Submitted:
20 June 2023
Posted:
21 June 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
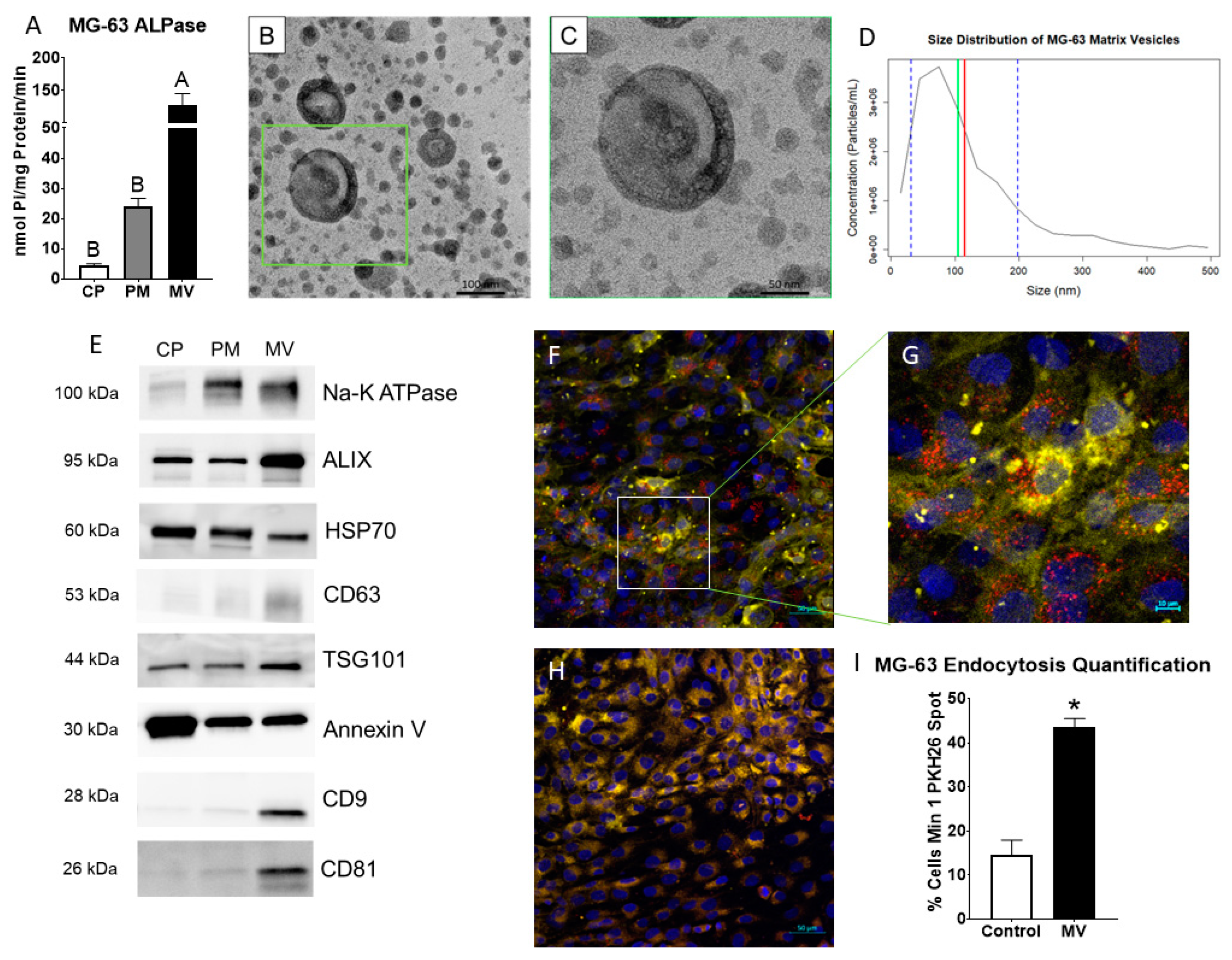
2.1. Matrix vesicles display exosomal characteristics.
2.2. MVs are endocytosed by MG-63 cells.
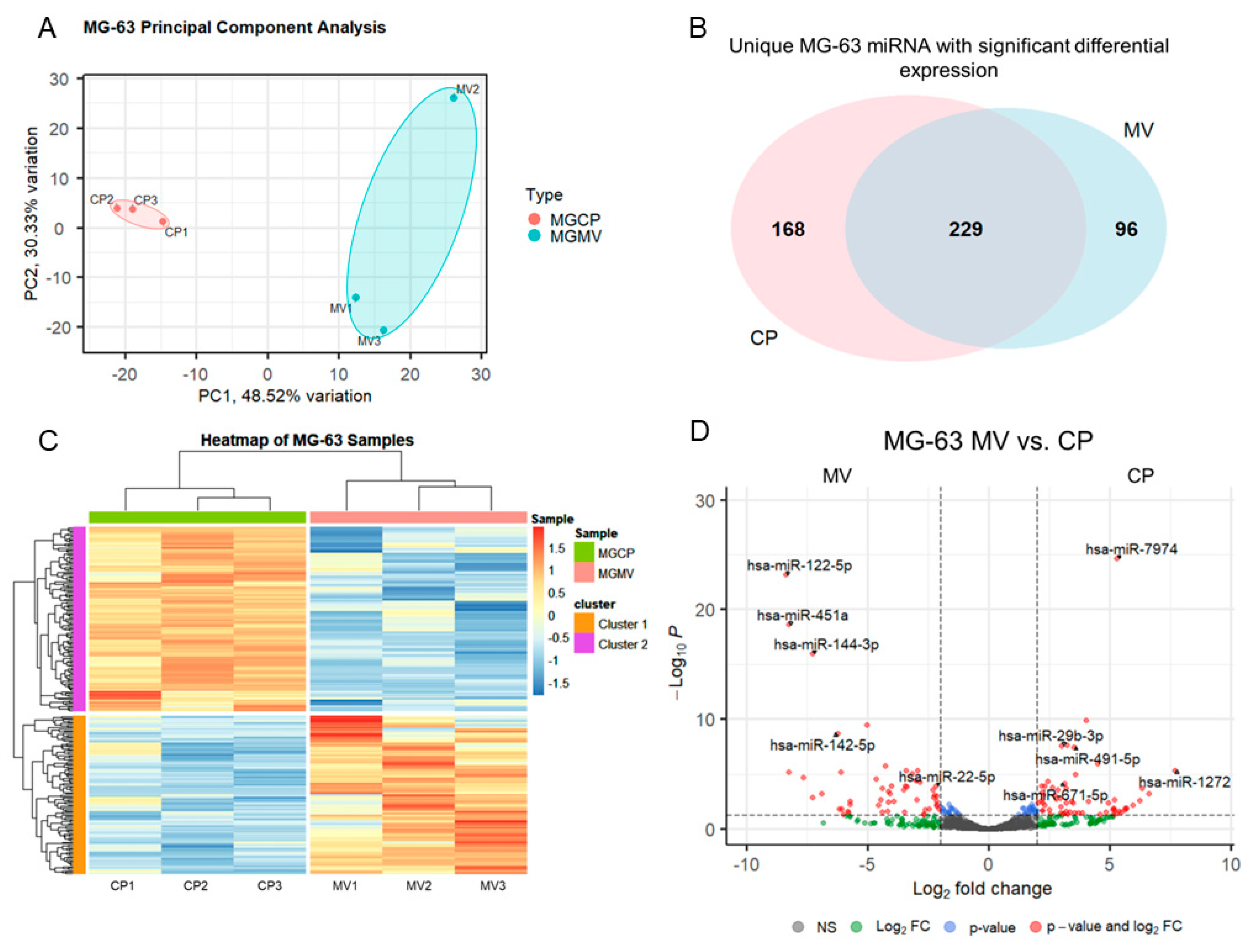
2.3. microRNAs are selectively packaged into MG-63 MVs.
