Submitted:
25 June 2023
Posted:
26 June 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Clinical Examination
2.2. DNA Sequence Analysis
2.3. Functional Assay
2.3.1. Cell Lines and Culture
2.3.2. Antibodies
2.3.3. Plasmid Preparation
2.3.4. Establishment of Lentivirus and Stable Expression Cells
2.3.5. RNA Preparation and Quantitative RT-PCR
2.3.6. Transfection of Expression Plasmid
2.3.7. Reporter Assay
2.3.8. Cell Fractionation and Pull-Down Assay
2.3.9. Western Blot and Slot Blot
3. Results
3.1. Genetic Findings and Clinical Features
| Nucleotide Change | Amino Acid Change | Polyphen2 HumDIV [13] (Cutoff=0.85) | GERP++ [14] (Cutoff=2) | REVEL [15] (Cutoff=0.5)* | M-CAP [16] (Cutoff=0.025) | CADD Phred [17] (Cutoff=15)** | SIFT [18] (Cutoff=0.05) | Pathogenicity (Evidenced Criteria Points)*** |
|---|---|---|---|---|---|---|---|---|
| c.58C>T | p.P20S | 0.980 | 4.48 | 0.927 | 0.939 | 23.900 | 0.010 | Pathogenic (PS=1, PM=2, PP=4) |
| c.150_151insA | p.G65Rfs*5 | Pathogenic (PVS=1, PS=1, PM=2, PP=1) |
||||||
| c.233T>A | p.V78E | 1.000 | 5.35 | 0.979 | 0.957 | 28.600 | 0.000 | Pathogenic (PS=1, PM=3, PP=4) |
| c.247G>T | p.V83F | 1.000 | 5.35 | 0.942 | 0.916 | 29.800 | 0.000 | Pathogenic (PS=1, PM=2, PP=4) |
| c.383G>A | p.R128H | 0.997 | 3.83 | 0.933 | 0.786 | 28.100 | 0.000 | Pathogenic (PS=2, PM=3, PP=3) |
| c.764A>T | p.Q255L | 0.999 | 5.47 | 0.984 | 0.822 | 34.000 | 0.000 | Pathogenic (PS=1, PM=2, PP=3) |
| c.767T>A | p.V256E | 1.000 | 5.53 | 0.976 | 0.824 | 32.000 | 0.000 | Pathogenic (PS=2, PM=3, PP=3) |
