Submitted:
27 May 2023
Posted:
30 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
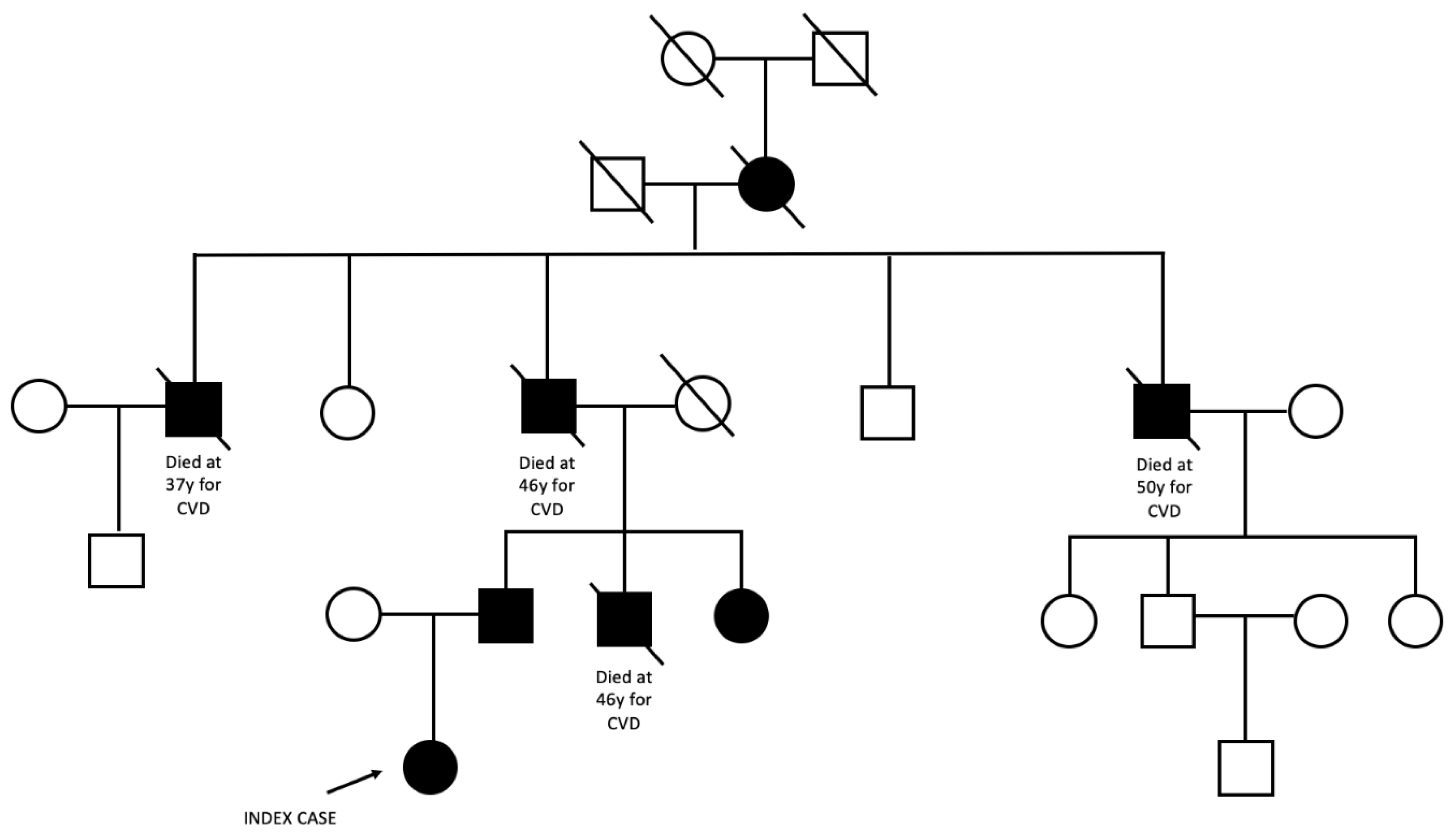
2.1. Case presentation and family history
2.2. NGS and Large Genomic Rearrangement (LGR) Detection
2.3. Analysis of Breakpoint Region
3. Results
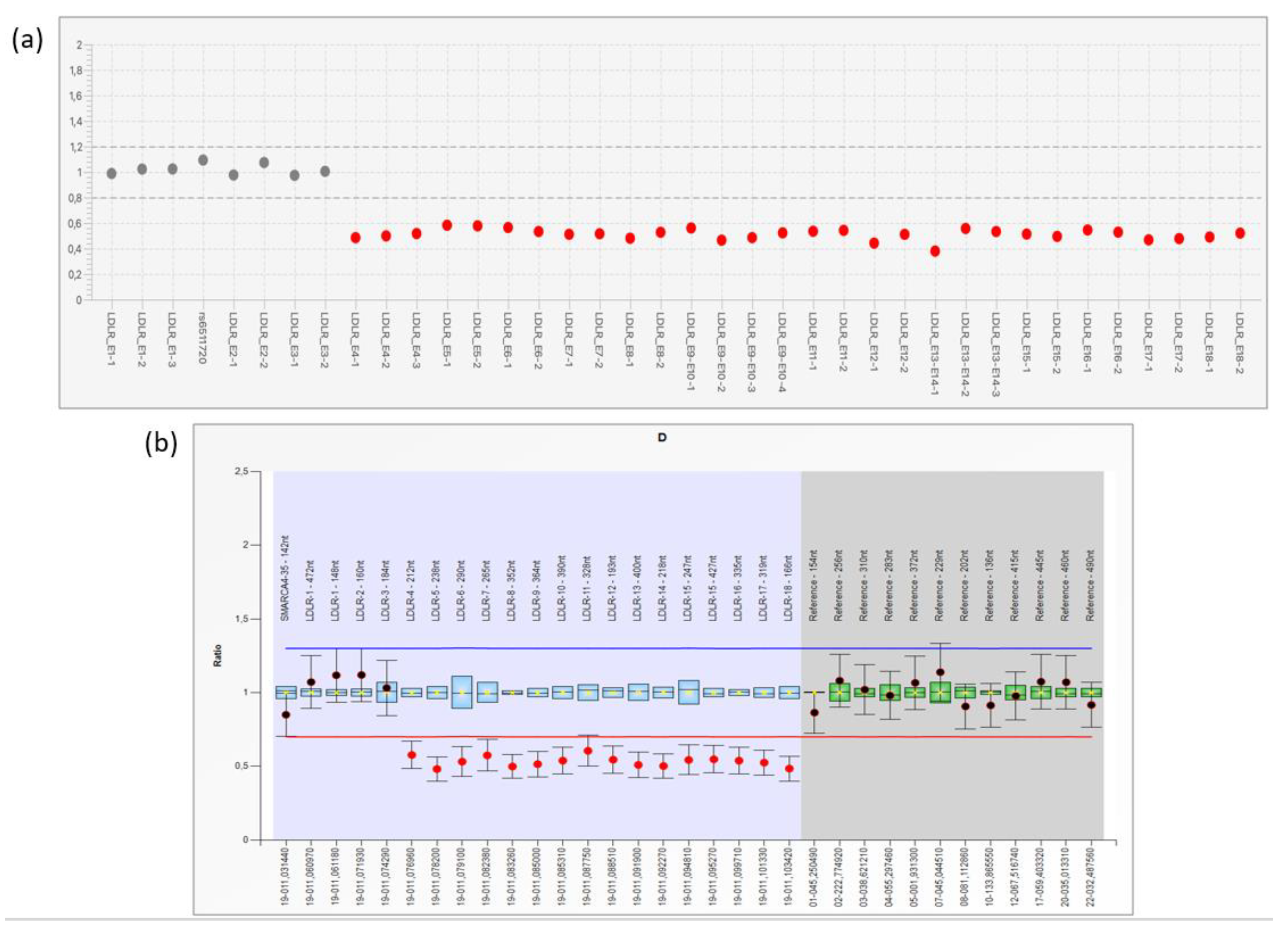
3.1. NGS Analysis and LRG Detection