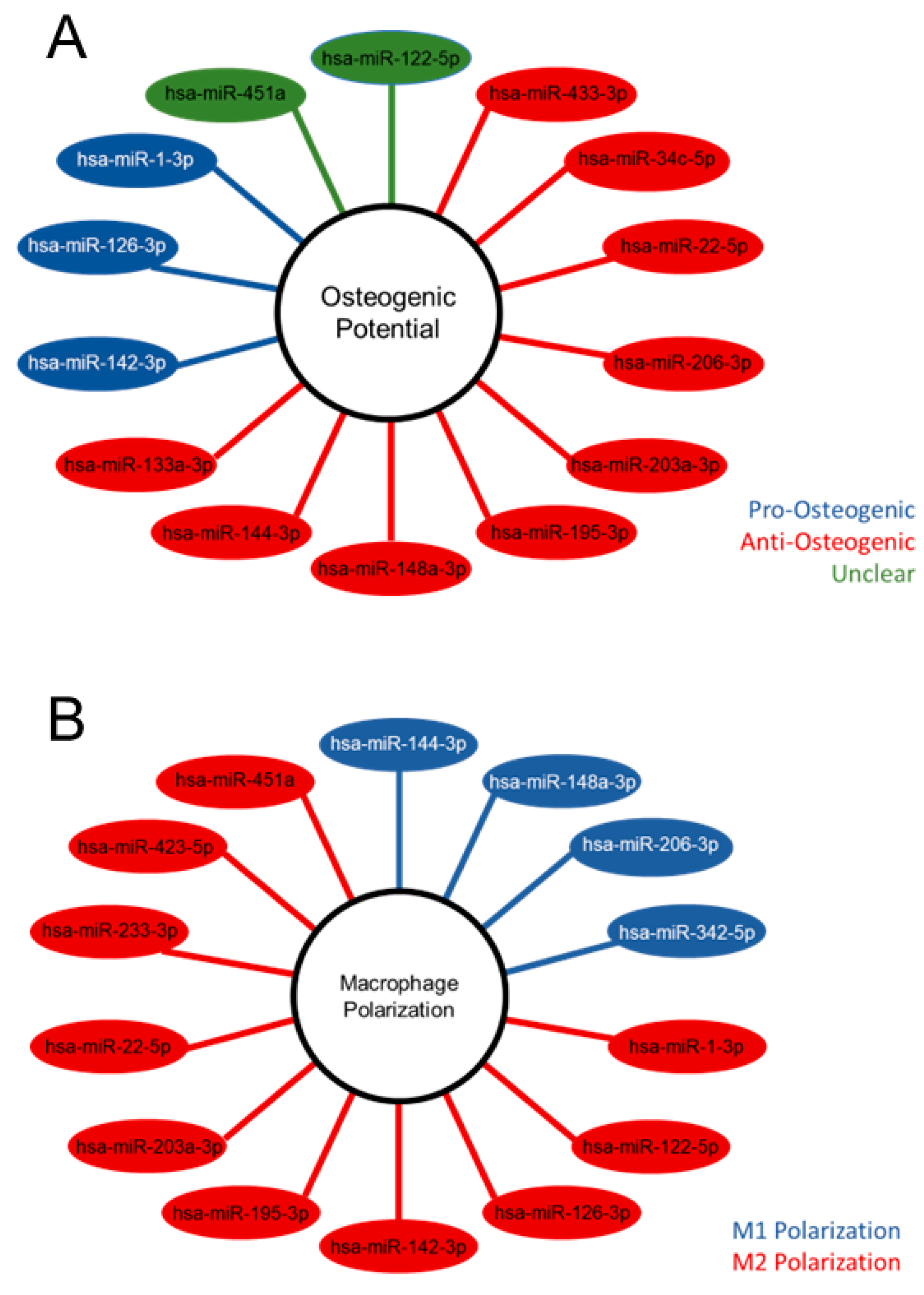
2.4. Differentially expressed MV miRNA indicate regulatory roles in osteoblasts and a regenerative role in macrophages.
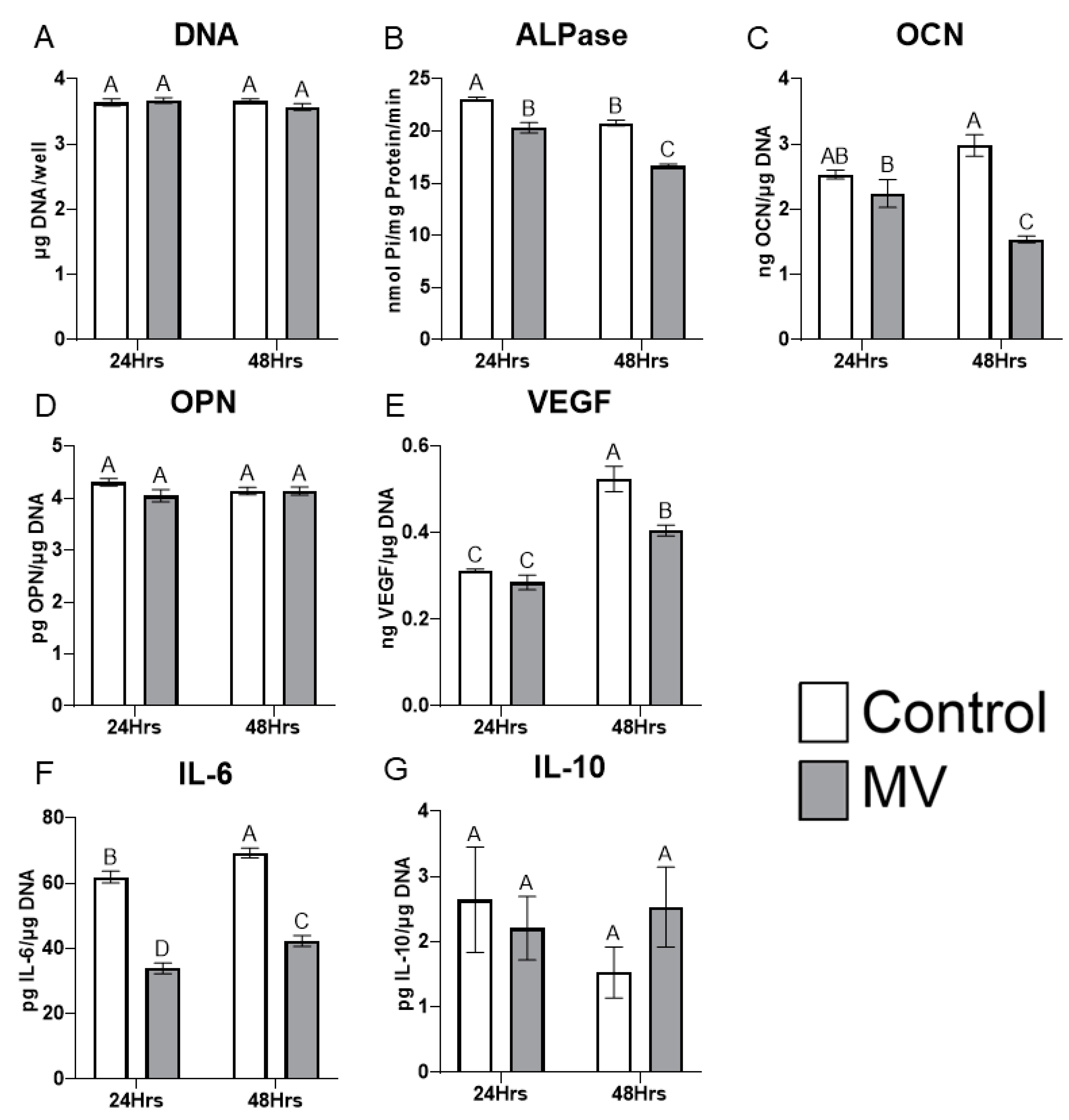
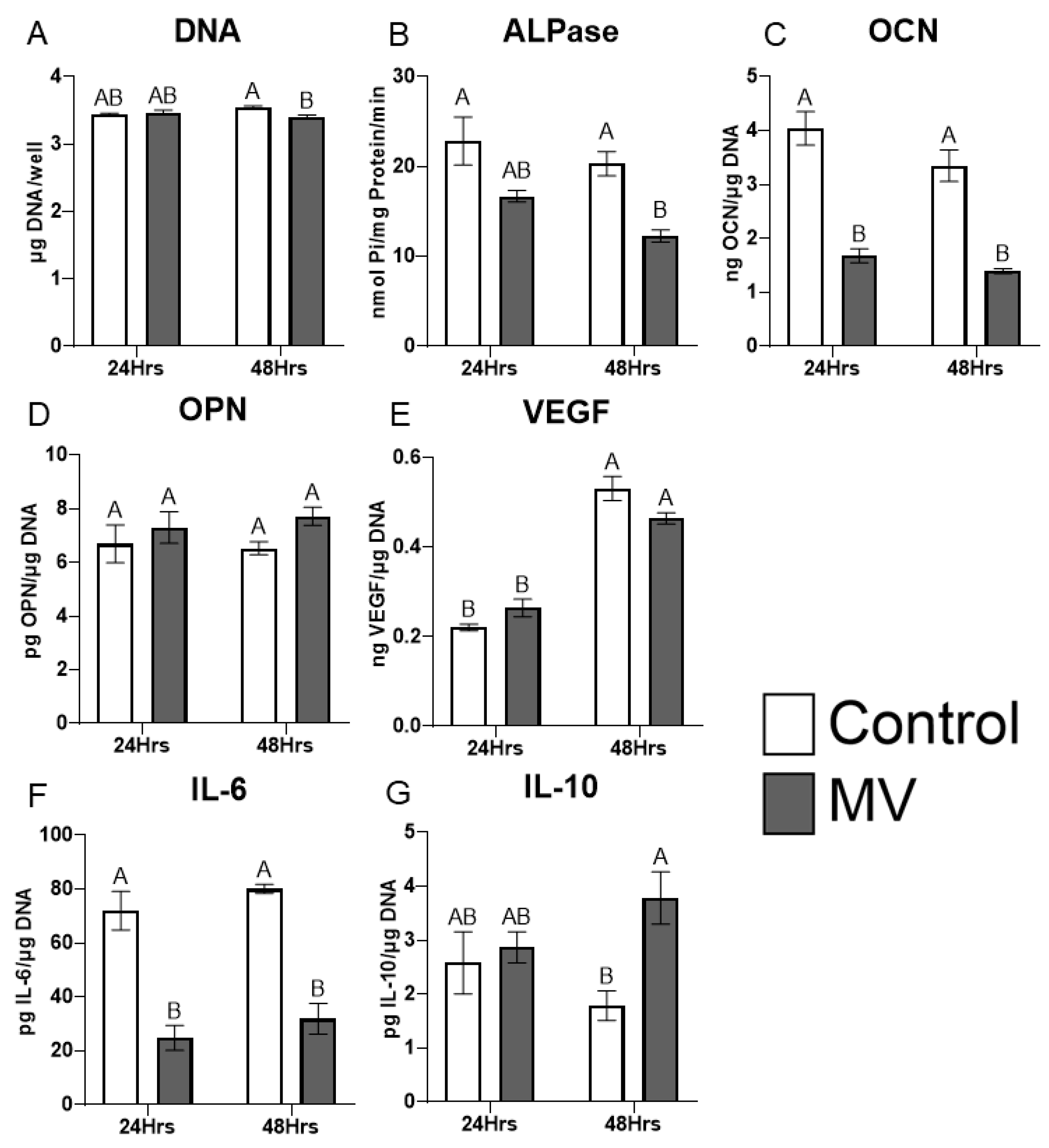
2.5. Treatment of MG-63 cells with matrix vesicles decreases markers of osteoblast differentiation.
2.6. MV composition is sensitive to culture conditions.
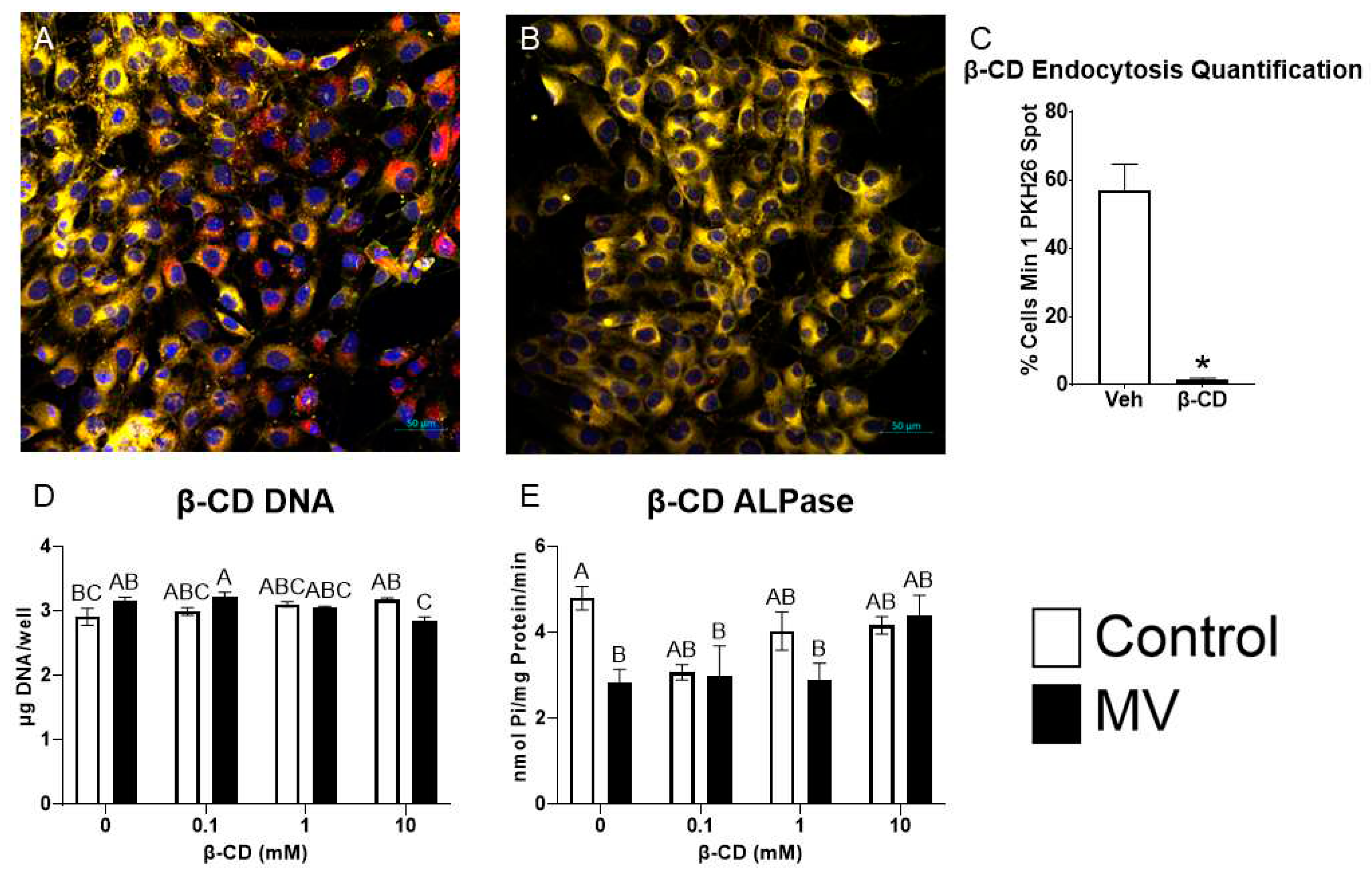
2.7. Inhibition of caveolae-mediated endocytosis prevents uptake and effect of MVs on osteoblast-like cells.
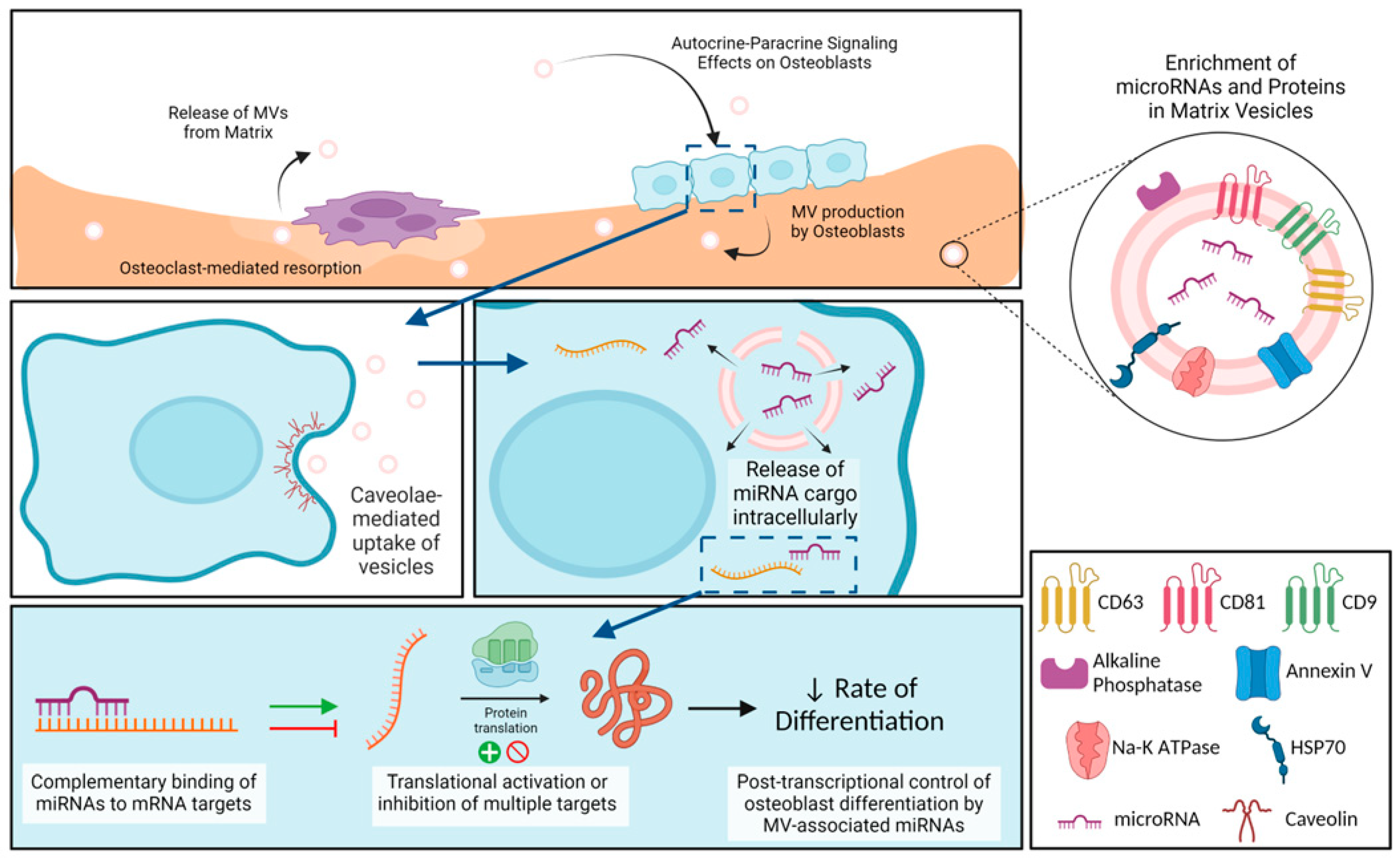
3. Discussion
4. Materials and Methods
4.1. Osteoblast Cultures
4.2. Cell Harvest and Isolation of Matrix Vesicle and Plasma Membrane Fractions
4.3. Alkaline Phosphatase Specific Activity Assay
4.4. Transmission Electron Microscopy
4.5. Nanoparticle Tracking Analysis
4.6. Western Blot
4.7. PKH26 Labeling and Confocal Microscopy
4.8. RNA Extraction and Detection
4.9. microRNA Sequencing (RNA-Seq)
4.10. Pathway Analysis of microRNA Associations
4.11. MV Treatment and Osteogenic Response
4.12. Methyl-β-cyclodextrin Endocytosis Inhibition Treatment
4.13. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Azoidis, I.; Cox, S.C.; Davies, O.G. The role of extracellular vesicles in biomineralisation: Current perspective and application in regenerative medicine. J. Tissue Eng. 2018, 9, 2041731418810130. [Google Scholar] [CrossRef] [PubMed]
- Anderson, H.C.; Garimella, R.; Tague, S.E. The role of matrix vesicles in growth plate development and biomineralization. Front. Biosci. 2005, 10, 822–837. [Google Scholar] [CrossRef]
- Dean, D.D.; Schwartz, Z.; Bonewald, L.; Muniz, O.E.; Morales, S.; Gomez, R.; Brooks, B.P.; Qiao, M.; Howell, D.S.; Boyan, B.D. Matrix Vesicles produced by osteoblast-like cells in culture become significantly enriched in proteoglycan-degrading metalloproteinases after addition of β-glycerophosphate and ascorbic acid. Calcif. Tissue Int. 1994, 54, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Rodriguez, N.E.; Zhao, J.; Ramey, A.N.; Hyzy, S.L.; Boyan, B.D.; Schwartz, Z. Selective enrichment of microRNAs in extracellular matrix vesicles produced by growth plate chondrocytes. Bone 2016, 88, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Nahar, N.N.; Missana, L.R.; Garimella, R.; Tague, S.E.; Anderson, H.C. Matrix vesicles are carriers of bone morphogenetic proteins (BMPs), Vascular endothelial growth factor (VEGF), and noncollagenous matrix proteins. J. Bone Miner. Metab. 2008, 26, 514–519. [Google Scholar] [CrossRef]
- Boyan, B.D.; Schwartz, Z.; Swain, L.D.; Khare, A. Role of lipids in calcification of cartilage. Anat. Rec. 1989, 224, 211–219. [Google Scholar] [CrossRef]
- Dean, D.D.; Boyan, B.D.; Muniz, O.E.; Howell, D.S.; Schwartz, Z. Vitamin D metabolites regulate matrix vesicle metalloproteinase content in a cell maturation-dependent manner. Calcif. Tissue Int. 1996, 59, 109–116. [Google Scholar] [CrossRef]
- Sylvia, V.L.; Schwartz, Z.; Holmes, S.C.; Dean, D.D.; Boyan, B.D. 24,25-(OH)2D3 regulation of matrix vesicle protein kinase C occurs both during biosynthesis and in the extracellular matrix. Calcif. Tissue Int. 1997, 61, 313–321. [Google Scholar] [CrossRef]
- Schwartz, Z.; Sylvia, V.L.; Larsson, D.; Nemere, I.; Casasola, D.; Dean, D.D.; Boyan, B.D. 1α,25(OH)2D3 regulates chondrocyte matrix vesicle protein kinase C (PKC) directly via G-protein-dependent mechanisms and indirectly via incorporation of PKC during matrix vesicle biogenesis. J. Biol. Chem. 2002, 277, 11828–11837. [Google Scholar] [CrossRef]
- Boyan, B.D.; Schwartz, Z.; Park-Snyder, S.; Dean, D.D.; Yang, F.; Twardzik, D.; Bonewald, L.F. Latent transforming growth factor-β is produced by chondrocytes and activated by extracellular matrix vesicles upon exposure to 1,25-(OH)2D3. J. Biol. Chem. 1994, 269, 28374–28381. [Google Scholar] [CrossRef]