| c.1032+5G>A | - | Pathogenic (PVS=1, PM=2, PP=2) |
||||||
| c.1095T>G | p.N365K | 0.736 | NA | 0.499 | 0.145 | 8.266 | 0.020 | Pathogenic (PS=2, PM=2, PP=2) |
| PAX6 P20S FW | GGGCGGTCACTGCCGGACTCCACCC |
| PAX6 P20S RV | GCAGTGACCGCCCGTTGACAAAGAC |
| PAX6 V78E FW | CGAGAGAAGCGACTCCAGAAGTTGT |
| PAX6 V78E RV | GTCGCTTCTCTCGGTTTACTACCAC |
| PAX6 V83F FW | CCAGAATTTGTAAGCAAAATAGCCC |
| PAX6 V83F RV | TTACAAATTCTGGAGTCGCTACTCT |
| PAX6 R128H FW | TTCTTCACAACCTGGCTAGCGAAAA |
| PAX6 R128H RV | AGGTTGTGAAGAACTCTGTTTATTG |
| PAX6 Q255L FW | GAATACTGGTATGGTTTTCTAATCG |
| PAX6 Q255L RV | CATACCAGTATTCTTGCTTCAGGTA |
| PAX6 V256E FW | TACAGGAATGGTTTTCTAATCGAAG |
| PAX6 V256E RV | AACCATTCCTGTATTCTTGCTTCAG |
| PAX6 N365K FW | GGTGAAGGGGCGGAGTTATGATACC |
| PAX6 N365K RV | CCGCCCCTTCACCGAAGGGCTGGTG |
| 5aCON FW 5-biotin | (Biothin)ATCTGAACATGCTCAGTGAATGTTCATTGACTCTC |
| 5aCON RV | GAGAGTCAATGAACATTCACTGAGCATGTTCAGAT |
| TA FW | (EcoRI) gaattcGAAAAACTGAGGAATCAGAGAAG |
3.2. Functional Assays
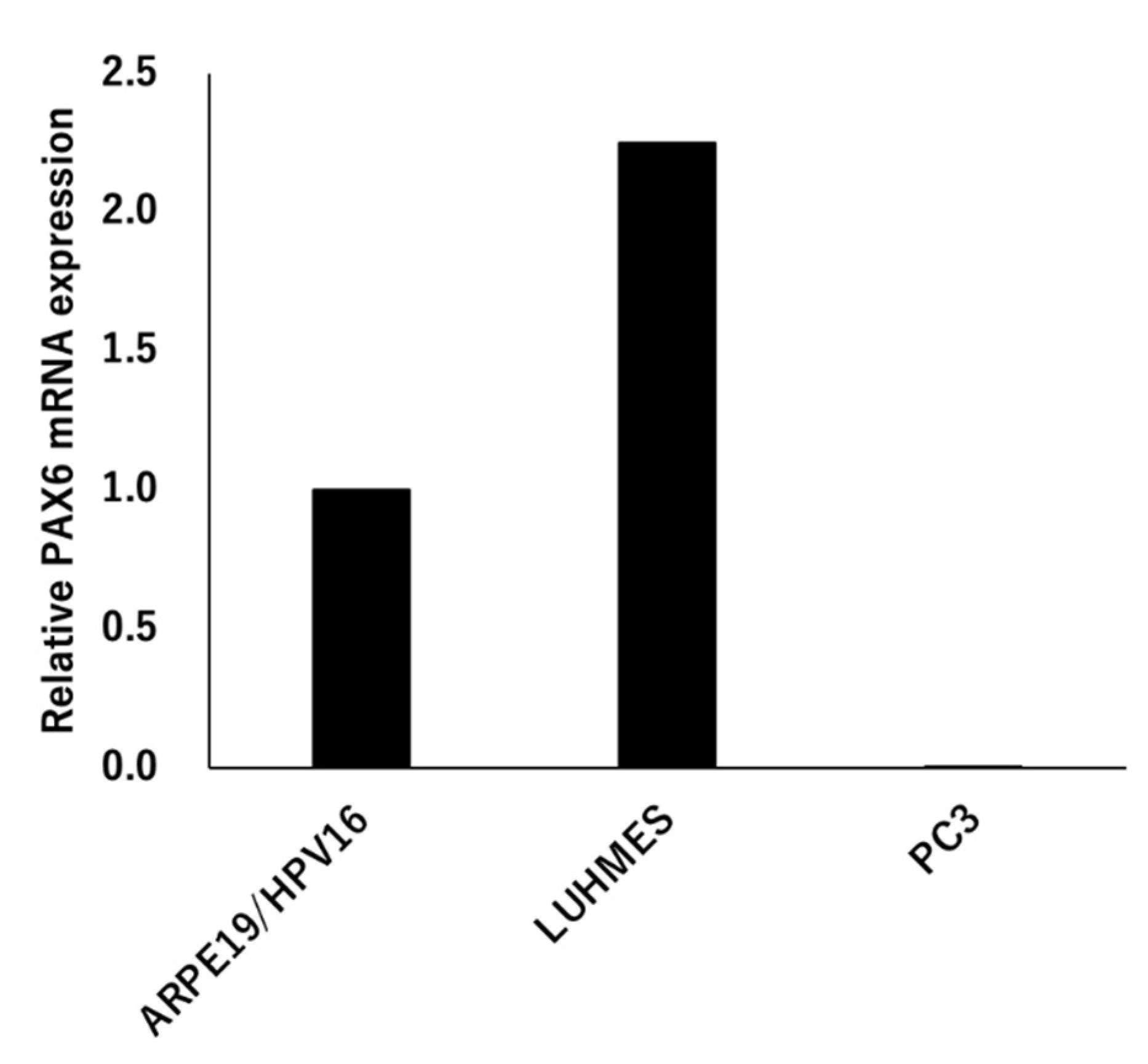
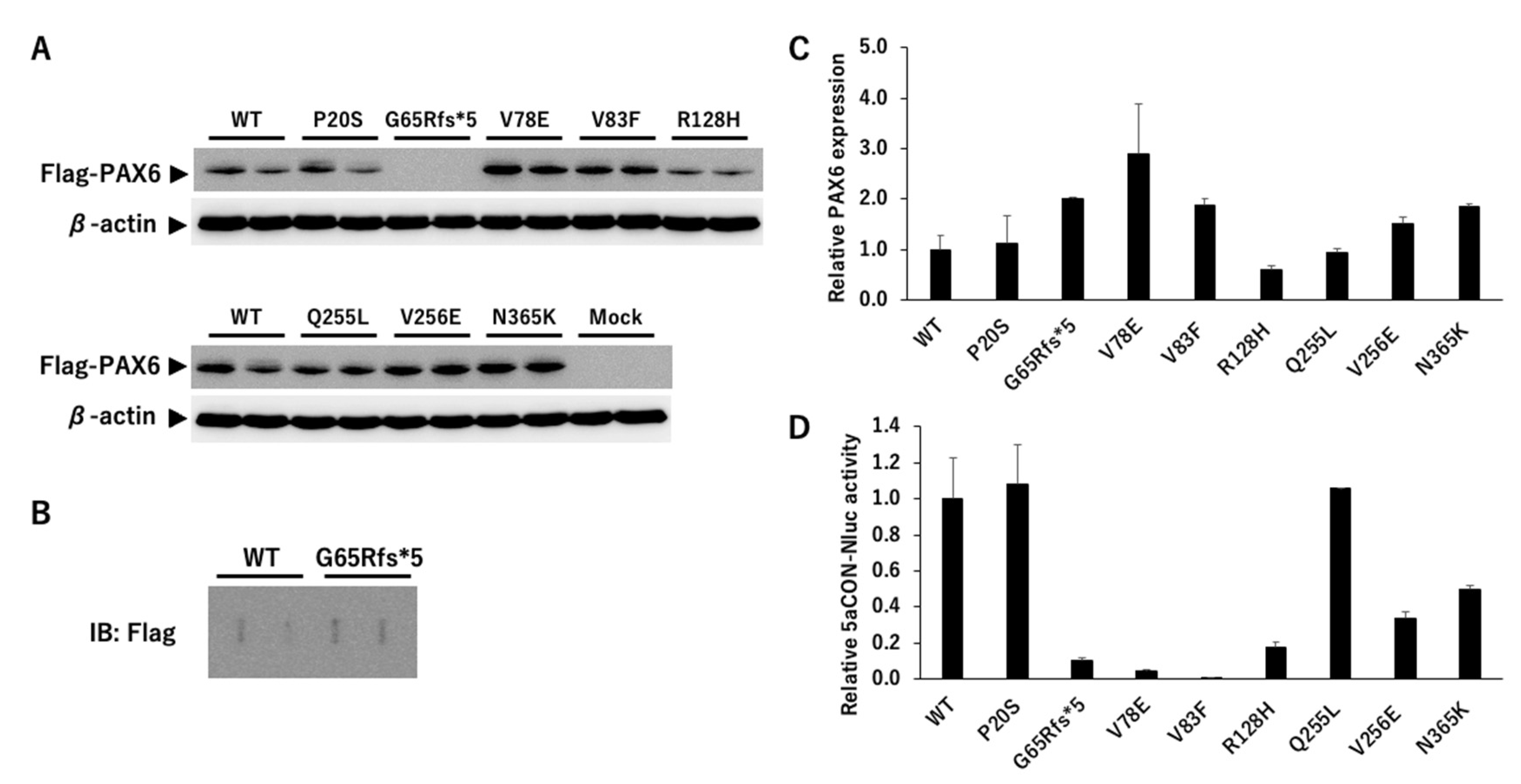
3.2.1. Expression Analysis of PAX6
3.2.2. PAX6 Mutants and Transcriptional Activity via 5aCON
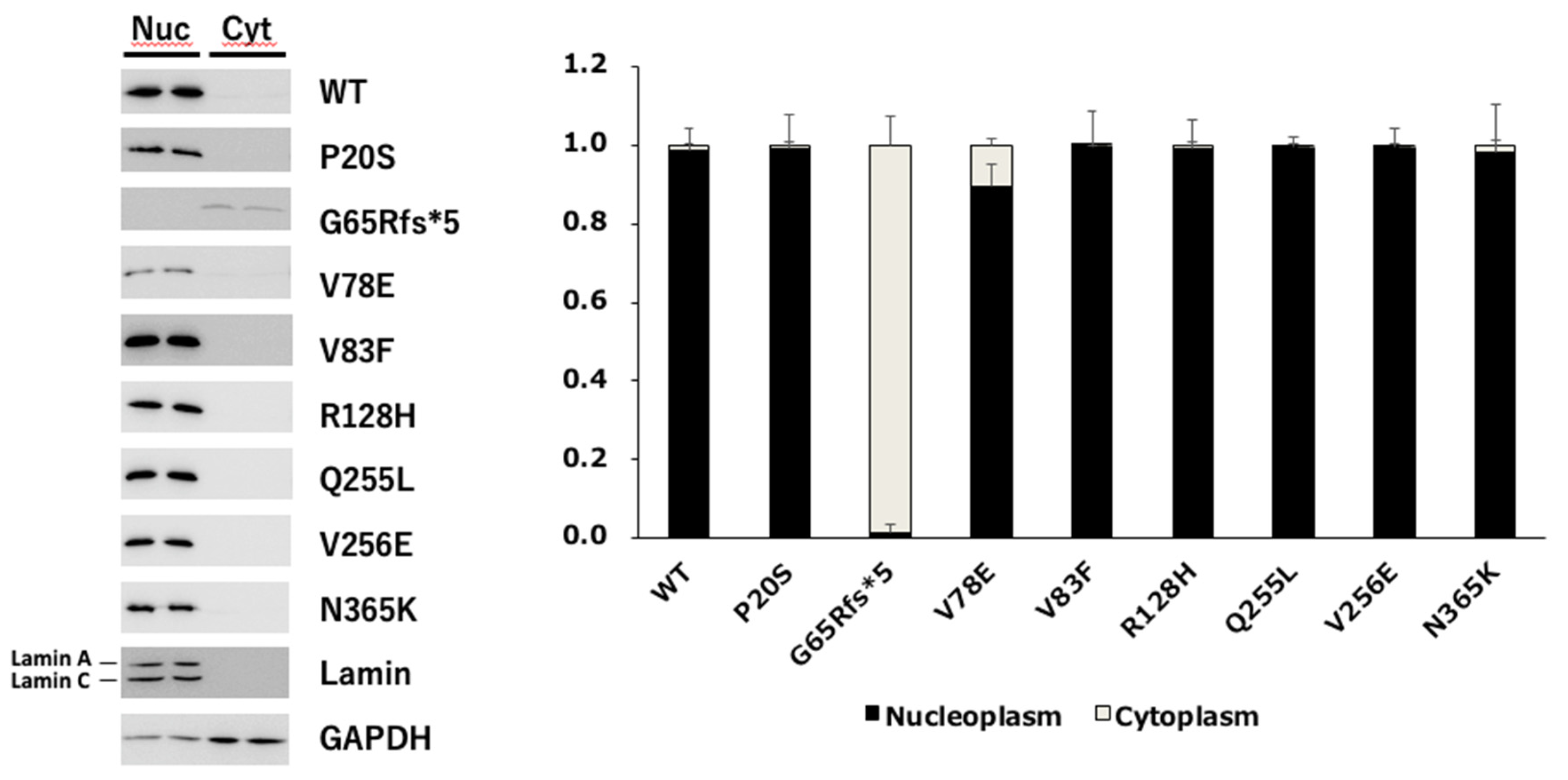
3.2.3. Nuclear Translocation Analysis of PAX6 Mutants
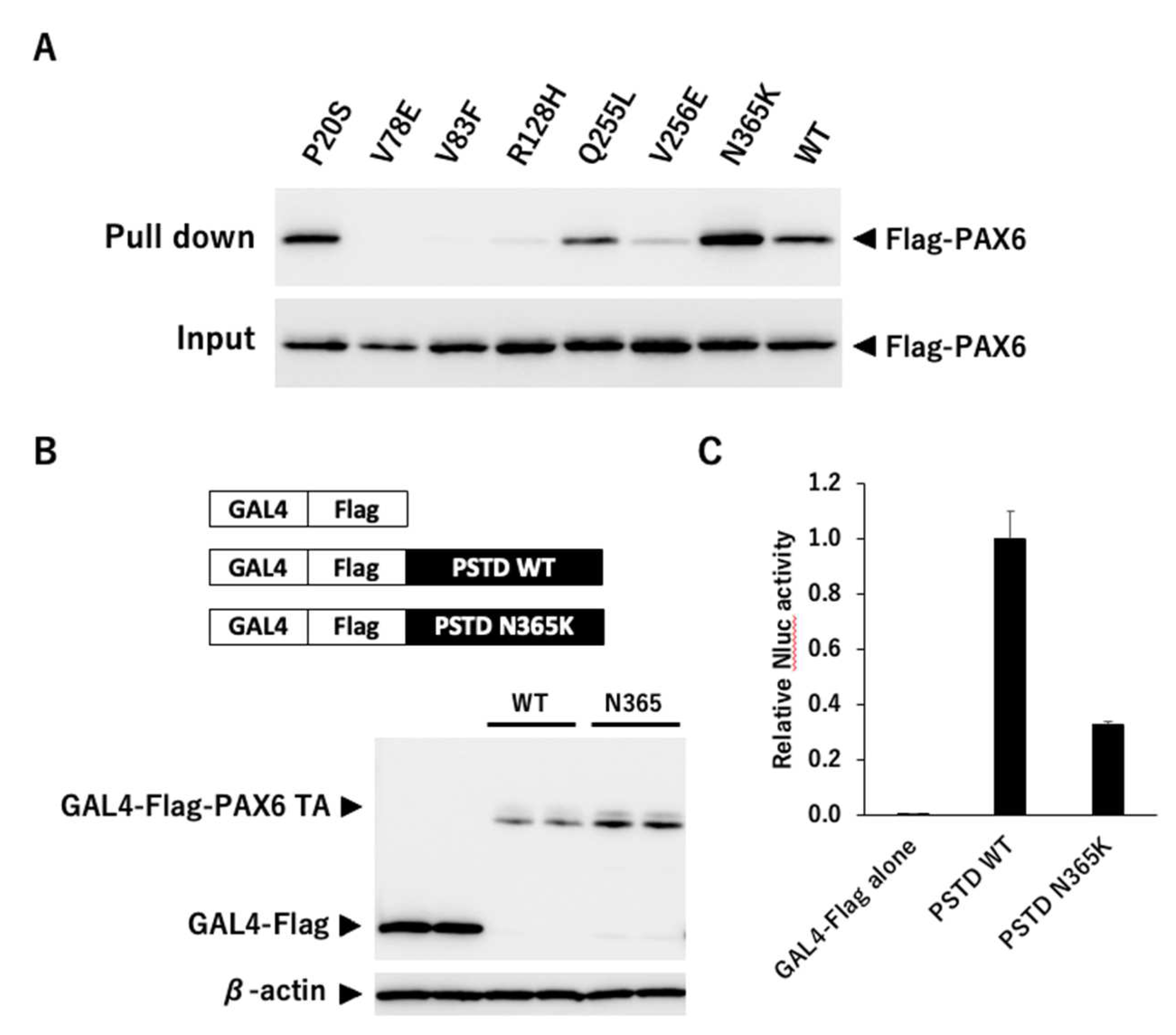
3.2.4. Binding Analysis of PAX6 Mutants and 5aCON
3.2.5. Transcription Activity Analysis using Gal4 System
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kondo, H., Foveal hypoplasia and optical coherence tomographic imaging. Taiwan J Ophthalmol 2018, 8, (4), 181-188. [CrossRef]
