Submitted:
05 July 2023
Posted:
06 July 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset
- Control group (MK): This group was injected intraperitoneally with physiological saline as a carrier solvent on the first day of the experiment.
- Nephrotoxicity group (M): This group was given a single dose of 20 mg/kg MTX intraperitoneal on the first day of the experiment.
2.2. Random Forest Method
2.3. Data Analysis and Modeling Tasks
2.4. Histopathological and Immunohistochemical Analyses
2.4.1. Histopathological analyses
2.4.2. Immunohistochemical analyses
2.5. Genomic analyses
2.5.1. Total Rna Isolation and Quality Control from Harvested Tissues
2.5.2. Preparing and sequencing NGS libraries for lncRNA sequences
3. Results
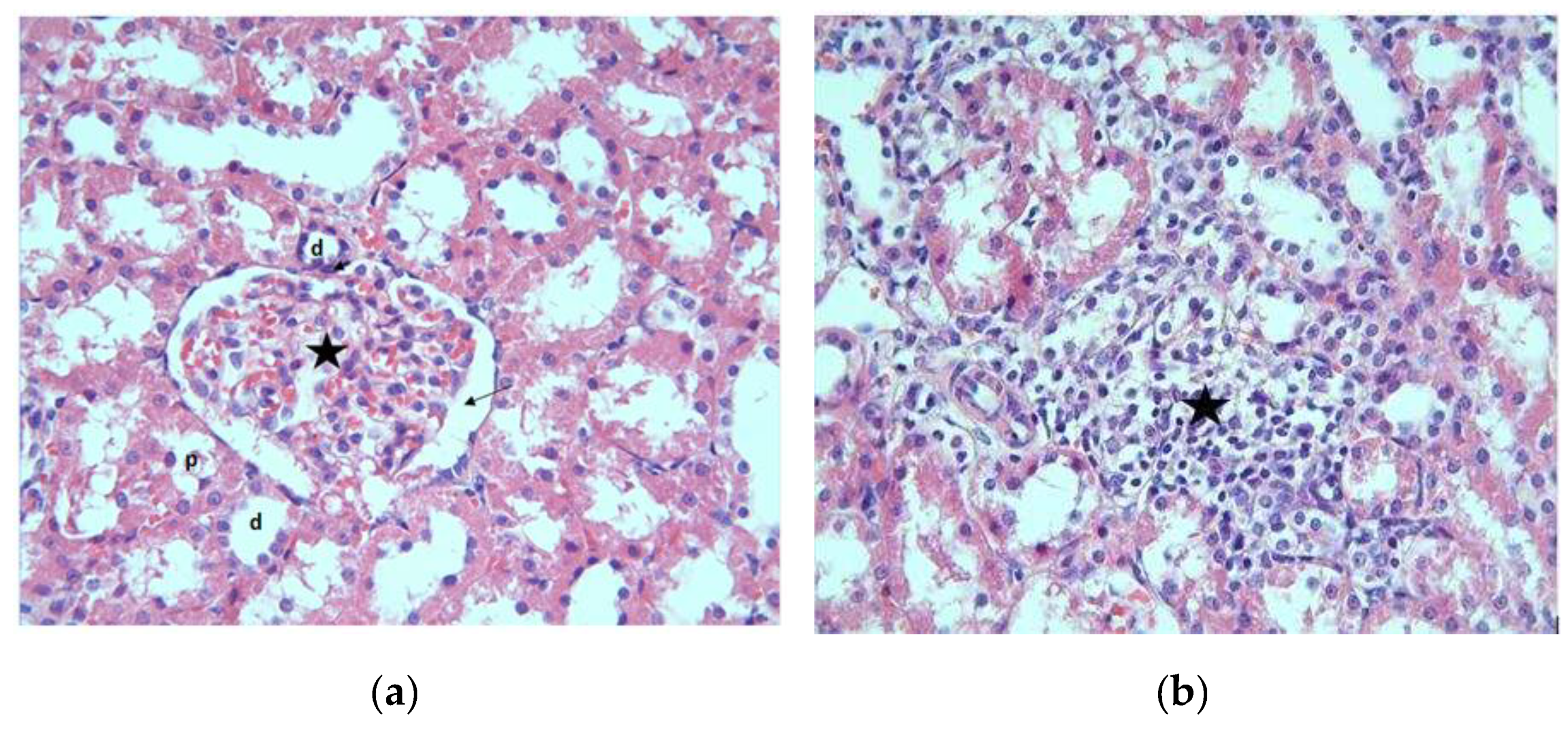
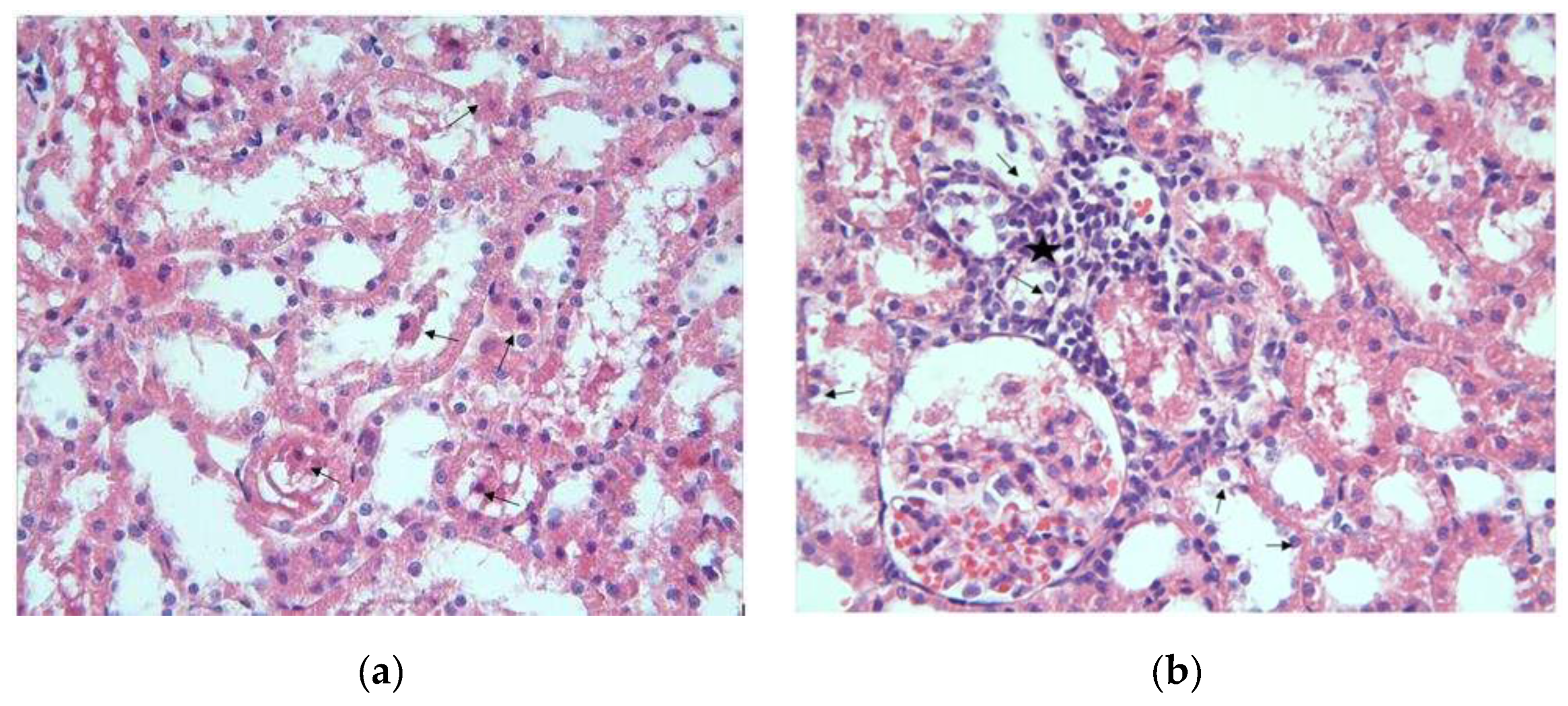
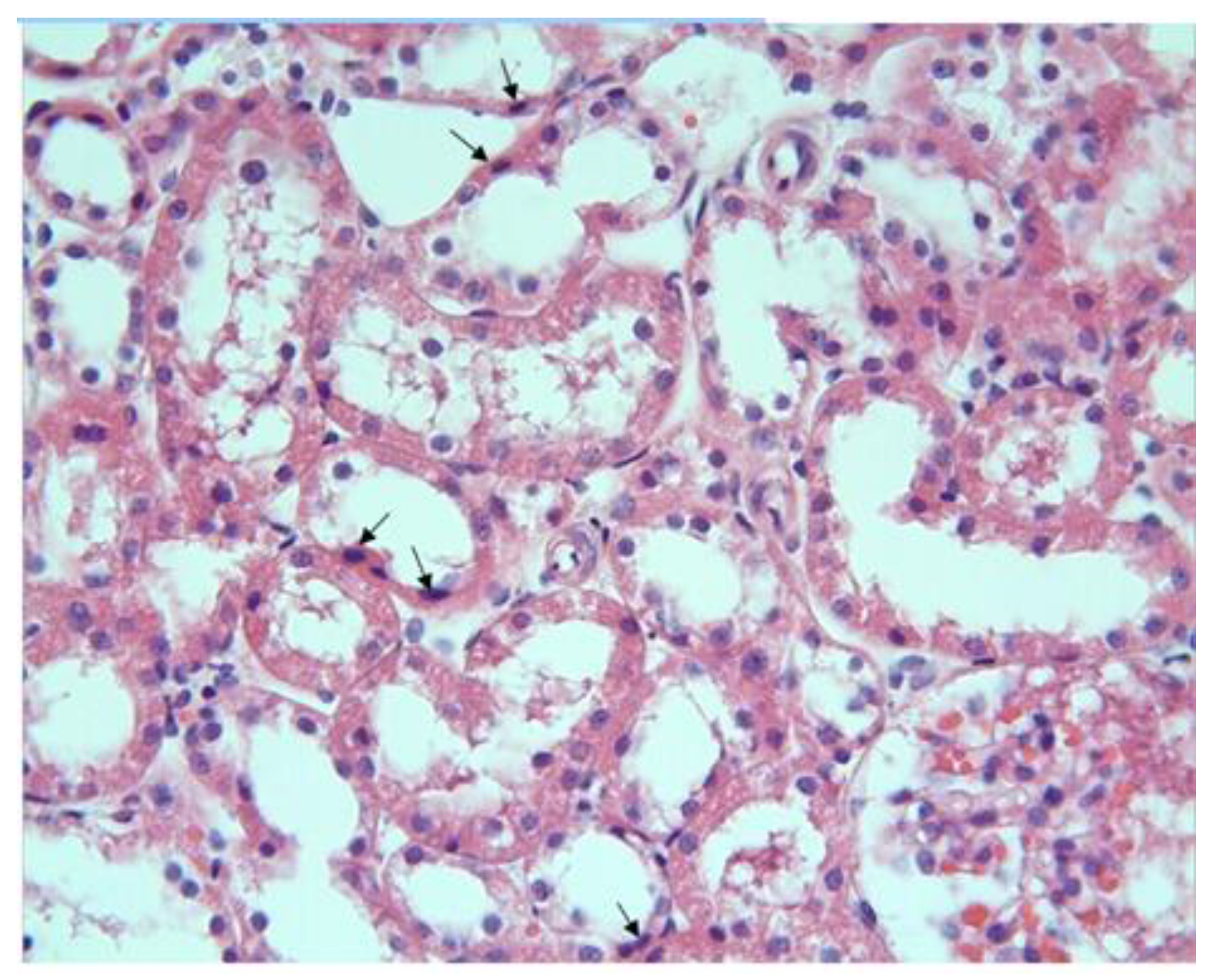
3.1. Histopathological Results
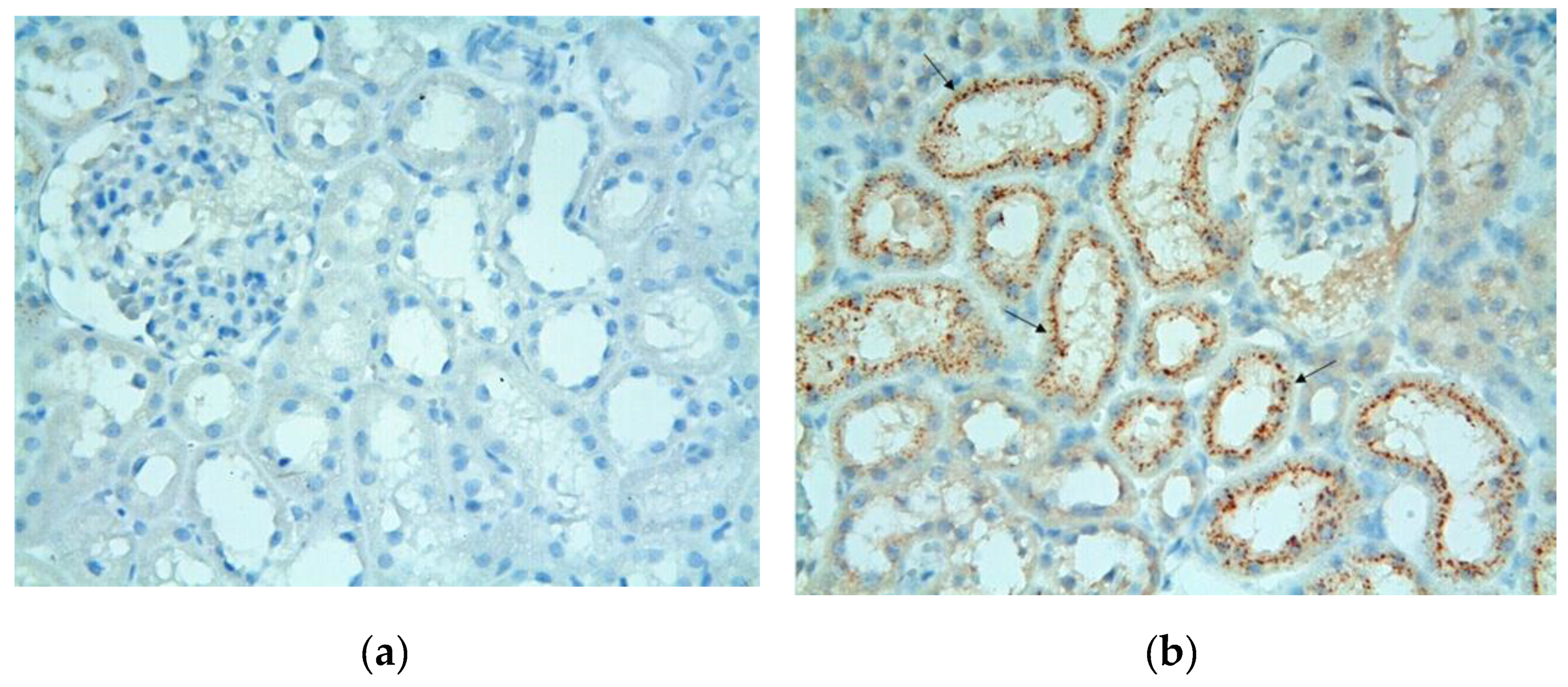
3.2. Immunohistochemical Results
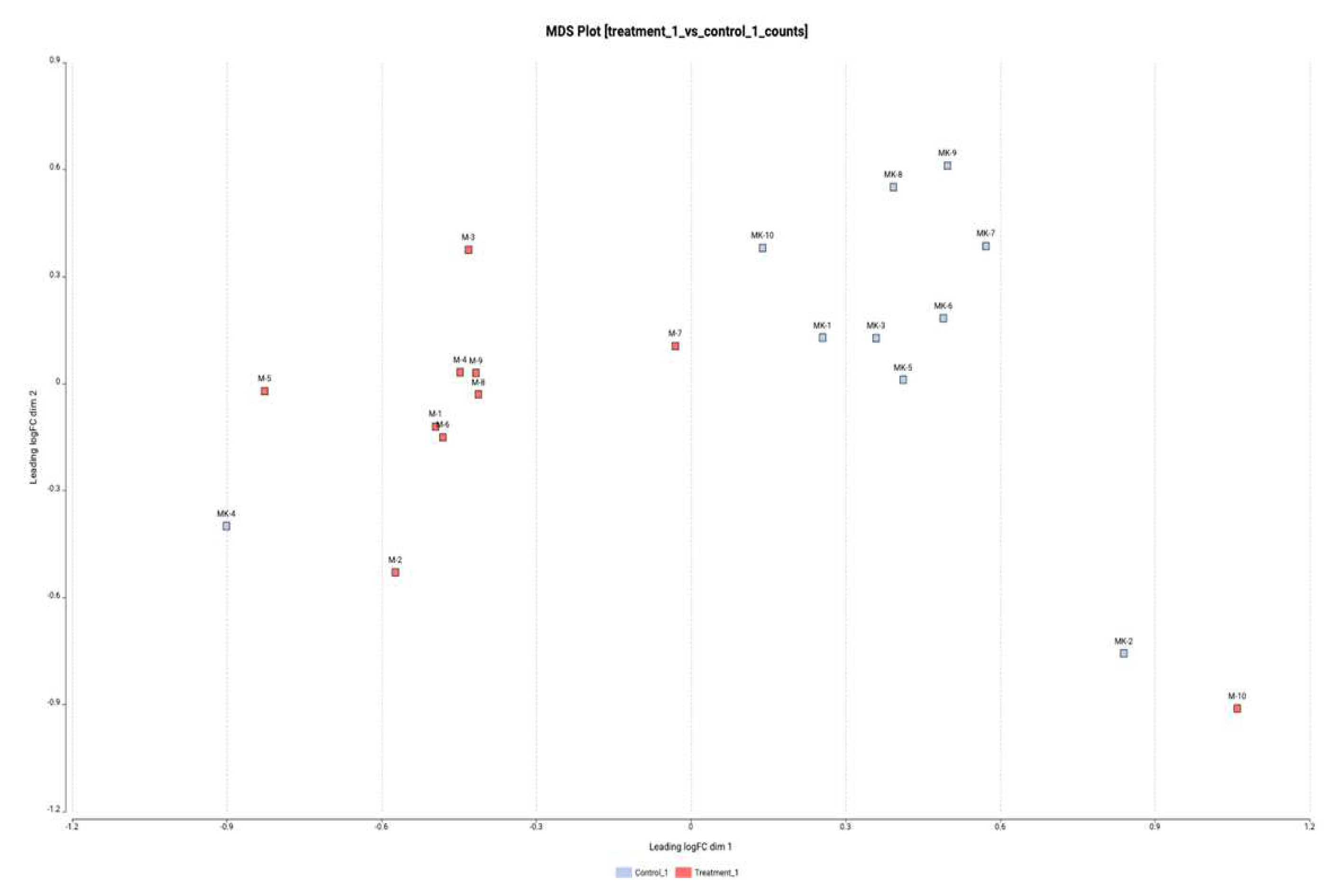
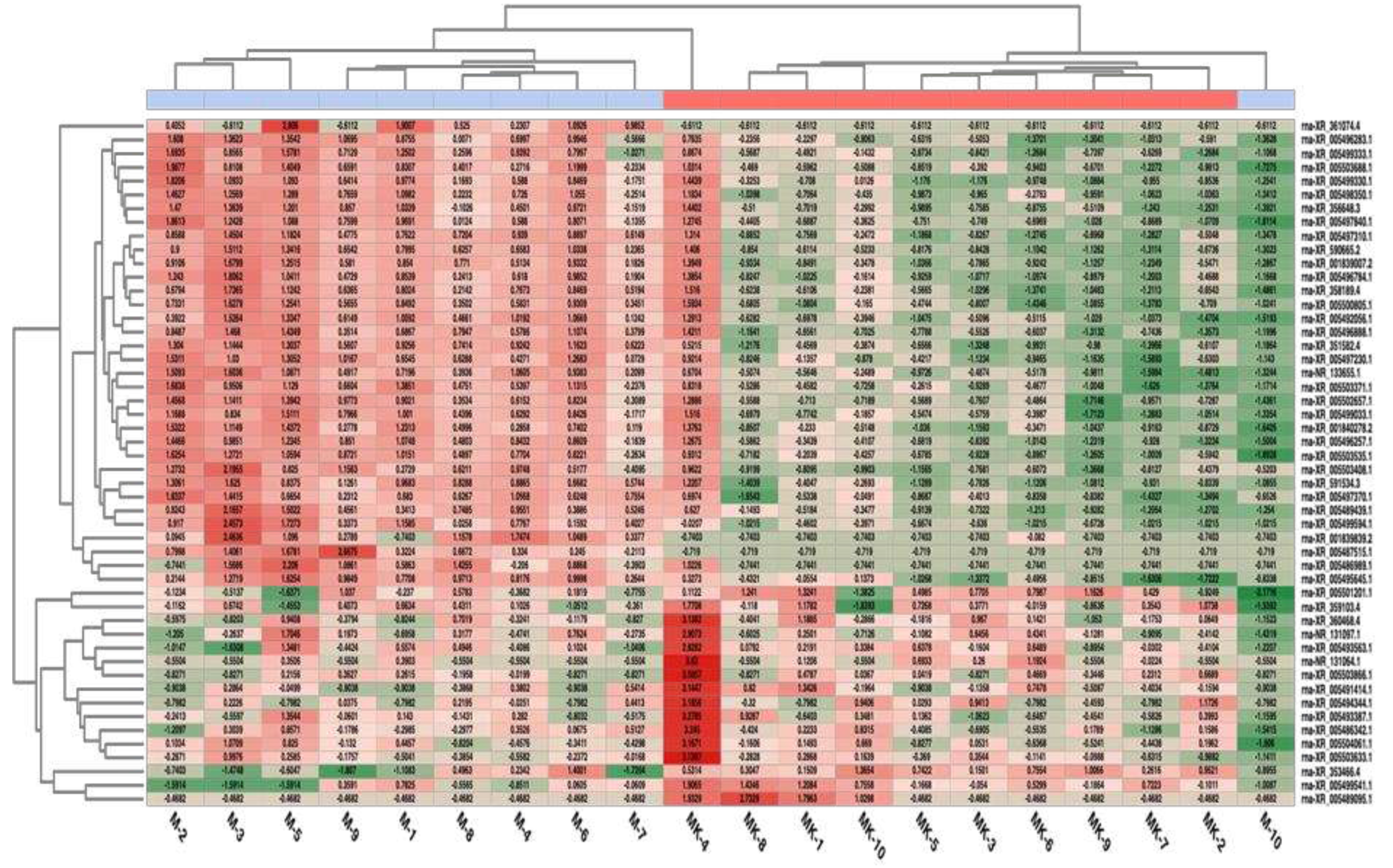
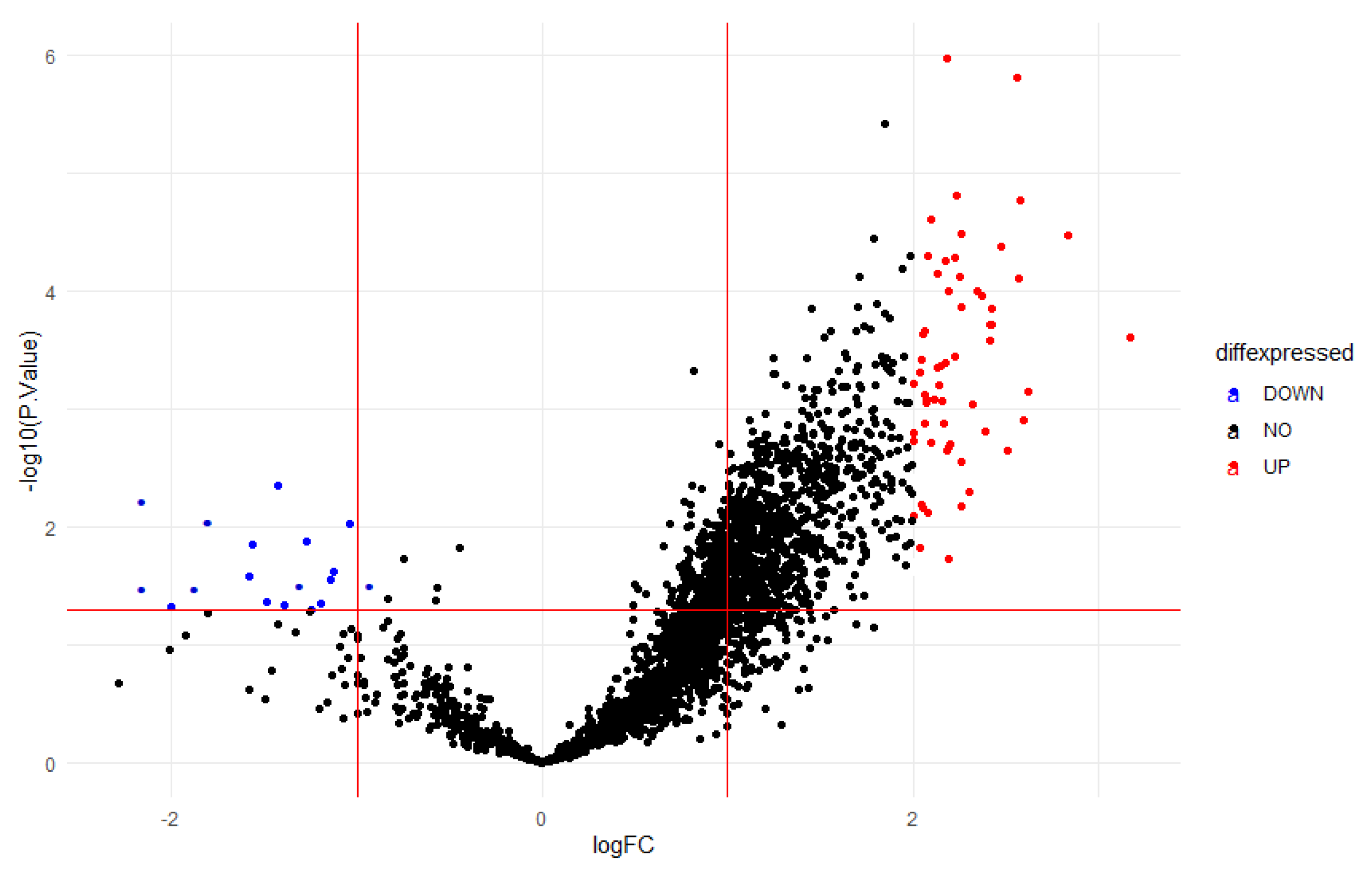
3.3. Differential Expression Results
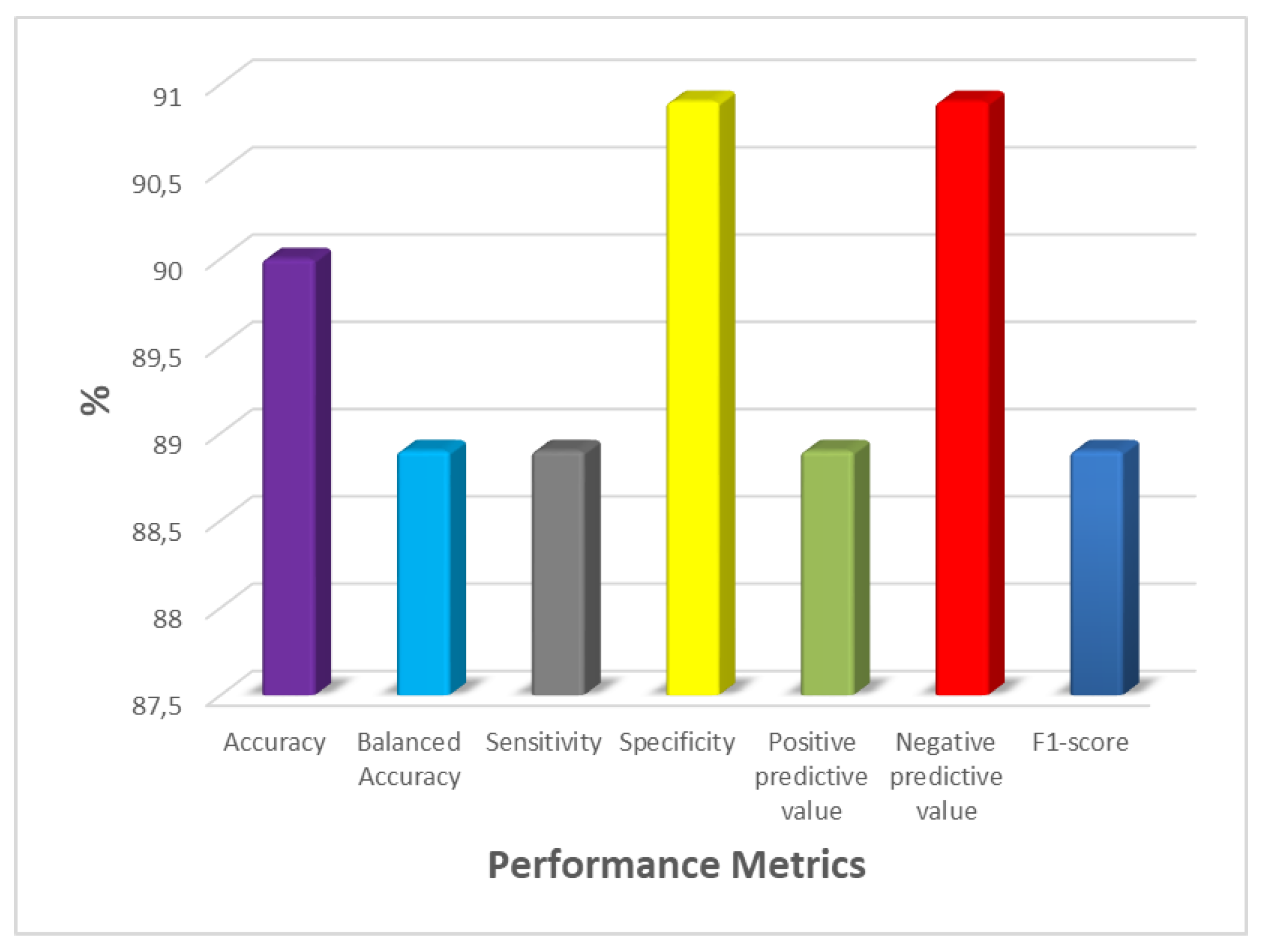
3.4. Biostatistics Analysis and Modeling Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Maurya, H.; Kumar, T.; Kumar, S. Anatomical