- Mohr, A.M.; Mott, J.L. Overview of microRNA biology. Semin. Liver Dis. 2015, 35, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.X.; Rothenberg, M.E. MicroRNA. J. Allergy Clin. Immunol. 2018, 141, 1202–1207. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; McClure, M.J.; Zhao, J.; Ramey, A.N.; Asmussen, N.; Hyzy, S.L.; Schwartz, Z.; Boyan, B.D. MicroRNA contents in matrix vesicles produced by growth plate chondrocytes are cell maturation dependent. Sci. Rep. 2018, 8, 3609. [Google Scholar] [CrossRef] [PubMed]
- Asmussen, N.C.; Cohen, D.J.; Lin, Z.; McClure, M.J.; Boyan, B.D.; Schwartz, Z. Specific microRNAs found in extracellular matrix vesicles regulate proliferation and differentiation in growth plate chondrocytes. Calcif. Tissue Int. 2021. [Google Scholar] [CrossRef] [PubMed]
- Bonewald, L.F.; Schwartz, Z.; Swain, L.D.; Boyan, B.D. Stimulation of matrix vesicle enzyme activity in osteoblast-like cells by 1,25(OH)2D3 and transforming growth factor beta (TGF Beta). Bone Miner. 1992, 17, 139–144. [Google Scholar] [CrossRef]
- Hasegawa, T. Ultrastructure and biological function of matrix vesicles in bone mineralization. Histochem. Cell Biol. 2018, 149, 289–304. [Google Scholar] [CrossRef]
- Boyan, B.D.; Schwartz, Z.; Swain, L.D. Matrix vesicles as a marker of endochondral ossification. Connect. Tissue Res. 1990, 24, 67–75. [Google Scholar] [CrossRef]
- Bonucci, E. Bone mineralization. Front. Biosci. 2012, 17, 100. [Google Scholar] [CrossRef]
- Cui, Y.; Luan, J.; Li, H.; Zhou, X.; Han, J. Exosomes derived from mineralizing osteoblasts promote ST2 cell osteogenic differentiation by alteration of microRNA expression. FEBS Lett. 2016, 590, 185–192. [Google Scholar] [CrossRef]
- Qin, Y.; Wang, L.; Gao, Z.; Chen, G.; Zhang, C. Bone marrow stromal/stem cell-derived extracellular vesicles regulate osteoblast activity and differentiation in vitro and promote bone regeneration in vivo. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef]
- Xu, J.; Wang, Y.; Hsu, C.Y.; Gao, Y.; Meyers, C.A.; Chang, L.; Zhang, L.; Broderick, K.; Ding, C.; Peault, B.; et al. Human perivascular stem cell-derived extracellular vesicles mediate bone repair. Elife 2019, 8, 1–23. [Google Scholar] [CrossRef]
- Yoshiko, Y.; Minamizaki, T. Emerging roles of microRNAs as extracellular vesicle cargo secreted from osteoblasts. J. Oral Biosci. 2020, 62, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Bjørge, I.M.; Kim, S.Y.; Mano, J.F.; Kalionis, B.; Chrzanowski, W. Extracellular vesicles, exosomes and shedding vesicles in regenerative medicine-a new paradigm for tissue repair. Biomater. Sci. 2018, 6, 60–78. [Google Scholar] [CrossRef]
- Gu, H.; Shi, S.; Xiao, F.; Huang, Z.; Xu, J.; Chen, G.; Zhou, K.; Lu, L.; Yin, X. miR-1-3p regulates the differentiation of mesenchymal stem cells to prevent osteoporosis by targeting secreted frizzled-related protein 1. Bone 2020, 137, 115444. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.G.; Konicke, K.; Kristianto, J.; Gustavson, A.; Garbo, R.; Wang, X.; Yuan, B.; Blank, R.D. Endothelin signaling regulates mineralization and posttranscriptionally regulates SOST in TMOb cells via miR 126-3p. Physiol. Rep. 2017, 5, e13088. [Google Scholar] [CrossRef] [PubMed]
- Bedene, A.; Mencej Bedrač, S.; Ješe, L.; Marc, J.; Vrtačnik, P.; Preželj, J.; Kocjan, T.; Kranjc, T.; Ostanek, B. miR-148a the epigenetic regulator of bone homeostasis is increased in plasma of osteoporotic postmenopausal women. Wien. Klin. Wochenschr. 2016, 128, 519–526. [Google Scholar] [CrossRef]
- Ji, Y.; Fang, Q.Y.; Wang, S.N.; Zhang, Z.W.; Hou, Z.J.; Li, J.N.; Fu, S.Q. Lnc-RNA BLACAT1 regulates differentiation of bone marrow stromal stem cells by targeting miR-142-5pin osteoarthritis. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 2893–2901. [Google Scholar] [CrossRef]