- Lima Cunha, D.; Arno, G.; Corton, M.; Moosajee, M., The Spectrum of PAX6 Mutations and Genotype-Phenotype Correlations in the Eye. Genes (Basel) 2019, 10, (12). [CrossRef]
- Cvekl, A.; Callaerts, P., PAX6: 25th anniversary and more to learn. Exp Eye Res 2017, 156, 10-21. [CrossRef]
- Azuma, N.; Tadokoro, K.; Asaka, A.; Yamada, M.; Yamaguchi, Y.; Handa, H.; Matsushima, S.; Watanabe, T.; Kohsaka, S.; Kida, Y.; Shiraishi, T.; Ogura, T.; Shimamura, K.; Nakafuku, M., The Pax6 isoform bearing an alternative spliced exon promotes the development of the neural retinal structure. Hum Mol Genet 2005, 14, (6), 735-45. [CrossRef]
- Jun, S.; Desplan, C., Cooperative interactions between paired domain and homeodomain. Development 1996, 122, (9), 2639-50. [CrossRef]
- Tang, H. K.; Singh, S.; Saunders, G. F., Dissection of the transactivation function of the transcription factor encoded by the eye developmental gene PAX6. J Biol Chem 1998, 273, (13), 7210-21. [CrossRef]
- Epstein, J. A.; Glaser, T.; Cai, J.; Jepeal, L.; Walton, D. S.; Maas, R. L., Two independent and interactive DNA-binding subdomains of the Pax6 paired domain are regulated by alternative splicing. Genes Dev 1994, 8, (17), 2022-34. [CrossRef]
- Azuma, N.; Nishina, S.; Yanagisawa, H.; Okuyama, T.; Yamada, M., PAX6 missense mutation in isolated foveal hypoplasia. Nat Genet 1996, 13, (2), 141-2. [CrossRef]
