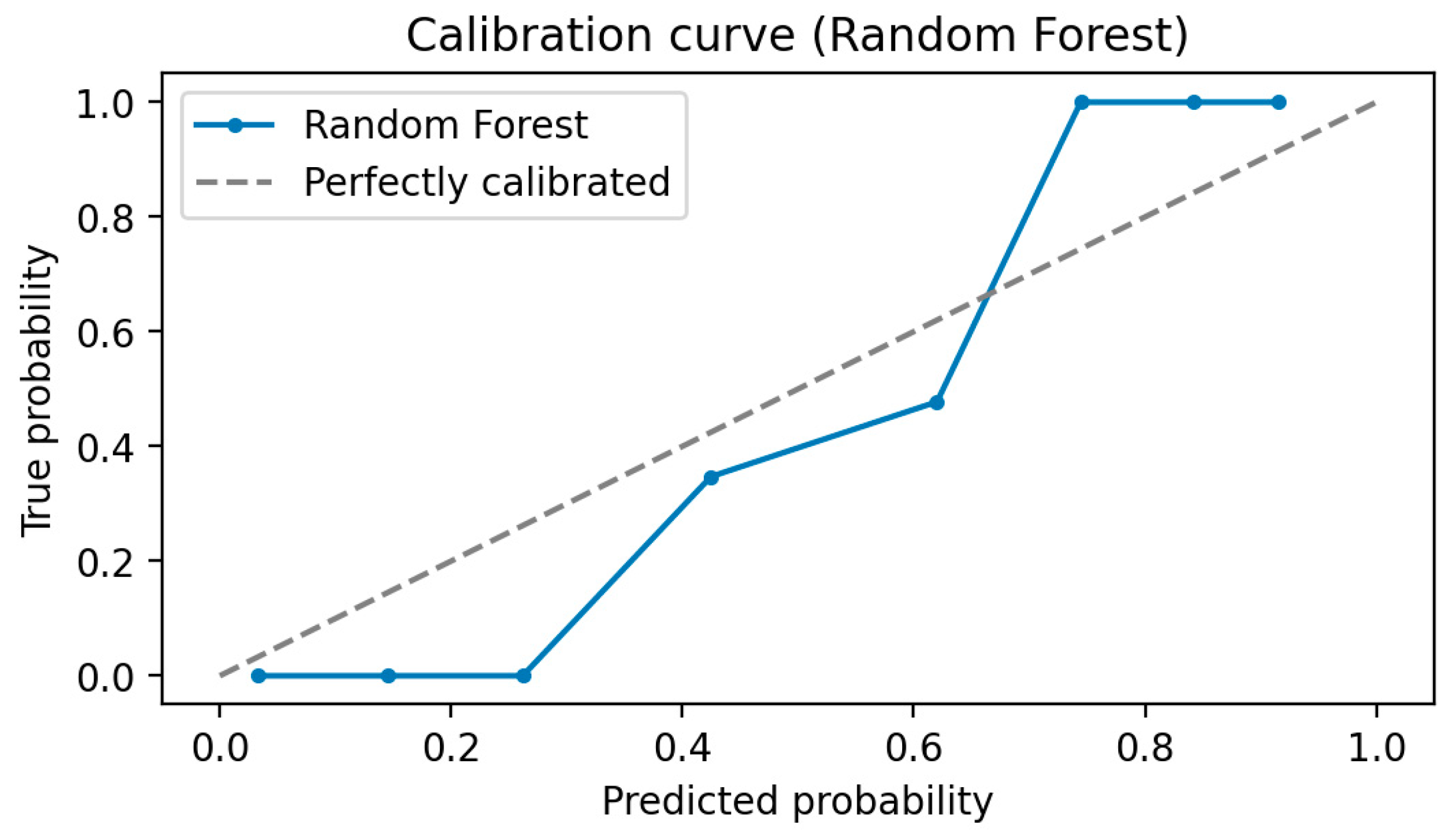
- In the fifth step, predictive models were built to diagnose ME/CFS patients. For this purpose, Gaussian Naive Bayes (GNB), Gradient Boosting Classifier (GBC), Logistic regression (LR), and Random Forest Classifier (RFC) algorithms were constructed. Performance of the models was evaluated via area under (AUC) Receiver operating characteristic (ROC) Curve, Brier score, accuracy, precision, recall, and F1-score. While the primary purpose of the methodology is biomarker discovery and diagnosis of ME/CFS, an important secondary purpose is to provide users with indicative probability scores. Therefore, we evaluated the quality of the probabilities via a calibration curve and by calculating the Brier score.