- Tu, M.; Tang, J.; He, H.; Cheng, P.; Chen, C. miR-142-5p promotes bone repair by maintaining osteoblast activity. J. Bone Miner. Metab. 2017, 35, 255–264. [Google Scholar] [CrossRef]
- Zhao, R.; Zhu, Y.; Sun, B. Exploration of the effect of mmu-miR-142-5p on osteoblast and the mechanism. Cell Biochem. Biophys. 2015, 71, 255–260. [Google Scholar] [CrossRef]
- Wang, Q.; Li, Y.; Zhang, Y.; Ma, L.; Lin, L.; Meng, J.; Jiang, L.; Wang, L.; Zhou, P.; Zhang, Y. LncRNA MEG3 inhibited osteogenic differentiation of bone marrow mesenchymal stem cells from postmenopausal osteoporosis by targeting miR-133a-3p. Biomed. Pharmacother. 2017, 89, 1178–1186. [Google Scholar] [CrossRef]
- Li, M.; Shen, Y.-J.; Chai, S.; Bai, Y.-L.; Li, Z.-H. miR-133a-3p inhibits the osteogenic differentiation of bone marrow mesenchymal stem cells by regulating ankyrin repeat domain 44. Gen. Physiol. Biophys. 2021, 40, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Bourgery, M.; Ekholm, E.; Hiltunen, A.; Heino, T.J.; Pursiheimo, J.P.; Bendre, A.; Yatkin, E.; Laitala, T.; Määttä, J.; Säämänen, A.M. signature of circulating small non-coding RNAs during early fracture healing in mice. Bone Reports 2022, 17, 101627. [Google Scholar] [CrossRef]
- Peng, J.; Zhan, Y.; Zong, Y. METTL3-Mediated LINC00657 promotes osteogenic differentiation of mesenchymal stem cells via miR-144-3p/BMPR1B axis. Cell Tissue Res. 2022, 388, 301–312. [Google Scholar] [CrossRef]
- Li, N.; Liu, L.; Liu, Y.; Luo, S.; Song, Y.; Fang, B. miR-144-3p suppresses osteogenic differentiation of BMSCs from patients with aplastic anemia through repression of TET2. Mol. Ther. - Nucleic Acids 2020, 19, 619–626. [Google Scholar] [CrossRef]
- Sun, Z.; Wu, F.; Yang, Y.; Liu, F.; Mo, F.; Chen, J.; Wang, G.; Zhang, B. miR-144-3p inhibits bmsc proliferation and osteogenic differentiation via targeting FZD4 in steroid-associated osteonecrosis. Curr. Pharm. Des. 2019, 25, 4806–4812. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.X.; Zhang, J.F.; Xu, J.; Xu, L.L.; Wu, T.Y.; Wang, B.; Pan, X.H.; Li, G. MicroRNA-144-3p inhibits bone formation in distraction osteogenesis through targeting Connexin 43. Oncotarget 2017, 8, 89913–89922. [Google Scholar] [CrossRef]
- Liu, N.; Sun, Y. MicroRNA-148a-3p-targeting P300 protects against osteoblast differentiation and osteoporotic bone reconstruction. Regen. Med. 2021, 16, 435–449. [Google Scholar] [CrossRef]
- Tian, L.; Zheng, F.; Li, Z.; Wang, H.; Yuan, H.; Zhang, X.; Ma, Z.; Li, X.; Gao, X.; Wang, B. miR-148a-3p regulates adipocyte and osteoblast differentiation by targeting lysine-specific demethylase 6b. Gene 2017, 627, 32–39. [Google Scholar] [CrossRef]
- Almeida, M.I.; Silva, A.M.; Vasconcelos, D.M.; Almeida, C.R.; Caires, H.; Pinto, M.T.; Calin, G.A.; Santos, S.G.; Barbosa, M.A. miR-195 in human primary mesenchymal stromal/stem cells regulates proliferation, osteogenesis and paracrine effect on angiogenesis. Oncotarget 2016, 7, 7–22. [Google Scholar] [CrossRef] [PubMed]
- Fan, F.Y.; Deng, R.; Qiu, L.; Wen, Q.; Zeng, Y.; Gao, L.; Zhang, C.; Kong, P.; Zhong, J.; Zeng, N.; et al. miR-203a-3p.1 is involved in the regulation of osteogenic differentiation by directly targeting Smad9 in MM-MSCs. Oncol. Lett. 2019, 18, 6339–6346. [Google Scholar] [CrossRef]
- Zhang, B.; Huo, S.; Cen, X.; Pan, X.; Huang, X.; Zhao, Z. CircAKT3 positively regulates osteogenic differentiation of human dental pulp stromal cells via miR-206/CX43 axis. Stem Cell Res. Ther. 2020, 11, 531. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Chang, B.; Zheng, G.; Du, S.; Li, X. quercetin stimulates osteogenic differentiation of bone marrow stromal cells through miRNA-206/Connexin 43 pathway. Am. J. Transl. Res. 2020, 12, 2062–2070. [Google Scholar] [PubMed]
