Submitted:
09 February 2023
Posted:
13 February 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
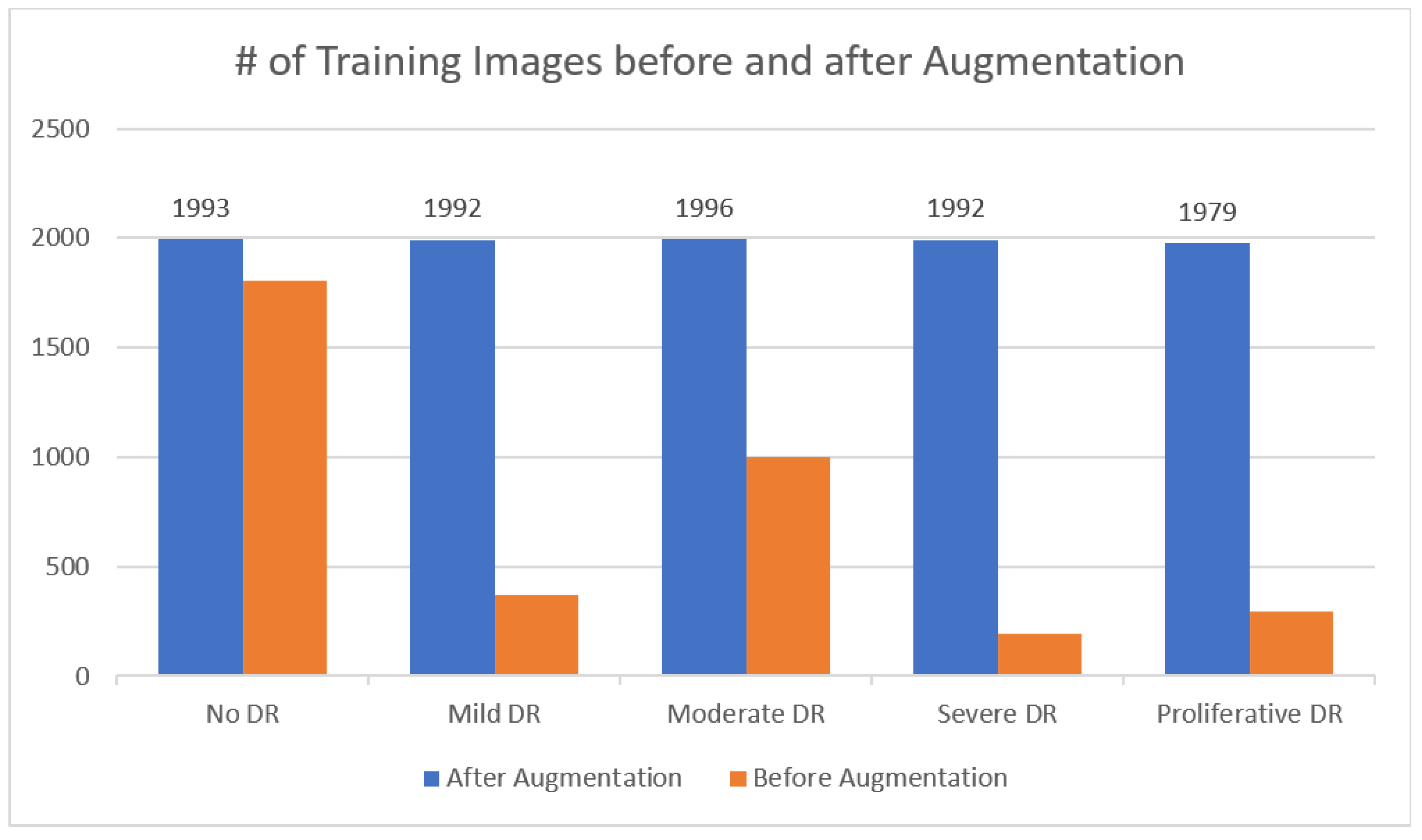
- By utilizing the techniques of augmentation, we ensured that the APTOS dataset contained a constant number of records.
- Accuracy (Acc), Confusion matrix (CM), Specificity, Sensitivity, top n accuracy, and the F1-score (F1c) are the indicators utilized in a comprehensive comparison research to establish the viability of the suggested system.
- Using the DensNet-121 weight-tuning algorithm, pre-trained network is fine-tuned using the APTOS data set.
- By employing a diverse training technique supported by multiple permutations of training strategies, the overall dependability of the proposed method is improved, and overfitting is avoided (e.g., data augmentation (DA), batch size, validation patience, and learning rate).
