Submitted:
17 February 2023
Posted:
20 February 2023
You are already at the latest version
Abstract
Keywords:
Introduction
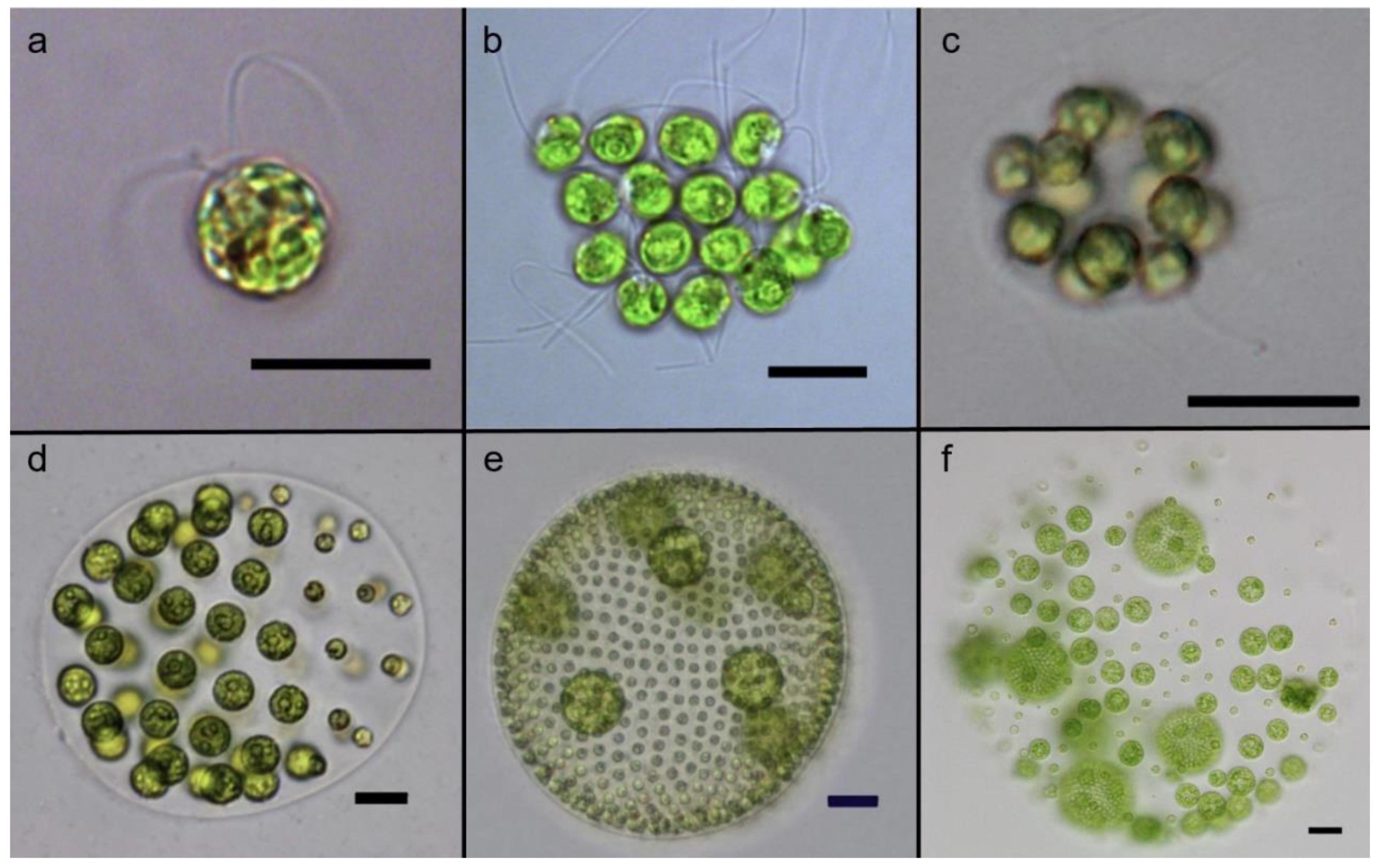
The volvocine model system
regA gene structure and function
Evolution of the VARL gene family
SAND domain-containing sequences beyond volvocine algae
Functional Evolution of regA Through Co-option of a Life History Trade-off Gene
Life history trade-offs in single celled organisms
V. carteri regA retained the ancestral environmental regulation
Co-option of an environmentally regulated gene into a developmental master regulator
The general role of stress and life history trade-off genes in the re-organization of fitness during the evolution of multicellularity
Future directions
Summary
- The evolution of cellular differentiation is a key event during the transition from single-celled to multicellular life. Specialized germ cells and somatic cells reorganize the two essential components of fitness between different cell types, thereby transferring fitness from the cell-level to the multicellular level. Understanding the genetic basis for the evolution of cellular differentiation during the unicellular-to-multicellular transitions is a major challenge of evolutionary biology. Co-option of life history trade-off genes present in unicellular organisms that differentially affect survival and reproductive functions in response to the environment is one route for the evolution of genes controlling cellular differentiation. The regA-like gene family of the volvocine green algae is an unrivaled model system to study this co-option due to the recent origin of multicellularity in this clade and presence of extant relatives at different levels of multicellular complexity.