- Chen, Y.; Yang, Y.R.; Fan, X.L.; Lin, P.; Yang, H.; Chen, X.Z.; Xu, X.D. miR-206 inhibits osteogenic differentiation of bone marrow mesenchymal stem cells by targetting glutaminase. Biosci. Rep. 2019, 39, BSR20181108. [Google Scholar] [CrossRef] [PubMed]
- Inose, H.; Ochi, H.; Kimura, A.; Fujita, K.; Xu, R.; Sato, S.; Iwasaki, M.; Sunamura, S.; Takeuchi, Y.; Fukumoto, S.; et al. A microRNA regulatory mechanism of osteoblast differentiation. Proc. Natl. Acad. Sci. U. S. A. 2009, 106, 20794–20799. [Google Scholar] [CrossRef]
- Li, W.; Li, L.; Cui, R.; Chen, X.; Hu, H.; Qiu, Y. Bone marrow mesenchymal stem cells derived exosomal Lnc TUG1 promotes bone fracture recovery via miR-22-5p/Anxa8 axis. Hum. Cell 2023, 36, 1041–1053. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, Y.; Li, Y.; Wen, T. MALAT1 enhanced the proliferation of human osteoblasts treated with ultra-high molecular weight polyethylene by targeting VEGF via miR-22-5p. Int. J. Mol. Med. 2018, 41, 1536–1546. [Google Scholar] [CrossRef]
- Wei, J.; Shi, Y.; Zheng, L.; Zhou, B.; Inose, H.; Wang, J.; Guo, X.E.; Grosschedl, R.; Karsenty, G. miR-34s inhibit osteoblast proliferation and differentiation in the mouse by targeting SATB2. J. Cell Biol. 2012, 197, 509–521. [Google Scholar] [CrossRef]
- Garcia, J.; Smith, S.S.; Karki, S.; Drissi, H.; Hrdlicka, H.H.; Youngstrom, D.W.; Delany, A.M. miR-433-3p suppresses bone formation and mRNAs critical for osteoblast function in mice. J. Bone Miner. Res. 2021, 36, 1808–1822. [Google Scholar] [CrossRef]
- Kim, E.J.; Kang, I.H.; Lee, J.W.; Jang, W.G.; Koh, J.T. miR-433 mediates ERRγ-suppressed osteoblast differentiation via direct targeting to Runx2 mRNA in C3H10T1/2 cells. Life Sci. 2013, 92, 562–568. [Google Scholar] [CrossRef]
- Liao, W.; Ning, Y.; Xu, H.J.; Zou, W.Z.; Hu, J.; Liu, X.Z.; Yang, Y.; Li, Z.H. BMSC-derived exosomes carrying microRNA-122-5p promote proliferation of osteoblasts in osteonecrosis of the femoral head. Clin. Sci. 2019, 133, 1955–1975. [Google Scholar] [CrossRef]
- Lu, X.D.; Han, W.X.; Liu, Y.X. Suppression of miR-451a accelerates osteogenic differentiation and inhibits bone loss via Bmp6 signaling during osteoporosis. Biomed. Pharmacother. 2019, 120, 109378. [Google Scholar] [CrossRef] [PubMed]
- Karvande, A.; Kushwaha, P.; Ahmad, N.; Adhikary, S.; Kothari, P.; Tripathi, A.K.; Khedgikar, V.; Trivedi, R. Glucose dependent miR-451a expression contributes to parathyroid hormone mediated osteoblast differentiation. Bone 2018, 117, 98–115. [Google Scholar] [CrossRef] [PubMed]
- Shi, R.; Jin, Y.; Zhao, S.; Yuan, H.; Shi, J.; Zhao, H. Hypoxic ADSC-derived exosomes enhance wound healing in diabetic mice via delivery of circ-Snhg11 and induction of M2-like macrophage polarization. Biomed. Pharmacother. 2022, 153, 113463. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.W.; Hu, Y.R.; Zhao, J.Y.; Li, S.F.; Ma, X.; Wu, S.G.; Lu, J.B.; Qiu, Y.R.; Sha, Y.H.; Wang, Y.C.; et al. An agomir of miR-144-3p accelerates plaque formation through impairing reverse cholesterol transport and promoting pro-inflammatory cytokine production. PLoS One 2014, 9, e94997. [Google Scholar] [CrossRef]
- Bai, X.; Li, J.; Li, L.; Liu, M.; Liu, Y.; Cao, M.; Tao, K.; Xie, S.; Hu, D. Extracellular vesicles from adipose tissue-derived stem cells affect Notch-miR148a-3p axis to regulate polarization of macrophages and alleviate sepsis in mice. Front. Immunol. 2020, 11, 1391. [Google Scholar] [CrossRef]
- Huang, F.; Zhao, J.L.; Wang, L.; Gao, C.C.; Liang, S.Q.; An, D.J.; Bai, J.; Chen, Y.; Han, H.; Qin, H.Y. miR-148a-3p mediates Notch signaling to promote the differentiation and M1 activation of macrophages. Front. Immunol. 2017, 8, 1327. [Google Scholar] [CrossRef]
