Submitted:
09 March 2023
Posted:
17 March 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
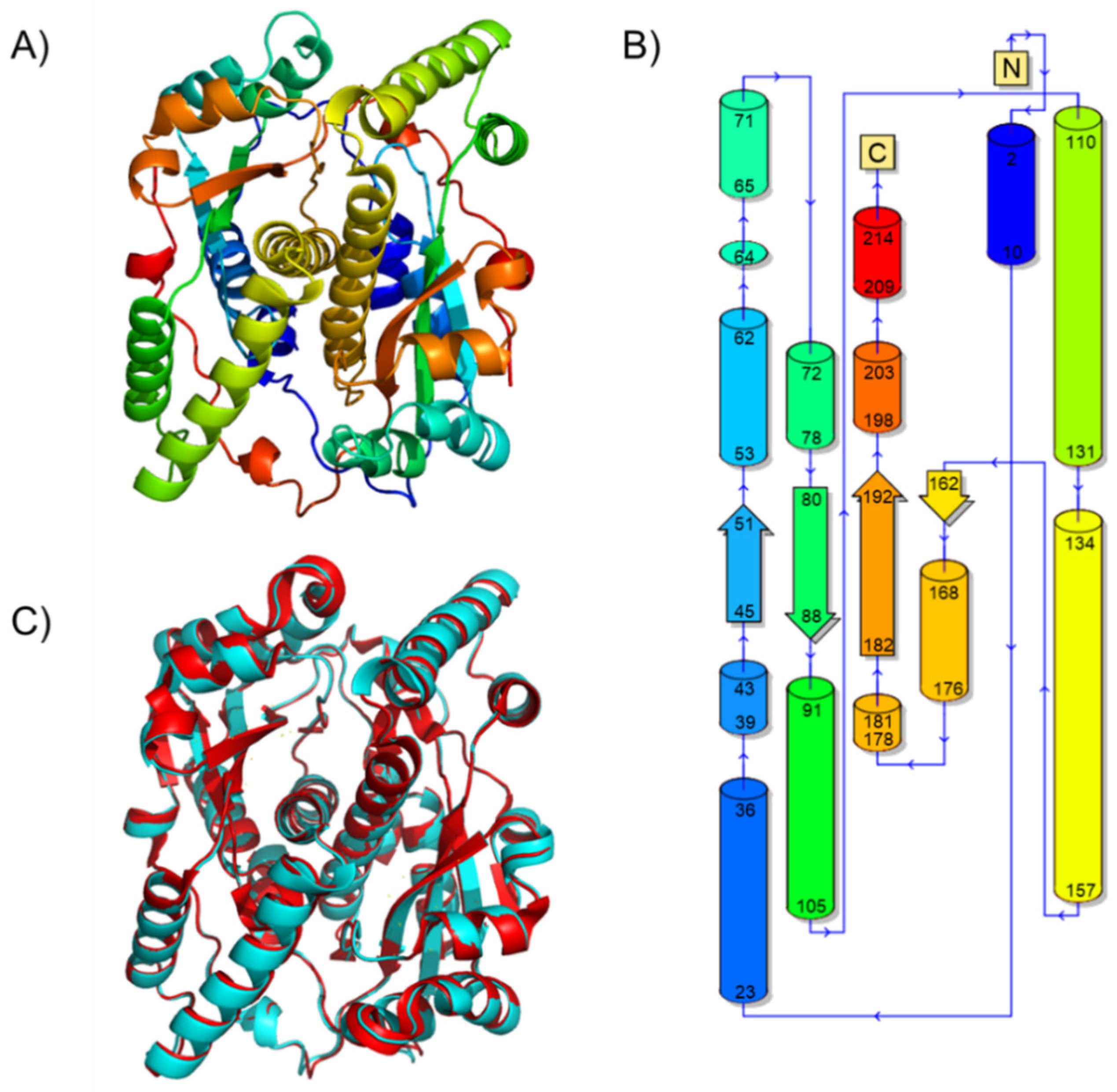
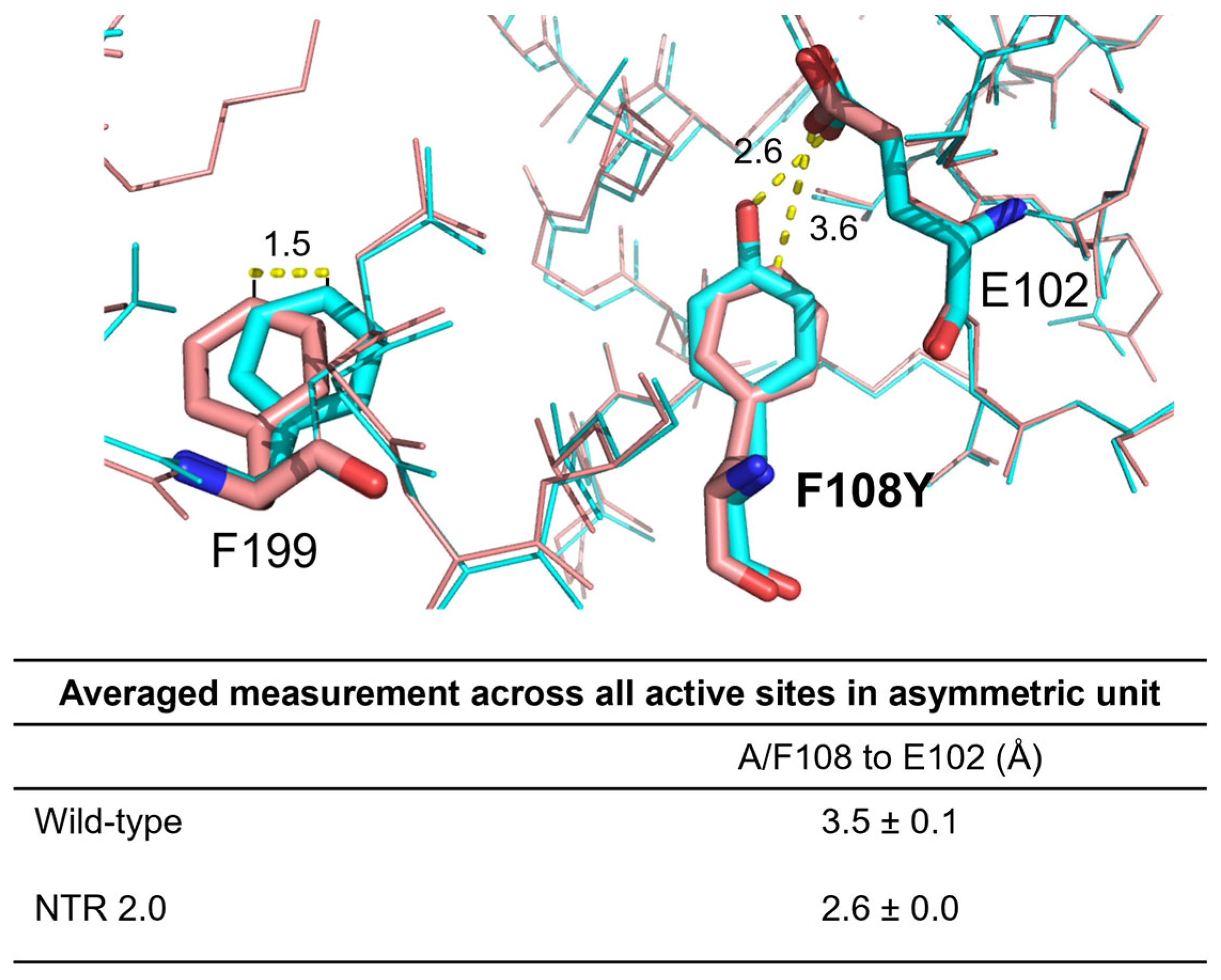
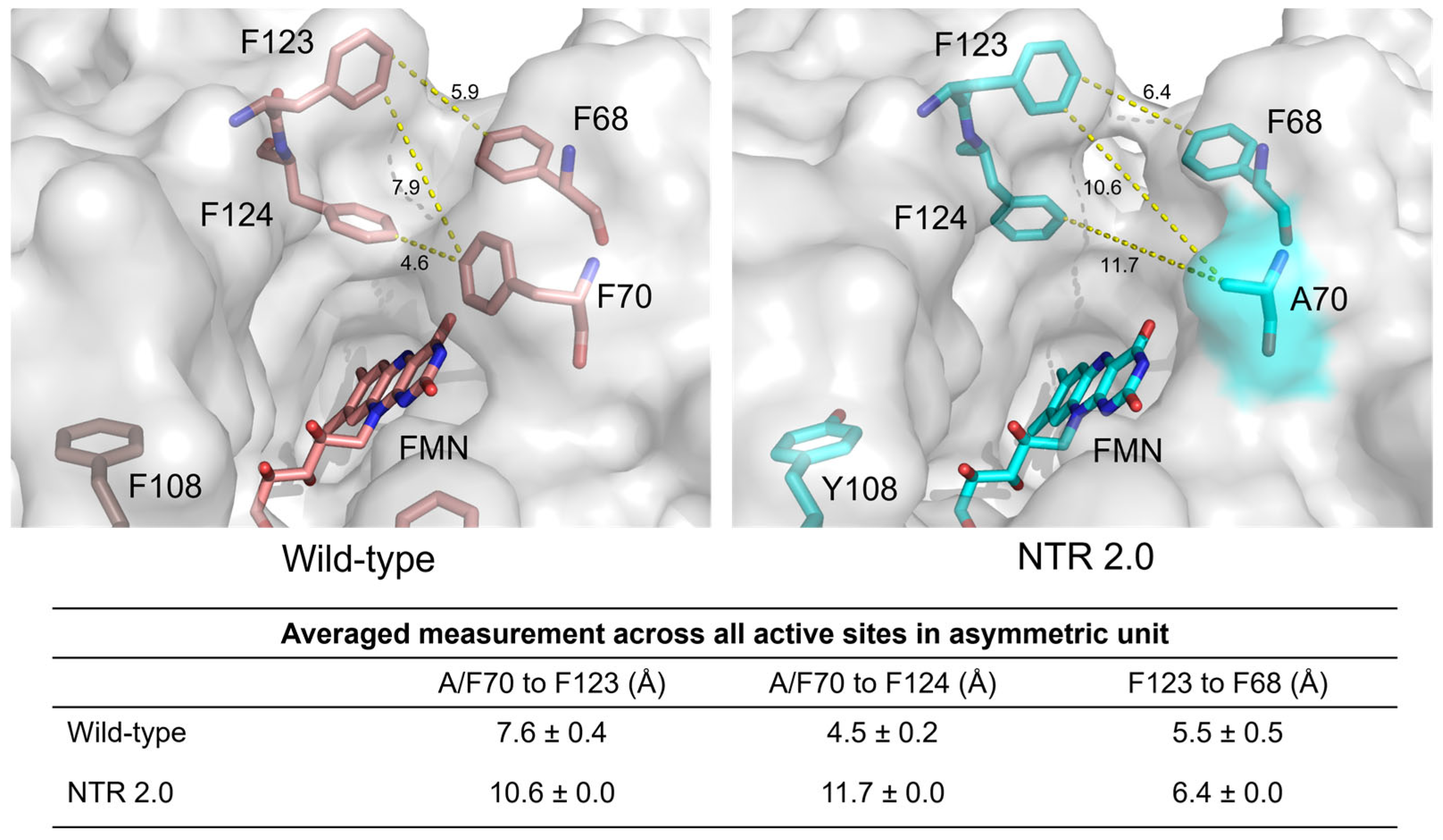
2.1. The Structure of Homodimeric NTR 2.0 and Comparison to Unmodified V. vulnificus NfsB
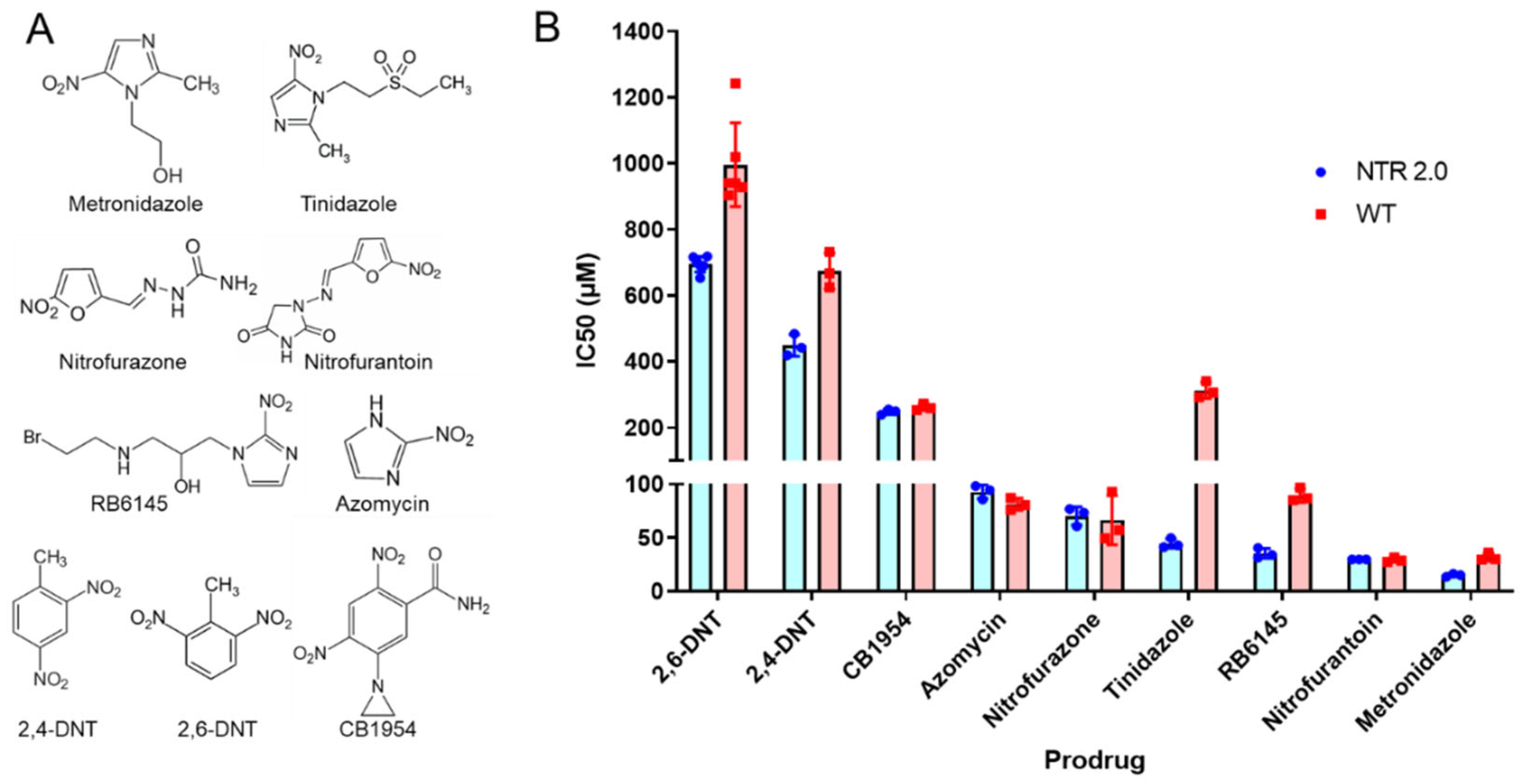
2.2. Surveying NTR 2.0 Activity with Alternative Nitroaromatic Substrates
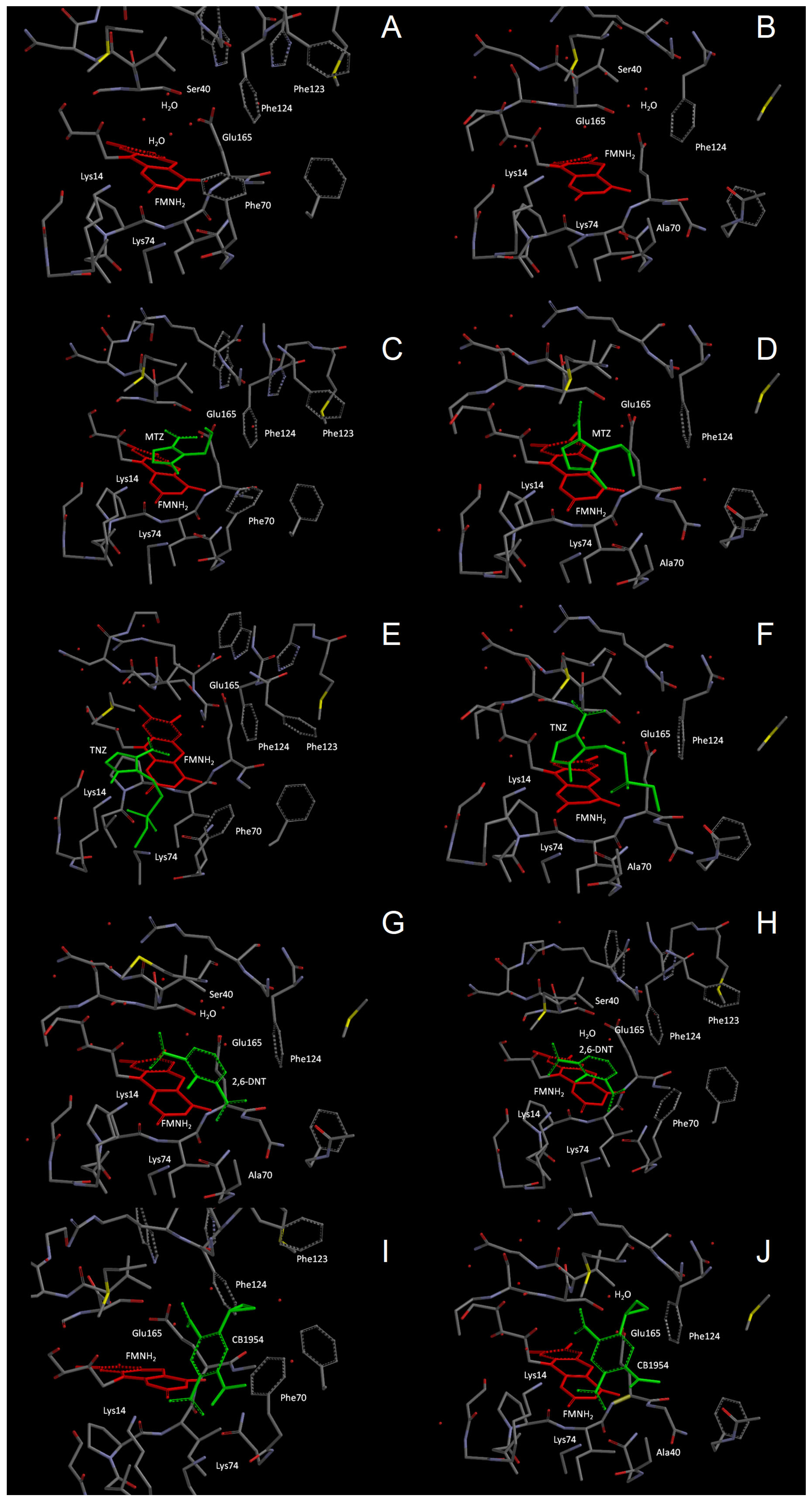
2.3. Molecular Modelling of Prodrug Binding—NTR 2.0 vs. Unmodified V. vulnificus NfsB
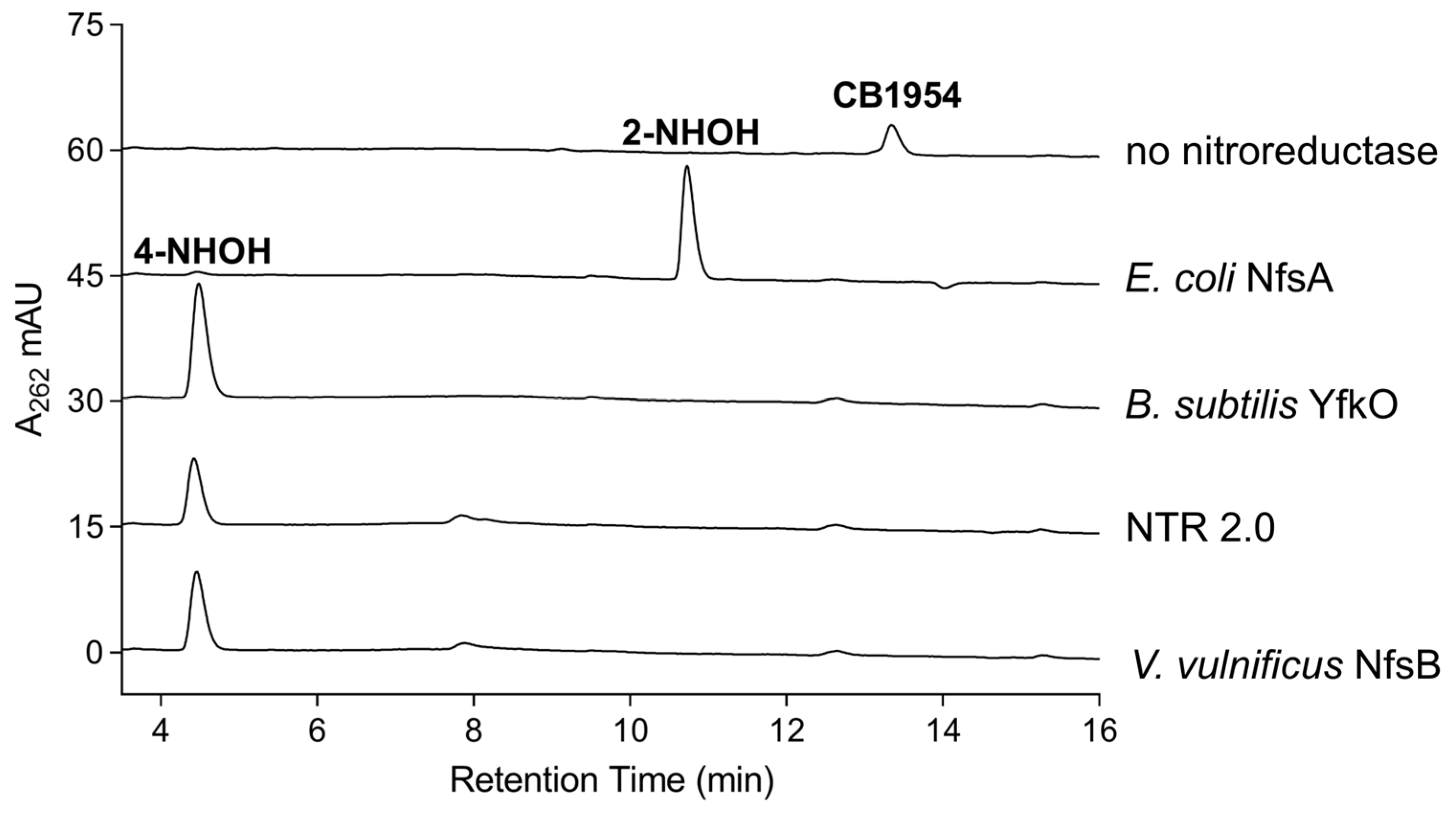
2.4. Analysis of CB1954 Reduction Products
3. Materials and Methods
3.1. Bacterial Strains, Media and Growth Conditions
3.2. Cloning, Expression and Purification of NTR 2.0
3.3. Crystallisation