- Schroeder, H. W.; Orth, U.; Meyer-Konig, E.; Gal, A., [Hereditary foveal hypoplasia - clinical differentiation]. Klin Monbl Augenheilkd 2003, 220, (8), 559-62. [CrossRef]
- Hingorani, M.; Williamson, K. A.; Moore, A. T.; van Heyningen, V., Detailed ophthalmologic evaluation of 43 individuals with PAX6 mutations. Invest Ophthalmol Vis Sci 2009, 50, (6), 2581-90. [CrossRef]
- Matsushita, I.; Morita, H.; Kondo, H., Autosomal dominant foveal hypoplasia without visible macular abnormalities and PAX6 mutations. Jpn J Ophthalmol 2020, 64, (6), 635-641. [CrossRef]
- Suga, A.; Yoshitake, K.; Minematsu, N.; Tsunoda, K.; Fujinami, K.; Miyake, Y.; Kuniyoshi, K.; Hayashi, T.; Mizobuchi, K.; Ueno, S.; Terasaki, H.; Kominami, T.; Nao, I. N.; Mawatari, G.; Mizota, A.; Shinoda, K.; Kondo, M.; Kato, K.; Sekiryu, T.; Nakamura, M.; Kusuhara, S.; Yamamoto, H.; Yamamoto, S.; Mochizuki, K.; Kondo, H.; Matsushita, I.; Kameya, S.; Fukuchi, T.; Hatase, T.; Horiguchi, M.; Shimada, Y.; Tanikawa, A.; Yamamoto, S.; Miura, G.; Ito, N.; Murakami, A.; Fujimaki, T.; Hotta, Y.; Tanaka, K.; Iwata, T., Genetic characterization of 1210 Japanese pedigrees with inherited retinal diseases by whole-exome sequencing. Hum Mutat 2022, 43, (12), 2251-2264. [CrossRef]
- Adzhubei, I. A.; Schmidt, S.; Peshkin, L.; Ramensky, V. E.; Gerasimova, A.; Bork, P.; Kondrashov, A. S.; Sunyaev, S. R., A method and server for predicting damaging missense mutations. Nat Methods 2010, 7, (4), 248-9. [CrossRef]
- Davydov, E. V.; Goode, D. L.; Sirota, M.; Cooper, G. M.; Sidow, A.; Batzoglou, S., Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput Biol 2010, 6, (12), e1001025. [CrossRef]