- Wei, Y.; Nazari-Jahantigh, M.; Chan, L.; Zhu, M.; Heyll, K.; Corbalán-Campos, J.; Hartmann, P.; Thiemann, A.; Weber, C.; Schober, A. The microRNA-342-5p fosters inflammatory macrophage activation through an Akt1- and microRNA-155-dependent pathway during atherosclerosis. Circulation 2013, 127, 1609–1619. [Google Scholar] [CrossRef]
- Liu, N.; Wang, X.; Steer, C.J.; Song, G. MicroRNA-206 promotes the recruitment of CD8+ T cells by driving M1 polarisation of kupffer cells. Gut 2022, 71, 1642–1655. [Google Scholar] [CrossRef]
- Wu, J.; Liao, Y.; Li, D.; Zhu, Z.; Zhang, L.; Wu, Z.; He, P.; Wang, L. Extracellular vesicles derived from Trichinella Spiralis larvae promote the polarization of macrophages to M2b type and inhibit the activation of fibroblasts. Front. Immunol. 2022, 13, 974332. [Google Scholar] [CrossRef]
- Li, K.; Yan, G.; Huang, H.; Zheng, M.; Ma, K.; Cui, X.; Lu, D.; Zheng, L.; Zhu, B.; Cheng, J.; et al. Anti-inflammatory and immunomodulatory effects of the extracellular vesicles derived from human umbilical cord mesenchymal stem cells on osteoarthritis via M2 macrophages. J. Nanobiotechnology 2022, 20, 38. [Google Scholar] [CrossRef]
- Coulson, D.J.; Bakhashab, S.; Latief, J.S.; Weaver, J.U. miR-126, IL-7, CXCR1/2 receptors, inflammation and circulating endothelial progenitor cells: The study on targets for treatment pathways in a model of subclinical cardiovascular disease (Type 1 diabetes mellitus). J. Transl. Med. 2021, 19, 140. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Ming, J.; Li, Y.; Li, B.; Deng, M.; Ma, Y.; Chen, Z.; Zhang, Y.; Li, J.; Liu, S. Exosomes derived from miR-126-3p-overexpressing synovial fibroblasts suppress chondrocyte inflammation and cartilage degradation in a rat model of osteoarthritis. Cell Death Discov. 2021, 7, 37. [Google Scholar] [CrossRef] [PubMed]
- Njock, M.S.; O’Grady, T.; Nivelles, O.; Lion, M.; Jacques, S.; Cambier, M.; Herkenne, S.; Muller, F.; Christian, A.; Remacle, C.; et al. Endothelial extracellular vesicles promote tumour growth by tumour-associated macrophage reprogramming. J. Extracell. Vesicles 2022, 11, e12228. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Chen, M.; Ouyang, L.; Wang, Q.; Guo, Y.; Huang, L.; Jiang, S. miR-142-5p and miR-130a-3p regulate pulmonary macrophage polarization and asthma airway remodeling. Immunol. Cell Biol. 2020, 98, 715–725. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Tong, L.; Liu, F.; Liu, A.; Zeng, S.; Xiong, Q.; Yang, Z.; He, X.; Sun, Y.; Xu, C. Tumor-infiltrating M2 macrophages driven by specific genomic alterations are associated with prognosis in bladder cancer. Oncol. Rep. 2019, 42, 581–594. [Google Scholar] [CrossRef]
- Mao, M.; Xu, Y.; Zhang, X.Y.; Yang, L.; An, X. Bin; Qu, Y.; Chai, Y.N.; Wang, Y.R.; Li, T.T.; Ai, J. MicroRNA-195 prevents hippocampal microglial/macrophage polarization towards the M1 phenotype induced by chronic brain hypoperfusion through regulating CX3CL1/CX3CR1 signaling. J. Neuroinflammation 2020, 17, 244. [Google Scholar] [CrossRef]
- Bras, J.P.; Silva, A.M.; Calin, G.A.; Barbosa, M.A.; Santos, S.G.; Almeida, M.I. miR-195 inhibits macrophages pro-inflammatory profile and impacts the crosstalk with smooth muscle cells. PLoS One 2017, 12, e0188530. [Google Scholar] [CrossRef]
- Youn, G.S.; Park, J.K.; Lee, C.Y.; Jang, J.H.; Yun, S.H.; Kwon, H.Y.; Choi, S.Y.; Park, J. MicroRNA-22 negatively regulates LPS-induced inflammatory responses by targeting HDAC6 in macrophages. BMB Rep. 2020, 53, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhong, Y.; Zhou, W.; Song, Y.; Li, W.; Jin, Q.; Gao, T.; Zhang, L.; Xie, M. Low-intensity pulsed ultrasound (LIPUS) enhances the anti-inflammatory effects of bone marrow mesenchymal stem cells (BMSCs)-derived extracellular vesicles. Cell. Mol. Biol. Lett. 2023, 28, 9. [Google Scholar] [CrossRef]