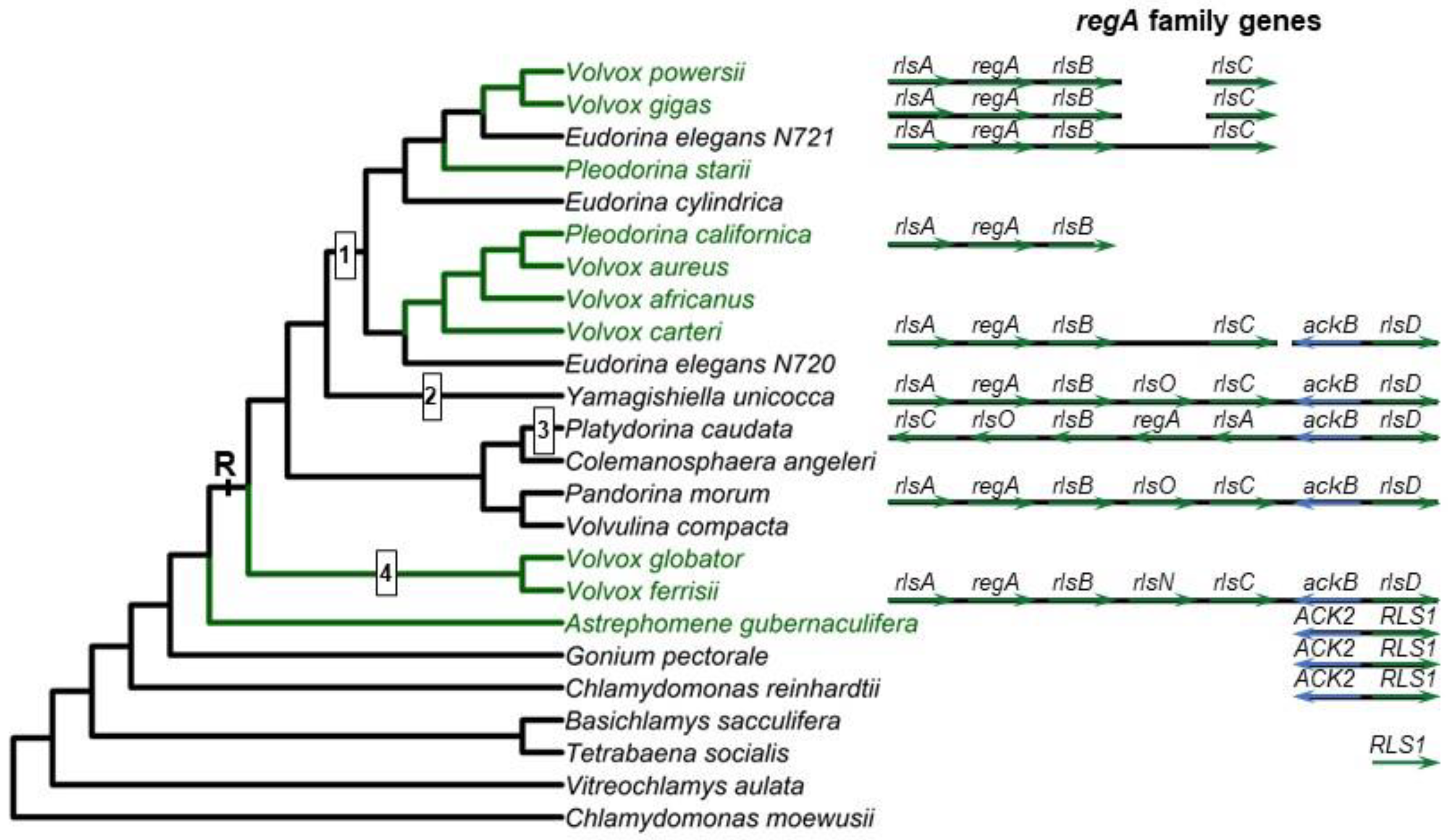
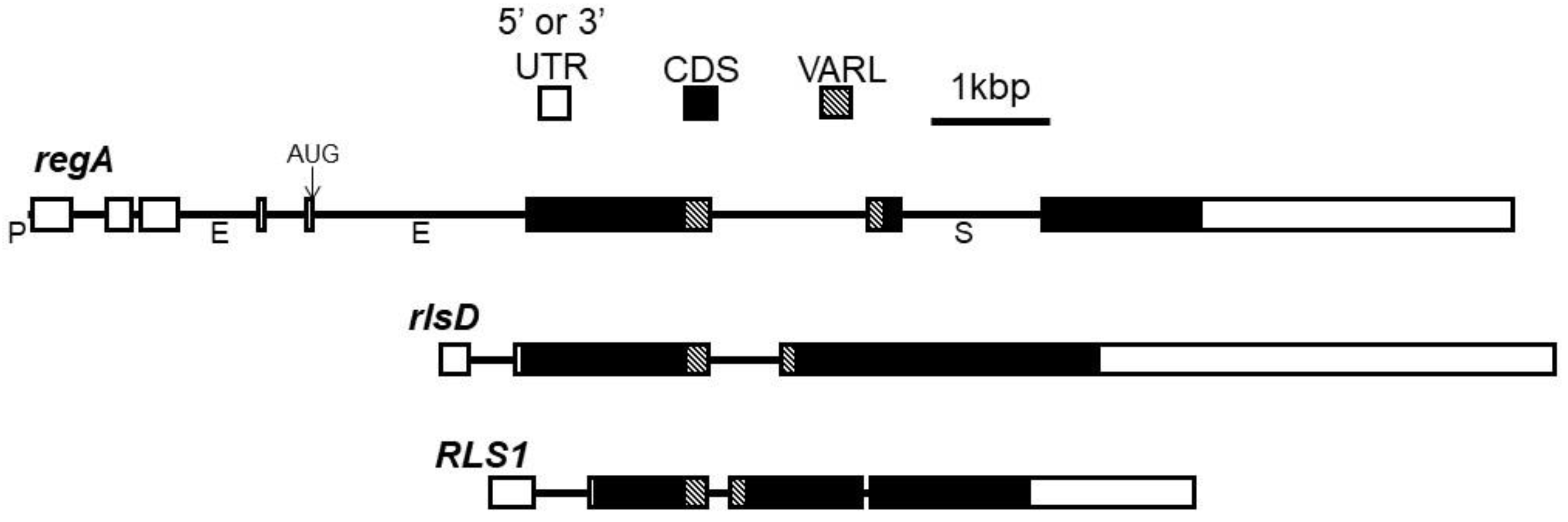
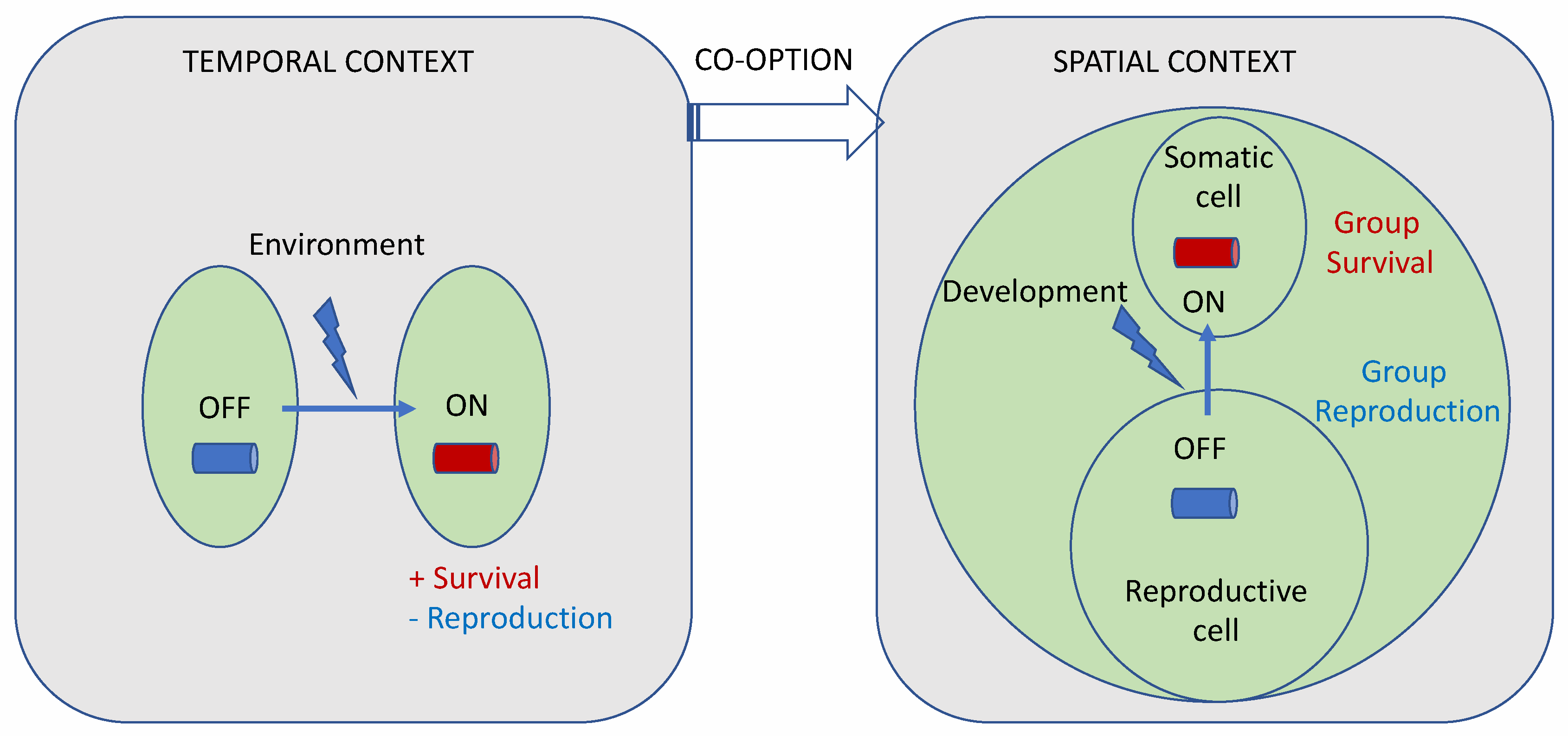
- The regA gene from V. carteri f. nagariensis is the type-gene for the VARL gene family specific to the volvocine algae lineage. The common ancestor of V. carteri and C. reinhardtii likely had several VARL gene family members, one of which was RLS1. The RLS1 gene duplicated several times to give rise to the regA gene cluster in the common ancestor of the Volvocaceae, setting the stage for the functional co-option of regA during the evolution of cellular differentiation as well as other lineage-specific changes to regA cluster genes (Figure 2). The co-option of RLS1’s functions into a regA-like gene responsible for somatic cell differentiation likely involved the simulation of the ancestral environmentally induced signal into a developmental context (Figure 9).