- Ioannidis, N. M.; Rothstein, J. H.; Pejaver, V.; Middha, S.; McDonnell, S. K.; Baheti, S.; Musolf, A.; Li, Q.; Holzinger, E.; Karyadi, D.; Cannon-Albright, L. A.; Teerlink, C. C.; Stanford, J. L.; Isaacs, W. B.; Xu, J.; Cooney, K. A.; Lange, E. M.; Schleutker, J.; Carpten, J. D.; Powell, I. J.; Cussenot, O.; Cancel-Tassin, G.; Giles, G. G.; MacInnis, R. J.; Maier, C.; Hsieh, C. L.; Wiklund, F.; Catalona, W. J.; Foulkes, W. D.; Mandal, D.; Eeles, R. A.; Kote-Jarai, Z.; Bustamante, C. D.; Schaid, D. J.; Hastie, T.; Ostrander, E. A.; Bailey-Wilson, J. E.; Radivojac, P.; Thibodeau, S. N.; Whittemore, A. S.; Sieh, W., REVEL: An Ensemble Method for Predicting the Pathogenicity of Rare Missense Variants. Am J Hum Genet 2016, 99, (4), 877-885. [CrossRef]
- Jagadeesh, K. A.; Wenger, A. M.; Berger, M. J.; Guturu, H.; Stenson, P. D.; Cooper, D. N.; Bernstein, J. A.; Bejerano, G., M-CAP eliminates a majority of variants of uncertain significance in clinical exomes at high sensitivity. Nat Genet 2016, 48, (12), 1581-1586. [CrossRef]
- Kircher, M.; Witten, D. M.; Jain, P.; O'Roak, B. J.; Cooper, G. M.; Shendure, J., A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet 2014, 46, (3), 310-5. [CrossRef]
- Kumar, P.; Henikoff, S.; Ng, P. C., Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 2009, 4, (7), 1073-81. [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W. W.; Hegde, M.; Lyon, E.; Spector, E.; Voelkerding, K.; Rehm, H. L., Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015, 17, (5), 405-24. [CrossRef]