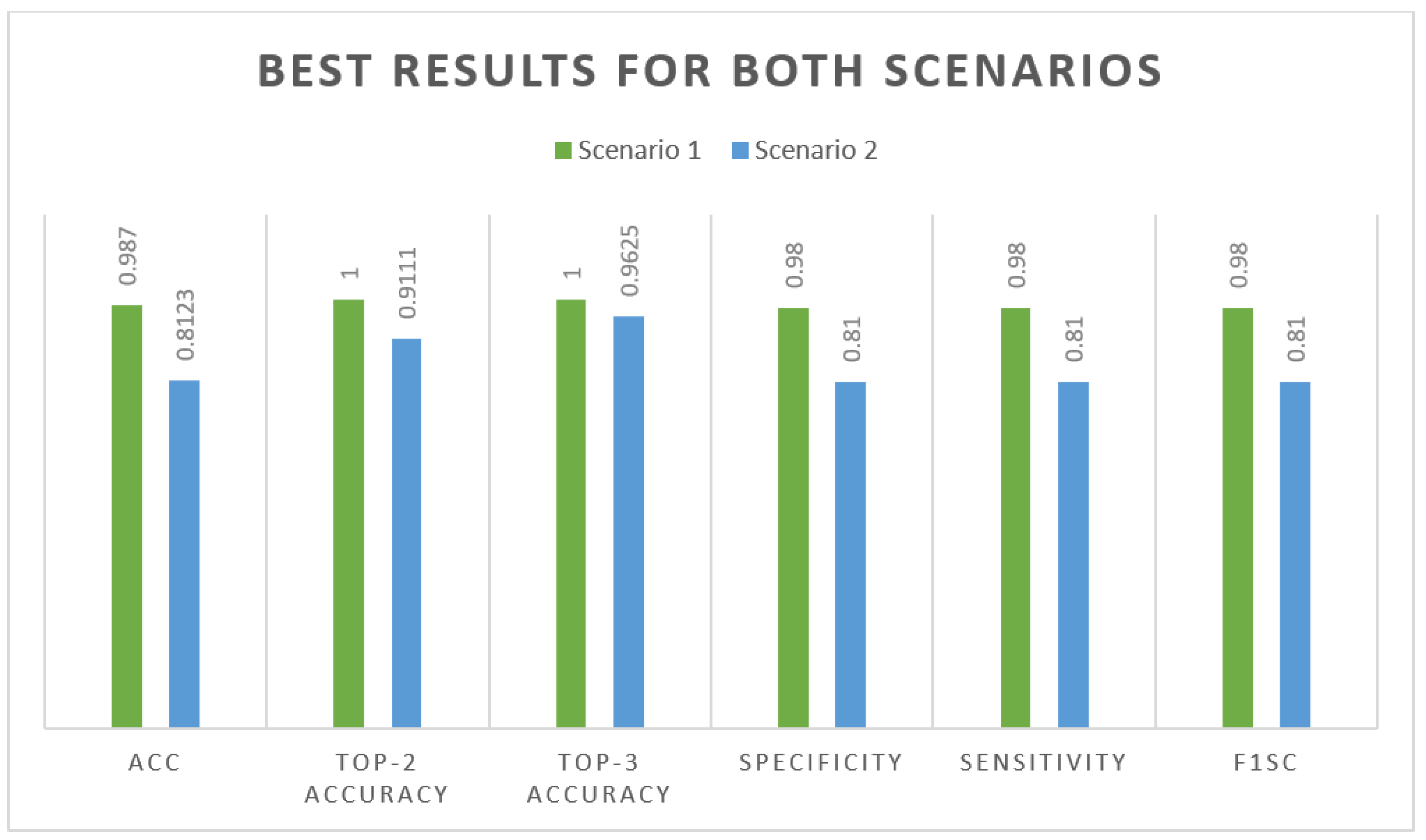
- The APTOS dataset was utilized in both the training and evaluation phases of model construction. By utilizing severe 80:20 hold-out validation, the model obtained a stunning 98.7% accuracy of classification with enhancement approaches and 81.23% without enhancement techniques.
2. Related Work
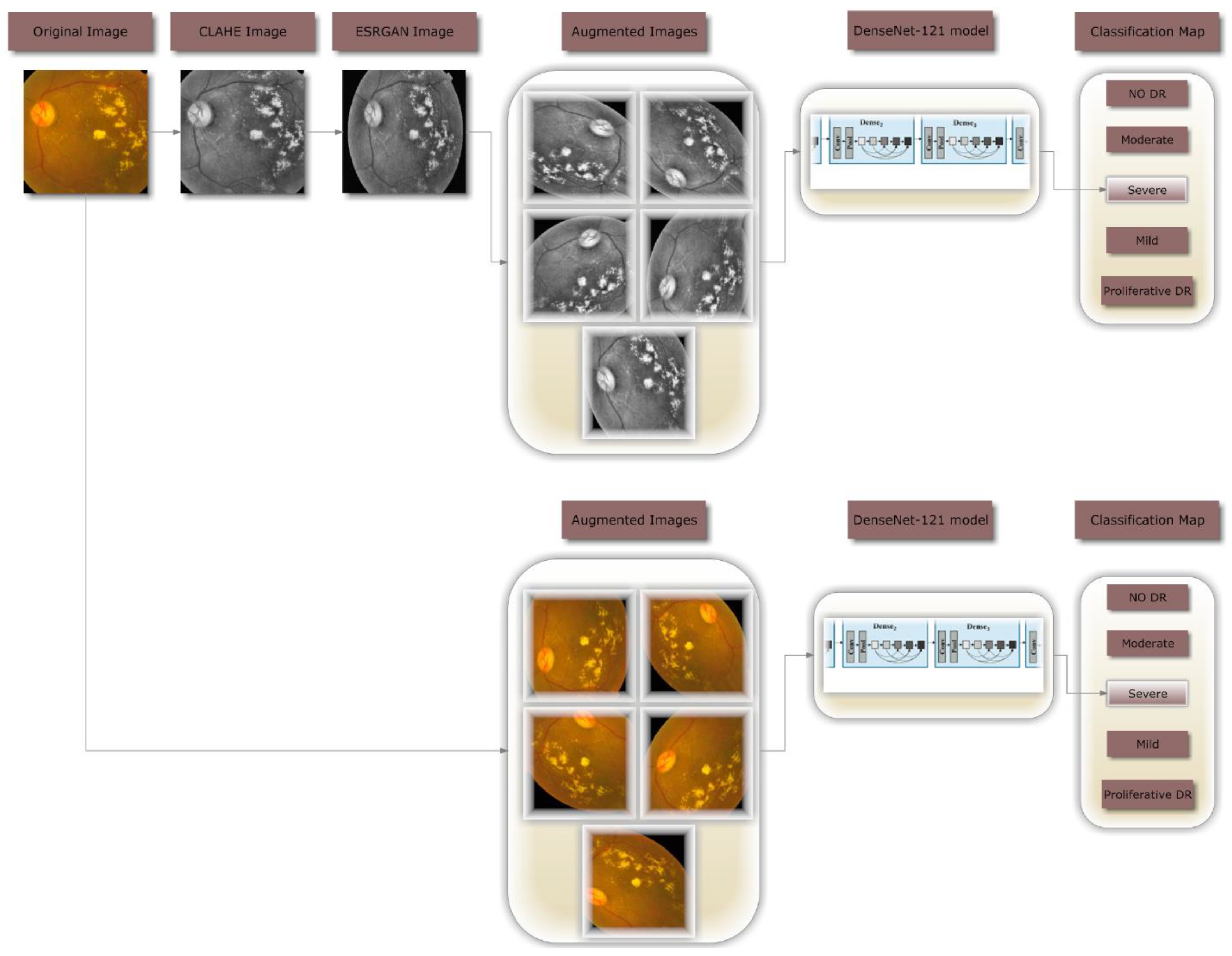
3. Research Methodology
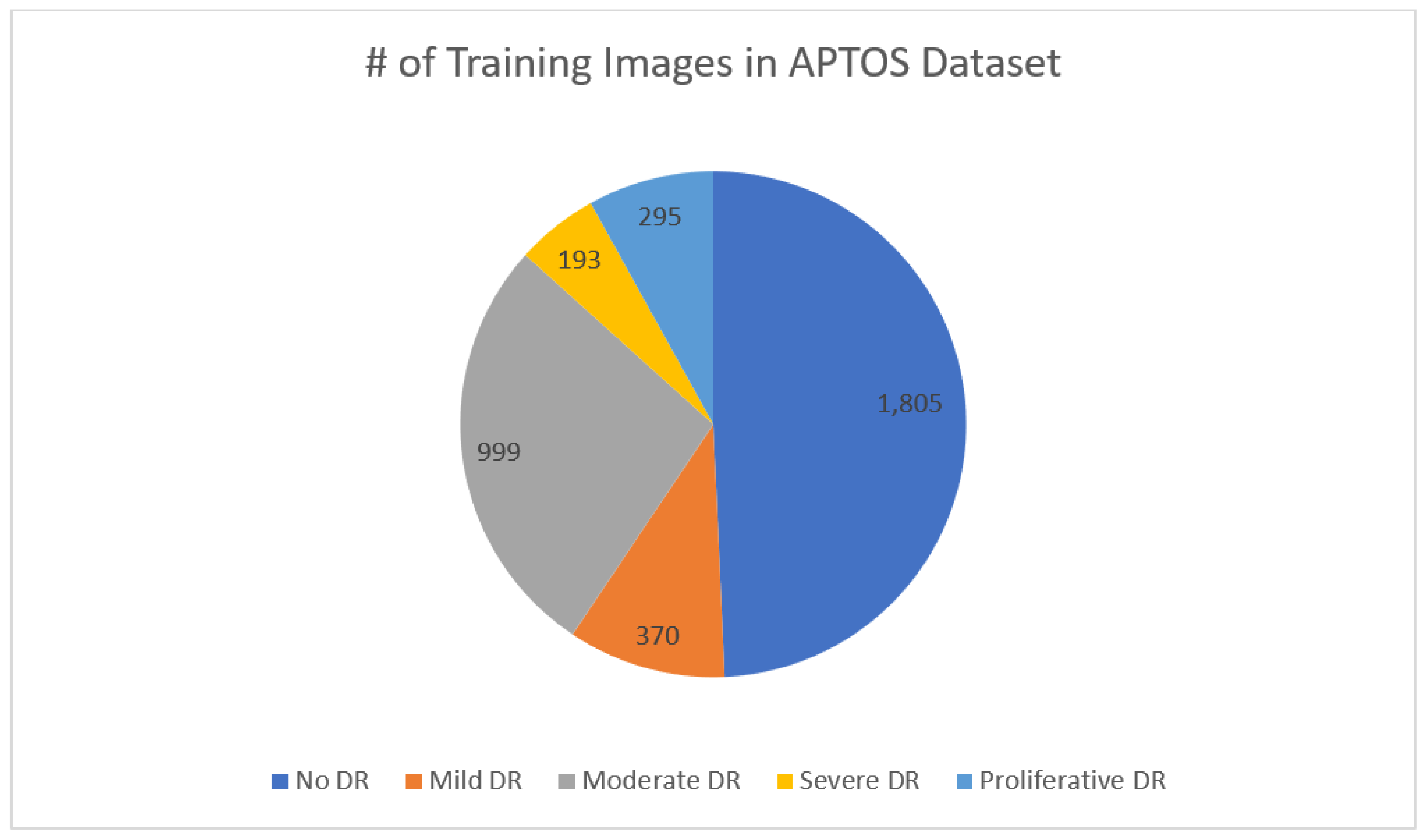
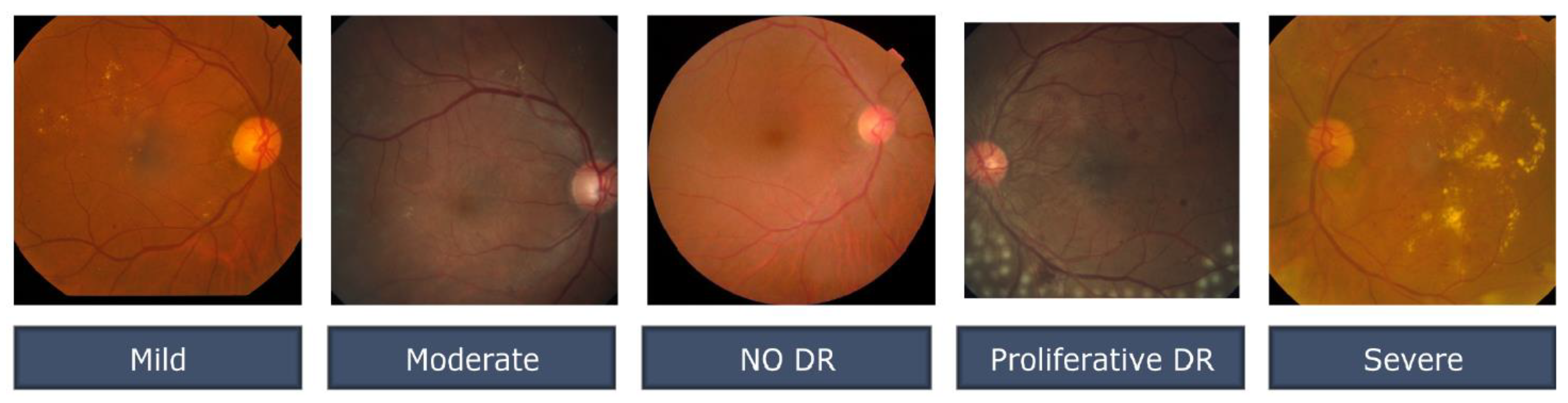
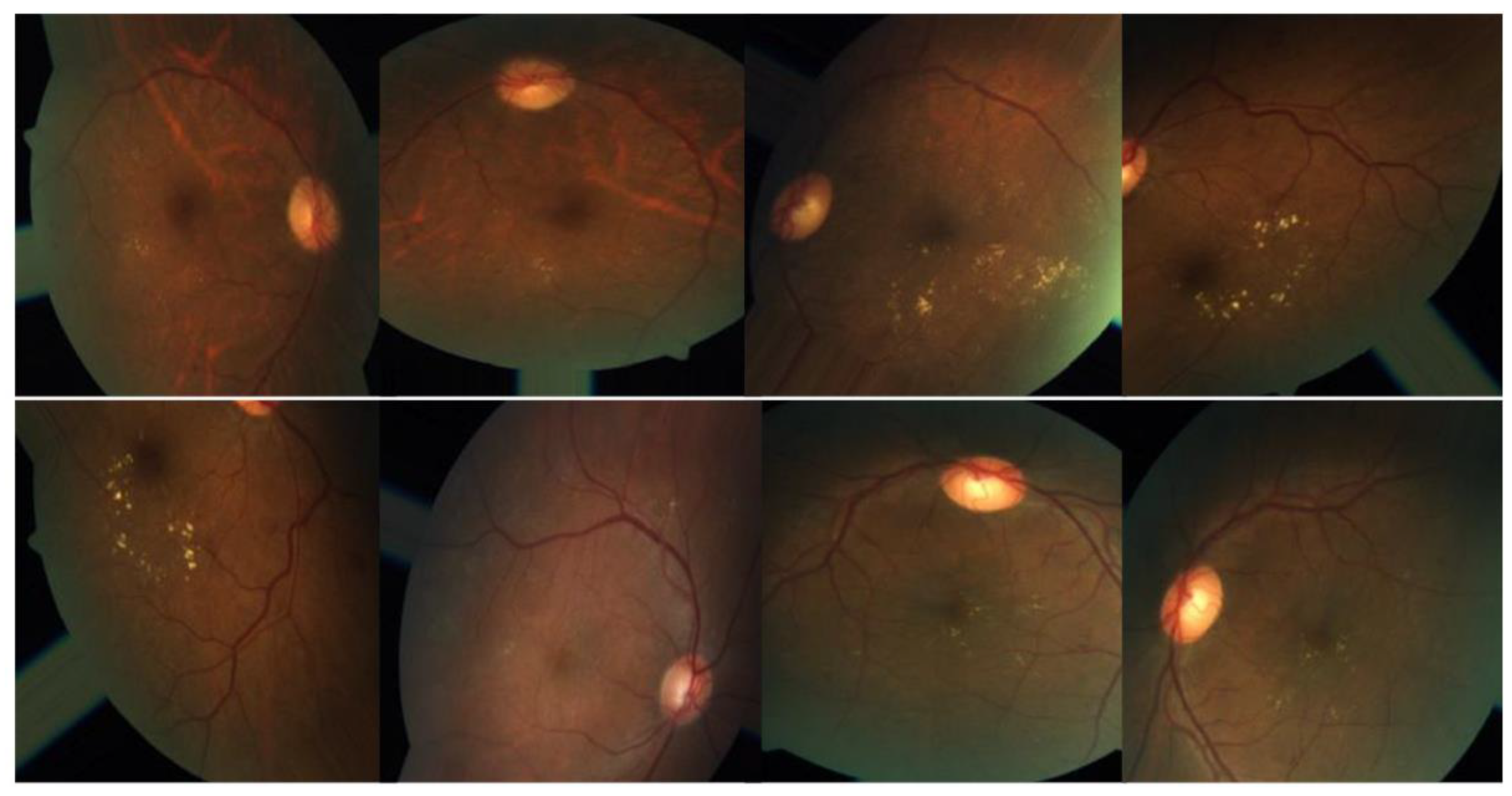
3.1. Data set Description.
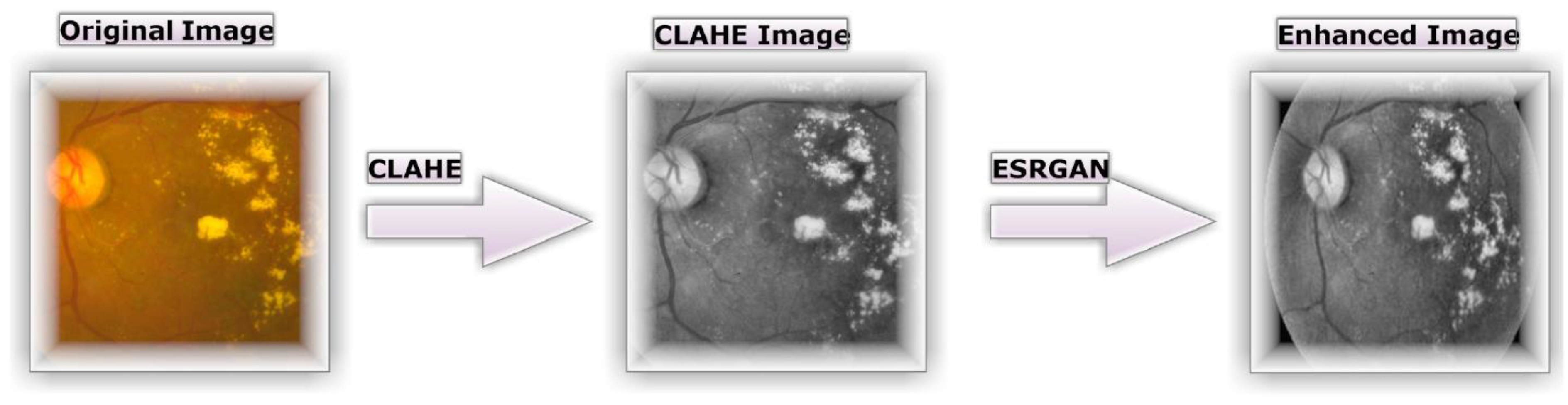
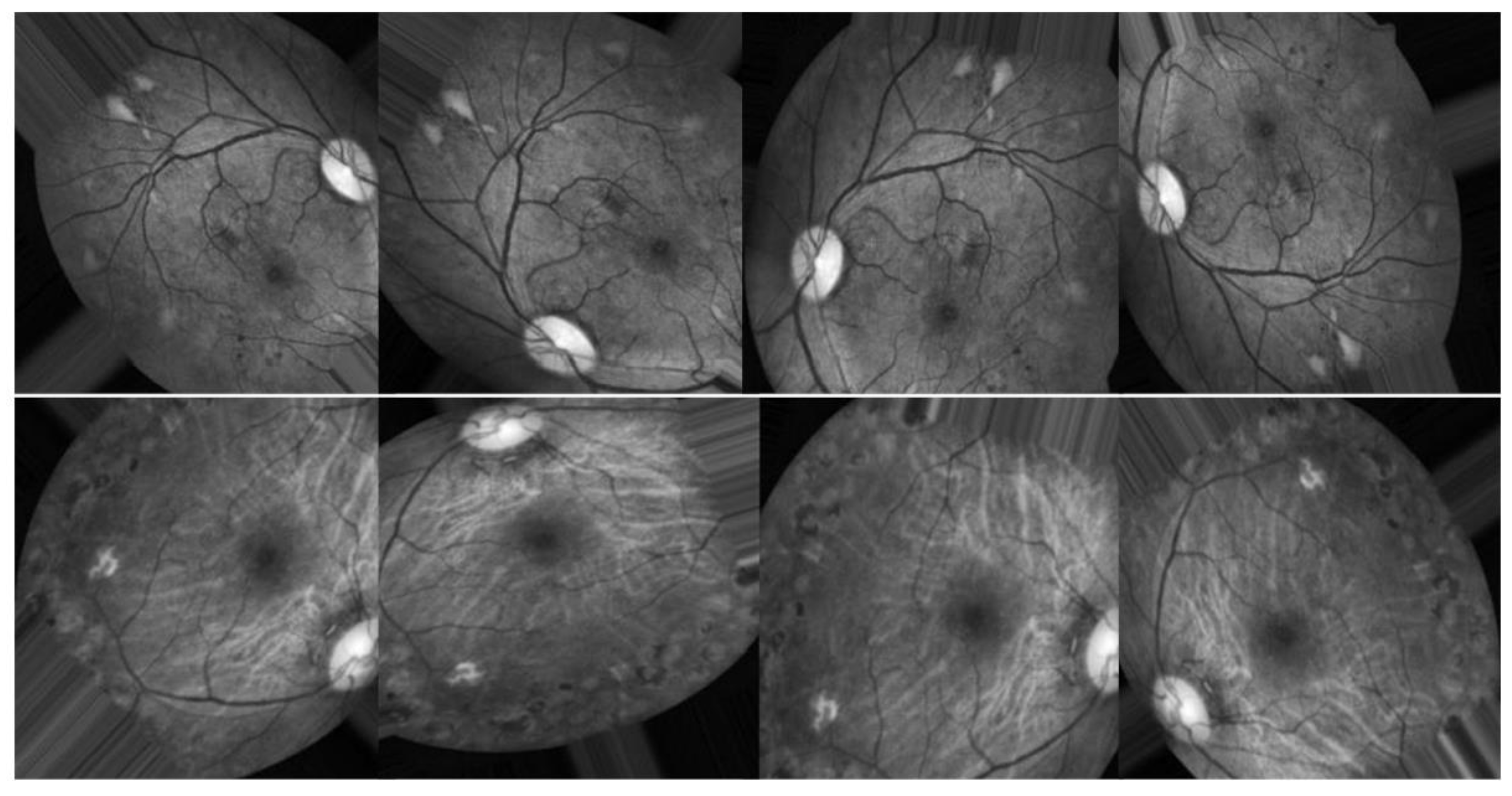
3.2. Preprocessing using CLAHE and ESRGAN
3.3. Data Augmentation
4. Experimental Results
4.1. Instruction and Setup of DenseNet-121
4.2. Performance appraisal
4.3. Performance of DenseNet-121 Model Outcomes:
4.4. Evaluation Considering a Variety of Other Methodologies
4.5. Discussion
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Association, A.D. Diagnosis and classification of diabetes mellitus. Diabetes Care 2014, 37 (Suppl. 1), S81–S90. [Google Scholar] [CrossRef] [PubMed]
- Al-Antary, M.T.; Arafa, Y. Multi-scale attention network for diabetic retinopathy classification. IEEE Access 2021, 9, 54190–54200. [Google Scholar] [CrossRef]
- Humayun, M.; Alsayat, A. Prediction Model for Coronavirus Pandemic Using Deep Learning. Comput. Syst. Sci. Eng. 2022, 40, 947–961. [Google Scholar] [CrossRef]
- Alex, S.A.; Jhanjhi, N.Z.; Humayun, M.; Ibrahim, A.O.; Abulfaraj, A.W. Deep LSTM Model for Diabetes Prediction with Class Balancing by SMOTE. Electronics 2022, 11, 2737. [Google Scholar] [CrossRef]
- Alwakid, G.; Gouda, W.; Humayun, M. Deep Learning-based prediction of Diabetic Retinopathy using CLAHE and ESRGAN for Enhancement. Preprints 2023, 2023020097. [Google Scholar] [CrossRef]
- Pandey, S.K.; Sharma, V. World diabetes day 2018: battling the emerging epidemic of diabetic retinopathy. Indian J. Ophthalmol. 2018, 66, 1652. [Google Scholar] [CrossRef] [PubMed]
- Kobat, S.G.; et al. Automated Diabetic Retinopathy Detection Using Horizontal and Vertical Patch Division-Based Pre-Trained DenseNET with Digital Fundus Images. Diagnostics 2022, 12, 1975. [Google Scholar] [CrossRef]
- Singh, A.; et al. Mechanistic Insight into Oxidative Stress-Triggered Signaling Pathways and Type 2 Diabetes. Molecules 2022, 27, 950. [Google Scholar] [CrossRef]
- Wykoff, C.C.; et al. Risk of blindness among patients with diabetes and newly diagnosed diabetic retinopathy. Diabetes Care 2021, 44, 748–756. [Google Scholar] [CrossRef]
- Alyoubi, W.L.; Shalash, W.M.; Abulkhair, M.F. Diabetic retinopathy detection through deep learning techniques: A review. Inform. Med. Unlocked 2020, 20, 100377. [Google Scholar] [CrossRef]
- Humayun, M.; Khalil, M.I.; Almuayqil, S.N.; Jhanjhi, N.Z. Framework for Detecting Breast Cancer Risk Presence Using Deep Learning. Electronics 2023, 12, 403. [Google Scholar] [CrossRef]
- Gouda, W.; et al. Detection of COVID-19 Based on Chest X-rays Using Deep Learning. Healthcare 2022, 10, 343. [Google Scholar] [CrossRef]
- Alruwaili, M.; Gouda, W. Automated Breast Cancer Detection Models Based on Transfer Learning. Sensors 2022, 22, 876. [Google Scholar] [CrossRef]
- APTOS 2019 Blindness Detection; Kaggle: San Francisco, CA, USA, 2019.