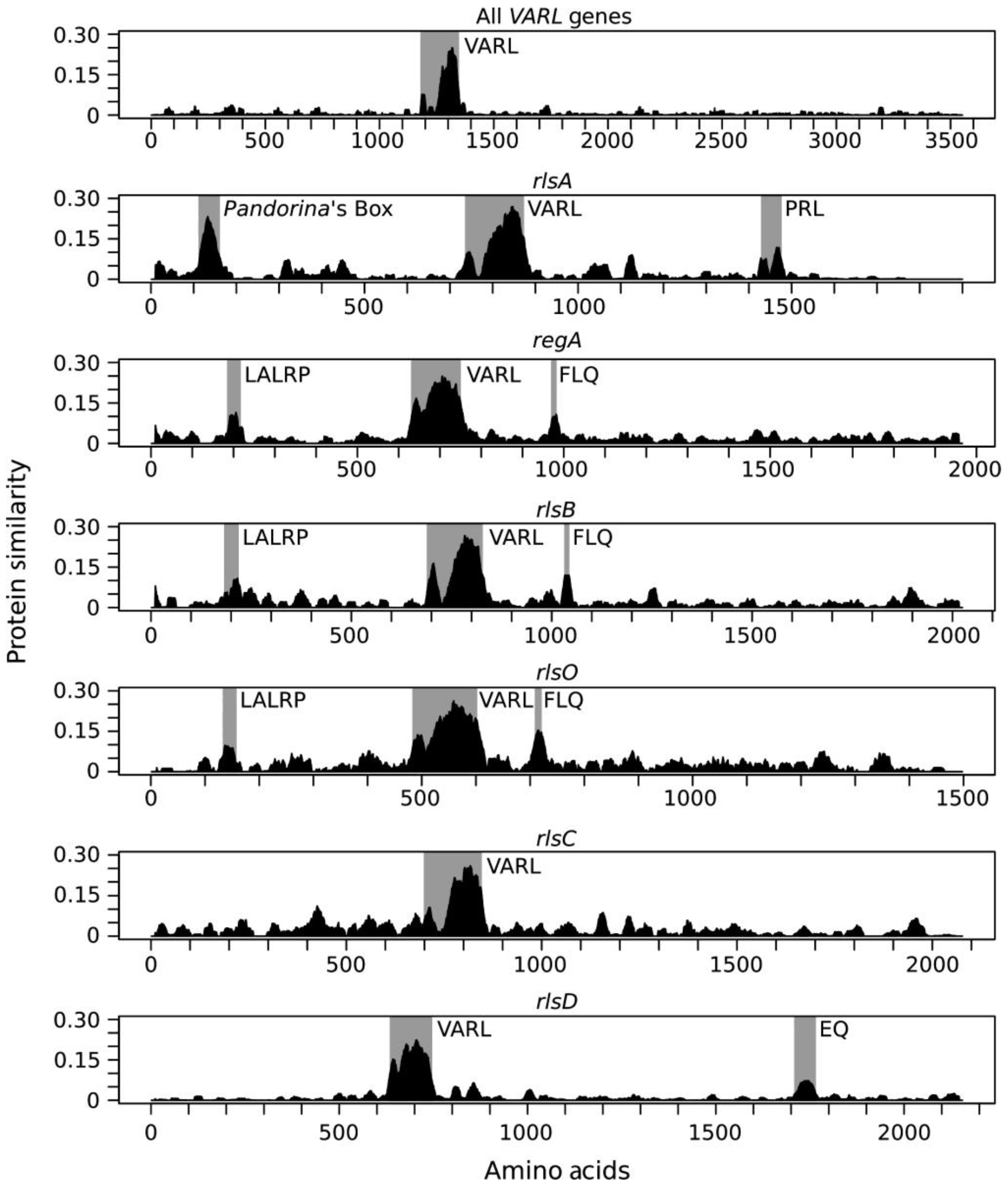
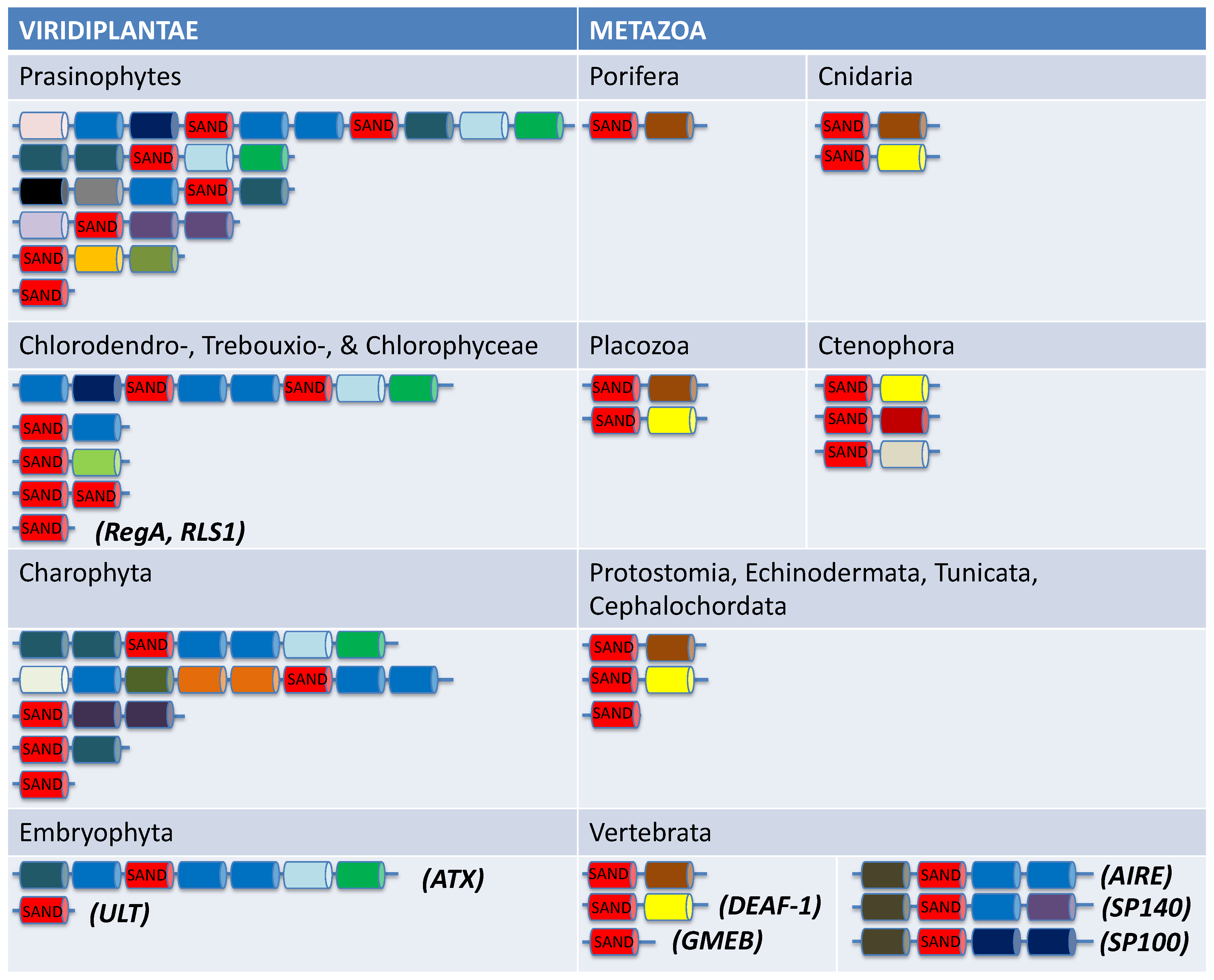
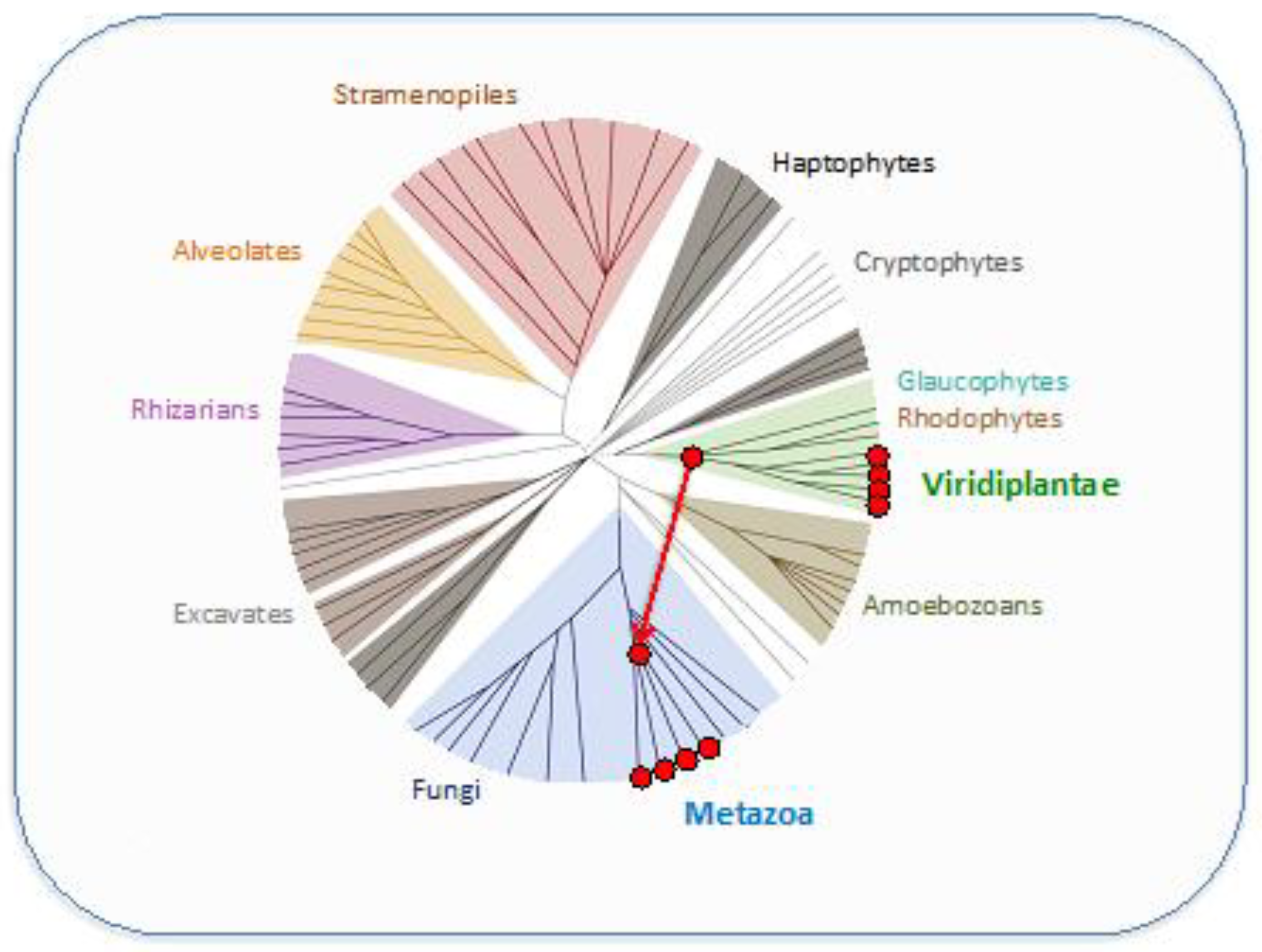
- The defining feature of all VARL genes is the presence of the VARL domain which contains a conserved SAND domain. The SAND domain is found in other green algae, land plants, and animals but appears to be missing from other eukaryotic lineages. This indicates the possibility that the SAND domain was horizontally transferred between green algae and animals early in eukaryotic evolution where it appears to have been co-opted multiple times independently in a variety of developmentally important functions.
- Overall, we argue that the co-option of life history trade-off genes during the transition to multicellularity underlies the re-organization of fitness between soma and germ to optimize various aspects of fitness at the multicellular level and contribute to the stability of the multicellular individual. Although all organisms must have such stress response trade-off genes, it remains to be determined whether or how fitness reorganization and co-option of life history genes during the evolution of specialized cell types applies to other lineages that evolved multicellularity independently.