- Shimizu, N.; Watanabe, H.; Kubota, J.; Wu, J.; Saito, R.; Yokoi, T.; Era, T.; Iwatsubo, T.; Watanabe, T.; Nishina, S.; Azuma, N.; Katada, T.; Nishina, H., Pax6-5a promotes neuronal differentiation of murine embryonic stem cells. Biol Pharm Bull 2009, 32, (6), 999-1003. [CrossRef]
- Murakami, M.; Izumi, H.; Kurita, T.; Koi, C.; Morimoto, Y.; Yoshino, K., UBE2L6 is Involved in Cisplatin Resistance by Regulating the Transcription of ABCB6. Anticancer Agents Med Chem 2020, 20, (12), 1487-1496. [CrossRef]
- Yahalom, C.; Blumenfeld, A.; Hendler, K.; Wussuki-Lior, O.; Macarov, M.; Shohat, M.; Khateb, S., Mild aniridia phenotype: an under-recognized diagnosis of a severe inherited ocular disease. Graefes Arch Clin Exp Ophthalmol 2018, 256, (11), 2157-2164. [CrossRef]
- Thomas, S.; Thomas, M. G.; Andrews, C.; Chan, W. M.; Proudlock, F. A.; McLean, R. J.; Pradeep, A.; Engle, E. C.; Gottlob, I., Autosomal-dominant nystagmus, foveal hypoplasia and presenile cataract associated with a novel PAX6 mutation. Eur J Hum Genet 2014, 22, (3), 344-9. [CrossRef]
- Rufai, S. R.; Thomas, M. G.; Purohit, R.; Bunce, C.; Lee, H.; Proudlock, F. A.; Gottlob, I., Can Structural Grading of Foveal Hypoplasia Predict Future Vision in Infantile Nystagmus?: A Longitudinal Study. Ophthalmology 2020, 127, (4), 492-500. [CrossRef]
- Azuma, N.; Yamaguchi, Y.; Handa, H.; Hayakawa, M.; Kanai, A.; Yamada, M., Missense mutation in the alternative splice region of the PAX6 gene in eye anomalies. Am J Hum Genet 1999, 65, (3), 656-63. [CrossRef]