and physiological similarities of kidney in different experimental animals used for basic studies. J Clin Exp Nephrol 2018, 3. [Google Scholar] [CrossRef]
- Kwiatkowska, E.; Domański, L.; Dziedziejko, V.; Kajdy, A.; Stefańska, K.; Kwiatkowski, S. The Mechanism of Drug Nephrotoxicity and the Methods for Preventing Kidney Damage. Int. J. Mol. Sci. 2021, 22, 6109. [Google Scholar] [CrossRef]
- Sales, G.T.M.; Foresto, R.D. Drug-induced nephrotoxicity. Revista da Associação Médica Brasileira 2020, 66, s82–s90. [Google Scholar] [CrossRef] [PubMed]
- Hannoodee, M.; Mittal, M. Methotrexate. In StatPearls [internet]; StatPearls Publishing: 2022.
- Bhattacharya, S. Reactive Oxygen Species and Cellular Defense System. In Free Radicals in Human Health and Disease; Rani, V., Yadav, U.C.S., Eds.; Springer: New Delhi, India, 2015; pp. 17–29. [Google Scholar]
- Florea, A.-M.; Büsselberg, D. Cisplatin as an Anti-Tumor Drug: Cellular Mechanisms of Activity, Drug Resistance and Induced Side Effects. Cancers 2011, 3, 1351–1371. [Google Scholar] [CrossRef]
- Rawls, K.D.; Dougherty, B.V.; Vinnakota, K.C.; Pannala, V.R.; Wallqvist, A.; Kolling, G.L.; Papin, J.A. Predicting changes in renal metabolism after compound exposure with a genome-scale metabolic model. Toxicol. Appl. Pharmacol. 2020, 412, 115390–115390. [Google Scholar] [CrossRef]
- Blanchet, L.; Smolinska, A.; Attali, A.; Stoop, M.P.; Ampt, K.A.; van Aken, H.; Suidgeest, E.; Tuinstra, T.; Wijmenga, S.S.; Luider, T.; et al. Fusion of metabolomics and proteomics data for biomarkers discovery: case study on the experimental autoimmune encephalomyelitis. BMC Bioinform. 2011, 12, 254–254. [Google Scholar] [CrossRef] [PubMed]
- Agirbasli, D.; Isil Ulman, F. Coronary artery disease from a perspective of genomic risk score, ethical approaches and suggestions. Anadolu kardiyoloji dergisi-the anatolian journal of cardiology 2012, 12.