- Huang, G.; et al. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Reza, A.M. Realization of the contrast limited adaptive histogram equalization (CLAHE) for real-time image enhancement. J. VLSI Signal Process. Syst. Signal Image Video Technol. 2004, 38, 35–44. [Google Scholar] [CrossRef]
- Ledig, C.; et al. Photo-realistic single image super-resolution using a generative adversarial network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Gargeya, R.; Leng, T. Automated identification of diabetic retinopathy using deep learning. Ophthalmology, 2017, 124, 962–969. [Google Scholar] [CrossRef]
- Costa, P.; et al. A weakly-supervised framework for interpretable diabetic retinopathy detection on retinal images. IEEE Access 2018, 6, 18747–18758. [Google Scholar] [CrossRef]
- Wang, J.; Bai, Y.; Xia, B. Feasibility of diagnosing both severity and features of diabetic retinopathy in fundus photography. IEEE Access 2019, 7, 102589–102597. [Google Scholar] [CrossRef]
- Leeza, M.; Farooq, H. Detection of severity level of diabetic retinopathy using Bag of features model. IET Comput. Vis. 2019, 13, 523–530. [Google Scholar] [CrossRef]
- Gayathri, S.; et al. Automated binary and multiclass classification of diabetic retinopathy using haralick and multiresolution features. IEEE Access 2020, 8, 57497–57504. [Google Scholar] [CrossRef]
- Pires, R.; et al. A data-driven approach to referable diabetic retinopathy detection. Artif. Intell. Med. 2019, 96, 93–106. [Google Scholar] [CrossRef]
- Zhang, W.; et al. Automated identification and grading system of diabetic retinopathy using deep neural networks. Knowl.-Based Syst. 2019, 175, 12–25. [Google Scholar] [CrossRef]
- Math, L.; Fatima, R. Adaptive machine learning classification for diabetic retinopathy. Multimed. Tools Appl. 2021, 80, 5173–5186. [Google Scholar] [CrossRef]
- Maqsood, S.; Damaševičius, R.; Maskeliūnas, R. Hemorrhage detection based on 3D CNN deep learning framework and feature fusion for evaluating retinal abnormality in diabetic patients. Sensors 2021, 21, 3865. [Google Scholar] [CrossRef] [PubMed]
- Majumder, S.; Ullah, M.A. Feature extraction from dermoscopy images for melanoma diagnosis. SN Appl. Sci. 2019, 1, 1–11. [Google Scholar] [CrossRef]
- Majumder, S.; Ullah, M.A. A computational approach to pertinent feature extraction for diagnosis of melanoma skin lesion. Pattern Recognit. Image Anal. 2019, 29, 503–514. [Google Scholar] [CrossRef]
- Crane, A.; Dastjerdi, M. Effect of Simulated Cataract on the Accuracy of an Artificial Intelligence Algorithm in Detecting Diabetic Retinopathy in Color Fundus Photos. Investig. Ophthalmol. Vis. Sci. 2022, 63, 2100–F0089. [Google Scholar]