- Mishra, R.; Gorlov, I. P.; Chao, L. Y.; Singh, S.; Saunders, G. F., PAX6, paired domain influences sequence recognition by the homeodomain. J Biol Chem 2002, 277, (51), 49488-94. [CrossRef]
- Chauhan, B. K.; Yang, Y.; Cveklova, K.; Cvekl, A., Functional properties of natural human PAX6 and PAX6(5a) mutants. Invest Ophthalmol Vis Sci 2004, 45, (2), 385-92.Title of Site. Available online: URL (accessed on Day Month Year). [CrossRef]
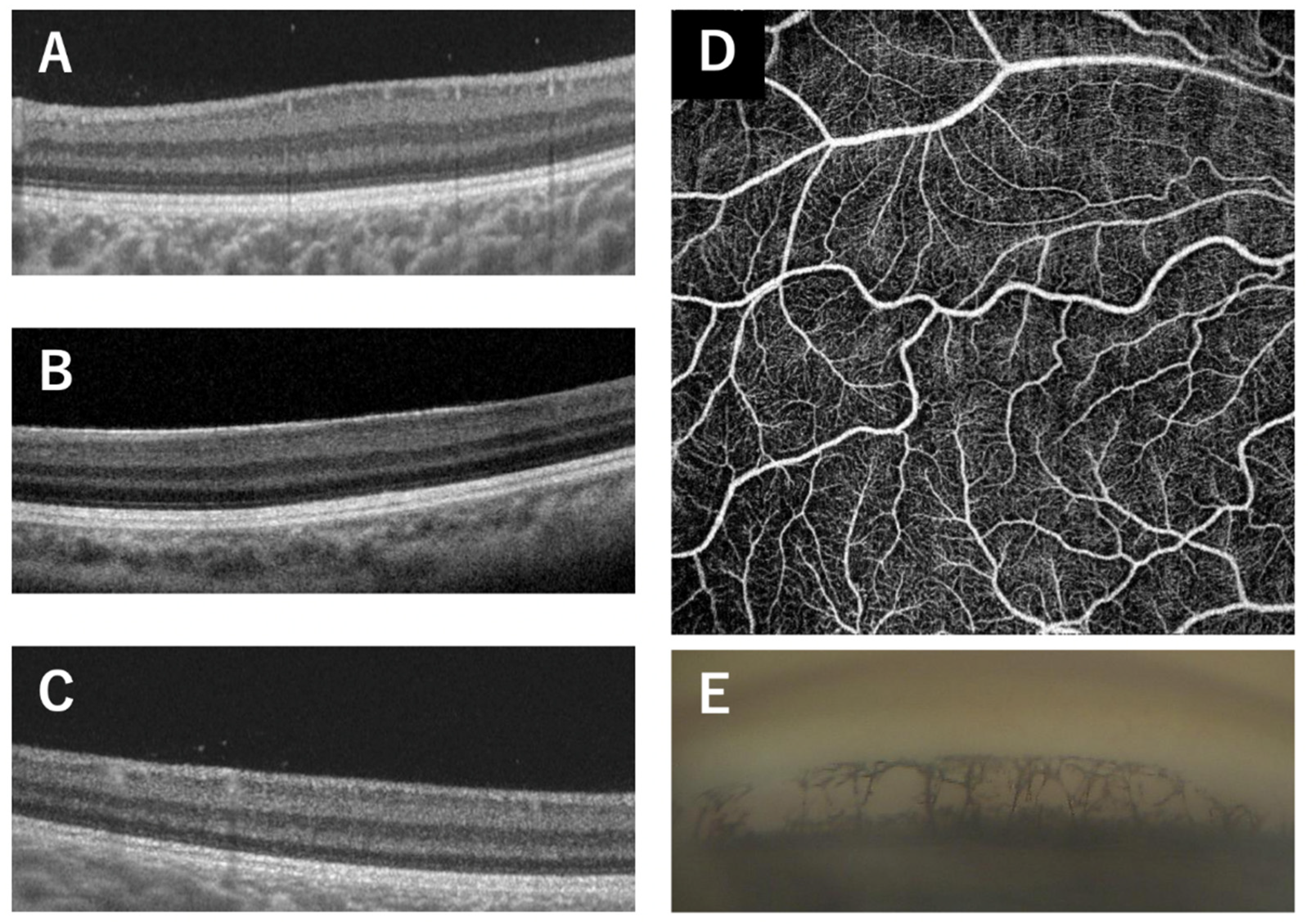
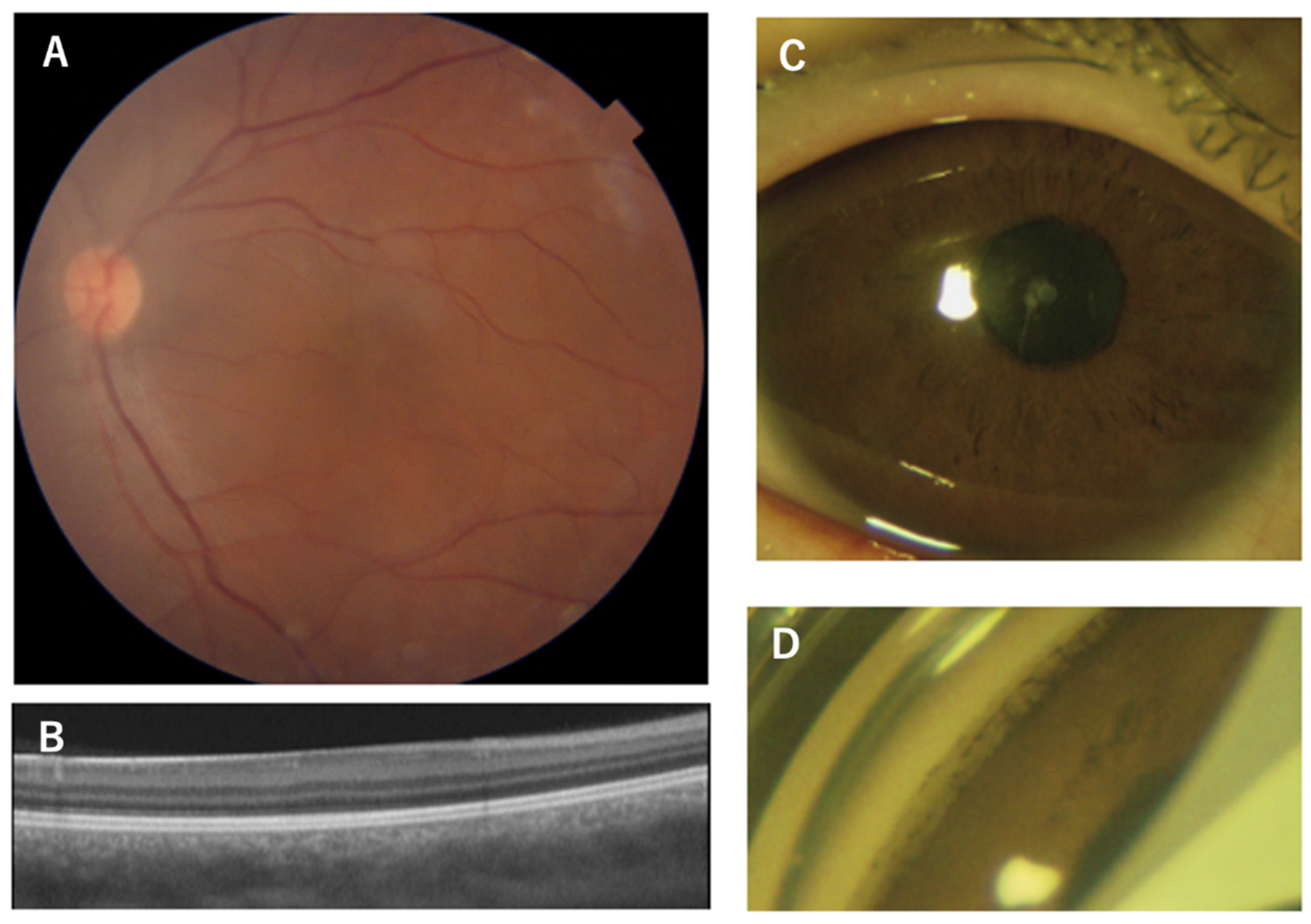
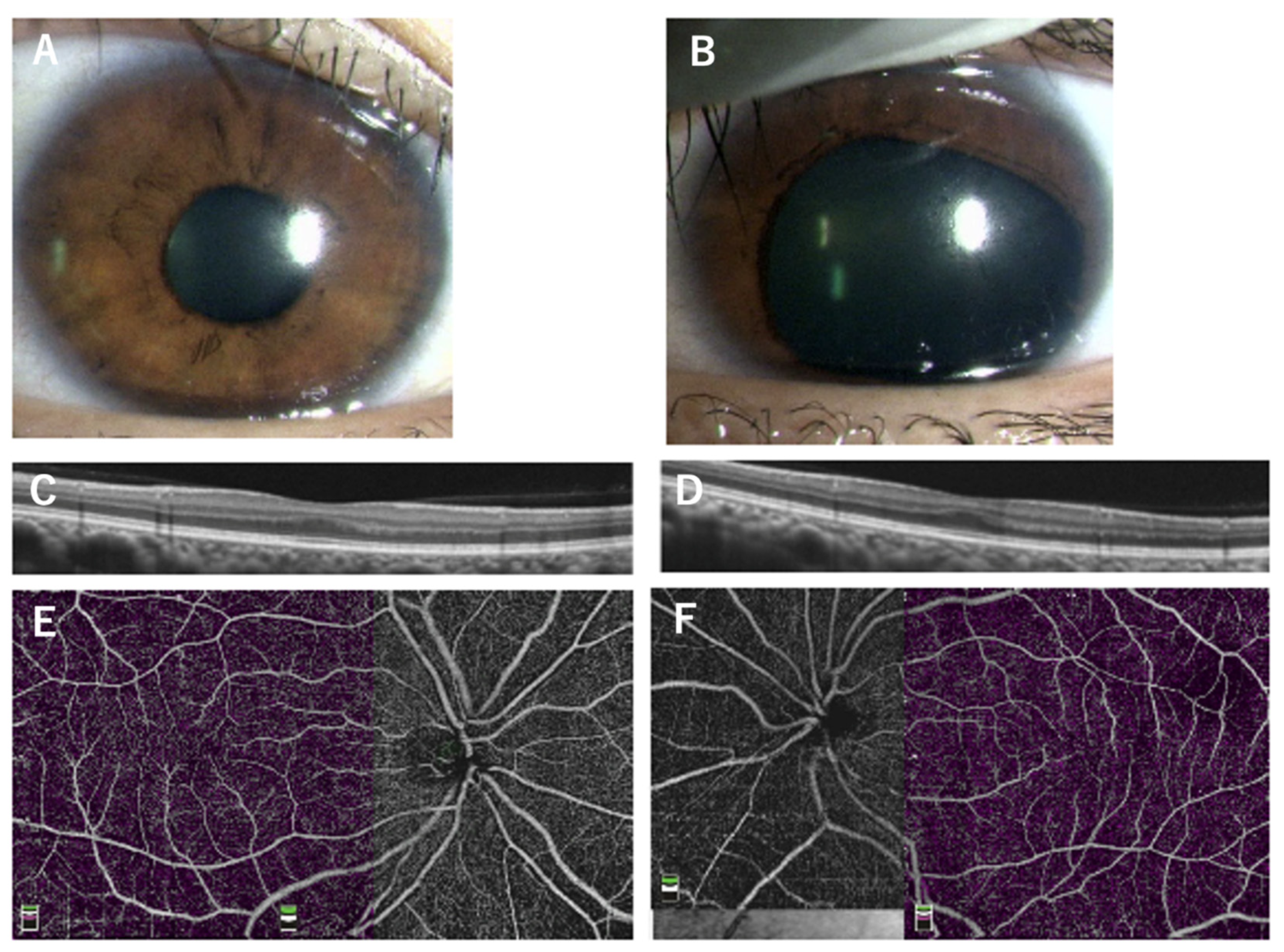
| Patient | Age, sex | Family No. | Diagnosis | Mutation (NM_000280.4) | PAX6 domain | logMAR visual acuity (R/L) | Refraction (R/L) | Nystagmus | Gonio dysgenesis | OCT grade* | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 13y, F | 1 (proband) | FVH1 | c.58C>T (p.P20S) | PD (NTS) | 0.5/0.2 | -3.75/-4.0 | present | present | 1b | [11] |
| 2 | 34y, F | 1 (mother) | FVH1 | c.58C>T (p.P20S) | 0.1/0.1 | -0.5/0 | absent | present | 1b | [11] | |
| 3 | 64y, M | 1 (grandfather) | FVH1 | c.58C>T (p.P20S) | 0.1/0 | NA | absent | present | 1b | [11] | |
| 4 | 55, M | 2 (proband) | aniridia, corneal opacity | c.150_151insA (p.G65Rfs*5) | PD (NTS) | NA | NA | NA | NA | NA | This study |
| 5 | 3y, F | 3 (proband) | FVH1 | c.233T>A (p.V78E) | PD (CTS) | 1.2/1.3 | -2.0/-2.0 | present | NA | 4 | This study |
| 6 | 33y, F | 3 (mother) | FVH1 | c.233T>A (p.V78E) | 1.0/1.0 | -10.25/-12.0 | present | present | 4 | This study | |