Author Contributions
Funding
Conflicts of Interest
References
- Alvarez-Curto, E., Rozen, D. E., Ritchie, A. V., Fouquet, C., Baldauf, S. L., & Schaap, P. (2005). Evolutionary origin of cAMP-based chemoattraction in the social amoebae. Proceedings of the National Academy of Sciences of the United States of America, 102(18), 6385–6390. [CrossRef]
- Arakaki, Y., Kawai-Toyooka, H., Hamamura, Y., Higashiyama, T., Noga, A., Hirono, M., Olson, B. J. S. C., & Nozaki, H. (2013). The simplest integrated multicellular organism unveiled. PloS One, 8(12), e81641. [CrossRef]
- Babinger, K., Hallmann, A., & Schmitt, R. (2006). Translational control of regA, a key gene controlling cell differentiation in Volvox carteri. Development (Cambridge, England), 133, 4045–4051. [CrossRef]
- Baran, G. (1984). Analysis of somatic cell differentiation in Volvox carteri f. nagariensis. University of Virginia.
- Barker, H. E., Smyth, G. K., Wettenhall, J., Ward, T. A., Bath, M. L., Lindeman, G. J., & Visvader, J. E. (2008). Deaf-1 regulates epithelial cell proliferation and side-branching in the mammary gland. BMC Developmental Biology, 8(1), 94. [CrossRef]
- Bochdanovits, Z., & De Jong, G. (2004). Antagonistic pleiotropy for life-history traits at the gene expression level. Proceedings of the Royal Society B: Biological Sciences, 271(SUPPL. 3), S75–S78. [CrossRef]
- Bonner, J. T. (1998). The origins of multicellularity. Integrative Biology: Issues, News, and Reviews, 1(1), 27–36. [CrossRef]
- Boraas, M. E., Seale, D. B., & Boxhorn, J. E. (1998). Phagotrophy by flagellate selects for colonial prey: A possible origin of multicellularity. Evolutionary Ecology, 12(1973), 153–164. [CrossRef]
- Bottomley, M. J. J., Collard, M. W. W., Huggenvik, J. I. I., Liu, Z., Gibson, T. J. J., & Sattler, M. (2001). The SAND domain structure defines a novel DNA-binding fold in transcriptional regulation. Nature Structural Biology, 8(7), 626–633. [CrossRef]
- Brunet, T., & King, N. (2017). The Origin of Animal Multicellularity and Cell Differentiation. Developmental Cell, 43(2), 124–140. [CrossRef]
- Buss, LeoW. (1987). The Evolution of Individuality. Princeton University Press.
- Cameron-Pack, M. E., König, S. G., Reyes-Guevara, A., Reyes-Prieto, A., & Nedelcu, A. M. (2022). A personal cost of cheating can stabilize reproductive altruism during the early evolution of clonal multicellularity. Biology Letters, 18(6), 20220059. [CrossRef]
- Carles, C. C. (2005). ULTRAPETALA1 encodes a SAND domain putative transcriptional regulator that controls shoot and floral meristem activity in Arabidopsis. Development, 132(5), 897–911. [CrossRef]
- Carles, C. C., & Fletcher, J. C. (2009). The SAND domain protein ULTRAPETALA1 acts as a trithorax group factor to regulate cell fate in plants. Genes and Development, 23(23), 2723–2728. [CrossRef]
- Chen, L.-Q., Luo, J.-H., Cui, Z.-H., Xue, M., Wang, L., Zhang, X.-Y., Pawlowski, W. P., & He, Y. (2017). ATX3, ATX4, and ATX5 Encode Putative H3K4 Methyltransferases and Are Critical for Plant Development. Plant Physiology, 174(3), 1795–1806. [CrossRef]
- Choi, G., Przybylska, M., & Straus, D. (1996). Three abundant germ line-specific transcripts in Volvox carteri encode photosynthetic proteins. Current Genetics, 30(4), 347–355. http://www.ncbi.nlm.nih.gov/pubmed/8781179.
- Coleman, A. W. (2012). A Comparative Analysis of the Volvocaceae (Chlorophyta). Journal of Phycology, 48(3), 491–513. [CrossRef]
- Davison, D. R., & Michod, R. E. (2021). Phenotypic Plasticity and Evolutionary Transitions in Individuality. In D. W. Pfennig (Ed.), Phenotypic Plasticity & Evolution: Causes, Consequences, Controversies (pp. 241–266). CRC Press.
- Duncan, L., Nishii, I., Harryman, A., Buckley, S., Howard, A., Friedman, N. R., & Miller, S. M. (2007). The VARL gene family and the evolutionary origins of the master cell-type regulatory gene, regA, in Volvox carteri. Journal of Molecular Evolution, 65(1), 1–11. [CrossRef]
- Duncan, L., Nishii, I., Howard, A., Kirk, D., & Miller, S. M. (2006). Orthologs and paralogs of regA, a master cell-type regulatory gene in Volvox carteri. Current Genetics, 50(1), 61–72. [CrossRef]
- Featherston, J., Arakaki, Y., Hanschen, E. R., Ferris, P. J., Michod, R. E., Olson, B. J. S. C., Nozaki, H., & Durand, P. M. (2018). The 4-celled tetrabaena socialis nuclear genome reveals the essential components for genetic control of cell number at the origin of multicellularity in the volvocine lineage. Molecular Biology and Evolution, 35(4), 855–870. [CrossRef]
- Ferenci, T. (2016). Trade-off Mechanisms Shaping the Diversity of Bacteria. Trends in Microbiology, 24(3), 209–223. [CrossRef]
- Flatt, T. (2020). Life-history evolution and the genetics of fitness components in drosophila melanogaster. Genetics, 214(1), 3–48. [CrossRef]
- Flatt, T., & Heyland, Andreas. (2011). Mechanisms of Life History Evolution: the Genetics and Physiology of Life History Traits and Trade-Offs. Oxford University Press.
- Fletcher, J. C. (2001). The ULTRAPETALA gene controls shoot and floral meristem size in Arabidopsis. Development (Cambridge, England), 128(8), 1323–1333.
- Grochau-Wright, Z. I. (2019). THE ORIGIN AND EVOLUTION OF THE REG CLUSTER IN THE VOLVOCINE GREEN ALGAE: A MODEL SYSTEM FOR THE EVOLUTION OF CELLULAR DIFFERENTIATION [PhD Dissertation]. University of Arizona.
- Grochau-Wright, Z. I., Ferris, P. J., Tumberger, J., Jiménez-Marin, B., Olson, B. J. S. C., & Michod, R. E. (2021). Characterization and Transformation of reg Cluster Genes in Volvox powersii Enable Investigation of Convergent Evolution of Cellular Differentiation in Volvox. Protist, 172, 125834. [CrossRef]
- Grochau-Wright, Z. I., Hanschen, E. R., Ferris, P. J., Hamaji, T., Nozaki, H., Olson, B. J. S. C., & Michod, R. E. (2017). Genetic Basis for Soma is Present in Undifferentiated Volvocine Green Algae. Journal of Evolutionary Biology, 30, 1205–1218.
