1. Introduction
Atypical hemolytic uremic syndrome (aHUS) is a rare, severe thrombotic microangiopathy characterized by hemolytic anemia, thrombocytopenia, and acute kidney injury. Diagnosis is difficult due to nonspecific markers and rapid progression, leading to multi-organ failure and increased mortality. [
1]
aHUS is linked to complement system dysregulation, involving alternative pathway mutations or regulator abnormalities. Activation can be triggered by factors like infection, pregnancy, or specific drugs, causing complement deposition, immune cell recruitment, and clot formation. [
2] Patients may have multiple genetic variations, [
3] and while plasma exchange is not consistently effective, anti-complement therapy is often needed. Around 35-50% of aHUS patients lack detectable pathogenic genes [
4,
5] but respond to anticomplement therapy. In the absence of a definitive biomarker for disease activity assessment, clinicians rely on symptom alleviation and hemolysis markers; however, this approach frequently results in merely maintaining pace with disease progression rather than achieving control. Consequently, identifying a molecular marker is of utmost importance. Given the complement's role in immune homeostasis, [
6] we hypothesize immune cell dysregulation contributes to aHUS. Using single-cell RNA sequencing (scRNA-seq), we analyzed peripheral blood mononuclear cells (PBMCs) from aHUS patients, aHUS family members, and healthy controls, comparing immune cell dysregulation and investigating disease activity and treatment differences. Our study is the first to examine immune cell regulation in aHUS, providing insights into pathogenesis and clinical implications.
2. Results
2.1. The demography of studied cases
Table 1 presents the clinical features of 13 aHUS patients, diagnosed by hemolytic anemia, thrombocytopenia, and acute kidney injury, excluding infectious, autoimmune, or malignant etiologies. Patients' ages spanned from 30-81 years with balanced gender distribution (7:6). All exhibited extrarenal manifestations, with 9 (69.2%) stable and 4 (30.8%) unstable cases. Treatment involved plasma exchange alone (5 cases, 38.5%) or combined with anti-complement therapy (8 cases, 61.5%).
2.2. The immunological landscape of immune cells from aHUS, aHUS family and healthy
We conducted scRNA-seq on PBMCs from aHUS patients (N=13), family members (N=3), and healthy controls (N=4) to examine immune cell heterogeneity in aHUS (
Figure 1a). After preprocessing and quality control, we obtained single-cell transcriptomes of 112,191, 24,848, and 37,539 immune cells from aHUS patients, family members, and healthy controls, respectively. This enabled distinguishing among groups, disease activity, and treatments (plasma exchange only, combined anti-complement therapy).
Using SCTransform normalization and robust principal component analysis (rPCA) in Seurat, we identified 32 PBMC cell subpopulations in aHUS patients. SingleR annotation predicted B cells, T cells, monocytes, macrophages, dendritic cells, NK cells, megakaryocytes, granulocytes, and progenitors (
Figure 1b). The analysis of 5 B cell subpopulations displayed high diversity in aHUS patients and families, influenced by disease activity and treatment (
Figure 1c, d, and e). Investigation of 16 T and NK cell subpopulations also revealed significant diversity (
Figure 1f), impacted by disease activity and treatment (
Figure 1g, and h). In 7 monocyte subpopulations, aHUS patients had more intermediate monocytes than healthy controls (
Figure 1i). Additionally, intermediate monocytes increased in combined treatment compared to plasma exchange alone (
Figure 1j), with higher numbers in unstable aHUS group (
Figure 1k).
2.3. Cell populations in PBMCs
2.3.1. Comparing aHUS patients, aHUS family, and healthy controls
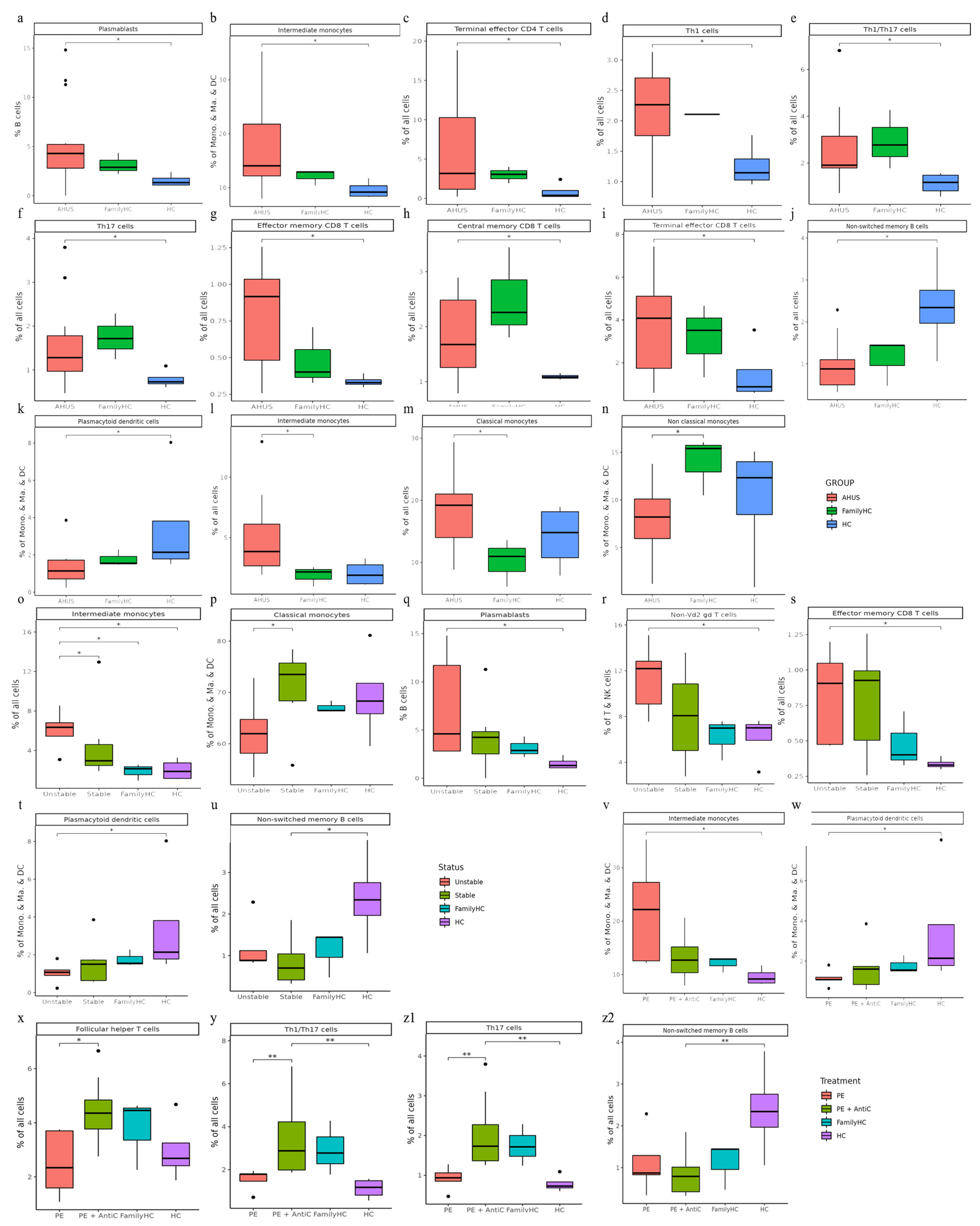
Utilizing Wilcoxon rank sum tests, we discovered significant increases in various immune cell populations, such as plasmablasts, intermediate monocytes, terminal effector CD4 T cells, Th1 cells, Th1/Th17 cells, Th17 cells, effector memory CD8 T cells, central memory CD8 T cells, and terminal effector CD8 T cells in aHUS patients compared to controls (
P <0.05,
Figure 2a, b, c, d, e, f, g, h, and i). Notably, aHUS families showed intermediate cell population levels between aHUS patients and healthy controls for plasmablasts, intermediate monocytes, terminal effector CD4 T cells, Th1 cells, effector memory CD8 T cells, and terminal effector CD8 T cells.