3.4. X-ray Data Collection
3.5. Structure Solution and Refinement
3.6. Bacterial Cytotoxicity Assays
3.7. HPLC Identification of CB1954 Nitroreduction Products
3.8. Theoretical Modelling
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Ju KS, Parales RE. Nitroaromatic compounds, from synthesis to biodegradation. Microbiol Mol Biol Rev. 2010;74(2):250-72. [CrossRef]
- Williams EM, Little RF, Mowday AM, Rich MH, Chan-Hyams JV, Copp JN, et al. Nitroreductase gene-directed enzyme prodrug therapy: insights and advances toward clinical utility. Biochem J. 2015;471(2):131-53. [CrossRef]
- White DT, Mumm JS. The nitroreductase system of inducible targeted ablation facilitates cell-specific regenerative studies in zebrafish. Methods. 2013;62(3):232-40. [CrossRef]
- Zhang L, Chen C, Fu J, Lilley B, Berlinicke C, Hansen B, et al. Large-scale phenotypic drug screen identifies neuroprotectants in zebrafish and mouse models of retinitis pigmentosa. eLife. 2021;10:e57245. [CrossRef]
- Pisharath H, Parsons MJ. Nitroreductase-mediated cell ablation in transgenic zebrafish embryos. Methods Mol Biol. 2009;546:133-43. [CrossRef]
- Sharrock AV, Mulligan TS, Hall KR, Williams EM, White DT, Zhang L, et al. NTR 2.0: a rationally engineered prodrug-converting enzyme with substantially enhanced efficacy for targeted cell ablation. Nature Methods. 2022;19(2):205-15. [CrossRef]
- Mathias JR, Zhang Z, Saxena MT, Mumm JS. Enhanced cell-specific ablation in zebrafish using a triple mutant of Escherichia coli nitroreductase. Zebrafish. 2014;11(2):85-97. [CrossRef]
