Submitted:
25 May 2023
Posted:
26 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Imaging Datasets
2.1.1. RIDER Dataset
2.1.2. HN1 dataset
2.2. Texture Features Analyzed
2.3. Experimental Set-up and Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Aerts, H.J.W.L.; Velazquez, E.R.; Leijenaar, R.T.H.; Parmar, C.; Grossmann, P.; Carvalho, S.; et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nature Communications 2014, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, B.; Brierley, J.; Byrd, D.; Bosman, F.; Kehoe, S.; Kossary, C.; et al. The TNM classification of malignant tumours—towards common understanding and reasonable expectations. Lancet Oncol 2017, 18, 849–51. [Google Scholar] [CrossRef]
- Chalmers, I.; Glasziou, P. Avoidable waste in the production and reporting of research evidence. The Lancet 2009, 374, 86–9. [Google Scholar] [CrossRef]
- Traverso, A.; Wee, L.; Dekker, A. ; GilliesR. Repeatability and Reproducibility of Radiomic Features: A Systematic Review. International Journal of Radiation Oncology, Biology, Physics 2018, 102, 1143–58. [Google Scholar] [CrossRef] [PubMed]
- Balagurunathan, Y.; Gu, Y.; Wang, H.; Kumar, V.; Grove, O.; Hawkins, S.; et al. Reproducibility and Prognosis of Quantitative Features Extracted from CT Images. Translational Oncology 2014, 7, 72–87. [Google Scholar] [CrossRef] [PubMed]
- Coroller, T.P.; Agrawal, V.; Narayan, V.; Hou, Y.; Grossmann, P.; Lee, S.W.; et al. Radiomic phenotype features predict pathological response in non-small cell lung cancer. Radiother Oncol 2016, 119, 480–6. [Google Scholar] [CrossRef] [PubMed]
- Fave, X.; Mackin, D.; Yang, J.; Zhang, J.; Fried, D.; Balter, P.; et al. Can radiomics features be reproducibly measured from CBCT images for patients with non-small cell lung cancer? Med Phys 2015, 42, 6784–97. [Google Scholar] [CrossRef] [PubMed]
- Fave, X.; Zhang, L.; Yang, J.; Mackin, D.; Balter, P.; Gomez, D.; et al. Delta-radiomics features for the prediction of patient outcomes in non–small cell lung cancer. Scientific Reports 2017, 7, 588. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.C.; Donovan, E.M.; Nisbet, A.; South, C.P.; Alobaidli, S.; Ezhil, V.; et al. The stability of imaging biomarkers in radiomics: a framework for evaluation. Phys Med Biol 2019, 64, 165012. [Google Scholar] [CrossRef] [PubMed]
- Vallières, M.; Kay-Rivest, E.; Perrin, L.J.; Liem, X.; Furstoss, C.; Aerts, H.J.W.L.; et al. Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer. Scientific Reports 2017, 7, 10117. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; James, L.P.; Moskowitz, C.S.; Guo, P.; Ginsberg, M.S.; Lefkowitz, R.A.; et al. Evaluating variability in tumor measurements from same-day repeat CT scans of patients with non-small cell lung cancer. Radiology 2009, 252, 263–72. [Google Scholar] [CrossRef] [PubMed]
- Welcome to pyradiomics documentation! — pyradiomics v3.0.1.post15+g2791e23 documentation n.d. https://pyradiomics.readthedocs.io/en/latest/ (accessed January 27, 2023). 27 January.