Non-switch memory B cells and plasmacytoid dendritic cells were more abundant in controls than in aHUS patients and families (
P <0.05,
Figure 2j and k). Intermediate and classical monocytes were higher in patients compared to families (
P <0.05,
Figure 2l, and m), while non-classical monocytes were lower in patients compared to families (
P <0.05,
Figure 2n).
2.3.2. Comparing stable and unstable aHUS patients, aHUS family, and healthy controls
Intermediate monocytes were significantly enriched in the unstable aHUS group, followed by the stable group, aHUS family, and controls (
P <0.05,
Figure 2o). Conversely, classical monocytes were enriched in the stable group compared to the unstable group (
P <0.05,
Figure 2p). Plasmablasts, non-Vd2 gd T cells, and effector memory CD8 T cells increased in the unstable group, followed by the stable group, aHUS family, and controls, with significant differences between the control and unstable groups (
P <0.05,
Figure 2q, r, and s).
Plasmacytoid dendritic cells were more abundant in the healthy group, followed by the aHUS family, stable group, and unstable group (
P <0.05,
Figure 2t), with the unstable group showing significantly lower levels compared to the control group. For non-switched B cells, the stable group had significantly lower levels compared to the control group (
P <0.05,
Figure 2u).
2.3.3. Comparing different treatment in aHUS patients, aHUS family, and healthy controls
In this subgroup analysis of aHUS treatment, intermediate monocyte enrichment showed an increasing trend from the plasma exchange group to the combined plasma exchange with anti-complement therapy group, aHUS family group, and healthy control group (
Figure 2v). The difference was only statistically significant (p<0.05) between plasma exchange and healthy control groups, with no significant difference between the two treatment groups. Plasmacytoid dendritic cell abundance exhibited an increasing trend from the healthy control group to the aHUS family group, followed by the combined therapy group and plasma exchange group, with the highest levels in controls and the lowest levels in the plasma exchange group (
Figure 2w). The combined therapy group exhibited significant enrichment of follicular helper T cells, Th1/Th17 cells, and Th17 cells compared to the plasma exchange group (
Figure 2x, y, and z1,
P <0.05). Non-switched memory B cells were significantly less abundant in the combined therapy group compared to the control group (
Figure 2z2,
P <0.05).
2.4. Cell subclusters in PBMCs
2.4.1. Comparing aHUS patients, aHUS family, and healthy controls
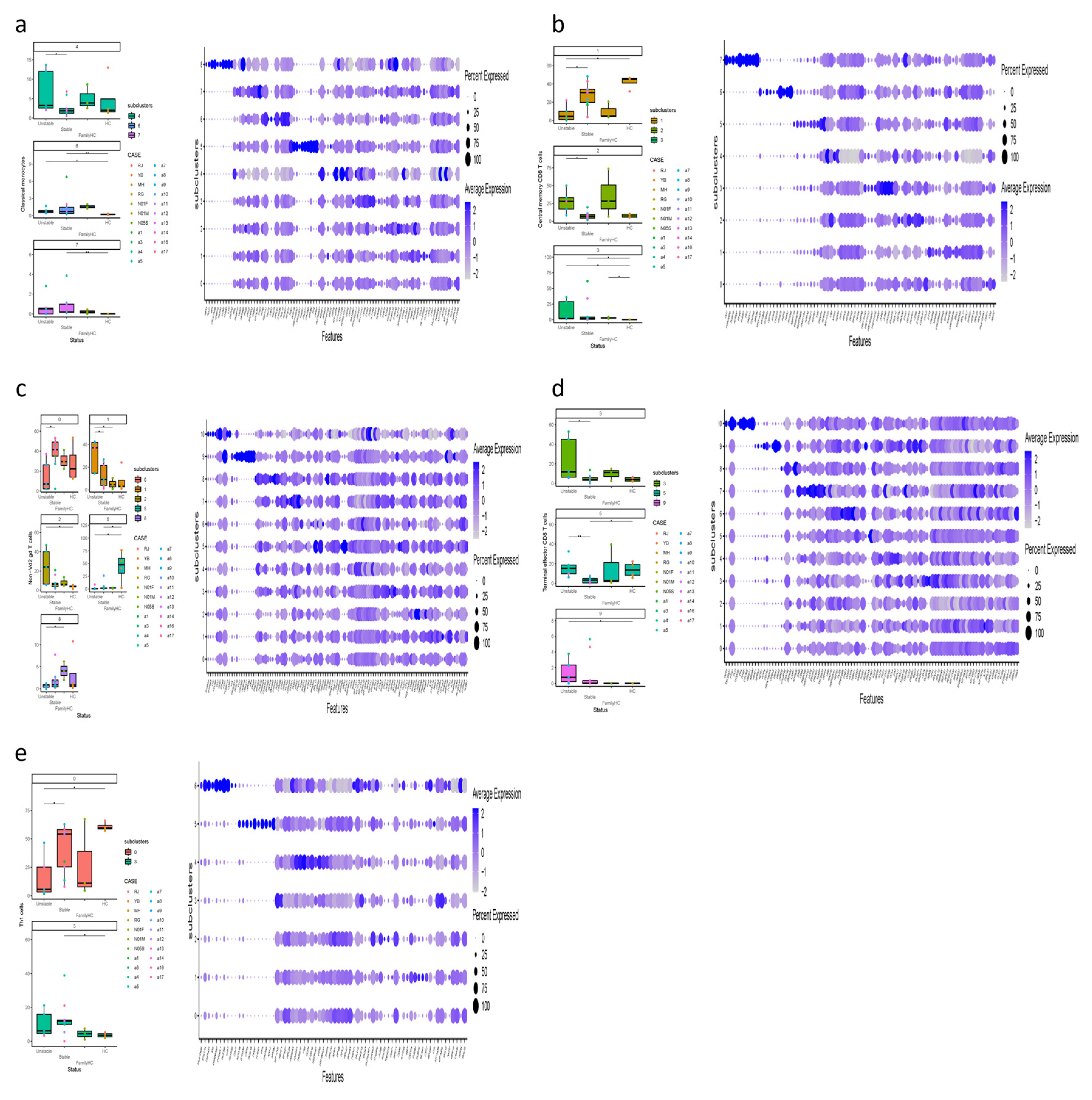
This study identified significant differences in immune cell subclusters among aHUS patients, aHUS family members, and healthy controls. In aHUS patients compared to healthy controls, we observed increased levels of classical monocytes (subclusters 6, 7) with higher RPS27 and IFI27 expression, central memory CD8 T cells (subcluster 3) with higher CXCR4 expression in patients and family members, non-Vd2 gd T cells (subcluster 4) with higher SYNE2 expression, Th1 T cells (subcluster 3) with higher MT-CYB expression, and Th17 cells (subcluster 4) with higher MT-ATP6 expression (
Figure 3a, b, c, d, and e).
Conversely, in healthy controls compared to aHUS patients, increased levels were found for central memory CD8 T cells (subcluster 1) with higher EIF3E expression, Th1 cells (subcluster 0) with higher RPS27 expression, non-classical monocytes (subcluster 5) with higher LYPD2 expression, terminal effector CD4 T cells (subcluster 3) with higher KLRD1 expression, and Th17 cells (subcluster 3) with higher ACTG1, CD52, and LGALS1 expression (
Figure 3b, d-g). Gene expression levels for each cell type were illustrated using dot plots, and findings are summarized in
Table 2.
2.4.2. Comparing stable and unstable aHUS patients, aHUS family, and healthy controls
Our study revealed significant differences in immune cell subclusters among unstable aHUS, stable aHUS, aHUS family members, and healthy controls. In unstable aHUS compared to stable aHUS, we observed increased classical monocytes (subcluster 4) with higher NEAT1, MT-ATP6, and MT-CYB expression, central memory CD8 T cells (subclusters 2) with elevated VIM expression, non-Vd2 gd T cells (subcluster 1) with increased ACTG1 expression, and terminal effector CD8 T cells (subclusters 3, 5) with elevated RPL13 and KLRB1 expression (
Figure 4a, b, c, and d).