- Nguyen, Q.; Carninci, P. Expression Specificity of Disease-Associated lncRNAs: Toward Personalized Medicine. Poxviruses 2015, 394, 237–258. [Google Scholar] [CrossRef]
- Zhao, H.; Shi, J.; Zhang, Y.; Xie, A.; Yu, L.; Zhang, C.; Lei, J.; Xu, H.; Leng, Z.; Li, T.; et al. LncTarD: a manually-curated database of experimentally-supported functional lncRNA–target regulations in human diseases. Nucleic Acids Res. 2019, 48, D118–D126. [Google Scholar] [CrossRef]
- Okuyan, H.M.; Dogan, S.; Terzi, M.Y.; Begen, M.A.; Turgut, F.H. Association of serum lncRNA H19 expression with inflammatory and oxidative stress markers and routine biochemical parameters in chronic kidney disease. Clin. Exp. Nephrol. 2021, 25, 522–530. [Google Scholar] [CrossRef] [PubMed]
- Ma, T.; Jia, H.; Ji, P.; He, Y.; Chen, L. Identification of the candidate lncRNA biomarkers for acute kidney injury: a systematic review and meta-analysis. Expert Rev. Mol. Diagn. 2021, 21, 77–89. [Google Scholar] [CrossRef]
- Battineni, G.; Sagaro, G.G.; Chinatalapudi, N.; Amenta, F. Applications of Machine Learning Predictive Models in the Chronic Disease Diagnosis. J. Pers. Med. 2020, 10, 21. [Google Scholar] [CrossRef]
- Kumar, N.; Das, N.N.; Gupta, D.; Gupta, K.; Bindra, J. Efficient Automated Disease Diagnosis Using Machine Learning Models. J. Heal. Eng. 2021, 2021, 1–13. [Google Scholar] [CrossRef]
- Chang, C.-H.; Lin, C.-H.; Lane, H.-Y. Machine Learning and Novel Biomarkers for the Diagnosis of Alzheimer’s Disease. Int. J. Mol. Sci. 2021, 22, 2761. [Google Scholar] [CrossRef] [PubMed]
- Tjoa, E.; Guan, C. A Survey on Explainable Artificial Intelligence (XAI): Toward Medical XAI. IEEE Trans. Neural Networks Learn. Syst. 2020, 32, 4793–4813. [Google Scholar] [CrossRef]
- Samek, W.; Müller, K.-R. Towards explainable artificial intelligence. Explainable AI: interpreting, explaining and visualizing deep learning 2019, 5-22.
- Shi, S.; Zhang, X.; Fan, W. A modified perturbed sampling method for local interpretable model-agnostic explanation. arXiv preprint arXiv:2002.07434 2020. arXiv:2002.07434 2020.
- Breiman, L. Random forests. Machine learning 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Prasad, A.M.; Iverson, L.R.; Liaw, A. Newer Classification and Regression Tree Techniques: Bagging and Random Forests for Ecological Prediction. Ecosystems 2006, 9, 181–199. [Google Scholar] [CrossRef]
- Panov, P.; Džeroski, S. Combining bagging and random subspaces to create better ensembles. In Proceedings of the International Symposium on Intelligent Data Analysis; 2007; pp. 118–129. [Google Scholar]
- Taslidere, E.; Dogan, Z.; Elbe, H.; Vardi, N.; Cetin, A.; Turkoz, Y. Quercetin protection against ciprofloxacin induced liver damage in rats. Biotech. Histochem. 2015, 91, 116–121. [Google Scholar] [CrossRef]
- Parlakpinar, H.; Ozhan, O.; Ermis, N.; Vardi, N.; Cigremis, Y.; Tanriverdi, L.H.; Colak, C.; Acet, A. Acute and Subacute Effects of Low Versus High Doses of Standardized Panax ginseng Extract on the Heart: An Experimental Study. Cardiovasc. Toxicol. 2019, 19, 306–320. [Google Scholar] [CrossRef] [PubMed]
- Napolitano, M.; Comegna, M.; Succoio, M.; Leggiero, E.; Pastore, L.; Faraonio, R.; Cimino, F.; Passaro, F. Comparative Analysis of Gene Expression Data Reveals Novel Targets of Senescence-Associated microRNAs. PLOS ONE 2014, 9, e98669. [Google Scholar] [CrossRef]
- Arrigoni, A.; Ranzani, V.; Rossetti, G.; Panzeri, I.; Abrignani, S.; Bonnal, R.J.; Pagani, M. Analysis RNA-seq and Noncoding RNA. Polycomb Group Proteins: Methods and Protocols 2016, 125-135.