- Finally, XAI approaches were applied to the proposed model to provide transparency and interpretability to the model and to explain the decisions made by the model. Through the use of XAI, we can grasp both the rationale and the process behind a particular decision made by the proposed model.
Feature selection
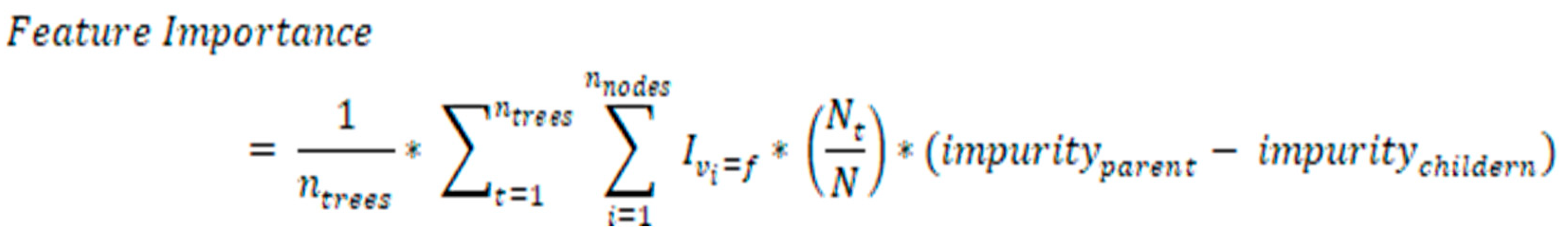
Validation Methods
The Bayesian Approach for hyper-parameter optimization
Classification models
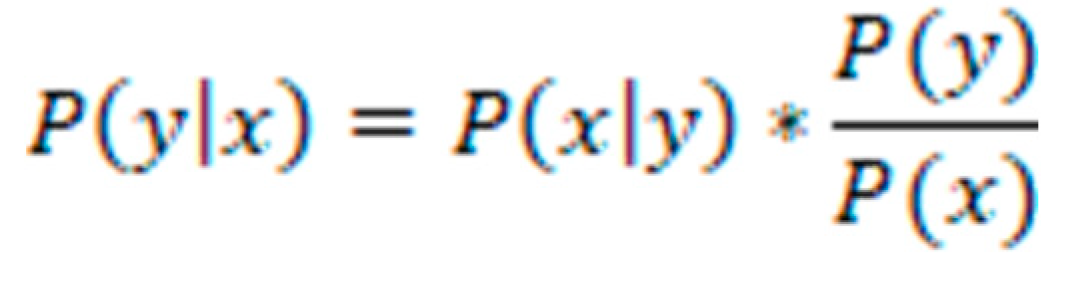
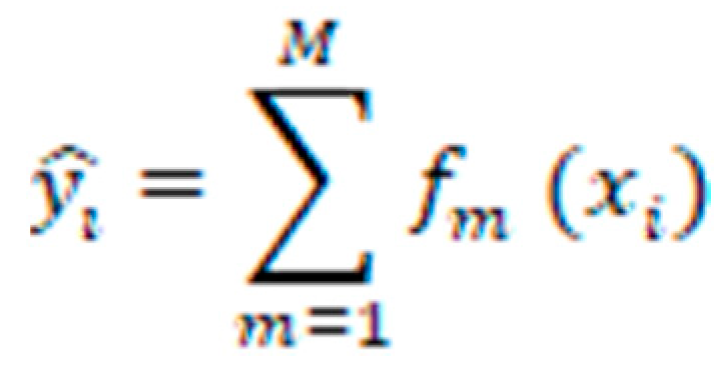
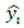 represents the predicted value for the i-th instance.