In contrast, subclusters that increased in stable aHUS compared to unstable aHUS include central memory CD8 T cells (subcluster 1) with higher RPL23 expression, non-Vd2 gd T cells (subcluster 0) with elevated GZMH expression, and Th1 cells (subcluster 0) with increased RPS27 and RPS4X expression (
Figure 4b, c, and e). These findings are summarized in
Table 3.
2.5. Trajectory analysis for B cell, T cell, and monocyte
2.5.1. Comparing aHUS patients, aHUS family, and healthy control
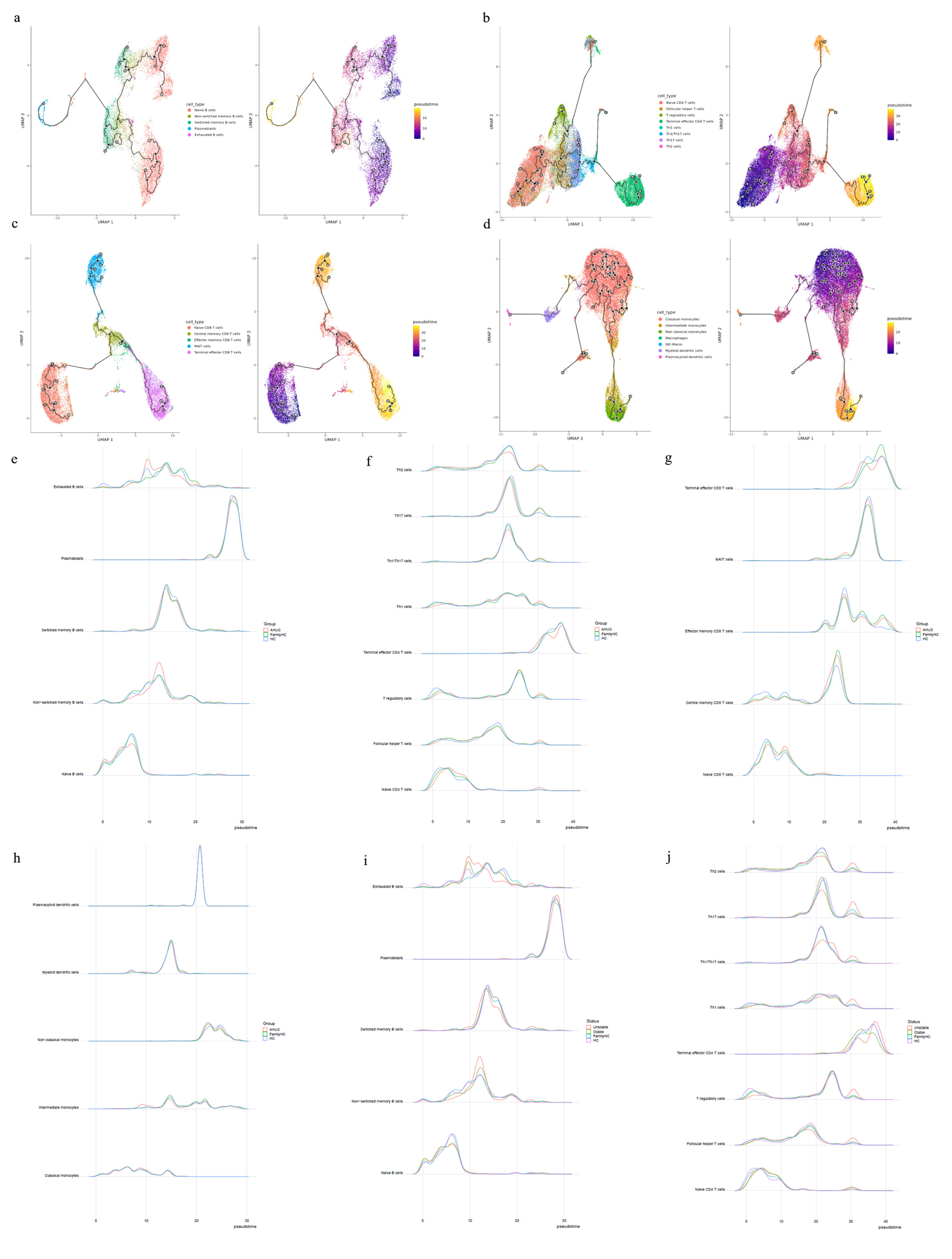
Cytopath analysis showed immune cell state dynamics in B cell, T cell, and monocyte trajectories (
Figure 5a, b, c, and d). Naïve B cells, exhausted B cells, and non-switched memory B cells in the aHUS group peaked at pseudotimes 0, 9, and 12, differing from healthy controls and aHUS nuclear families. From pseudotimes 5-10, naïve B cell abundance was highest in healthy controls, followed by aHUS families, and lowest in the aHUS group (
Figure 5e).
CD4 T cell trajectory analysis showed distinct abundance patterns for naïve CD4 T cells, T regulatory cells, Th1 cells, and Th1/Th17 cells in healthy controls, aHUS families, and the aHUS group across pseudotimes. During 20-25, Th2, Th17, and Th1/Th17 cells were most abundant in healthy controls, followed by aHUS families and the aHUS group (
Figure 5f). Terminal effector CD4 T cell abundance exhibited a similar pattern in 35-40. At pseudotime 30, Th2, Th1, T regulatory, and follicular T helper cells were most abundant in the aHUS group, then aHUS families and healthy controls.
In pseudotimes 0-5, naïve CD8 T cell abundance was highest in healthy controls, followed by aHUS families and patients (
Figure 5g). The trend reversed in 5-10. No significant differences in plasmacytoid dendritic cells, myeloid dendritic cells, non-classical monocytes, and classical monocytes were observed among aHUS patients, families, and healthy controls in 0-30. However, intermediate monocyte abundance in aHUS patients significantly increased during 7-10, not observed in families or healthy controls (
Figure 5h).
2.5.2. Comparing stable and unstable aHUS patients, aHUS family, and healthy controls
The unstable aHUS group showed a significant increase in exhausted B cells during pseudotime 7-13 compared to stable aHUS, aHUS families, and healthy controls, followed by a decline from 13-18. In this interval, switched memory B cell abundance was lowest in the unstable group compared to others. Non-switched memory B cells were more abundant in the unstable group, with the largest difference at pseudotime 10-15 (
Figure 5i).
Th2, Th17, and Th1/Th17 cells had the lowest abundance in the unstable aHUS group at pseudotime 18-22 but peaked at 28-32 for Th2, Th17, Th1/Th17, Th1, T regulatory, follicular helper T, and naïve CD4 T cells (
Figure 5j). The stable aHUS group showed a pattern more akin to the unstable group than to aHUS families and healthy controls.
2.6. Immune cell interactions in blood samples from aHUS, aHUS family and healthy control
To establish a comprehensive immune cell-complement pathway interaction network, we used the STRING database, integrating identified pathway interactions with aHUS-associated genes, including CFH, CD46, CFI, C3, CFB, THBD, CFHR1-5, DGKE, VTR, C2, C3AR1, C8B, C9, C4BPA, CFD, MASP1-2, MMACHC, PLG, WT1, VWF, CR1, CXCL12, C5, TLR4, CXCR4, HASP, KNK, INF2, EXOSC3, TSEN2, CD36, and VTN. [
2,
4,
5,
7,
8,
9,
10,
11,
12,
13,
14,
15,
16] Enriched pathways in aHUS patients include ALCAM-CD6, IL16-CD4, APP-CD40, CD86-CTLA4, CXC, and SELPLG (
Figure 6a, b, c, d, e, and f).
ALCAM-CD6 interactions had two patterns: one resembling healthy controls and another similar to aHUS families. Plasmacytoid dendritic cells exhibited increased outgoing signaling in pattern 1, while various T cells showed increased incoming signaling in pattern 2.
APP-CD40 interactions in aHUS patients were divided into two patterns, with pattern 2 similar to aHUS family and healthy controls. Classical monocytes, intermediate monocytes, megakaryocytes, and myeloid dendritic cells demonstrated increased outgoing signaling in pattern 1.