- Eren, E.; Alper, A.; Arıcan, A. Kanser tedavisinde kullanılan ilaçlar ve nefrotoksisite. Dokuz Eylül Üniversitesi Tıp Fakültesi Dergisi 2012, 26, 229–235. [Google Scholar]
- Petejova, N.; Martinek, A.; Zadrazil, J.; Teplan, V. Acute toxic kidney injury. Ren. Fail. 2019, 41, 576–594. [Google Scholar] [CrossRef]
- Al-Kuraishy, H.M.; Al-Naimi, M.S.; Rasheed, H.; Hussien, N.R.; Al-Gareeb, A. Nephrotoxicity: Role and significance of renal biomarkers in the early detection of acute renal injury. J. Adv. Pharm. Technol. Res. 2019, 10, 95–99. [Google Scholar] [CrossRef]
- Kim, S.-Y.; Moon, A.-R. Drug-Induced Nephrotoxicity and Its Biomarkers. Biomol. Ther. 2012, 20, 268–272. [Google Scholar] [CrossRef]
- Ferguson, M.A.; Vaidya, V.S.; Bonventre, J.V. Biomarkers of nephrotoxic acute kidney injury. Toxicology 2008, 245, 182–193. [Google Scholar] [CrossRef] [PubMed]
- Bonventre, J.V.; Vaidya, V.S.; Schmouder, R.; Feig, P.; Dieterle, F. Next-generation biomarkers for detecting kidney toxicity. Nat. Biotechnol. 2010, 28, 436–440. [Google Scholar] [CrossRef]
| Variables | Mean ± Standard Deviation |
|---|---|
| Rat weight starting (g) | 249.15±22.32 |
| Rat weight end (g) | 252.1±24.05 |
| Kidney weight (g) | 0.968±0.1 |
| Variables | Control | Nephrotoxicity |
|---|---|---|
| Rat weight starting (g) | 245.3±24.02 | 253±21.01 |
| Rat weight end (g) | 252±24.03 | 252.2±25.37 |
| Kidney weight (g) | 0.97±0.08 | 0.96±0.12 |
| Gene Name | Chromosome | ID | Group | |||||
|---|---|---|---|---|---|---|---|---|
| M | MK | LogFC | p | |||||
| Mean ± SD | Median (Min-Max) | Mean ± SD | Median (Min-Max) | |||||
| LOC102555118 | NC_051337.1 | rna-XR_351582.4 | 226.4±116.41 | 248(42-447) | 35.6±16.85 | 34(17-66) | 1.616 | 0.001* |
| LOC106736471 | NC_051345.1 | rna-NR_133655.1 | 102.4±76.73 | 88(5-257) | 11.9±9.48 | 10.5(1-34) | 2.198 | 0.005* |
| LOC103691349 | NC_051336.1 | rna-XR_590665.2 | 281.2±123.78 | 294(49-470) | 68.4±78.82 | 46(26-290) | 1.247 | 0.001** |
| LOC108351528 | NC_051342.1 | rna-XR_001839007.2 | 454.2±191.95 | 486.5(96-661) | 117.1±118.09 | 80.5(55-449) | 1.118 | 0.001** |
| LOC120098801 | NC_051336.1 | rna-XR_005497310.1 | 166±92.76 | 164.5(29-370) | 38.6±36.34 | 26.5(13-139) | 1.187 | 0.001** |
| LOC120094778 | NC_051344.1 | rna-XR_005489439.1 | 140±90.62 | 125(28-296) | 28.9±14.03 | 30(9-51) | 1.248 | 0.004* |
| LOC120099280 | NC_051336.1 | rna-XR_005498350.1 | 109.6±68.4 | 96(13-206) | 21.6±21.84 | 15.5(7-82) | 1.488 | 0.002** |
| LOC120096007 | NC_051347.1 | rna-XR_005492056.1 | 134.4±91.26 | 123.5(17-332) | 32.6±28.96 | 25.5(6-111) | 1.087 | 0.004** |
| LOC120098788 | NC_051336.1 | rna-XR_005497230.1 | 27.3±13.61 | 29.5(3-52) | 4.6±4.62 | 2.5(0-15) | 1.751 | <0.001** |
| LOC120098190 | NC_051353.1 | rna-XR_005496257.1 | 85.5±54.7 | 70(9-172) | 19.5±18.58 | 16(4-70) | 1.277 | 0.004** |
| LOC108348888 | NC_051354.1 | rna-XR_005496888.1 | 71.2±32.64 | 75.5(12-112) | 17.1±20.59 | 11.5(3-74) | 1.250 | 0.002** |
| LOC103691816 | NC_051338.1 | rna-XR_591534.3 | 210.4±116.14 | 230.5(54-421) | 49.2±36.54 | 40.5(19-147) | 1.171 | 0.001** |
| LOC120098816 | NC_051355.1 | rna-XR_005497370.1 | 220.6±173.89 | 186(48-552) | 31.3±22.35 | 30(6-73) | 1.992 | 0.007* |
| LOC120096731 | NC_051349.1 | rna-XR_005493563.1 | 6.6±6.64 | 3.5(0-18) | 13.2±14.34 | 8(3-51) | -1.862 | 0.093** |