| 7 | 1y, M | 3 (brother) | FVH1 | c.233T>A (p.V78E) | 1.2/1.4 | +3.0/+3.75 | present | present | 4 | This study | |
| 8 | 6y, M | 4 (proband) | FVH1 | c.247G>T (p.V83F) | PD (CTS) | 0.5/0.4 | -7.125/-7.25 | absent | NA | 4 | This study |
| 9 | 10y, M | 4 (brother) | FVH1 | c.247G>T (p.V83F) | 0/0.1 | -4.5/-3.875 | absent | NA | 4 | This study | |
| 10 | 68y, M | 5 (proband) | FVH1 | c.383G>A (p.R128H) | PD (CTS) | 0.2/0.3 | NA | NA | NA | 3 | This study |
| 11 | 21y, M | 6 (proband) | FVH1 | c.764A>T (p.Q255L) | HD | 0.3/0.4 | +0.75/-1.0 | present | present | 4 | This study |
| 12 | 8y, F | 7 (proband) | aniridia, corneal opacity, secondary retinal detachments | c.767T>A (p.V256E) | HD | LP/1.7 | NA/+16.5 | NA | NA | NA | This study |
| 13 | 36y, F | 7 (mother) | unilateral partial aniridia | c.767T>A (p.V256E) | HD | NA | NA | absent | absent | 1b | This study |
| 14 | 6y, M | 8 (proband) | peripheral corneal opacity, mild iris anomaly, exotropia | c.1032+5G>A | PSTD | 0.1/0.1 | +0.25/+0.5 | absent | NA | 3 | This study |
| 15 | 41y, F | 8 (mother) | peripheral corneal opacity, mild iris anomaly | c.1032+5G>A | PSTD | 0.2/0.1 | +2.0/-1.25 | absent | NA | 3 | This study |
| 16 | 37y, M | 8 (uncle) | peripheral corneal opacity, mild iris anomaly | c.1032+5G>A | PSTD | 0.6/0.5 | -2.0/-4.0 | absent | NA | 3 | This study |
| 17 | 6y, F | 9 (proband) | FVH1 | c.1095T>G (p.N365K) | PSTD | -0.2/-0.1 | -5.5/-5.75 | absent | present | 1b | [11] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