IL16-CD4 interactions in aHUS patients had two patterns. Outgoing signaling from myeloid dendritic cells and plasma blasts was highest in pattern 1, while intermediate monocytes, myeloid dendritic cells, non-classical monocytes, and classical monocytes exhibited increased incoming signaling.
CD86-CTLA4 pathways in aHUS patients displayed distinct patterns with heightened outgoing signaling in non-classical monocytes. In the SELPLG pathway, cases a7 and a10 showed significantly increased outgoing signaling in megakaryocytes. CXC interactions in aHUS patients had three patterns. Patterns 1 and 2 demonstrated significantly reduced incoming signaling from non-switched memory B cells, central memory CD8 T cells, Vd2 gd T cells, Th1/Th17 cells, and Th1 cells compared to pattern 3 and the healthy control. Notably, case a10 exhibited significantly increased outgoing signaling from non-classical monocytes and incoming signaling from MAIT cells, unobserved in other participants.
3. Discussion
In our study, we analyzed cell subpopulations and found that aHUS patients had higher levels of plasmablasts, intermediate monocytes, terminal effector CD4 T cells, Th1 cells, effector memory CD8 T cells, and terminal effector CD8 T cells compared to aHUS families and healthy controls. In contrast, non-switch memory B cells and plasmacytoid dendritic cells were most abundant in healthy controls, followed by aHUS families and patients. Unstable aHUS group showed significantly higher intermediate monocyte abundance than stable aHUS, aHUS families, and healthy controls. We suggest intermediate monocytes as potential aHUS disease activity markers, as they are associated with cardiovascular and autoimmune diseases. [
17,
18,
19] These monocytes also interact with endothelial cells, indicating a potential contribution to aHUS pathogenesis and correlation with endothelial cells.
In this study, we identified significant gene upregulation in various immune cell subclusters in aHUS patients compared to healthy controls. Classical monocyte subclusters 6 and 7 showed upregulated RPS27 and IFI27 genes, while central memory CD8 T cells subcluster 3, non-Vd2 gd T cells subcluster 4, Th1 cells subcluster 3, and Th17 cells subcluster 4 exhibited upregulated CXCR4, SYNE2, MT-CYB, and MT-ATP6 genes, respectively.
We also observed distinct gene expression patterns between unstable and stable aHUS. Unstable aHUS exhibited increased expression of NEAT1, MT-ATP6, MT-CYB, VIM, ACTG1, RPL13, and KLRB1 genes in various immune cell subclusters, while stable aHUS showed upregulated RPS27, RPS4X, RPL23, and GZMH genes. These genes may serve as potential clinical markers for aHUS disease activity. Elevated mitochondria-related gene expression suggests cell metabolism's role in aHUS clinical course, warranting further investigation. Notably, these gene expression patterns were not observed in other autoimmune diseases like systemic lupus erythematosus [
20,
21,
22,
23] or immunoglobulin G4-related disease [
24], highlighting the unique immune cell profile in aHUS.
Our pseudotime trajectory analysis revealed a unique point where immune cell differentiation in aHUS patients diverged from healthy individuals. This divergence was also apparent in some aHUS family members, falling between aHUS patients and healthy controls.
In our cell-cell interaction analysis, we aimed to identify signaling pathway differences between healthy individuals and aHUS patients, ensuring observed complement and immune cell interactions were not comparable between groups. Results showed unique signaling patterns in aHUS patients, specifically in ALCAM-CD6, IL16-CD4, APP-CD40, CD86-CTLA4, CXC, and SELPLG pathways, indicating distinctions from healthy individuals. Moreover, MIF or BAFF pathways, common in SLE and IgG4-related diseases, weren't increased. Despite significant statistical differences, aHUS's rarity and limited case enrollment warrant further studies with expanded enrollment for follow-up research.
4. Materials and Methods
4.1. Patient recruitment
In this single-center Taiwanese study, peripheral blood samples from 13 adult aHUS patients, 3 unaffected family members, and 4 healthy subjects were analyzed using scRNA-seq. aHUS patients were classified into stable and unstable groups, receiving plasma exchange alone or combined with anti-complement therapy. Stable disease had stable TMA-related organ involvement and normal hemolysis markers.
4.2. Single cell RNA-seq and data analysis
scRNA-seq was performed as previously described. [
25] Sequencer raw data was processed using CellRanger v3.1.0 or v6.0.2 with the GRCh38-3.0.0 reference file. Cells were selected using Seurat v4.0.4 [
26] in R software(
www.R-project.org) based on criteria including detected genes, unique molecular identifiers (UMIs), mitochondrial gene read counts, and doublet identification using Scrublet v0.2.2. [
27] Data type conversion was performed with Scanpy v1.8.1 [
28] and SeuratDisk v0.0.0.9015 tools(
https://mojaveazure.github.io/seurat-disk/).
4.3. Single cell RNA-seq data integration and clustering
Using the SCTransform workflow, scRNA-seq datasets were integrated, scaled, and normalized, considering regression variables like cell cycle stage, mitochondrial reads, gene number, and UMI count. The top 3,000 variable genes were selected for PCA using SelectIntegrationFeatures. A reference-based integration workflow with rPCA was applied, using four healthy control samples as reference. The top 50 PCs from PCA were used for UMAP, and the FindNeighbors function constructed a nearest neighbors graph for clustering analysis, all provided by the Seurat package.
4.4. Cell type annotations
Using SingleR v1.4.1, [
29] a reference-based cell type annotation tool, cell types in the dataset were classified by comparing gene expression profiles and assigning nomenclature and cell ontology terms. Reference gene expression data were obtained from five functions provided by celldex v1.0.0 R package. MonacoImmuneData labels were selected first, followed by Macrophages M1 and M2 labels from BlueprintEncodeData. Lastly, cell types such as Macrophages (CL: 0000235), Lung Macro (CL: 0000583), INF-Macro (CL: 0000863), and Megakaryocyte (CL: 0000556) were identified using cell ontology terms.
4.5. Clustering analysis
Further clustering analysis was performed using the same parameters as before, with a resolution range of 0.1 to 0.5. The Seurat function FindAllMarkers was applied to identify expression markers for each cluster in each cell type.
4.5. Pseudotime estimation
Monocle 3 [
30] was utilized to construct cell trajectory paths for B cells, CD4+ T cells, CD8+ T cells, and monocyte lineages. This involved dimensionality reduction using PCA and UMAP, followed by Leiden clustering. [
31] Trajectory paths were built by connecting nearest neighbors in the UMAP graph, with the root node determined based on early-stage cell types, such as naïve B cells, naïve T cells, and monocytes. Cell type abundance along pseudotime from the trajectory path was also analyzed.
4.6. Cell-cell communication analysis
The CellChat v1.1.3 [
32] R package was used to infer the probability of ligand-receptor signaling communication among all cell types in scRNA-seq datasets. Heatmaps of cell-cell interaction probability were generated for each sample using the ComplexHeatmap [
33] R package and visualized with Morpheus (
https://software.broadinstitute.org/morpheus) for each signaling pathway.
4.7. Statistical analysis
The abundance of cell type and further cluster are presented as percentages. Continuous data were compared using Wilcoxon rank sum tests with the R package ggpubr (v0.4.0) (
https://rpkgs.datanovia.com/ggpubr), and confidence intervals were calculated using the R package asht (v0.9.7). [
34]
5. Conclusions
In this study, we present compelling evidence demonstrating that immune cell dysregulation plays a significant role in the pathogenesis and clinical manifestations of aHUS. Intermediate monocytes were identified as potential disease activity markers in unstable aHUS, while immune cell subclusters showed significant gene expression differences between unstable and stable aHUS, offering potential clinical markers. Pseudotime trajectory analysis revealed a divergence point for immune cell differentiation in aHUS patients versus healthy individuals. Furthermore, cell-cell interaction analysis showed distinct signaling pathways in aHUS patients. These findings could contribute to the development of new diagnostic and disease activity markers for aHUS and enhance our understanding of its molecular mechanisms.