represents the predicted value for the i-th instance.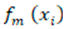 refers to the m-th weak classifier's prediction for the i-th instance.
refers to the m-th weak classifier's prediction for the i-th instance.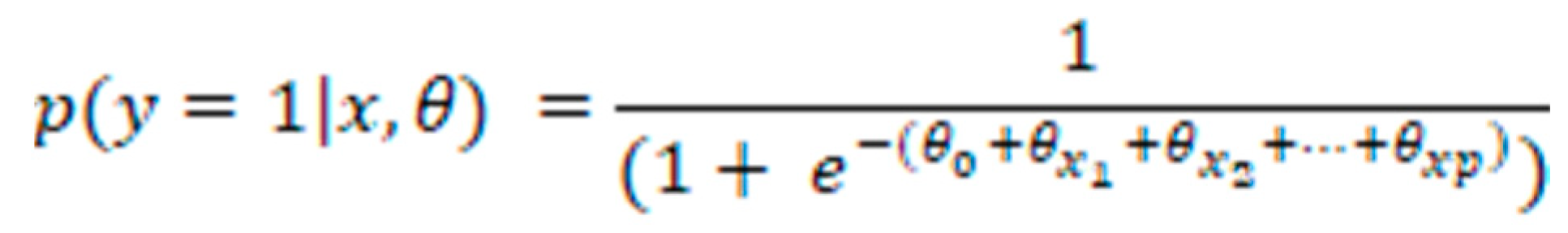
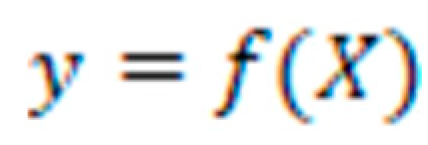
Performance Evaluation and Model Calibration
Performance Evaluation
Model Calibration
XAI Approach
3. Results
Feature Selection Results
Hyper-parameters Optimization Results
The Model Performance Results
| Attained performance using all input features | Attained performance using feature selection | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Technique | A (%) | P (%) | R (%) | F1 (%) | B | AUC (%) | A (%) | P (%) | R (%) | F1 (%) | B | AUC (%) |
| 80%-20% split validation | 80%-20% split validation | |||||||||||
| GNB | 73 | 72 | 73 | 72 | 0.27 | 67 | 73 | 72 | 73 | 72 | 0.27 | 67 |
| GBC | 36 | 39 | 36 | 37 | 0.63 | 33 | 73 | 75 | 73 | 73 | 0.27 | 73 |
| LR | 64 | 64 | 64 | 64 | 0.36 | 60 | 73 | 72 | 73 | 72 | 0.27 | 67 |
| RFC | 45 | 56 | 45 | 44 | 0.54 | 51 | 82 | 86 | 82 | 80 | 0.18 | 75 |
| Results with 5-folds cross validation | Results with 5-folds cross validation | |||||||||||
| GNB | 52 | 36 | 94 | 62 | 0.26 | 59 | 82 | 77 | 92 | 84 | 0.15 | 91 |