- Matthews BW. Solvent content of protein crystals. J Mol Biol. 1968;33(2):491-7. [CrossRef]
- Pitsawong W, Hoben JP, Miller AF. Understanding the broad substrate repertoire of nitroreductase based on its kinetic mechanism. J Biol Chem. 2014;289(22):15203-14. [CrossRef]
- Parkinson GN, Skelly JV, Neidle S. Crystal structure of FMN-dependent nitroreductase from Escherichia coli B: a prodrug-activating enzyme. J Med Chem. 2000;43(20):3624-31. [CrossRef]
- Valiauga B, Williams EM, Ackerley DF, Cenas N. Reduction of quinones and nitroaromatic compounds by Escherichia coli nitroreductase A (NfsA): characterization of kinetics and substrate specificity. Arch Biochem Biophys. 2017;614:14-22. [CrossRef]
- Williams EM. Development of bacterial nitroreductase enzymes for noninvasive imaging in cancer gene therapy : PhD Thesis. School of Biological Sciences, Victoria University of Wellington, Wellington.; 2013.
- Koder RL, Haynes CA, Rodgers ME, Rodgers DW, Miller A-F. Flavin thermodynamics explain the oxygen insensitivity of enteric nitroreductases. Biochemistry. 2002;41(48):14197-205. [CrossRef]
- Race PR, Lovering AL, Green RM, Ossor A, White SA, Searle PF, et al. Structural and mechanistic studies of Escherichia coli nitroreductase with the antibiotic nitrofurazone: reversed binding orientations in different redox states of the enzyme. J Biol Chem. 2005;280(14):13256-64. [CrossRef]
- Bauzá A, Deyà PM, Frontera A. Anion-π Interactions in Supramolecular Chemistry and Catalysis. In: Scheiner S, editor. Noncovalent Forces. Cham: Springer International Publishing; 2015. p. 471-500. [CrossRef]
- Kokkonen P, Bednar D, Pinto G, Prokop Z, Damborsky J. Engineering enzyme access tunnels. Biotechnology Advances. 2019;37(6):107386. [CrossRef]
- Ashkenazy H, Abadi S, Martz E, Chay O, Mayrose I, Pupko T, et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 2016;44(W1):W344-50. [CrossRef]
- Williams EM, Sharrock AV, Rylott EL, Bruce NC, MacKichan JK, Ackerley DF. A cofactor consumption screen identifies promising NfsB family nitroreductases for dinitrotoluene remediation. Biotechnol Lett. 2019;41(10):1155-62. [CrossRef]
- Valiauga B, Bagdžiūnas G, Sharrock AV, Ackerley DF, Čėnas N, editors. E. coli nitroreductase-A as a candidate for gene-directed prodrug therapy: potentiometric data and modes of substrate binding. SCT 29th Young Research Fellows Meeting; 2022 4-6 July; Nantes, France.