Author Contributions
Conceptualization, I.-R.C. and J.-G.C.; data curation, I.-R.C., S.-J.T., Y.-T.L., J.-C.Y., Y.-S.C. and J.-G.C.; formal analysis, I.-R.C., S.-J.T., Y.-T.L., J.-C.Y., Y.-S.C. and J.-G.C.; funding acquisition, I.-R.C.; investigation, I.-R.C. and J.-G.C.; methodology, I.-R.C., S.-J.T. and J.-G.C.; project administration, I.-R.C., S.-J.T. and J.-G.C.; resources, I.-R.C. and J.-G.C.; supervision, C.-C.H., G.-J.W., P.-C.L. and J.-G.C.; visualization, I.-R.C.; S.-J.T. and Y.-S.C.; writing—original draft, I.-R.C.; writing—review and editing, C.-C.H., P.-C.L. and J.-G.C. All authors have read and agreed to the published version of the manuscript.
Funding
Part of this research of I-Ru Chen was supported by the China Medical University Hospital, grant numbers DMR-110-036.
Institutional Review Board Statement
All the protocols for the trial was approved by the Human Research Ethics Committee of CMUH (CMUH111-REC2-048).
Informed Consent Statement
All the patients who participated in this study signed informed consent.
Data Availability Statement
Data will be made available upon request to the corresponding author.
Acknowledgments
The authors extend their appreciation to all the participants who generously provided blood samples for this study. The authors would like to thank the peer reviewers input in improving this manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| aHUS |
Atypical hemolytic uremic syndrome |
| PBMCs |
Peripheral blood mononuclear cells |
| scRNA-seq |
Single-cell RNA sequencing |
| PCA |
Principal Component Analysis |
| rPCA |
Robust Principal Component Analysis |
| UMAP |
Uniform manifold approximation and projection |
| NK |
Nature killer |
| Mono |
Monocytes |
| Ma |
Macrophages |
| DC |
Dendritic cells |
| STRING |
Search Tool for the Retrieval of Interacting Genes/Proteins |
References
- Han-Mou Tsai Atypical Hemolytic Uremic Syndrome: Beyond Hemolysis and Uremia. Am J Med. 2019, 132, 161−167. [CrossRef]
- Noris M,; Remuzzi G. Atypical hemolytic-uremic syndrome. N Engl J Med. 2009, 361, 1676-87.
- Bu F.; Maga T.; Meyer NC.; Wang K.; Thomas CP.; Nester CM.; Smith RJ. Comprehensive genetic analysis of complement and coagulation genes in atypical hemolytic uremic syndrome. J Am Soc Nephrol. 2014, 25, 55-64. [CrossRef]
- Afshar-Kharghan V. Atypical hemolytic uremic syndrome. Hematology Am Soc Hematol Educ Program. 2016, 2016, 217-225.
- Tseng MH.; Lin SH.; Tsai JD.; Wu MS.; Tsai IJ.; Chen YC.; Chang MC.; Chou WC.; Chiou YH.; Huang CC. Atypical hemolytic uremic syndrome: Consensus of diagnosis and treatment in Taiwan. J Formos Med Assoc. 2023, 122, 366-375. [CrossRef]
- Merle NS.; Noe R.; Halbwachs-Mecarelli L.; Fremeaux-Bacchi V.; Roumenina LT. Complement System Part II: Role in Immunity. Front Immunol. 2015, 6, 257. [CrossRef]
- Fremeaux-Bacchi V.; Fakhouri F.; Garnier A.; Bienaimé F.; Dragon-Durey MA.; Ngo S.; Moulin B.; Servais A.; Provot F.; Rostaing L, Burtey S.; Niaudet P.; Deschênes G.; Lebranchu Y.; Zuber J.; Loirat C. Genetics and outcome of atypical hemolytic uremic syndrome: A nationwide French series comparing children and adults. Clin J Am Soc Nephrol. 2013, 8, 554-62. [CrossRef]
- Liu D.; Ding Q.; Dai DF.; Padhy B.; Nayak MK.; Li C.; Purvis M.; Jin H.; Shu C.; Chauhan AK.; Huang CL.; Attanasio M. Loss of diacylglycerol kinase ε causes thrombotic microangiopathy by impairing endothelial VEGFA signaling. JCI Insight. 2021, 6, e146959. [CrossRef]
- Challis RC.; Ring T.; Xu Y.; Wong EK.; Flossmann O.; Roberts IS.; Ahmed S.; Wetherall M.; Salkus G.; Brocklebank V.; Fester J.; Strain L.; Wilson V.; Wood KM.; Marchbank KJ.; Santibanez-Koref M.; Goodship TH.; Kavanagh D. Thrombotic Microangiopathy in Inverted Formin 2-Mediated Renal Disease. J Am Soc Nephrol. 2017, 28, 1084-1091. [CrossRef]
- Van Quekelberghe C.; Latta K.; Kunzmann S.; Grohmann M.; Hansen M. Atypical hemolytic uremic syndrome induced by SARS-CoV2 infection in infants with EXOSC3 mutation. Pediatr Nephrol. 2022, 37, 2781-2784. [CrossRef]
- Canpolat N.; Liu D.; Atayar E.; Saygili S.; Kara NS.; Westfall TA.; Ding Q.; Brown BJ.; Braun TA.; Slusarski D.; Karli Oguz K.; Ozluk Y.; Tuysuz B.; Tastemel Ozturk T.; Sever L.; Sezerman OU.; Topaloglu R.; Caliskan S.; Attanasio M.; Ozaltin F. A splice site mutation in the TSEN2 causes a new syndrome with craniofacial and central nervous system malformations, and atypical hemolytic uremic syndrome. Clin Genet. 2022, 101, 346-358. [CrossRef]
- Sartain SE.; Turner NA.; Moake JL. Brain microvascular endothelial cells exhibit lower activation of the alternative complement pathway than glomerular microvascular endothelial cells. J Biol Chem. 2018, 293, 7195-7208. [CrossRef]
- Thergaonkar RW.; Narang A.; Gurjar BS.; Tiwari P.; Puraswani M.; Saini H.; Sinha A.; Varma B.; Mukerji M.; Hari P.; Bagga A. Targeted exome sequencing in anti-factor H antibody negative HUS reveals multiple variations. Clin Exp Nephrol. 2018, 22, 653-660. [CrossRef]
- Schramm EC.; Roumenina LT.; Rybkine T.; Chauvet S.; Vieira-Martins P.; Hue C.; Maga T.; Valoti E.; Wilson V.; Jokiranta S.; Smith RJ.; Noris M.; Goodship T.; Atkinson JP.; Fremeaux-Bacchi V. Mapping interactions between complement C3 and regulators using mutations in atypical hemolytic uremic syndrome. Blood. 2015, 125, 2359-69. [CrossRef]
- Urban A.; Kowalska D.; Stasiłojć G.; Kuźniewska A.; Skrobińska A.; Arjona E.; Alonso EC.; Fenollosa Segarra MÁ.; Jongerius I.; Spaapen R.; Satchell S.; Thiel M, Ołdziej S.; Rodriguez de Córdoba S.; Okrój M. Gain-of-Function Mutations R249C and S250C in Complement C2 Protein Increase C3 Deposition in the Presence of C-Reactive Protein. Front Immunol. 2021, 12, 724361. [CrossRef]
- Cheng C.; Chen L.; Wen S.; Lin Z.; Jiang X. Case Report: Denys-Drash Syndrome With WT1 Causative Variant Presenting as Atypical Hemolytic Uremic Syndrome. Front Pediatr. 2020, 8, 605889. [CrossRef]
- Zawada AM.; Rogacev KS.; Schirmer SH.; Sester M.; Böhm M.; Fliser D.; Heine GH. Monocyte heterogeneity in human cardiovascular disease. Immunobiology. 2012, 217, 1273-84. [CrossRef]