- Cheng, J.; Hao, J.; Jiang, X.; Ji, J.; Wu, T.; Chen, X.; Zhang, F. Ameliorative effects of miR-423-5p against polarization of microglial cells of the M1 phenotype by targeting a NLRP3 inflammasome signaling pathway. Int. Immunopharmacol. 2021, 99, 108006. [Google Scholar] [CrossRef]
- Li, R.; Li, D.; Wang, H.; Chen, K.; Wang, S.; Xu, J.; Ji, P. Exosomes from adipose-derived stem cells regulate M1/M2 macrophage phenotypic polarization to promote bone healing via miR-451a/MIF. Stem Cell Res. Ther. 2022, 13, 149. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhang, D.; Wang, H.; Ren, Q.; Li, B.; Wang, L.; Zheng, G. MiR-451a enhances the phagocytosis and affects both M1 and M2 polarization in macrophages. Cell. Immunol. 2021, 365, 104377. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, I.M.; Landis, W.J.; Risbud, M. V. Matrix vesicles: Are they anchored exosomes? Bone 2015, 79, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Leach, R.J.; Schwartz, Z.; Johnson-Pais, T.L.; Dean, D.D.; Luna, M.; Boyan, B.D. Osteosarcoma hybrids can preferentially target alkaline phosphatase activity to matrix vesicles: Evidence for independent membrane biogenesis. J. Bone Miner. Res. 1995, 10, 1614–1624. [Google Scholar] [CrossRef] [PubMed]
- Asmussen, N.; Lin, Z.; McClure, M.J.; Schwartz, Z.; Boyan, B.D. Regulation of extracellular matrix vesicles via rapid responses to steroid hormones during endochondral bone formation. Steroids 2019, 142, 43–47. [Google Scholar] [CrossRef]
- Bottini, M.; Mebarek, S.; Anderson, K.L.; Strzelecka-Kiliszek, A.; Bozycki, L.; Simão, A.M.S.; Bolean, M.; Ciancaglini, P.; Pikula, J.B.; Pikula, S.; et al. Matrix vesicles from chondrocytes and osteoblasts: Their biogenesis, properties, functions and biomimetic models. Biochim. Biophys. acta. Gen. Subj. 2018, 1862, 532–546. [Google Scholar] [CrossRef]
- Soe, Z.Y.; Park, E.J.; Shimaoka, M. Integrin regulation in immunological and cancerous cells and exosomes. Int. J. Mol. Sci. 2021, 22, 1–22. [Google Scholar] [CrossRef]
- Minamizaki, T.; Nakao, Y.; Irie, Y.; Ahmed, F.; Itoh, S.; Sarmin, N.; Yoshioka, H.; Nobukiyo, A.; Fujimoto, C.; Niida, S.; et al. The matrix vesicle cargo miR-125b accumulates in the bone matrix, inhibiting bone resorption in mice. Commun. Biol. 2020, 3, 1–2. [Google Scholar] [CrossRef]
- Rao, M.S.; Van Vleet, T.R.; Ciurlionis, R.; Buck, W.R.; Mittelstadt, S.W.; Blomme, E.A.G.; Liguori, M.J. Comparison of RNA-Seq and microarray gene expression platforms for the toxicogenomic evaluation of liver from short-term rat toxicity studies. Front. Genet. 2018, 9, 636. [Google Scholar] [CrossRef]
- Mizukami, Y.; Kawao, N.; Takafuji, Y.; Ohira, T.; Okada, K.; Jo, J.-I.; Tabata, Y.; Kaji, H. Matrix vesicles promote bone repair after a femoral bone defect in mice. PLoS One 2023, 18, e0284258. [Google Scholar] [CrossRef]
- Li, W.; Liu, Y.; Zhang, P.; Tang, Y.; Zhou, M.; Jiang, W.; Zhang, X.; Wu, G.; Zhou, Y. Tissue-engineered bone immobilized with human adipose stem cells-derived exosomes promotes bone regeneration. ACS Appl. Mater. Interfaces 2018, 10, 5240–5254. [Google Scholar] [CrossRef] [PubMed]
- Shapouri-Moghaddam, A.; Mohammadian, S.; Vazini, H.; Taghadosi, M.; Esmaeili, S.A.; Mardani, F.; Seifi, B.; Mohammadi, A.; Afshari, J.T.; Sahebkar, A. Macrophage plasticity, polarization, and function in health and disease. J. Cell. Physiol. 2018, 233, 6425–6440. [Google Scholar] [CrossRef] [PubMed]
- Costa Verdera, H.; Gitz-Francois, J.J.; Schiffelers, R.M.; Vader, P. Cellular uptake of extracellular vesicles Is mediated by clathrin-independent endocytosis and macropinocytosis. J. Control. Release 2017, 266, 100–108. [Google Scholar] [CrossRef]
- Svensson, K.J.; Christianson, H.C.; Wittrup, A.; Bourseau-Guilmain, E.; Lindqvist, E.; Svensson, L.M.; Mörgelin, M.; Belting, M. Exosome uptake depends on ERK1/2-Heat Shock Protein 27 signaling and lipid raft-mediated endocytosis negatively regulated by caveolin-1. J. Biol. Chem. 2013, 288, 17713–17724. [Google Scholar] [CrossRef]