| GBC | 48 | 47 | 35 | 37 | 0.34 | 52 | 95 | 94 | 99 | 95 | 0.05 | 98 |
| LR | 58 | 46 | 71 | 54 | 0.45 | 46 | 95 | 95 | 96 | 96 | 0.03 | 98 |
| RFC | 56 | 68 | 38 | 56 | 0.28 | 64 | 97 | 96 | 97 | 98 | 0.04 | 99 |
| Results with 1000 repetition bootstrap | Results with 1000 repetition bootstrap | |||||||||||
| GNB | 63 | 70 | 63 | 60 | 0.36 | 63 | 83 | 84 | 83 | 83 | 0.17 | 91 |
| GBC | 92 | 92 | 92 | 92 | 0.07 | 92 | 96 | 96 | 96 | 96 | 0.03 | 92 |
| LR | 96 | 96 | 96 | 96 | 0.04 | 95 | 96 | 96 | 96 | 96 | 0.04 | 99 |
| RFC | 90 | 90 | 90 | 90 | 0.09 | 90 | 98 | 98 | 98 | 98 | 0.01 | 99 |
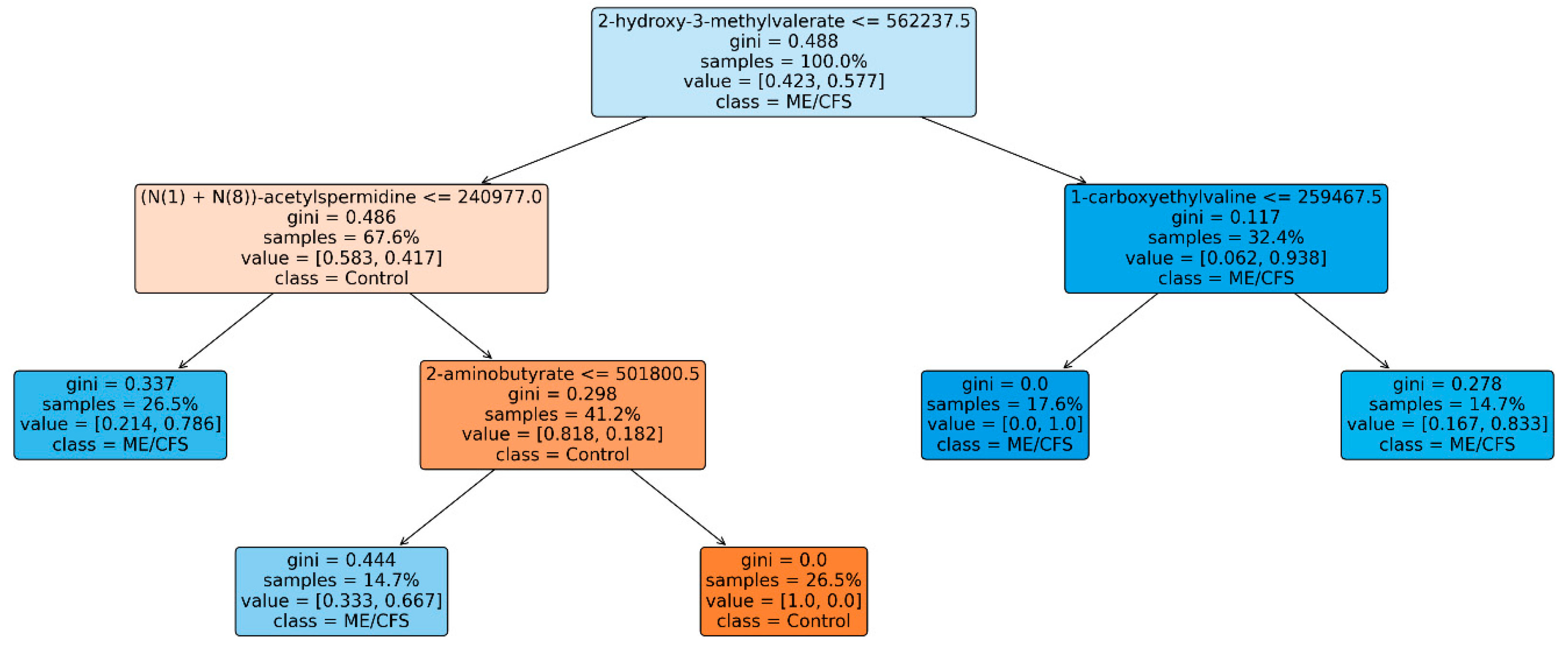
XAI Results
4. Discussion
5. Conclusions
6. Limitations and future works
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Toogood PL, Clauw DJ, Phadke S, Hoffman D. Myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS): Where will the drugs come from? Pharmacological Research. 2021, 165, 105465. [CrossRef]
- Germain A, Barupal DK, Levine SM, Hanson MR. Comprehensive circulatory metabolomics in ME/CFS reveals disrupted metabolism of acyl lipids and steroids. Metabolites. 2020, 10, 34. [CrossRef] [PubMed]
- Malato J, Graça L, Sepúlveda N. Impact of imperfect diagnosis in ME/CFS association studies. medRxiv. 2022, 2022.06. 08.22276100.