- Majumder, S.; Kehtarnavaz, N. Multitasking deep learning model for detection of five stages of diabetic retinopathy. IEEE Access 2021, 9, 123220–123230. [Google Scholar] [CrossRef]
- Yadav, S.; Awasthi, P.; Pathak, S. Retina Image and Diabetic Retinopathy: A Deep Learning Based Approach. Int. Res. J. Mod. Eng. Technol. Sci. 2022, 4, 3790–3794. [Google Scholar]
- Majumder, S.; Ullah, M.A.; Dhar, J.P. Melanoma diagnosis from dermoscopy images using artificial neural network. In Proceedings of the 2019 5th International Conference on Advances in Electrical Engineering (ICAEE), Dhaka, Bangladesh, 26–28 September 2019; IEEE: New York, NY, USA. [Google Scholar]
- Jolicoeur-Martineau, A. The relativistic discriminator: a key element missing from standard GAN. arXiv 2018, arXiv:1807.00734. [Google Scholar]
- Maqsood, Z.; Gupta, M.K. Automatic Detection of Diabetic Retinopathy on the Edge. In Cyber Security, Privacy and Networking; Springer: Singapore, 2022; pp. 129–139. [Google Scholar]
- Saranya, P.; et al. Red Lesion Detection in Color Fundus Images for Diabetic Retinopathy Detection. In Proceedings of the International Conference on Deep Learning, Computing and Intelligence; Springer: Singapore, 2022. [Google Scholar]
- Lahmar, C.; Idri, A. Deep hybrid architectures for diabetic retinopathy classification. Comput. Methods Biomech. Biomed. Eng. Imaging Vis. 2022, 11, 1–19. [Google Scholar] [CrossRef]
- Oulhadj, M.; et al. Diabetic retinopathy prediction based on deep learning and deformable registration. Multimed. Tools Appl. 2022, 81, 1–19. [Google Scholar] [CrossRef]
- Gangwar, A.K.; Ravi, V. Diabetic retinopathy detection using transfer learning and deep learning. In Evolution in Computational Intelligence; Springer: Singapore, 2021; pp. 679–689. [Google Scholar]
- Lahmar, C.; Idri, A. On the value of deep learning for diagnosing diabetic retinopathy. Health Technol. 2022, 12, 89–105. [Google Scholar] [CrossRef]
- Canayaz, M. Classification of diabetic retinopathy with feature selection over deep features using nature-inspired wrapper methods. Appl. Soft Comput. 2022, 128, 109462. [Google Scholar] [CrossRef]
- Escorcia-Gutierrez, J.; et al. Analysis of Pre-trained Convolutional Neural Network Models in Diabetic Retinopathy Detection through Retinal Fundus Images. In International Conference on Computer Information Systems and Industrial Management; Springer: Cham, Switzerland, 2022. [Google Scholar]