- Tian, T.; Zhu, Y.L.; Zhou, Y.Y.; Liang, G.F.; Wang, Y.Y.; Hu, F.H.; Xiao, Z.D. Exosome uptake through clathrin-mediated endocytosis and macropinocytosis and mediating miR-21 delivery. J. Biol. Chem. 2014, 289, 22258–22267. [Google Scholar] [CrossRef]
- Hazan-Halevy, I.; Rosenblum, D.; Weinstein, S.; Bairey, O.; Raanani, P.; Peer, D. Cell-specific uptake of mantle cell lymphoma-derived exosomes by malignant and non-malignant B-lymphocytes. Cancer Lett. 2015, 364, 59–69. [Google Scholar] [CrossRef]
- Doherty, G.J.; McMahon, H.T. Mechanisms of endocytosis. Annu. Rev. Biochem. 2009, 78, 857–902. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.Y.; Schwartz, Z.; Hummert, T.W.; Schraub, D.M.; Simpson, J.; Lankford, J.; Dean, D.D.; Cochran, D.L.; Boyan, B.D. Effect of titanium surface roughness on proliferation, differentiation, and protein synthesis of human osteoblast-like cells (MG63). J. Biomed. Mater. Res. 1995, 29, 389–401. [Google Scholar] [CrossRef]
- Schwartz, Z.; Lohmann, C.H.; Oefinger, J.; Bonewald, L.F.; Dean, D.D.; Boyan, B.D. Implant surface characteristics modulate differentiation behavior of cells in the osteoblastic lineage. Adv. Dent. Res. 1999, 13, 38–48. [Google Scholar] [CrossRef]
- Heremans, H.; Billiau, A.; Cassiman, J.J.; Mulier, J.C.; De Somer, P. In vitro cultivation of human tumor tissues II. Morphological and virological characterization of three cell lines. Oncol. 1978, 35, 246–252. [Google Scholar] [CrossRef]
- Czekanska, E.M.; Stoddart, M.J.; Richards, R.G.; Hayes, J.S. In search of an osteoblast cell model for in vitro research. Eur. Cells Mater. 2012, 24, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Fitzpatrick, D.F.; Davenport, G.R.; Forte, L.; Landon, E.J. Characterization of plasma membrane proteins in mammalian kidney. I. Preparation of a membrane fraction and separation of the protein. J. Biol. Chem. 1969, 244, 3561–3569. [Google Scholar] [CrossRef] [PubMed]
- Boyan, B.D.; Schwartz, Z.; Swain, L.D.; Carnes Jr., D. L.; Zislis, T. Differential expression of phenotype by resting zone and growth region costochondral chondrocytes in vitro. Bone 1988, 9, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Boyan, B.D.; Wong, K.L.; Wang, L.; Yao, H.; Guldberg, R.E.; Drab, M.; Jo, H.; Schwartz, Z. Regulation of growth plate chondrocytes by 1,25-dihydroxyvitamin D 3 requires caveolae and caveolin-1. J. Bone Miner. Res. 2006, 21, 1637–1647. [Google Scholar] [CrossRef] [PubMed]
| miRNA | Osteogenic Potential | References |
|---|---|---|
| hsa-miR-1-3p | Pro-Osteogenic | [24] |
| hsa-miR-126-3p | Pro-Osteogenic | [25,26] |
| hsa-miR-142-5p | Pro-Osteogenic | [27,28,29] |
| hsa-miR-133a-3p | Anti-Osteogenic | [30,31,32] |
| hsa-miR-144-3p | Anti-Osteogenic | [33,34,35,36] |
| hsa-miR-148a-3p | Anti-Osteogenic | [26,37,38] |
| hsa-miR-195-3p | Anti-Osteogenic | [39] |
| hsa-miR-203a-3p | Anti-Osteogenic | [40] |
| hsa-miR-206 | Anti-Osteogenic | [41,42,43,44] |
| hsa-miR-22-5p | Anti-Osteogenic | [45,46] |
| hsa-miR-34c-3p | Anti-Osteogenic | [47] |
| hsa-miR-433-3p | Anti-Osteogenic | [48,49] |
| hsa-miR-122-5p | Unclear | [14,50] |
| hsa-miR-451a | Unclear | [51,52] |
| miRNA | M1/M2 Polarization | References |
|---|---|---|
| hsa-miR-144-3p | M1 | [53,54] |
| hsa-miR-148a-3p | M1 | [55,56] |
| hsa-miR-342-5p | M1 | [57] |
| hsa-miR-206-3p | M1 | [58] |
| hsa-miR-1-3p | M2b | [59] |
| hsa-miR-122-5p | M2 | [60] |
| hsa-miR-126-3p | M2 | [61,62] |
| hsa-miR-142-5p | M2 | [63,64,65] |
| hsa-miR-195-3p | M2 | [66,67] |
| hsa-miR-22-5p | M2 | [68] |
| hsa-miR-223-3p | M2 | [65] |
| hsa-miR-328-5p | M2 | [69] |
| hsa-miR-423-5p | M2 | [70] |
| hsa-miR-451a | M2 | [71,72] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