- Vallières, M.; Kay-Rivest, E.; Perrin, L.; Liem, X.; Furstoss, C.; Khaouam, N.; et al. Data from Head-Neck-PET-CT 2017. [CrossRef]
- Mottola, M.; Ursprung, S.; Rundo, L.; Sanchez, L.E.; Klatte, T.; Mendichovszky, I.; et al. Reproducibility of CT-based radiomic features against image resampling and perturbations for tumour and healthy kidney in renal cancer patients. Sci Rep 2021, 11, 11542. [Google Scholar] [CrossRef] [PubMed]
- Hatt, M.; Vallieres, M.; Visvikis, D.; Zwanenburg, A. IBSI: an international community radiomics standardization initiative. J Nucl Med 2018, 59, 287–287. [Google Scholar]
- Win, T.; Miles, K.A.; Janes, S.M.; Ganeshan, B.; Shastry, M.; Endozo, R.; et al. Tumor heterogeneity and permeability as measured on the CT component of PET/CT predict survival in patients with non-small cell lung cancer. Clin Cancer Res 2013, 19, 3591–9. [Google Scholar] [CrossRef] [PubMed]
- Coroller, T.P.; Agrawal, V.; Huynh, E.; Narayan, V.; Lee, S.W.; Mak, R.H.; et al. Radiomic-Based Pathological Response Prediction from Primary Tumors and Lymph Nodes in NSCLC. Journal of Thoracic Oncology 2017, 12, 467–76. [Google Scholar] [CrossRef] [PubMed]
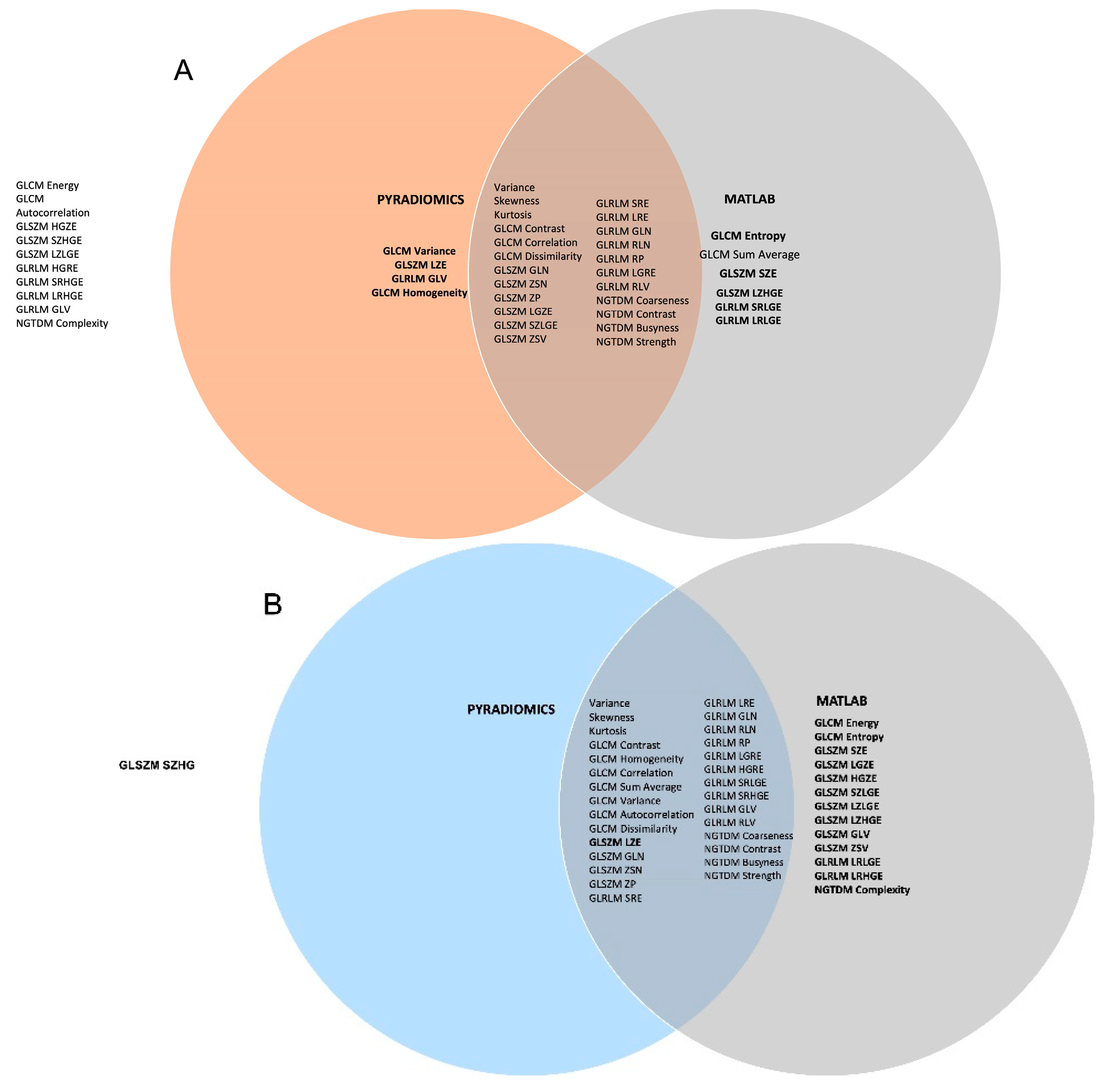
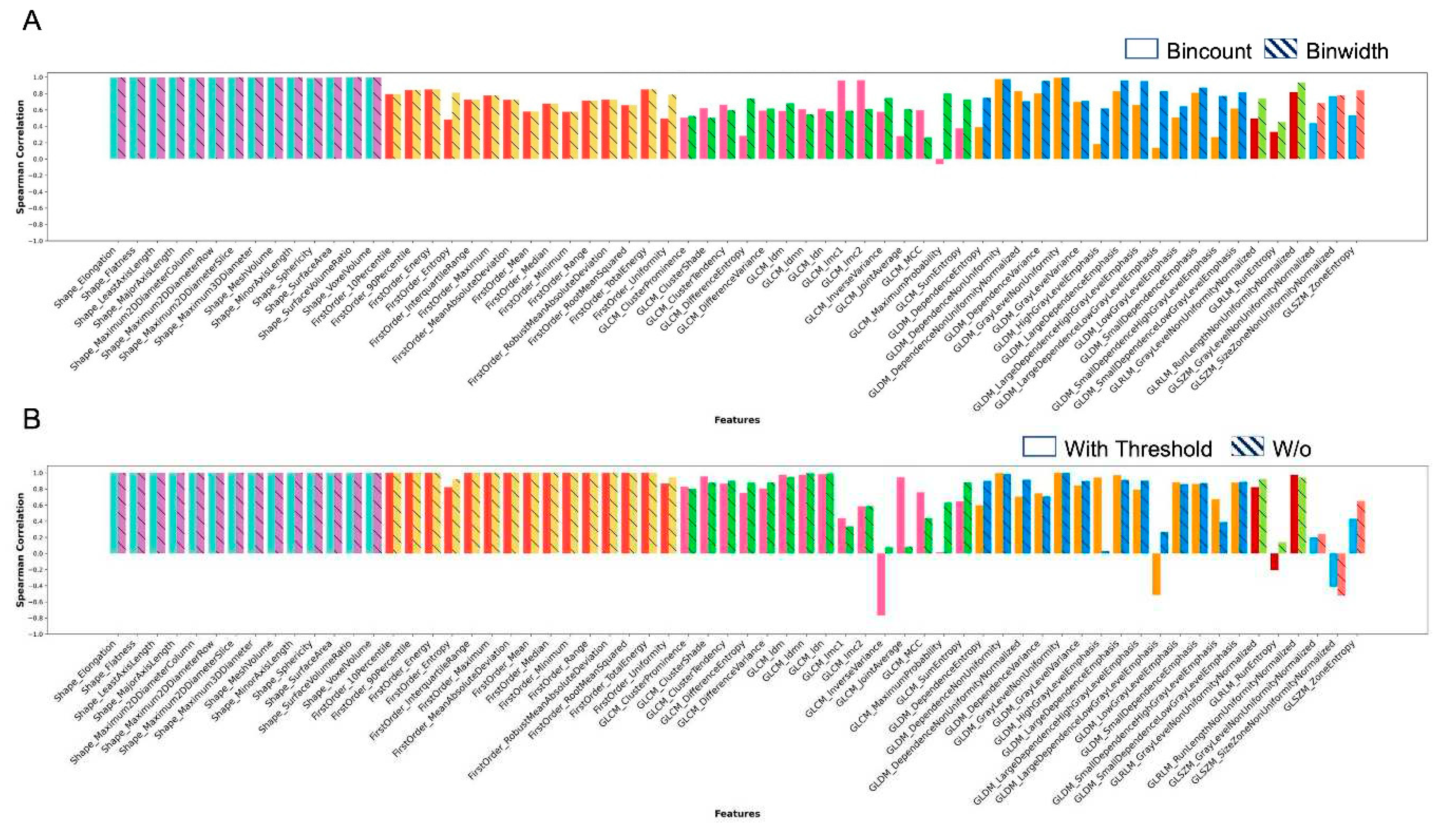
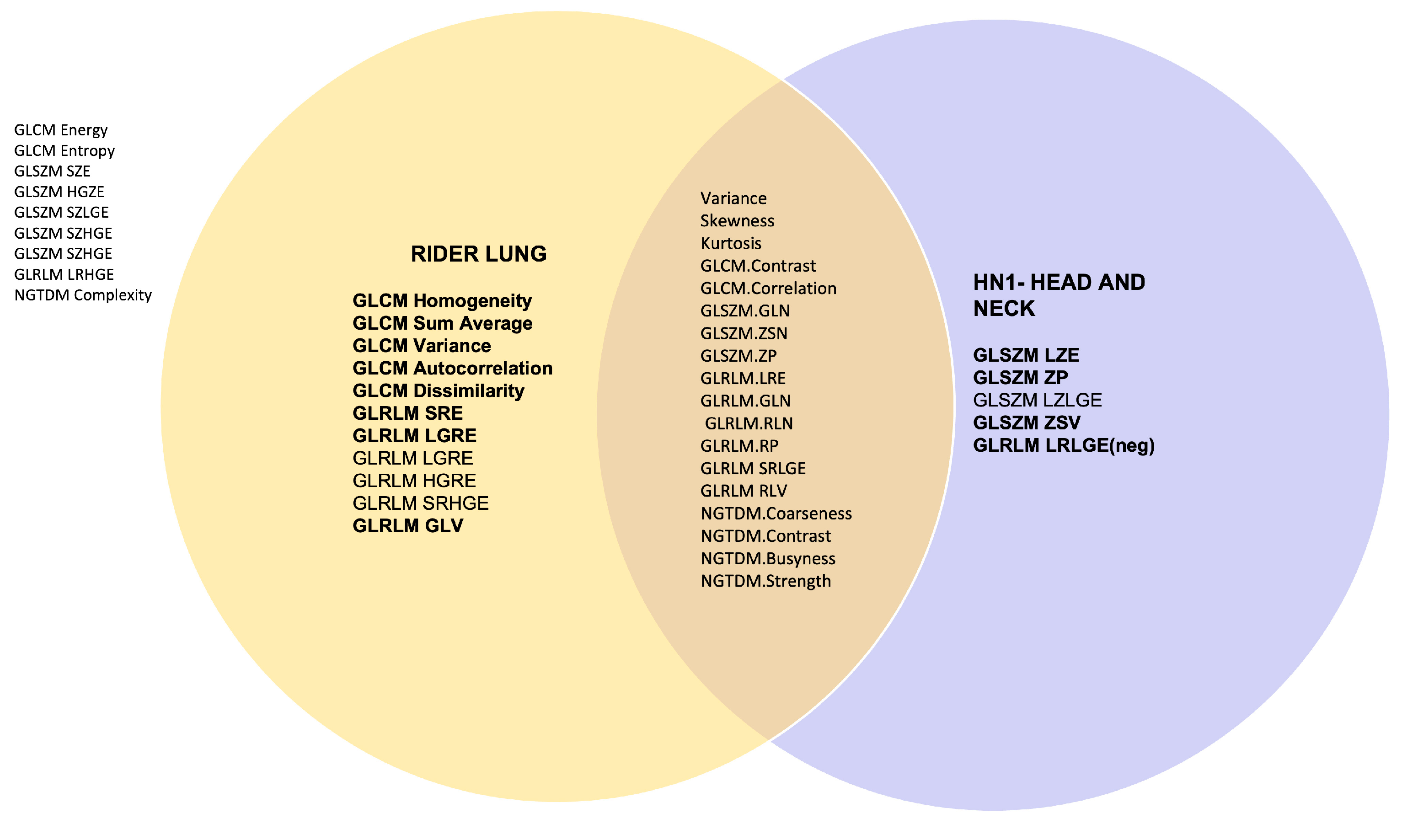
| Radiomics Feature | Matlab | Pyradiomics | Across scans 1 &2 and feature extraction implementations | All data | |||||||
| Scan 1 | Scan 2 | Scan 1 | Scan 2 | Scan 1 | Scan 2 | Scan 1 | Scan 2 | Scan 1 | Scan 2 | ||
| Threshold | W/o threshold | Threshold | W/o threshold | Threshold | W/o threshold | ||||||
| Variance | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| Skewness | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| Kurtosis | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLCM Energy | ü | ü | ü | ||||||||
| GLCM Contrast | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLCM Entropy | ü | ü | ü | ü | |||||||
| GLCM Homogeneity | ü | ü | ü | ü | ü | ü | ü | ü | |||