- Valdez AR, Hancock EE, Adebayo S, Kiernicki DJ, Proskauer D, Attewell JR, et al. Estimating prevalence, demographics, and costs of ME/CFS using large scale medical claims data and machine learning. Frontiers in pediatrics. 2019, 6, 412. [CrossRef]
- Faro M, Sàez-Francás N, Castro-Marrero J, Aliste L, de Sevilla TF, Alegre J. Gender differences in chronic fatigue syndrome. Reumatología clínica (English edition). 2016, 12, 72–77.
- Marshall-Gradisnik S, Eaton-Fitch N. Understanding myalgic encephalomyelitis. Science. 2022, 377, 1150–1151. [CrossRef]
- Malkova A, Shoenfeld Y. Autoimmune autonomic nervous system imbalance and conditions: Chronic fatigue syndrome, fibromyalgia, silicone breast implants, COVID and post-COVID syndrome, sick building syndrome, post-orthostatic tachycardia syndrome, autoimmune diseases and autoimmune/inflammatory syndrome induced by adjuvants. Autoimmunity reviews. 2022, 103230.
- Dehhaghi M, Panahi HKS, Kavyani B, Heng B, Tan V, Braidy N, et al. The role of kynurenine pathway and NAD+ metabolism in myalgic encephalomyelitis/chronic fatigue syndrome. Aging and disease. 2022, 13, 698. [CrossRef] [PubMed]
- Nunes JM, Kell DB, Pretorius E. Cardiovascular and haematological pathology in myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS): A role for viruses. Blood reviews. 2023, 101075.
- Hornig M, Montoya JG, Klimas NG, Levine S, Felsenstein D, Bateman L, et al. Distinct plasma immune signatures in ME/CFS are present early in the course of illness. Science advances. 2015, 1, e1400121. [CrossRef]
- Shan ZY, Barnden LR, Kwiatek RA, Bhuta S, Hermens DF, Lagopoulos J. Neuroimaging characteristics of myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS): a systematic review. Journal of translational medicine. 2020, 18, 1–11.
- Navaneetharaja N, Griffiths V, Wileman T, Carding SR. A role for the intestinal microbiota and virome in myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS)? Journal of clinical medicine. 2016, 5, 55. [CrossRef]
- Maes M, Leunis J-C, Geffard M, Berk M. Evidence for the existence of Myalgic Encephalomyelitis/Chronic Fatigue Syndrome (ME/CFS) with and without abdominal discomfort (irritable bowel) syndrome. Neuroendocrinol Lett. 2014, 35, 445–453.
- Germain A, Giloteaux L, Moore GE, Levine SM, Chia JK, Keller BA, et al. Plasma metabolomics reveals disrupted response and recovery following maximal exercise in myalgic encephalomyelitis/chronic fatigue syndrome. JCI insight. 2022, 7.
- Steyerberg EW, Harrell Jr FE, Borsboom GJ, Eijkemans M, Vergouwe Y, Habbema JDF. Internal validation of predictive models: efficiency of some procedures for logistic regression analysis. Journal of clinical epidemiology. 2001, 54, 774–781.
- Levman J, Ewenson B, Apaloo J, Berger D, Tyrrell PN. Error Consistency for Machine Learning Evaluation and Validation with Application to Biomedical Diagnostics. Diagnostics. 2023, 13, 1315. [CrossRef]
- Zhang X, Liu C-A. Model averaging prediction by K-fold cross-validation. Journal of Econometrics. 2023, 235, 280–301. [CrossRef]
- Diniz, MA. Statistical methods for validation of predictive models. Journal of Nuclear Cardiology. 2022, 29, 3248–3255. [Google Scholar] [CrossRef]
- Zhang J, Wang Q, Shen W. Hyper-parameter optimization of multiple machine learning algorithms for molecular property prediction using hyperopt library. Chinese Journal of Chemical Engineering. 2022, 52, 115–125. [CrossRef]
- Wu J, Chen X-Y, Zhang H, Xiong L-D, Lei H, Deng S-H. Hyperparameter optimization for machine learning models based on Bayesian optimization. Journal of Electronic Science and Technology. 2019, 17, 26–40.