- Day MA, Jarrom D, Christofferson AJ, Graziano AE, Anderson JLR, Searle PF, et al. The structures of E. coli NfsA bound to the antibiotic nitrofurantoin; to 1,4-benzoquinone and to FMN. Biochem J. 2021;478(13):2601-17. [CrossRef]
- Helsby NA, Ferry DM, Patterson AV, Pullen SM, Wilson WR. 2-Amino metabolites are key mediators of CB1954 and SN23862 bystander effects in nitroreductase GDEPT. Br J Cancer. 2004;90(5):1084-92. [CrossRef]
- Knox RJ, Friedlos F, Marchbank T, Roberts JJ. Bioactivation of CB 1954: reaction of the active 4-hydroxylamino derivative with thioesters to form the ultimate DNA-DNA interstrand crosslinking species. Biochem Pharmacol. 1991;42(9):1691-7. [CrossRef]
- Helsby NA, Wheeler SJ, Pruijn FB, Palmer BD, Yang S, Denny WA, et al. Effect of nitroreduction on the alkylating reactivity and cytotoxicity of the 2,4-dinitrobenzamide-5-aziridine CB1954 and the corresponding nitrogen mustard SN23862: distinct mechanisms of bioreductive activation. Chem Res Toxicol. 2003;16(4):469-78. [CrossRef]
- Knox RJ, Friedlos F, Jarman M, Roberts JJ. A new cytotoxic, DNA interstrand crosslinking agent, 5-(aziridin-1-yl)-4-hydroxylamino-2-nitrobenzamide, is formed from 5-(aziridin-1-yl)-2,4-dinitrobenzamide (CB 1954) by a nitroreductase enzyme in Walker carcinoma cells. Biochem Pharmacol. 1988;37(24):4661-9. [CrossRef]
- Osswald A, Sun Z, Grimm V, Ampem G, Riegel K, Westendorf AM, et al. Three-dimensional tumor spheroids for in vitro analysis of bacteria as gene delivery vectors in tumor therapy. Microbial Cell Factories. 2015;14(1):199. [CrossRef]
- Anlezark GM, Vaughan T, Fashola-Stone E, Michael NP, Murdoch H, Sims MA, et al. Bacillus amyloliquefaciens orthologue of Bacillus subtilis ywrO encodes a nitroreductase enzyme which activates the prodrug CB1954. Microbiology (Reading, England). 2002;148:297-306. [CrossRef]
- Vass SO, Jarrom D, Wilson WR, Hyde EI, Searle PF. E. coli NfsA: an alternative nitroreductase for prodrug activation gene therapy in combination with CB1954. Br J Cancer. 2009;100(12):1903-11. [CrossRef]
- Prosser GA, Copp JN, Mowday AM, Guise CP, Syddall SP, Williams EM, et al. Creation and screening of a multi-family bacterial oxidoreductase library to discover novel nitroreductases that efficiently activate the bioreductive prodrugs CB1954 and PR-104A. Biochem Pharmacol. 2013;85(8):1091-103. [CrossRef]