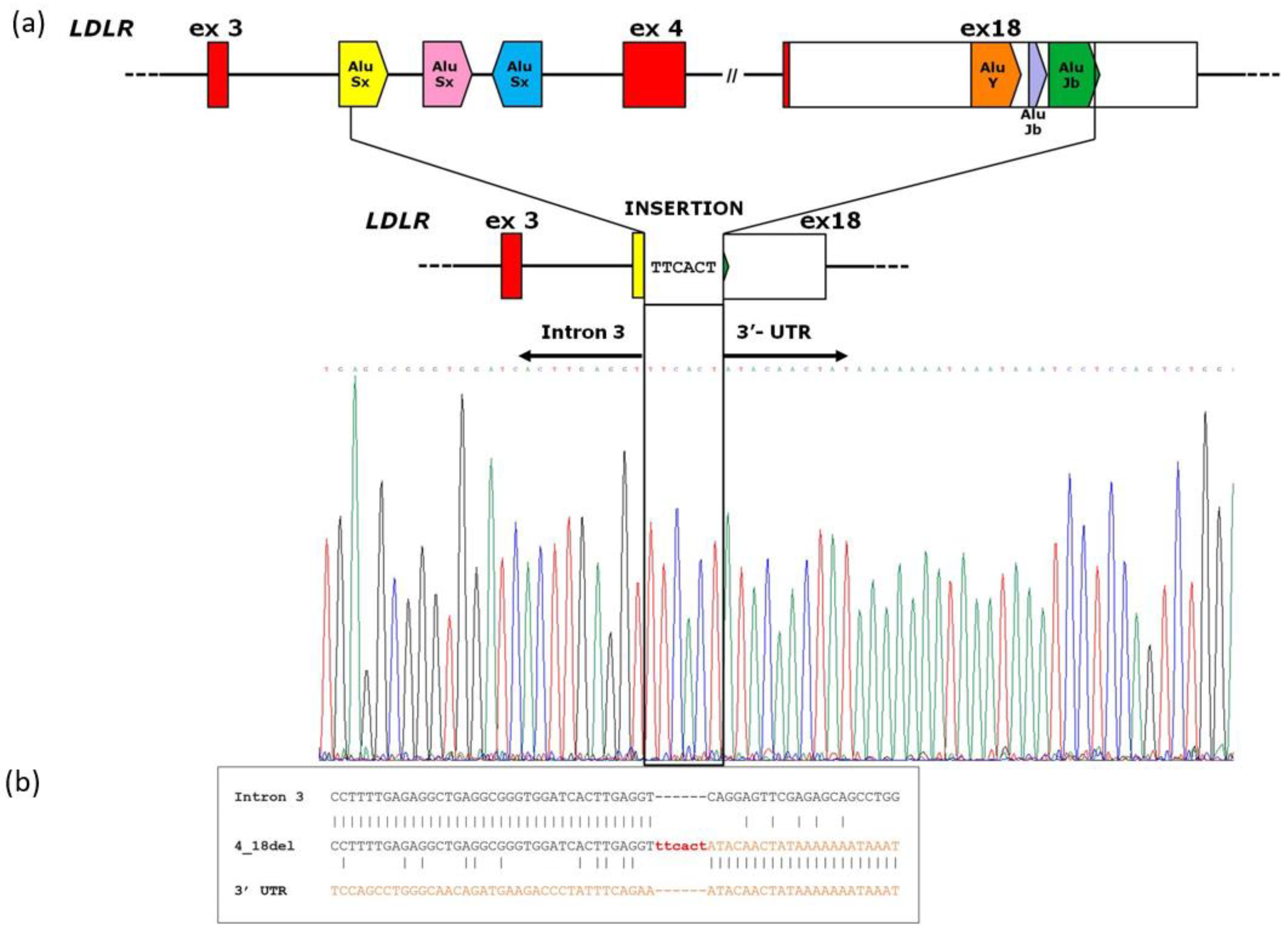
3.2. Analysis of Breakpoint Region
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Beheshti, S.O. , Madsen C.M., Varbo A., Nordestgaard B.G.. Worldwide Prevalence of Familial Hypercholesterolemia: Meta-Analyses of 11 Million Subjects. J. Am. Coll. Cardiol. 2020;75:2553–2566. [CrossRef]
- Vallejo-Vaz, A.J. , Stevens C.A.T., Lyons A.R.M., Dharmayat K.I., Freiberger T., Hovingh G.K., Mata P., Raal F.J., Santos R.D., Soran H., et al. Global Perspective of Familial Hypercholesterolaemia: A Cross-Sectional Study from the EAS Familial Hypercholesterolaemia Studies Collaboration (FHSC) Lancet. 2021;398:1713–1725.