- Jones, DR. A taxonomy of global optimization methods based on response surfaces. Journal of global optimization. 2001, 21, 345–383. [Google Scholar] [CrossRef]
- Yagin FH, Gülü M, Gormez Y, Castañeda-Babarro A, Colak C, Greco G, et al. Estimation of Obesity Levels with a Trained Neural Network Approach optimized by the Bayesian Technique. Applied Sciences. 2023, 13, 3875. [CrossRef]
- Mansourian P, Zhang N, Jaekel A, Zamanirafe M, Kneppers M. Anomaly Detection for Connected Autonomous Vehicles Using LSTM and Gaussian Naïve Bayes. International Conference on Wireless and Satellite Systems: Springer; 2023. p. 31-43.
- Maniruzzaman M, Rahman MJ, Ahammed B, Abedin MM, Suri HS, Biswas M, et al. Statistical characterization and classification of colon microarray gene expression data using multiple machine learning paradigms. Computer methods and programs in biomedicine. 2019, 176, 173–193. [CrossRef]
- Iqbal A, Barua K. A real-time emotion recognition from speech using gradient boosting. 2019 international conference on electrical, computer and communication engineering (ECCE): IEEE; 2019. p. 1-5.
- Alshboul O, Shehadeh A, Almasabha G, Almuflih AS. Extreme gradient boosting-based machine learning approach for green building cost prediction. Sustainability. 2022, 14, 6651. [CrossRef]
- Shah K, Patel H, Sanghvi D, Shah M. A comparative analysis of logistic regression, random forest and KNN models for the text classification. Augmented Human Research. 2020, 5, 1–16.
- Muharemi F, Logofătu D, Leon F. Machine learning approaches for anomaly detection of water quality on a real-world data set. Journal of Information and Telecommunication. 2019, 3, 294–307. [CrossRef]
- Ilyas H, Ali S, Ponum M, Hasan O, Mahmood MT, Iftikhar M, et al. Chronic kidney disease diagnosis using decision tree algorithms. BMC nephrology. 2021, 22, 1–11.
- Sattari MT, Apaydin H, Shamshirband S. Performance evaluation of deep learning-based gated recurrent units (GRUs) and tree-based models for estimating ETo by using limited meteorological variables. Mathematics. 2020, 8, 972. [CrossRef]
- Daneshvar D, Behnood A. Estimation of the dynamic modulus of asphalt concretes using random forests algorithm. International Journal of Pavement Engineering. 2022, 23, 250–260. [CrossRef]
- Yacouby R, Axman D. Probabilistic extension of precision, recall, and f1 score for more thorough evaluation of classification models. Proceedings of the first workshop on evaluation and comparison of NLP systems2020. p. 79-91.
- Bowers AJ, Zhou X. Receiver operating characteristic (ROC) area under the curve (AUC): A diagnostic measure for evaluating the accuracy of predictors of education outcomes. Journal of Education for Students Placed at Risk (JESPAR). 2019, 24, 20–46. [CrossRef]
- Nahm, FS. Receiver operating characteristic curve: overview and practical use for clinicians. Korean journal of anesthesiology. 2022, 75, 25–36. [Google Scholar] [CrossRef]
- Muschelli III, J. ROC and AUC with a binary predictor: a potentially misleading metric. Journal of classification. 2020, 37, 696–708. [Google Scholar] [CrossRef]
- Huang Y, Li W, Macheret F, Gabriel RA, Ohno-Machado L. A tutorial on calibration measurements and calibration models for clinical prediction models. Journal of the American Medical Informatics Association. 2020, 27, 621–633. [CrossRef]
- Liu J, Wang C, Yan R, Lu Y, Bai J, Wang H, et al. Machine learning-based prediction of postpartum hemorrhage after vaginal delivery: combining bleeding high risk factors and uterine contraction curve. Archives of Gynecology and Obstetrics. 2022, 306, 1015–1025. [CrossRef]
- Borys K, Schmitt YA, Nauta M, Seifert C, Krämer N, Friedrich CM, et al. Explainable AI in medical imaging: An overview for clinical practitioners–Beyond saliency-based XAI approaches. European journal of radiology. 2023, 110786.