- Grosberg, R. K., & Strathmann, R. R. (2007). The Evolution of Multicellularity: A Minor Major Transition? Annual Review of Ecology, Evolution, and Systematics, 38(1), 621–654. [CrossRef]
- Grossman, A. (2000). Acclimation of Chlamydomonas reinhardtii to its nutrient environment. In Protist (Vol. 151, Issue 3, pp. 201–224). Elsevier GmbH. [CrossRef]
- Hamaji, T., Kawai-Toyooka, H., Uchimura, H., Suzuki, M., Noguchi, H., Minakuchi, Y., Toyoda, A., Fujiyama, A., Miyagishima, S., Umen, J. G., & Nozaki, H. (2018). Anisogamy evolved with a reduced sex-determining region in volvocine green algae. Communications Biology, 1(1), 1–7. [CrossRef]
- Hanschen, E. R., Davison, D. R., Grochau-Wright, Z. I., & Michod, R. E. (2017). Evolution of individuality: a case study in the volvocine green algae. Philosophy, Theory, and Practice in Biology, 9(January), 3. [CrossRef]
- Hanschen, E. R., Ferris, P. J., & Michod, R. E. (2014). Early Evolution of the Genetic Basis for Soma in the Volvocaceae. Evolution, 68(7), 2014–2025.
- Hanschen, E. R., Marriage, T. N., Ferris, P. J., Hamaji, T., Toyoda, A., Fujiyama, A., Neme, R., Noguchi, H., Minakuchi, Y., Suzuki, M., Kawai-Toyooka, H., Smith, D. R., Luria, V., Karger, A., Kirschner, M. W., Sparks, H., Anderson, J., Bakaric, R., Durand, P. M., … Olson, B. J. S. C. (2016). The Gonium pectorale genome demonstrates co-option of cell cycle regulation during the evolution of multicellularity. Nature Communications, 7, 11370. [CrossRef]
- Harryman, A. (2012). Investigating the roles of regA and related genes in the evolution of multicellularity in the volvocine green algae [Doctor of Philosophy]. University of Maryland, Baltimore County.
- Herron, M. D., Borin, J. M., Boswell, J. C., Walker, J., Chen, I. K., Knox, C. A., Boyd, M., Rosenzweig, F., & Ratcliff, W. C. (2019). De novo origins of multicellularity in response to predation. Scientific Reports, 1–9. [CrossRef]
- Herron, M. D., Conlin, P. L., & Ratcliff, W. C. (2022). The Evolution of Multicellularity. CRC Press. [CrossRef]
- Herron, M. D., Hackett, J. D., Aylward, F. O., & Michod, R. E. (2009). Triassic origin and early radiation of multicellular volvocine algae. Proceedings of the National Academy of Sciences of the United States of America, 106(9), 3254–3258. [CrossRef]
- Herron, M. D., & Michod, R. E. (2008). Evolution of complexity in the volvocine algae: transitions in individuality through Darwin’s eye. Evolution; International Journal of Organic Evolution, 62(2), 436–451. [CrossRef]
- Hood, W. R., Zhang, Y., Mowry, A. V, Hyatt, H. W., & Kavazis, A. N. (2018). Life History Trade-offs within the Context of Mitochondrial Hormesis. Integrative and Comparative Biology, 58(3), 567–577. [CrossRef]
- Hughes, K. A., & Leips, J. (2017). Pleiotropy, constraint, and modularity in the evolution of life histories: insights from genomic analyses. Annals of the New York Academy of Sciences, 1389(1), 76–91. [CrossRef]
- Huskey, R. J., & Griffin, B. E. (1979). Genetic Control of Somatic Cell Differentiation in Volvox. Developmental Biology, 72, 226–235.
- Huskey, R. J., Griffin, B. E., Cecil, P. O., & Callahan, A. M. (1979). A Preliminary Genetic Investigation of Volvox carteri. Genetics, 91(2), 229–244. [CrossRef]
- Jimenez Marin, L. B. (2023). Gene loss, co-option and the evolution of developmental complexity in the volvocine algae [PhD Dissertation]. Kansas State University.
- Keeling, P. J., & Burki, F. (2019). Progress towards the Tree of Eukaryotes. Current Biology, 29(16), R808–R817. [CrossRef]
- Keeling, P. J., & Palmer, J. D. (2008). Horizontal gene transfer in eukaryotic evolution. Nature Reviews Genetics, 9(8), 605–618. [CrossRef]
- Kirk, D. L. (1998). Volvox: Molecular-Genetic Origins of Multicellularity and Cellular Differentiation. Cambridge University Press.
- Kirk, D. L. (2001). Germ-soma differentiation in Volvox. Developmental Biology, 238(2), 213–223. [CrossRef]
- Kirk, D. L. (2005). A twelve-step program for evolving multicellularity and a division of labor. BioEssays: News and Reviews in Molecular, Cellular and Developmental Biology, 27(3), 299–310. [CrossRef]
- Kirk, M. M., Ransick, A., McRae, S. E., & Kirk, D. L. (1993). The Relationship between Cell Size and Cell Fate in Volvox carteri. The Journal of Cell Biology, 123(1), 191–208.
- Kirk, M. M., Stark, K., Miller, S. M., Müller, W., Taillon, B. E., Gruber, H., Schmitt, R., & Kirk, D. L. (1999). regA, a Volvox gene that plays a central role in germ-soma differentiation, encodes a novel regulatory protein. Development, 126(4), 639–647. http://www.ncbi.nlm.nih.gov/pubmed/9895312.
- Klein, B., Wibberg, D., & Hallmann, A. (2017). Whole transcriptome RNA-Seq analysis reveals extensive cell type-specific compartmentalization in Volvox carteri. BMC Biology, 15(1), 1–22. [CrossRef]
- Knoll, A. H. (2011). The Multiple Origins of Complex Multicellularity. Http://Dx.Doi.Org/10.1146/Annurev.Earth.031208.100209, 39, 217–239. [CrossRef]
- Koch, R. E., Buchanan, K. L., Casagrande, S., Crino, O., Dowling, D. K., Hill, G. E., Hood, W. R., McKenzie, M., Mariette, M. M., Noble, D. W. A., Pavlova, A., Seebacher, F., Sunnucks, P., Udino, E., White, C. R., Salin, K., & Stier, A. (2021). Integrating Mitochondrial Aerobic Metabolism into Ecology and Evolution. Trends in Ecology and Evolution, 36(4), 321–332. [CrossRef]
- König, S. G., & Nedelcu, A. M. (2020). The genetic basis for the evolution of soma: Mechanistic evidence for the co-option of a stress-induced gene into a developmental master regulator. Proceedings of the Royal Society B: Biological Sciences, 287(1940). [CrossRef]
- Koshikawa, S., Giorgianni, M. W., Vaccaro, K., Kassner, V. A., Yoder, J. H., Werner, T., & Carroll, S. B. (2015). Gain of cis-regulatory activities underlies novel domains of wingless gene expression in Drosophila. Proceedings of the National Academy of Sciences of the United States of America, 112(24), 7524–7529. [CrossRef]
- Koufopanou, V. (1994). The Evolution of Soma in the Volvocales. The American Naturalist, 143(5), 907–931.