- Mach, F. , Baigent C., Catapano A.L., Koskinas K.C., Casula M., Badimon L., Chapman M.J., de Backer G.G., Delgado V., Ference B.A., et al. 2019 ESC/EAS Guidelines for the Management of Dyslipidaemias: Lipid Modification to Reduce Cardiovascular Risk. Eur. Heart J. 2020;41:111–188. [CrossRef]
- Haralambos, K. , Ashfield-Watt P., McDowell I.F. Diagnostic scoring for familial hypercholesterolaemia in practice. Curr Opin Lipidol. 2016 Aug;27(4):367-74. [CrossRef]
- Sturm A.C., Knowles J.W., Gidding S.S. et al Convened by the Familial Hypercholesterolemia Foundation. Clinical Genetic Testing for Familial Hypercholesterolemia: JACC Scientific Expert Panel. J Am Coll Cardiol. 2018 Aug 7;72(6):662-680. [CrossRef]
- Vallejo-Vaz A.J., De Marco M., Stevens C.A.T. et al Overview of the current status of familial hypercholesterolaemia care in over 60 countries - The EAS Familial Hypercholesterolaemia Studies Collaboration (FHSC). Atherosclerosis. 2018 Oct;277:234-255. [CrossRef]
- Moffa, S. , Onori M.E., De Paolis E., Ricciardi Tenore C., Perrucci A., Pontecorvi A., Giaccari A., Urbani A., Minucci A. A novel low-density lipoprotein receptor variant in a Ukrainian patient: a case report and overview of the disease-causing low-density lipoprotein receptor variants associated to familial hypercholesterolemia. Mol Biol Rep. 2022 Feb;49(2):1623-1630. [CrossRef]
- Iacocca, M.A. , Hegele R.A. Role of DNA Copy Number Variation in Dyslipidemias. Curr. Opin. Lipidol. 2018;29:125–132. [CrossRef]
- Amsellem, S. , Briffaut D., Carrié A., Rabés J.P., Girardet J.P., Fredenrich A., Moulin P., Krempf M., Reznik Y., Vialettes B., et al. Intronic Mutations Outside of Alu-Repeat-Rich Domains of the LDL Receptor Gene Are a Cause of Familial Hypercholesterolemia. Hum. Genet. 2002;111:501–510. [CrossRef]
- Defesche, J.C. , Lansberg P.J., Umans-Eckenhausen M.A.W. Advanced method for the identification of patients with inherited hypercholesterolemia. Semin Vasc Med. 2004 Feb;4(1):59-65. [CrossRef]
- Moffa, S. , Mazzuccato G., De Bonis M., De Paolis E., Onori M.E., Pontecorvi A., Urbani A., Giaccari A., Capoluongo E., Minucci A. Identification of two novel LDLR variants by Next Generation Sequencing. Ann Ist Super Sanita. 2020 Jan-Mar;56(1):122-127. [CrossRef]
- https://www. repeatmasker.org. Accessed 11 May 2023.
- Lander, E.S. , Linton L.M., Birren B., Nusbaum C., Zody M.C., et al. International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature. 2001 Feb 15;409(6822):860-921. [CrossRef]
- Venter, J.C. , Adams M.D., Myers E.W., Li P.W., Mural R.J., Sutton G.G., et al. The sequence of the human genome. Science. 2001 Feb 16;291(5507):1304-51. [CrossRef]
- Hobbs, H.H. , Russell D.W., Brown M.S., Goldstein J.L. The LDL receptor locus in familial hypercholesterolemia: mutational analysis of a membrane protein. Annu Rev Genet. 1990;24:133-70. [CrossRef]
- Goldmann, R. , Tichý L., Freiberger T., Zapletalová P., Letocha O., Soska V., Fajkus J., Fajkusová L. Genomic characterization of large rearrangements of the LDLR gene in Czech patients with familial hypercholesterolemia. BMC Med Genet. 2010 Jul 27;11:115. [CrossRef]
- Dron, J.S. , Wang J., McIntyre A.D., Iacocca M.A., Robinson J.F., Ban M.R., Cao H., Hegele R.A. Six years' experience with LipidSeq: clinical and research learnings from a hybrid, targeted sequencing panel for dyslipidemias. BMC Med Genomics. 2020 Feb 10;13(1):23. [CrossRef]
- Iacocca, M. A, Wang J., Dron J.S., Robinson J.F., McIntyre A.D., Cao H., Hegele R.A. Use of next-generation sequencing to detect LDLR gene copy number variation in familial hypercholesterolemia. J Lipid Res. 2017 Nov;58(11):2202-2209.
- Concolino, P. , Rizza R., Mignone F., Costella A., Guarino D., Carboni I., Capoluongo E., Santonocito C., Urbani A., Minucci A. A comprehensive BRCA1/2 NGS pipeline for an immediate Copy Number Variation (CNV) detection in breast and ovarian cancer molecular diagnosis. Clin Chim Acta. 2018 May;480:173-179. [CrossRef]
- Lazarte, J. , Berberich A.J, Wang J., Hegele R.A. A cautionary tale: Is this APOB whole-gene duplication actually pathogenic? J Clin Lipidol. 2020 Sep-Oct;14(5):631-63. [CrossRef]
- Iacocca, M.A. , Wang J., Sarkar S., Dron J.S., Lagace T., McIntyre A.D., Lau P., Robinson J.F., Yang P., et al. Whole-Gene Duplication of PCSK9 as a Novel Genetic Mechanism for Severe Familial Hypercholesterolemia. Can J Cardiol. 2018 Oct;34(10):1316-1324. [CrossRef]
| Lipid Profile | Years | Normal range | ||
|---|---|---|---|---|
| 1994 | 2019 | 2022 | ||
| Total Cholesterol (mg/dl) | 374 | 357 | 321 | 130-200 mg/dl |
| HDL | 93 | 68 | 66 | female>45; male>40 |
| Triglycerides (mg/dl) | 281 | 124 | 95 | 20-170 mg/dl |
| LDL (mg/dl) | 225 | 264 | 236 | <130 mg/dl |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