- Yagin FH, Cicek İB, Alkhateeb A, Yagin B, Colak C, Azzeh M, et al. Explainable artificial intelligence model for identifying COVID-19 gene biomarkers. Computers in Biology and Medicine. 2023, 154, 106619. [CrossRef]
- Khanna VV, Chadaga K, Sampathila N, Prabhu S, Bhandage V, Hegde GK. A distinctive explainable machine learning framework for detection of polycystic ovary syndrome. Applied System Innovation. 2023, 6, 32. [CrossRef]
- Chatterjee J, Dethlefs N. Scientometric review of artificial intelligence for operations & maintenance of wind turbines: The past, present and future. Renewable and Sustainable Energy Reviews. 2021, 144, 111051.
- Tanaka M, Tajima S, Mizuno K, Ishii A, Konishi Y, Miike T, et al. Frontier studies on fatigue, autonomic nerve dysfunction, and sleep-rhythm disorder. The Journal of Physiological Sciences. 2015, 65, 483–498. [CrossRef]
- Yamano E, Watanabe Y, Kataoka Y. Insights into metabolite diagnostic biomarkers for myalgic encephalomyelitis/chronic fatigue syndrome. International Journal of Molecular Sciences. 2021, 22, 3423. [CrossRef] [PubMed]
- Group ICFSS. The chronic fatigue syndrome: A comprehensive approach to its definition and study. Annals of Internal Medicine. 1994, 121, 953–959. [CrossRef] [PubMed]
- Armstrong CW, McGregor NR, Lewis DP, Butt HL, Gooley PR. The association of fecal microbiota and fecal, blood serum and urine metabolites in myalgic encephalomyelitis/chronic fatigue syndrome. Metabolomics. 2017, 13, 1–13.
- Tomas C, Newton J. Metabolic abnormalities in chronic fatigue syndrome/myalgic encephalomyelitis: a mini-review. Biochemical Society Transactions. 2018, 46, 547–553. [CrossRef]
- Huth TK, Eaton-Fitch N, Staines D, Marshall-Gradisnik S. A systematic review of metabolomic dysregulation in chronic fatigue syndrome/myalgic encephalomyelitis/systemic exertion intolerance disease (CFS/ME/SEID). Journal of translational medicine. 2020, 18, 1–14.
- Jason LA, Boulton A, Porter NS, Jessen T, Njoku MG, Friedberg F. Classification of myalgic encephalomyelitis/chronic fatigue syndrome by types of fatigue. Behavioral Medicine. 2010, 36, 24–31. [CrossRef] [PubMed]
- Nagy-Szakal D, Barupal DK, Lee B, Che X, Williams BL, Kahn EJ, et al. Insights into myalgic encephalomyelitis/chronic fatigue syndrome phenotypes through comprehensive metabolomics. Scientific reports. 2018, 8, 10056. [CrossRef] [PubMed]
- Petrick LM, Shomron N. AI/ML-driven advances in untargeted metabolomics and exposomics for biomedical applications. Cell Reports Physical Science. 2022, 3.

| Technique | Optimized Parameter Value |
|---|---|
| GNB | var_smoothing=1e-9. |
| GBC | n_estimators=3, learning_rate=1.0, max_depth=1, random_state=0. |
| LR | random_state=0, max_iter=30, solver='liblinear'. |
| RFC | max_depth=26, min_samples_leaf=5, min_samples_split=3, n_estimators= 12. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