- Prosser GA, Patterson AV, Ackerley DF. uvrB gene deletion enhances SOS chromotest sensitivity for nitroreductases that preferentially generate the 4-hydroxylamine metabolite of the anti-cancer prodrug CB1954. J Biotechnol. 2010;150(1):190-4. [CrossRef]
- Copp JN, Hanson-Manful P, Ackerley DF, Patrick WM. Error-prone PCR and effective generation of gene variant libraries for directed evolution. Methods Mol Biol. 2014;1179:3-22. [CrossRef]
- Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, et al. Overview of the CCP4 suite and current developments. Acta Crystallographica Section D: Biological Crystallography. 2011;67(4):235-42. [CrossRef]
- Leslie AG, Powell HR. Processing diffraction data with mosflm. Evolving methods for macromolecular crystallography: Springer; 2007. p. 41-51. [CrossRef]
- Evans PR, Murshudov GN. How good are my data and what is the resolution? Acta Crystallographica Section D: Biological Crystallography. 2013;69(7):1204-14. [CrossRef]
- McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ. Phaser crystallographic software. Journal of applied crystallography. 2007;40(4):658-74. [CrossRef]
- Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta Crystallographica Section D: Biological Crystallography. 2010;66(4):486-501. [CrossRef]
- Radveikienė I, Palinauskas D, Ragauskaitė E, Bagdžiūnas G. Self-assembled cyclodextrins-based nanostructures on indium-tin-oxide for a detection of catecholamine neurotransmitters. Applied Surface Science. 2022;600:154170. [CrossRef]
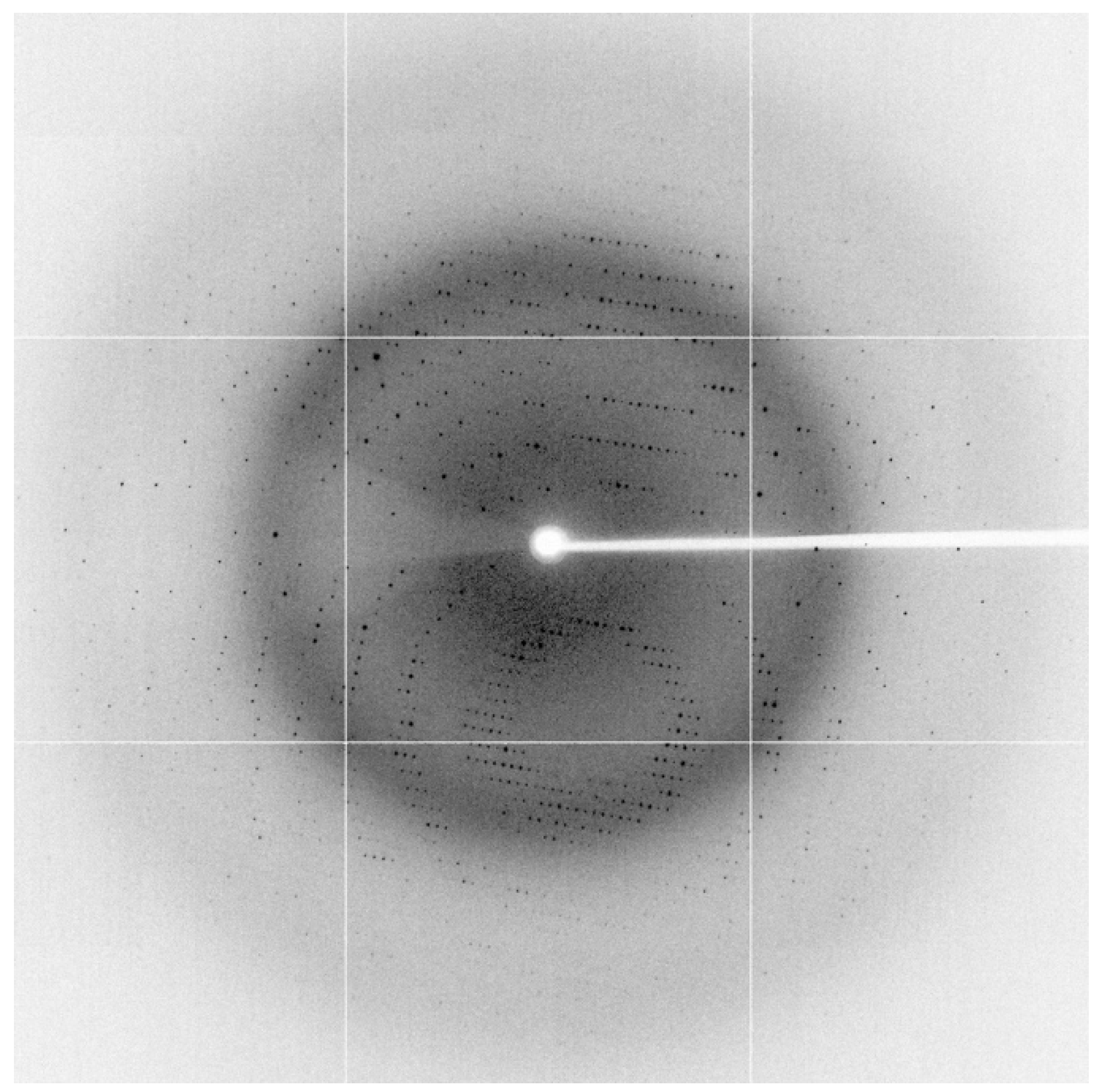


| Wavelength (Å) | 0.9537 |
| Space group | C 2 2 21 |
| Unit-cell parameters (Å, °) | a = 68.43, b = 137.36, c = 107.04, α/β/γ = 90/90/90 |
| Resolution range (Å) | 68.68-1.85 (1.89-1.85) |
| Measured reflections | 624970 |
| Unique reflections | 43344 |
| Multiplicity | 14.4 |
| Temperature (K) | 100 |
| Matthews coefficient (Å3 Da−1) | 2.32 |
| Solvent content (%) | 47 |
| No. of molecules in ASU | 2 |
| Completeness (%) | 99.9 (99.7) |
| Mean I/σ(I) | 18.3 (3.7) |
| Rmerge† (%) | 12 (79.2) |
| CC(1/2) | 0.999 |
| Wilson B factor (Å2) | 10.275 |
| Rfactor | 0.165 |
| Rfree | 0.196 |
| Protein atoms | 3342 |
| Other ions/molecules | 7 |
| Number of waters | 184 |
| B factors (protein) | 18.5 (Chain A)/19.8 (Chain B) |
| B factors (water) | 23.9 |
| RMSD | |
| Bond angles (°) | 1.6771 |
| Bond lengths (Å) | 0.0120 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