- Kulkarni, M., Shakes, D. C., Guevel, K., & Smith, H. E. (2012). SPE-44 Implements Sperm Cell Fate. PLoS Genetics, 8(4), e1002678. [CrossRef]
- Lang, G. I., Murray, A. W., & Botstein, D. (2009). The cost of gene expression underlies a fitness trade-off in yeast. Proceedings of the National Academy of Sciences, 106(14), 5755–5760. [CrossRef]
- Lee, C., Raffaghello, L., Brandhorst, S., Safdie, F. M., Bianchi, G., Martin-Montalvo, A., Pistoia, V., Wei, M., Hwang, S., Merlino, A., Emionite, L., de Cabo, R., & Longo, V. D. (2012). Fasting Cycles Retard Growth of Tumors and Sensitize a Range of Cancer Cell Types to Chemotherapy. Science Translational Medicine, 4(124), 124ra27-124ra27. [CrossRef]
- Leroi, A. M. (2001). Molecular signals versus the Loi de Balancement. In Trends in Ecology and Evolution (Vol. 16, Issue 1, pp. 24–29). Elsevier Ltd. [CrossRef]
- Lindsey, C. R., Rosenzweig, F., & Herron, M. D. (2021). Phylotranscriptomics points to multiple independent origins of multicellularity and cellular differentiation in the volvocine algae. BMC Biology, 19(1). [CrossRef]
- Matt, G. Y., & Umen, J. G. (2018). Cell-Type Transcriptomes of the Multicellular Green Alga Volvox carteri Yield Insights into the Evolutionary Origins of Germ and Somatic Differentiation Programs. G3: Genes|Genomes|Genetics, 8, 531–550. [CrossRef]
- Maynard Smith, J., & Szathmáry, E. (1995). The Major Transitions in Evolution. Oxford University Press.
- Meissner, M., Stark, K., Cresnar, B., Kirk, D. L., & Schmitt, R. (1999). Volvox germline-specific genes that are putative targets of RegA repression encode chloroplast proteins. Current Genetics, 36(6), 363–370. http://www.ncbi.nlm.nih.gov/pubmed/10654090.
- Michod, R. E. (1999). Darwinian Dynamics. Princeton University Press.
- Michod, R. E. (2006a). The group covariance effect and fitness trade-offs during evolutionary transitions in individuality. PNAS, 103(24), 9113–9117.
- Michod, R. E. (2006b). On the transfer of fitness from the cell to the multicellular organism. Biology & Philosophy, 20(5), 967–987. [CrossRef]
- Michod, R. E., & Nedelcu, A. M. (2003). On the reorganization of fitness during evolutionary transitions in individuality. Integrative and Comparative Biology, 43(1), 64–73. [CrossRef]
- Monaghan, P., Metcalfe, N. B., & Torres, R. (2009). Oxidative stress as a mediator of life history trade-offs: Mechanisms, measurements and interpretation. In Ecology Letters (Vol. 12, Issue 1, pp. 75–92). Ecol Lett. [CrossRef]
- Nakagawa, T. T. K. U. T. N. Y. (2008). GMEB1, a novel endogenous caspase inhibitor, prevents hypoxia- and oxidative stress-induced neuronal apoptosis. Neuroscience Letters, 438, 34–37. [CrossRef]
- Nedelcu, A. M. (2006). Evidence for p53-like-mediated stress responses in green algae. FEBS Letters, 580(13), 3013–3017. [CrossRef]
- Nedelcu, A. M. (2009). Environmentally induced responses co-opted for reproductive altruism. Biology Letters, 5(6), 805–808. [CrossRef]
- Nedelcu, A. M. (2019). Independent evolution of complex development in animals and plants: deep homology and lateral gene transfer. Development Genes and Evolution, 25–34. [CrossRef]
- Nedelcu, A. M., & Michod, R. E. (2003). Evolvability, modularity, and individuality during the transition to multicellularity in volvocalean green algae. In G. Schlosser & G. P. Wagner (Eds.), Modularity in development and evolution. University of Chicago Press.
- Nedelcu, A. M., & Michod, R. E. (2006). The evolutionary origin of an altruistic gene. Molecular Biology and Evolution, 23(8), 1460–1464. [CrossRef]
- Nedelcu, A. M., & Michod, R. E. (2020). Stress Responses Co-Opted for Specialized Cell Types During the Early Evolution of Multicellularity. BioEssays, 42(5), 2000029. [CrossRef]
- Nozaki, H., Itoh, M., Watanabe, M. M., & Kuroiwa, T. (1996). Ultrastructure of the vegetative colonies and systematic position of basichlamys (Volvocales, Chlorophyta). European Journal of Phycology, 31(1), 67–72. [CrossRef]
- Nozaki, H., Mahakham, W., Athibai, S., Yamamoto, K., Takusagawa, M., Misumi, O., Herron, M. D., Rosenzweig, F., & Kawachi, M. (2017). Rediscovery of the species of ‘ancestral Volvox’’: morphology and phylogenetic position of Pleodorina sphaerica (Volvocales, Chlorophyceae) from Thailand.’ Phycologia, 56(4), 469–475. [CrossRef]
- Nozaki, H., Misawa, K., Kajita, T., Kato, M., Nohara, S., & Watanabe, M. M. (2000). Origin and evolution of the colonial volvocales (Chlorophyceae) as inferred from multiple, chloroplast gene sequences. Molecular Phylogenetics and Evolution, 17(2), 256–268. [CrossRef]
- Olson, B. J. S. C., & Nedelcu, A. M. (2016). Co-option during the evolution of multicellularity and developmental complexity in the volvocine green algae. Current Opinion in Genetics and Development, 39, 107–115. [CrossRef]
- Ortega Escalante, J. A. (2018). Investigation of Volvox carteri Cell Differentiation and Its Evolution Through Functional Analysis of regA and regA Homologues [PhD Dissertation]. University of Maryland, Baltimore County.
- Pfannschmidt, T., Bräutigam, K., Wagner, R., Dietzel, L., Schröter, Y., Steiner, S., & Nykytenko, A. (2009). Potential regulation of gene expression in photosynthetic cells by redox and energy state: approaches towards better understanding. Annals of Botany, 103(4), 599. [CrossRef]
- Radhakrishnan, K., Bhagya, K. P., Kumar, A. T., Devi, A. N., Sengottaiyan, J., & Kumar, P. G. (2016). Autoimmune Regulator ( AIRE ) Is Expressed in Spermatogenic Cells, and It Altered the Expression of Several Nucleic-Acid-Binding and Cytoskeletal Proteins in Germ Cell 1 Spermatogonial (GC1-spg) Cells. Molecular & Cellular Proteomics, 15(8), 2686–2698. [CrossRef]
- Raffaghello, L., Lee, C., Safdie, F. M., Wei, M., Madia, F., Bianchi, G., & Longo, V. D. (2008). Starvation-dependent differential stress resistance protects normal but not cancer cells against high-dose chemotherapy. Proceedings of the National Academy of Sciences, 105(24), 8215–8220. [CrossRef]
- Ransick, A. (1991). Reproductive cell specification during Volvox obversus development. Developmental Biology, 143(1), 185–198. http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WDG-4DKTPB0-J&_user=441497&_coverDate=01/31/1991&_rdoc=1&_fmt=high&_orig=search&_sort=d&_docanchor=&view=c&_acct=C000020998&_version=1&_urlVersion=0.
- Ransick, A. (1993). Specification of reproductive cells in Volvox. In A. Spradling (Ed.), Evolutionary Conservation of Developmental Mechanisms: 50th Symposium of the Society for Developmental Biology, Marquette University, June 20–23, 1991. (pp. 55–70). Wiley-Liss.