- Wong KL.; Yeap WH.; Tai JJ.; Ong SM.; Dang TM.; Wong SC. The three human monocyte subsets: Implications for health and disease. Immunol Res. 2012, 53, 41-57. [CrossRef]
- Ziegler-Heitbrock L.; Ancuta P.; Crowe S.; Dalod M.; Grau V.; Hart DN.; Leenen PJ.; Liu YJ.; MacPherson G.; Randolph GJ.; Scherberich J.; Schmitz J.; Shortman K.; Sozzani S.; Strobl H.; Zembala M.; Austyn JM.; Lutz MB. Nomenclature of monocytes and dendritic cells in blood. Blood. 2010, 116, 74-80. [CrossRef]
- Perez RK.; Gordon MG.; Subramaniam M.; Kim MC.; Hartoularos GC.; Targ S.; Sun Y.; Ogorodnikov A.; Bueno R.; Lu A.; Thompson M.; Rappoport N.; Dahl A.; Lanata CM.; Matloubian M.; Maliskova L.; Kwek SS.; Li T.; Slyper M.; Waldman J.; Dionne D.; Rozenblatt-Rosen O.; Fong L.; Dall'Era M.; Balliu B.; Regev A.; Yazdany J.; Criswell LA.; Zaitlen N.; Ye CJ. Single-cell RNA-seq reveals cell type-specific molecular and genetic associations to lupus. Science. 2022 , 376, eabf1970. [CrossRef]
- Nehar-Belaid D.; Hong S.; Marches R.; Chen G.; Bolisetty M.; Baisch J.; Walters L.; Punaro M.; Rossi RJ.; Chung CH.; Huynh RP.; Singh P.; Flynn WF.; Tabanor-Gayle JA.; Kuchipudi N.; Mejias A.; Collet MA.; Lucido AL.; Palucka K.; Robson P.; Lakshminarayanan S.; Ramilo O.; Wright T.; Pascual V.; Banchereau JF. Mapping systemic lupus erythematosus heterogeneity at the single-cell level. Nat Immunol. 2020, 21, 1094-1106. [CrossRef]
- Li Y.; Ma C.; Liao S.; Qi S.; Meng S.; Cai W.; Dai W.; Cao R.; Dong X.; Krämer BK.; Yun C.; Hocher B.; Hong X.; Liu D.; Tang D.; He J.; Yin L.; Dai Y. Combined proteomics and single cell RNA-sequencing analysis to identify biomarkers of disease diagnosis and disease exacerbation for systemic lupus erythematosus. Front Immunol. 2022, 13, 969509. [CrossRef]
- Zhang Y.; Lee TY. Revealing the Immune Heterogeneity between Systemic Lupus Erythematosus and Rheumatoid Arthritis Based on Multi-Omics Data Analysis. Int J Mol Sci. 2022, 23, 5166. [CrossRef]
- Wu X.; Peng Y.; Li J.; Zhang P.; Liu Z.; Lu H.; Peng L.; Zhou J.; Fei Y.; Zeng X.; Zhao Y.; Zhang W. Single-Cell Sequencing of Immune Cell Heterogeneity in IgG4-Related Disease. Front Immunol. 2022, 13, 904288. [CrossRef]
- Chang JG.; Tu SJ.; Huang CM.; Chen YC.; Chiang HS.; Lee YT.; Yen JC.; Lin CL.; Chung CC.; Liu TC.; Chang YS. Single-cell RNA sequencing of immune cells in patients with acute gout. Sci Rep. 2022, 12, 22130. [CrossRef]
- Hao Y.; Hao S.; Andersen-Nissen E.; Mauck WM 3rd.; Zheng S.; Butler A.; Lee MJ.; Wilk AJ.; Darby C.; Zager M.; Hoffman P.; Stoeckius M.; Papalexi E.; Mimitou EP.; Jain J.; Srivastava A.; Stuart T.; Fleming LM.; Yeung B.; Rogers AJ.; McElrath JM.; Blish CA.; Gottardo R.; Smibert P.; Satija R. Integrated analysis of multimodal single-cell data. Cell. 2021, 184, 3573-3587. [CrossRef]
- Wolock SL.; Lopez R.; Klein AM. Scrublet: Computational Identification of Cell Doublets in Single-Cell Transcriptomic Data. Cell Syst. 2019, 8, 281-291. [CrossRef]
- Wolf FA.; Angerer P.; Theis FJ. SCANPY: Large-scale single-cell gene expression data analysis. Genome Biol. 2018, 19, 15. [CrossRef]
- Aran D.; Looney A.; Liu L.; Wu E.; Fong V.; Hsu A.; Chak S.; Naikawadi RP.; Wolters PJ.; Abate AR.; Butte AJ.; Bhattacharya M. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol. 2019 , 20, 163-172. [CrossRef]
- Cao J.; Spielmann M.; Qiu X.; Huang X.; Ibrahim DM.; Hill AJ.; Zhang F.; Mundlos S.; Christiansen L.; Steemers FJ.; Trapnell C.; Shendure J. The single-cell transcriptional landscape of mammalian organogenesis. Nature. 2019, 566, 496-502. [CrossRef]
- Traag VA.; Waltman L.; van Eck NJ. From Louvain to Leiden: Guaranteeing well-connected communities. Sci Rep. 2019, 9, 5233. [CrossRef]
- Jin S.; Guerrero-Juarez CF.; Zhang L.; Chang I.; Ramos R.; Kuan CH.; Myung P.; Plikus MV.; Nie Q. Inference and analysis of cell-cell communication using CellChat. Nat Commun. 2021, 12, 1088. [CrossRef]
- Gu Z.; Eils R.; Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016, 32, 2847-9. [CrossRef]
- Fay MP.; Malinovsky Y. Confidence intervals of the Mann-Whitney parameter that are compatible with the Wilcoxon-Mann-Whitney test. Stat Med. 2018, 37, 3991-4006. [CrossRef]
Figure 1.
The immune cell phenotype of aHUS patients, aHUS family, and healthy controls were investigated using scRNA-seq analysis of PBMCs. The study design is presented in (a). UMAP coordinates showing the distribution of immune cells in PBMCs are presented in (b). The relative abundance of subpopulations of B cells is shown in (c), while the impact of aHUS disease activity on B cell subpopulations is presented in (d), and the influence of aHUS treatment on B cell subpopulations is shown in (e). Similarly, the diversity of T and nature killer (NK) cells is shown in (f), and the impact of aHUS disease activity on T and NK cells is presented in (g), while the influence of aHUS treatment on T and NK cells is shown in (h). The analysis of monocyte series, including monocytes (Mono.), macrophages (Ma.), and dendritic cells (DC), is presented in (i), while the impact of aHUS disease activity on these cells is presented in (j), and the influence of aHUS treatment on these cells is shown in (k).
Figure 1.
The immune cell phenotype of aHUS patients, aHUS family, and healthy controls were investigated using scRNA-seq analysis of PBMCs. The study design is presented in (a). UMAP coordinates showing the distribution of immune cells in PBMCs are presented in (b). The relative abundance of subpopulations of B cells is shown in (c), while the impact of aHUS disease activity on B cell subpopulations is presented in (d), and the influence of aHUS treatment on B cell subpopulations is shown in (e). Similarly, the diversity of T and nature killer (NK) cells is shown in (f), and the impact of aHUS disease activity on T and NK cells is presented in (g), while the influence of aHUS treatment on T and NK cells is shown in (h). The analysis of monocyte series, including monocytes (Mono.), macrophages (Ma.), and dendritic cells (DC), is presented in (i), while the impact of aHUS disease activity on these cells is presented in (j), and the influence of aHUS treatment on these cells is shown in (k).