- Thomas, N.M.; Albert Jerome, S. Grading and Classification of Retinal Images for Detecting Diabetic Retinopathy Using Convolutional Neural Network. In Advances in Electrical and Computer Technologies; Springer: Singapore, 2022; pp. 607–614. [Google Scholar]
- Salluri, D.K.; Sistla, V.; Kolli, V.K.K. HRUNET: Hybrid Residual U-Net for automatic severity prediction of Diabetic Retinopathy. Comput. Methods Biomech. Biomed. Eng. Imaging Vis. 2022, 11, 1–12. [Google Scholar] [CrossRef]
- Deshpande, A.; Pardhi, J. Automated detection of Diabetic Retinopathy using VGG-16 architecture. Int. Res. J. Eng. Technol. 2021, 8, 2936–2940. [Google Scholar]
- Macsik, P.; et al. Local Binary CNN for Diabetic Retinopathy Classification on Fundus Images. Acta Polytech. Hung. 2022, 19, 27–45. [Google Scholar] [CrossRef]
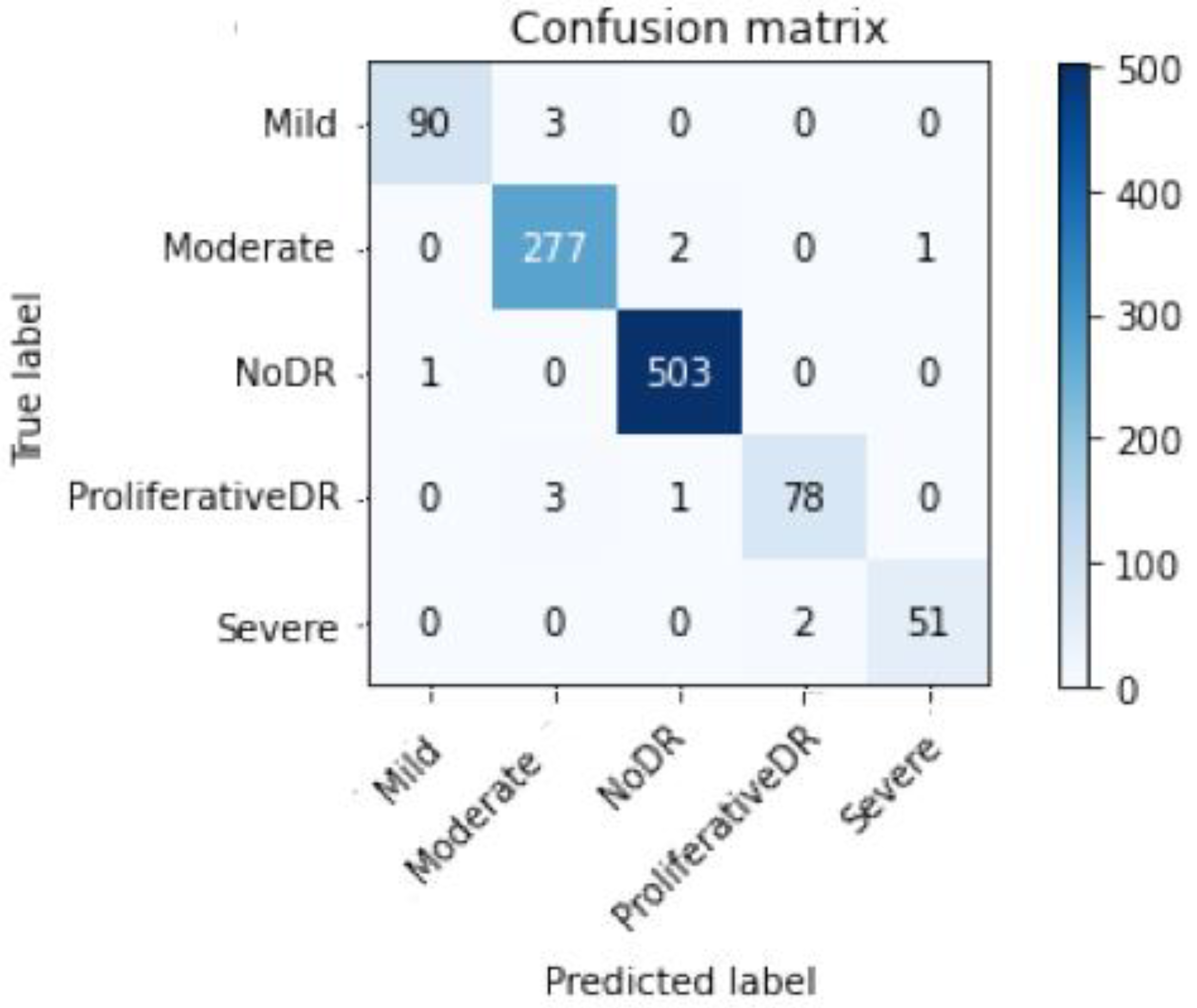
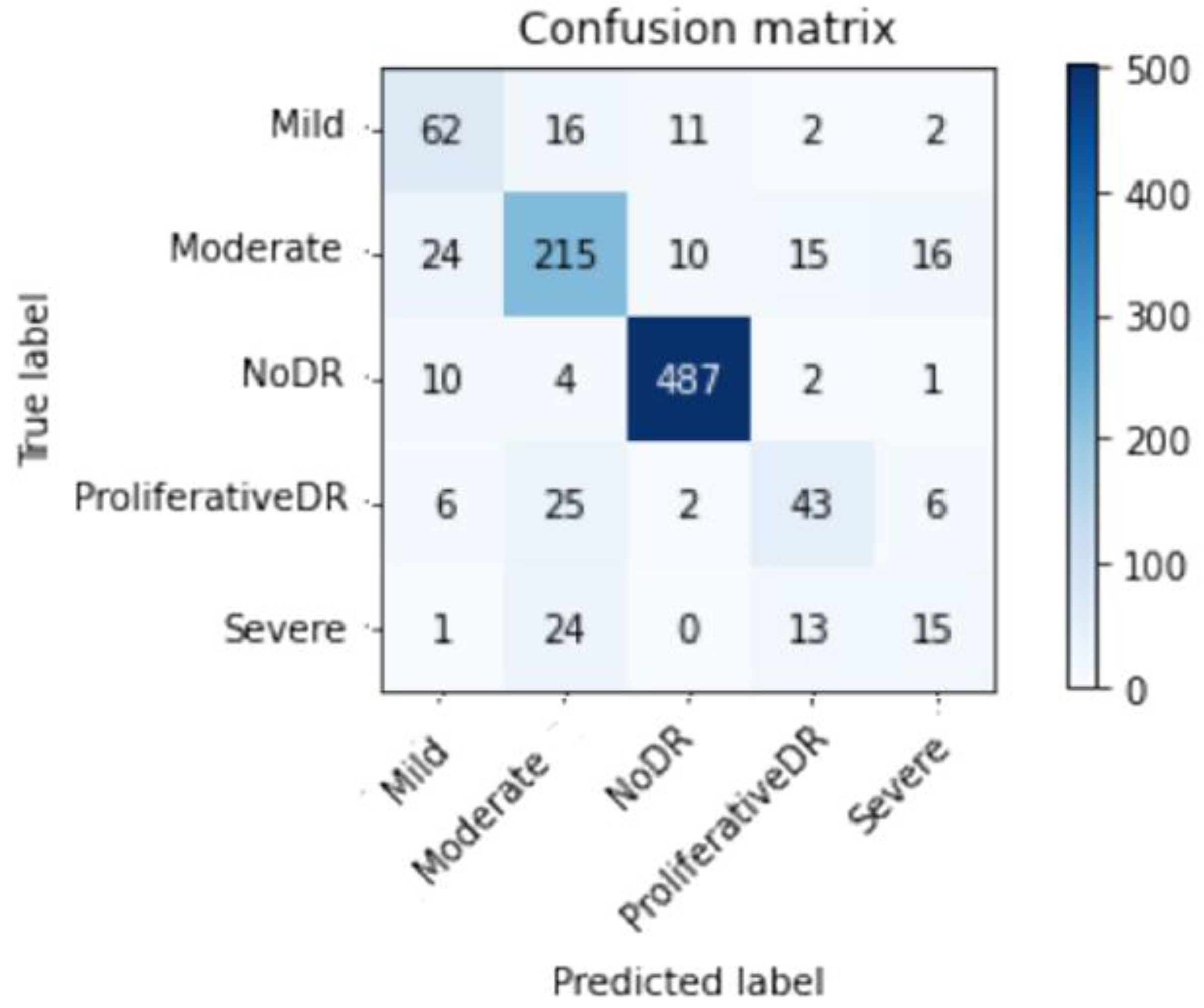
| Top-2 Accuracy | Top-3 Accuracy | Acc | Specificity | Sensitivity | F1sc |
|---|---|---|---|---|---|
| 1.0000 | 1.0000 | 0.987 | 0.98 | 0.98 | 0.98 |
| Top-2 Accuracy | Top-3 Accuracy | Acc | Specificity | Sensitivity | F1sc |
|---|---|---|---|---|---|
| 0.9111 | 0.9625 | 0.8123 | 0.81 | 0.81 | 0.81 |
| Specificity | Sensitivity | F1sc | Total images | |
|---|---|---|---|---|
| Mild DR | 0.95 | 1.00 | 0.97 | 93 |
| Moderate DR | 1.00 | 0.97 | 0.99 | 280 |
| No DR | 1.00 | 0.99 | 0.99 | 504 |
| Proliferative DR | 0.973 | 0.98 | 0.96 | 82 |
| Severe DR | 0.90 | 0.97 | 0.93 | 53 |
| Average | 0.98 | 0.98 | 0.98 | 1012 |
| Specificity | Sensitivity | F1sc | Total images | |
|---|---|---|---|---|
| Mild DR | 0.60 | 0.67 | 0.63 | 93 |
| Moderate DR | 0.76 | 0.77 | 0.76 | 280 |
| No DR | 0.95 | 0.97 | 0.96 | 504 |
| Proliferative DR | 0.57 | 0.52 | 0.55 | 82 |
| Severe DR | 0.38 | 0.28 | 0.32 | 53 |
| Average | 0.81 | 0.81 | 0.81 | 1012 |
| Ref# | Technique | Accuracy |
|---|---|---|
| [34] | EfficientNet-B6 | 86.03% |
| [35] | SVM | 94.5% |
| [36] | SVM classifier and MobileNet_V2 for feature extraction | 88.80% |
| [37] | Densenet-121, Xception, Inception-v3, Resnet-50 | 85.28% |
| [38] | Inception-ResNet-v2 | 72.33% |
| [39] | MobileNet_V2 | 93.09% |
| [40] | EfficientNet and DenseNet | 96.32% |
| [41] | VGG16 | 96.86% |
| [42] | CNN | 85% |
| [43] | Hybrid Residual U-Net | 94% |
| [29] | Inception-ResNet-v2 | 97.0%, |
| [44] | VGG-16 | 74.58% |
| [31] | VGG16 | 73.26% |
| DenseNet121 | 96.11% | |
| [45] | LBCNN | 97.41% |
| [3] | Inception-v3 | 88.1% |
| [7] | DenseNet201 | 93.85% |
| [2] | MSA-Net | 84.6% |
| Proposed Methodology | DenseNet-121 ( without using CLAHE + ESRGAN) Scenario 2 | 81.23% |
| Proposed Methodology | DenseNet-121 (using CLAHE + ESRGAN) Scenario 1 | 98.7% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