- Ratcliff, W. C., Denison, R. F., Borrello, M., & Travisano, M. (2012). Experimental evolution of multicellularity. Proceedings of the National Academy of Sciences, 109(5), 1595–1600. [CrossRef]
- Ratcliff, W. C., Herron, M. D., Howell, K., Pentz, J. T., Rosenzweig, F., & Travisano, M. (2013). Experimental evolution of an alternating uni- and multicellular life cycle in Chlamydomonas reinhardtii. Nature Communications, 4(May), 2742. [CrossRef]
- Roff, D. A., & Fairbairn, D. J. (2007). The evolution of trade-offs: Where are we? Journal of Evolutionary Biology, 20(2), 433–447. [CrossRef]
- Rose, C. J., Hammerschmidt, K., & Rainey, P. B. (2020). Experimental evolution of nascent multicellularity: Recognizing a Darwinian transition in individuality. BioRxiv. [CrossRef]
- Saggere, R. M. S., Lee, C. W. J., Chan, I. C. W., Durnford, D. G., & Nedelcu, A. M. (2022). A life-history trade-off gene with antagonistic pleiotropic effects on reproduction and survival in limiting environments. Proceedings of the Royal Society B, 289(1967). [CrossRef]
- Schaap, P. (2016). Evolution of developmental signalling in Dictyostelid social amoebas. In Current Opinion in Genetics and Development (Vol. 39, pp. 29–34). Elsevier Ltd. [CrossRef]
- Schaller, C. E., Wang, C. L., Beck-engeser, G., Goss, L., Scott, H. S., Anderson, M. S., & Wabl, M. (2008). Expression of Aire and the Early Wave of Apoptosis in Spermatogenesis. The Journal of Immunology, 180(3), 1338–1343. [CrossRef]
- Sessoms, A. H., & Huskey, R. J. (1973). Genetic Control of Development in Volvox: Isolation and Characterization of Morphogenetic Mutants. Proceedings of the National Academy of Sciences, 70(5), 1335–1338.
- Sharpe, S. C., Eme, L., Brown, M. W., & Roger, A. J. (2014). Timing the Origins of Multicellular Eukaryotes Through Phylogenomics and Relaxed Molecular Clock Analyses. In A. M. Nedelcu & I. Ruiz-Trillo (Eds.), Evolutionary Transitions to Multicellular Life (pp. 3–30). Springer.
- Stark, K., Kirk, D. L., & Schmitt, R. (2001). Two enhancers and one silencer located in the introns of regA control somatic cell differentiation in Volvox carteri. Genes & Development, 15, 1449–1460. [CrossRef]
- Starr, R. C. (1970). Control of differentiation in Volvox. Dev. Biol., 4, 59–100.
- Stearns, S. C. (1989). Trade-Offs in Life-History Evolution. Functional Ecology, 3(3), 259–268. [CrossRef]
- Strand, D. D., Livingston, A. K., Satoh-Cruz, M., Koepke, T., Enlow, H. M., Fisher, N., Froehlich, J. E., Cruz, J. A., Minhas, D., Hixson, K. K., Kohzuma, K., Lipton, M., Dhingra, A., & Kramer, D. M. (2017). Defects in the Expression of Chloroplast Proteins Leads to H2O2 Accumulation and Activation of Cyclic Electron Flow around Photosystem I. Frontiers in Plant Science, 7, 2073. [CrossRef]
- Tam, L., & Kirk, D. L. (1991). Identification of Cell-Type-Specific Characterization of their Expression Genes of Volvox carteri and during the Asexual Life Cycle. Developmental Biology, 145, 51–66.
- Umen, J. G. (2020). Volvox and volvocine green algae. EvoDevo, 11(1), 7–15. [CrossRef]
- Veraksa, A., Kennison, J., & McGinnis, W. (2002). DEAF-1 function is essential for the early embryonic development of Drosophila. Genesis, 33(2), 67–76. [CrossRef]
- Villellas, J., & García, M. B. (2018). Life-history trade-offs vary with resource availability across the geographic range of a widespread plant. Plant Biology, 20(3), 483–489. [CrossRef]
- Wagner, G. P., Erkenbrack, E. M., & Love, A. C. (2019). Stress-Induced Evolutionary Innovation: A Mechanism for the Origin of Cell Types. BioEssays, 41, 1800188. [CrossRef]
- Wenger, J. W., Piotrowski, J., Nagarajan, S., Chiotti, K., Sherlock, G., & Rosenzweig, F. (2011). Hunger Artists: Yeast Adapted to Carbon Limitation Show Trade-Offs under Carbon Sufficiency. PLOS Genetics, 7(8), e1002202. [CrossRef]
- Wolf, J. B., Howie, J. A., Parkinson, K., Gruenheit, N., Melo, D., Rozen, D., & Thompson, C. R. L. (2015). Fitness Trade-offs Result in the Illusion of Social Success. Current Biology: CB, 25(8), 1086–1090. [CrossRef]
- Wykoff, D. D., Davies, J. P., Melis, A., & Grossman, A. R. (1998). The regulation of photosynthetic electron transport during nutrient deprivation in Chlamydomonas reinhardtii. Plant Physiology, 117(1), 129–139. [CrossRef]
- Yamamoto, K., Hamaji, T., Kawai-Toyooka, H., Matsuzaki, R., Takahashi, F., Nishimura, Y., Kawachi, M., Noguchi, H., Minakuchi, Y., Umen, J. G., Toyoda, A., & Nozaki, H. (2021). Three genomes in the algal genus Volvox reveal the fate of a haploid sex-determining region after a transition to homothallism. PNAS, 118(21), e2100712118. [CrossRef]
- Yamashita, S., Arakaki, Y., Kawai-Toyooka, H., Noga, A., Hirono, M., & Nozaki, H. (2016). Alternative evolution of a spheroidal colony in volvocine algae: developmental analysis of embryogenesis in Astrephomene (Volvocales, Chlorophyta). BMC Evolutionary Biology, 16(1), 243. [CrossRef]
- Yamashita, S., Yamamoto, K., Matsuzaki, R., Suzuki, S., Yamaguchi, H., Hirooka, S., Minakuchi, Y., Miyagishima, S. ya, Kawachi, M., Toyoda, A., & Nozaki, H. (2021). Genome sequencing of the multicellular alga Astrephomene provides insights into convergent evolution of germ-soma differentiation. Scientific Reports, 11(1). [CrossRef]
- Zera, A. J., & Harshman, L. G. (2001). The Physiology of Life History Trade-Offs in Animals. Annual Review of Ecology and Systematics, 32(1), 95–126. [CrossRef]
- Zhang, J. (2003). Evolution by gene duplication: An update. In Trends in Ecology and Evolution (Vol. 18, Issue 6, pp. 292–298). Elsevier Ltd. [CrossRef]


| 1 | Huskey & Griffin, (1979) originally described a second regB locus based on linkage group analysis of regenerator mutants, but reexamination of regB mutants from members of the same research lab determined that regB mutants are not regenerator mutants and have a different mutant phenotype (Baran, 1984). Thus, in retrospect, all regenerator mutants can be mapped to the regA locus (D. L. Kirk, 1998). |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