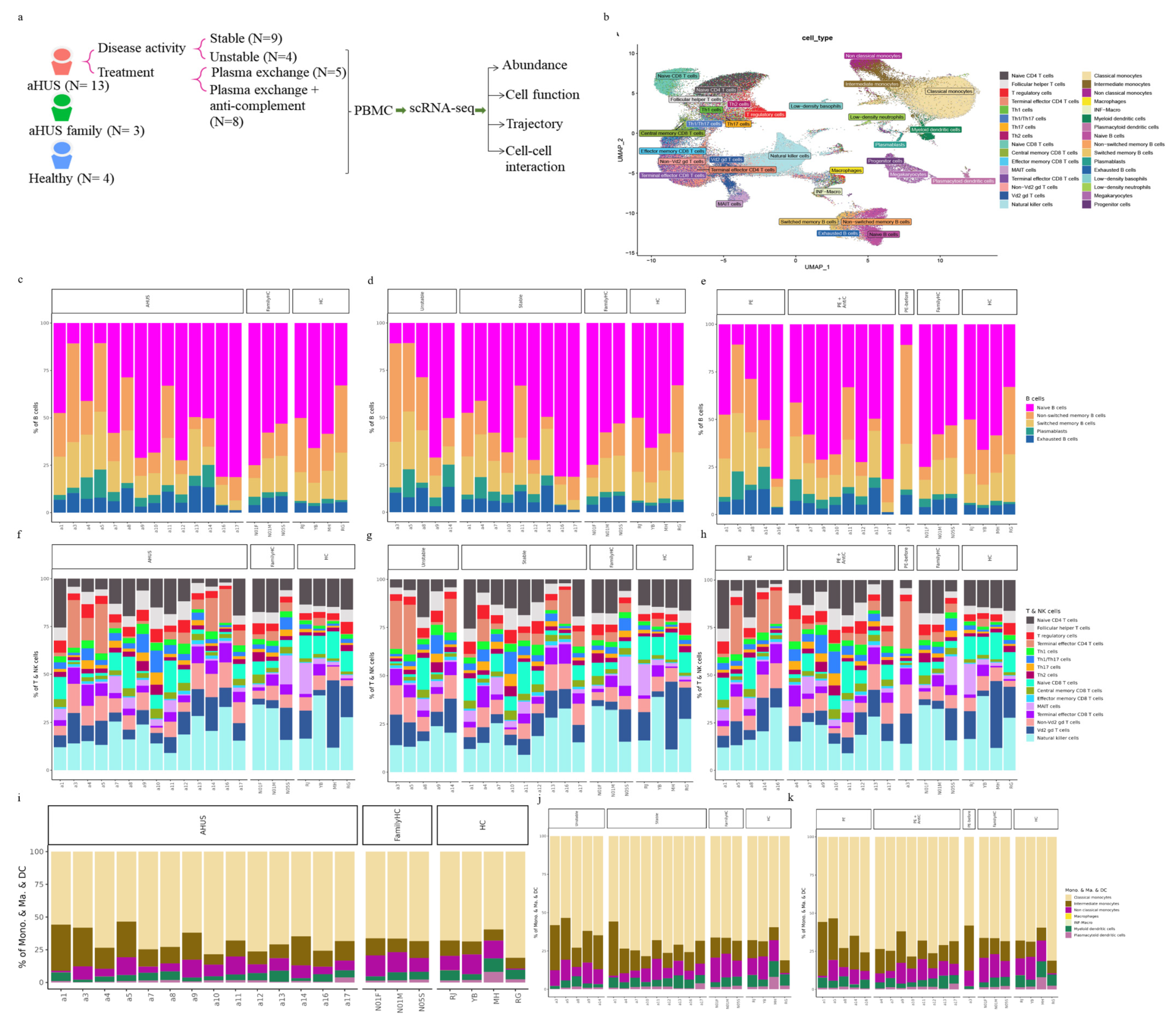
Figure 2.
Significant abundance of cell subpopulations between PBMCs from aHUS, aHUS family and healthy subjects. (a-n) Significant abundance of cell subpopulations in PBMCs from aHUS in unstable and stable disease activity, aHUS family and healthy subjects. (o-u) Significant abundance of cell subpopulations compared between PBMCs from aHUS treatment with plasma exchange alone, treatment combine with plasma exchange and anti-complement therapy, aHUS family and healthy subjects. (v-z2) *P < 0.05; **P < 0.01.
Figure 2.
Significant abundance of cell subpopulations between PBMCs from aHUS, aHUS family and healthy subjects. (a-n) Significant abundance of cell subpopulations in PBMCs from aHUS in unstable and stable disease activity, aHUS family and healthy subjects. (o-u) Significant abundance of cell subpopulations compared between PBMCs from aHUS treatment with plasma exchange alone, treatment combine with plasma exchange and anti-complement therapy, aHUS family and healthy subjects. (v-z2) *P < 0.05; **P < 0.01.
Figure 3.
(a-g) The figure presents boxplots displaying the subcluster significant abundance of various immune cell types (classical monocytes, central memory CD8 T cells, non-Vd2 gd T cells, Th1 cells, Th17 cells, non-classical monocytes, and terminal effector CD4 T cells) between PBMCs of individuals with aHUS, aHUS family, and healthy subjects. Dot plots of the gene expression profiles of the top 10 marker genes in each subcluster are also provided. Statistically significant differences are indicated by *P < 0.05 and **P < 0.01.
Figure 3.
(a-g) The figure presents boxplots displaying the subcluster significant abundance of various immune cell types (classical monocytes, central memory CD8 T cells, non-Vd2 gd T cells, Th1 cells, Th17 cells, non-classical monocytes, and terminal effector CD4 T cells) between PBMCs of individuals with aHUS, aHUS family, and healthy subjects. Dot plots of the gene expression profiles of the top 10 marker genes in each subcluster are also provided. Statistically significant differences are indicated by *P < 0.05 and **P < 0.01.
Figure 4.
(a-e) The figure displays boxplots of subcluster significant abundance for classical monocytes, central memory CD8 T cells, non-Vd2 gd T cells, terminal effector CD8 T cells, and Th1 cells between PBMCs of individuals with unstable and stable disease activity of aHUS, aHUS family, and healthy subjects, along with dot plots of the gene expression profiles of the top 10 marker genes in each subcluster. Statistically significant differences are indicated by *P < 0.05 and **P < 0.01.
Figure 4.
(a-e) The figure displays boxplots of subcluster significant abundance for classical monocytes, central memory CD8 T cells, non-Vd2 gd T cells, terminal effector CD8 T cells, and Th1 cells between PBMCs of individuals with unstable and stable disease activity of aHUS, aHUS family, and healthy subjects, along with dot plots of the gene expression profiles of the top 10 marker genes in each subcluster. Statistically significant differences are indicated by *P < 0.05 and **P < 0.01.
Figure 5.
The trajectories for B cells (a), T cells (b, c), and monocytes (d) with different state dynamics of the immune cells. (e) The pseudotime interval difference and abundance of B cells, while (f) and (g) show the pseudotime interval difference and abundance of T cells. (h) The pseudotime interval difference and abundance of monocytes. The pseudotime interval difference and abundance of B cells (i) and T cells (j), respectively, among the unstable aHUS group, stable aHUS group, aHUS family, and healthy subjects.
Figure 5.
The trajectories for B cells (a), T cells (b, c), and monocytes (d) with different state dynamics of the immune cells. (e) The pseudotime interval difference and abundance of B cells, while (f) and (g) show the pseudotime interval difference and abundance of T cells. (h) The pseudotime interval difference and abundance of monocytes. The pseudotime interval difference and abundance of B cells (i) and T cells (j), respectively, among the unstable aHUS group, stable aHUS group, aHUS family, and healthy subjects.