| LOC120098521 | NC_051354.1 | rna-XR_005496784.1 | 362.1±181.28 | 349.5(74-587) | 88.8±100.42 | 58(33-369) | 1.249 | 0.001** |
| LOC120102202 | NC_051339.1 | rna-XR_005503371.1 | 84.1±63.57 | 73(13-208) | 15.6±11.47 | 13.5(3-37) | 1.559 | 0.008* |
| LOC102549457 | NC_051346.1 | rna-XR_358189.4 | 77.7±42.9 | 75.5(8-154) | 21.1±26.93 | 12.5(4-96) | 1.078 | 0.007** |
| LOC120102261 | NC_051339.1 | rna-XR_005503535.1 | 215.2±138.84 | 176.5(16-442) | 47±27.2 | 38.5(26-116) | 1.205 | 0.003** |
| LOC120100781 | NC_051337.1 | rna-XR_005500805.1 | 51.1±23.38 | 49(11-82) | 14.4±20.5 | 8(2-71) | 1.114 | 0.002** |
| LOC108348808 | NC_051353.1 | rna-XR_005496283.1 | 42.2±26.1 | 37.5(5-84) | 9.1±5.61 | 9(2-19) | 1.287 | 0.003* |
| LOC103691306 | NC_051336.1 | rna-XR_005499594.1 | 6.2±4.47 | 5.5(0-12) | 0.6±0.52 | 1(0-1) | 2.178 | 0.001** |
| LOC102552040 | NC_051344.1 | rna-XR_001839839.2 | 3.8±4.49 | 3(0-15) | 0.1±0.32 | 0(0-1) | 3.296 | 0.002** |
| LOC120099889 | rna-XR_005499330.1 | 282.2±232.78 | 197(41-831) | 68.2±86.7 | 35(24-308) | 1.431 | 0.002** | |
| NC_051336.1 | ||||||||
| LOC120099800 | NC_051336.1 | rna-XR_005499033.1 | 53.7±33.95 | 45(5-102) | 14.2±19.85 | 9.5(1-69) | 1.176 | 0.004** |
| LOC120097836 | NC_051352.1 | rna-XR_005495645.1 | 32.8±16.73 | 28.5(13-62) | 7.7±4.32 | 8.5(1-14) | 1.089 | 0.001* |
| LOC120102212 | NC_051339.1 | rna-XR_005503408.1 | 18.5±10.54 | 14.5(8-42) | 4±2.31 | 3.5(2-10) | 1.313 | <0.001** |
| LOC102555751 | NC_051355.1 | rna-XR_005497840.1 | 54.9±45.9 | 41(1-162) | 12.1±12.54 | 8.5(3-47) | 1.431 | 0.008** |
| LOC120102327 | NC_051339.1 | rna-XR_005503688.1 | 50.7±46.08 | 41.5(1-165) | 9.8±8.04 | 7.5(3-30) | 1.612 | 0.005** |
| LOC120099962 | NC_051336.1 | rna-XR_005499541.1 | 1±0.94 | 1(0-3) | 2.3±0.82 | 2(1-4) | 2.047 | 0.005** |
| LOC108352129 | NC_051345.1 | rna-XR_001840278.2 | 26±18.34 | 21(0-59) | 5.8±6.94 | 3(2-25) | 1.282 | 0.008** |
| LOC102554372 | NC_051339.1 | rna-XR_353438.4 | 48.4±27.61 | 49.5(3-84) | 12.1±6.05 | 11.5(4-21) | 1.037 | 0.002* |
| Metric | Value (%) (95% CI) |
|---|---|
| B-Acc | 88.9 (76.7-100) |
| Acc | 90 (76.9-100) |
| Sp | 90.9 (58.7-99.8) |
| Se | 88.9 (51.8-99.7) |
| Npv | 90.9 (58.7-99.8) |
| Ppv | 88.9 (51.8-99.7) |
| F1-score | 88.9 (75.1-100) |
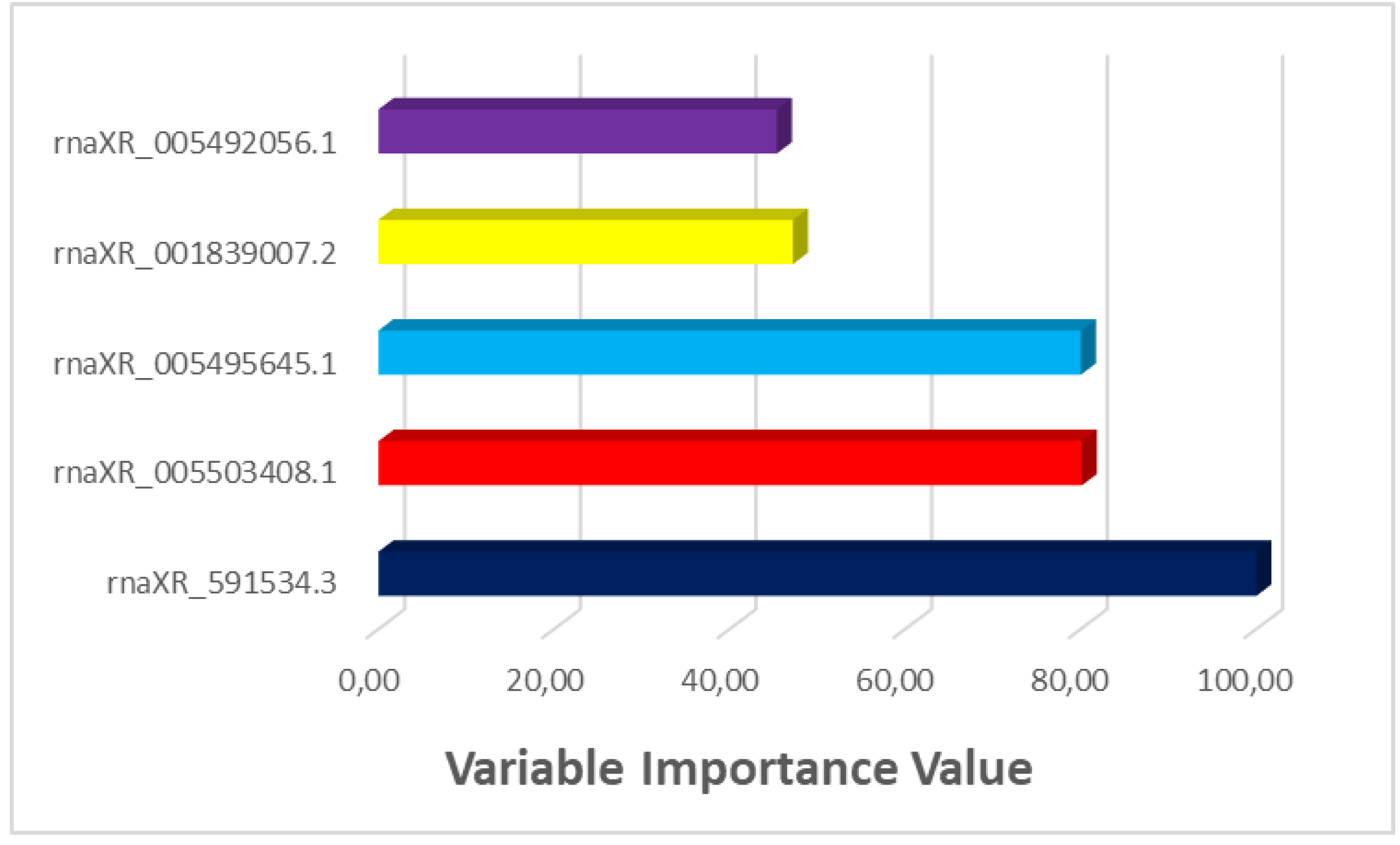
| Gene Name | Variable Importance Value |
|---|---|
| rnaXR_591534.3 | 100 |
| rnaXR_005503408.1 | 80.127 |
| rnaXR_005495645.1 | 80.02 |
| rnaXR_001839007.2 | 47.205 |
| rnaXR_005492056.1 | 45.374 |
| rnaXR_351582.4 | 42.972 |
| rnaXR_001840278.2 | 42.9 |
| rnaXR_005496784.1 | 41.422 |
| rnaXR_005498350.1 | 39.116 |
| rnaXR_005503371.1 | 38.433 |
| rnanr_133655.1 | 38.301 |
| rnaXR_005497370.1 | 35.986 |
| rnaXR_005500805.1 | 33.445 |
| rnaXR_005496283.1 | 31.788 |
| rnaXR_353438.4 | 30.313 |
| rnaXR_005499330.1 | 29.65 |
| rnaXR_005497310.1 | 29.435 |
| rnaXR_005503535.1 | 29.232 |
| rnaXR_358189.4 | 27.716 |
| rnaXR_005499033.1 | 24.311 |
| rnaXR_005496888.1 | 24.018 |
| rnaXR_590665.2 | 23.715 |
| rnaXR_005497840.1 | 23.365 |
| rnaXR_005503688.1 | 19.988 |
| rnaXR_005499541.1 | 18.123 |
| rnaXR_005496257.1 | 17.68 |
| rnaXR_005499594.1 | 17.632 |
| rnaXR_005497230.1 | 15.566 |
| rnaXR_005493563.1 | 8.101 |
| rnaXR_001839839.2 | 5.695 |
| rnaXR_005489439.1 | 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