| GLCM Correlation | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLCM Sum Average | ü | ü | ü | ü | ü | ||||||
| GLCM Variance | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLCM Autocorrelation | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLCM Dissimilarity | ü | ü | ü | ü | ü | ü | |||||
| GLSZM SZE | ü | ü | ü | ü | |||||||
| GLSZM LZE | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | |
| GLSZM GLN | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLSZM ZSN | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLSZM ZP | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLSZM LGZE | ü | ü | ü | ü | ü | ü | |||||
| GLSZM HGZE | ü | ü | |||||||||
| GLSZM SZLGE | ü | ü | ü | ü | ü | ü | ü | ||||
| GLSZM SZHGE | ü | ||||||||||
| GLSZM LZLGE | ü | ü | |||||||||
| GLSZM LZHGE | ü | ü | ü | ü | ü | ||||||
| GLSZM GLV | ü | ü | |||||||||
| GLSZM ZSV | ü | ü | ü | ü | ü | ||||||
| GLRLM SRE | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLRLM LRE | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLRLM GLN | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLRLM RLN | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLRLM RP | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| GLRLM LGRE | ü | ü | ü | ü | ü | ü | ü | ü | ü | ||
| GLRLM HGRE | ü | ü | ü | ü | ü | ||||||
| GLRLM SRLGE | ü | ü | ü | ü | ü | ü | ü | ü | ü | ||
| GLRLM SRHGE | ü | ü | ü | ü | ü | ||||||
| GLRLM LRLGE | ü | ü | ü | ü | |||||||
| GLRLM LRHGE | ü | ü | |||||||||
| GLRLM GLV | ü | ü | ü | ü | ü | ü | ü | ü | |||
| GLRLM RLV | ü | ü | ü | ü | ü | ü | ü | ü | |||
| NGTDM Coarseness | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| NGTDM Contrast | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| NGTDM Busyness | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
| NGTDM Complexity | ü | ü | |||||||||
| NGTDM Strength | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü | ü |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