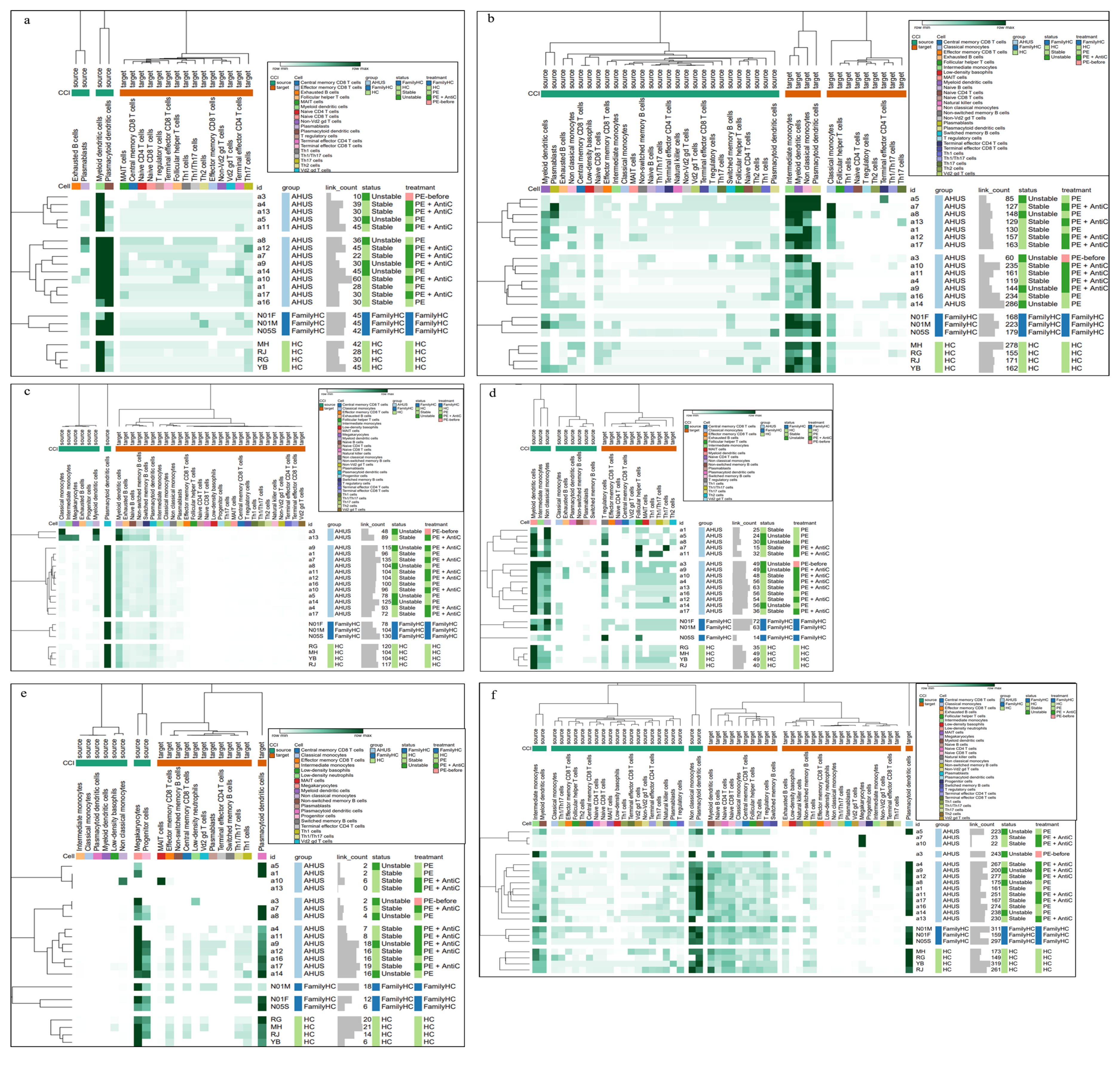
Figure 6.
Cell-cell interaction signaling of ALCAM-CD6 (a), IL16-CD4 (b), APP-CD40 (c), CD86-CTLA4 (d), CXC (e), and SELPLG (f) among individuals with aHUS with varying disease activity, treatment, aHUS family members, and healthy controls.
Figure 6.
Cell-cell interaction signaling of ALCAM-CD6 (a), IL16-CD4 (b), APP-CD40 (c), CD86-CTLA4 (d), CXC (e), and SELPLG (f) among individuals with aHUS with varying disease activity, treatment, aHUS family members, and healthy controls.
Table 1.
Characteristics of 13 aHUS patients.
Table 1.
Characteristics of 13 aHUS patients.
| Case |
Age |
Gender |
TMA involvement organ |
Treatment |
Disease activity |
| a1 |
39 |
M |
Kidney、Brain、Lung、Heart |
PE |
Stable |
| a3/a5 |
81 |
M |
Kidney、Heart |
a3 before 1st PE
a5 after PE |
Unstable
|
| a4 |
66 |
M |
Kidney、Brain、Heart |
PE + anti C5 |
Stable |
| a7 |
36 |
F |
Kidney、Heart、Pancreas、Eye |
PE + anti C5 |
Stable |
| a8 |
33 |
F |
Kidney、Brain、Lung、Heart |
PE |
Unstable |
| a9 |
62 |
F |
Kidney、Brain、Heart |
PE + anti C5 |
Unstable |
| a10 |
39 |
M |
Kidney、Brain、Lung、Heart、Eye |
PE + anti C5 |
Stable |
| a11 |
30 |
F |
Kidney、Brain、Lung、Heart、Eye、Bowel |
PE + anti C5 |
Stable |
| a12 |
42 |
F |
Kidney、Brain、Lung、Heart、Pancreas、Liver、Eye、Skin |
PE + anti C5 |
Stable |
| a13 |
53 |
M |
Kidney、Brain、Heart |
PE + anti C5 |
Stable |
| a14 |
70 |
M |
Kidney、Brain、Heart |
PE |
Unstable |
| a16 |
62 |
F |
Kidney、Heart |
PE |
Stable |
| a17 |
49 |
F |
Kidney、Brain、Lung、Heart、Pancreas、Liver、Eye |
PE+ anti C5 |
Stable |
Table 2.
Significantly difference in Cell subclusters in aHUS and healthy controls.
Table 2.
Significantly difference in Cell subclusters in aHUS and healthy controls.
| Significantly Increased Immune Cell Subclusters in aHUS Patients Compared to Healthy Controls with Correlated Gene Expression Increasing |
| Cell Subclusters |
|
P value |
Higher expression levels of gene |
| Classical monocyte subclusters 6 |
|
P < 0.01 |
RPS27 |
| Classical monocyte subclusters 7 |
|
P < 0.05 |
IFI27 |
| Central memory CD8 T cells subcluster 3 |
|
P < 0.05 |
CXCR4 |
| Non-Vd2 gd T cells subcluster 4 |
|
P < 0.05 |
SYNE2 |
| Th1 cells subcluster 3 |
|
P < 0.05 |
MT-CYB |
| Th17 cells subcluster 4 |
|
P < 0.05 |
MT-ATP6 |
| Significantly Increased Immune Cell Subclusters in Healthy Controls Compared to aHUS Patients with Correlated Gene Expression Increasing |
| Cell Subclusters |
|
P value |
Higher expression levels of gene |
| Central memory CD8 T cells subcluster 1 |
|
P < 0.05 |
EIF3E |
| Th1 cells subcluster 0 |
|
P < 0.05 |
RPS27 |
| Non classical monocytes subcluster 5 |
|
P< 0.01 |
LYPD2 |
| Terminal effector CD4 T cells subcluster 3 |
|
P< 0.01 |
KLRD1 |
| Th17 cells subcluster 3 |
|
P < 0.05 |
ACTG1, CD52 and LGALS1 |
Table 3.
Significantly difference in Cell subclusters in aHUS disease activity.
Table 3.
Significantly difference in Cell subclusters in aHUS disease activity.
| Significantly Increased Immune Cell Subclusters in unstable aHUS Patients Compared to stable aHUS Patients with Correlated Gene Expression Increasing |
|---|
| Cell Subclusters |
|
P value |
Higher expression levels of gene |
| Classical monocyte subclusters 4 |
|
P < 0.05 |
NEAT1, MT-ATP6 and MT-CYB |
| Central memory CD8 T cells subcluster 2 |
|
P < 0.05 |
VIM |
| Non-Vd2 gd T cells subcluster 1 |
|
P < 0.05 |
ACTG1 |
| Terminal effector CD8 T cells subcluster 3 |
|
P < 0.05 |
RPL13 |
| Terminal effector CD8 T cells subcluster 5 |
|
P < 0.01 |
KLRB1 |
| Significantly Increased Immune Cell Subclusters in stable aHUS Patients Compared to unstable aHUS Patients with Correlated Gene Expression Increasing |
| Cell Subclusters |
|
P value |
Higher expression levels of gene |
| Central memory CD8 T cells subcluster 1 |
|
P < 0.05 |
RPL23 |
| Non-Vd2 gd T cell subcluster 0 |
|
P < 0.05 |
GZMH |
| Th1 cells subcluster 0 |
|
P < 0.05 |
RPS27, RPS4X |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).