Submitted:
01 February 2023
Posted:
06 February 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
- By employing the technique of augmentation, we ensured that the APTOS dataset contained a consistent amount of data.
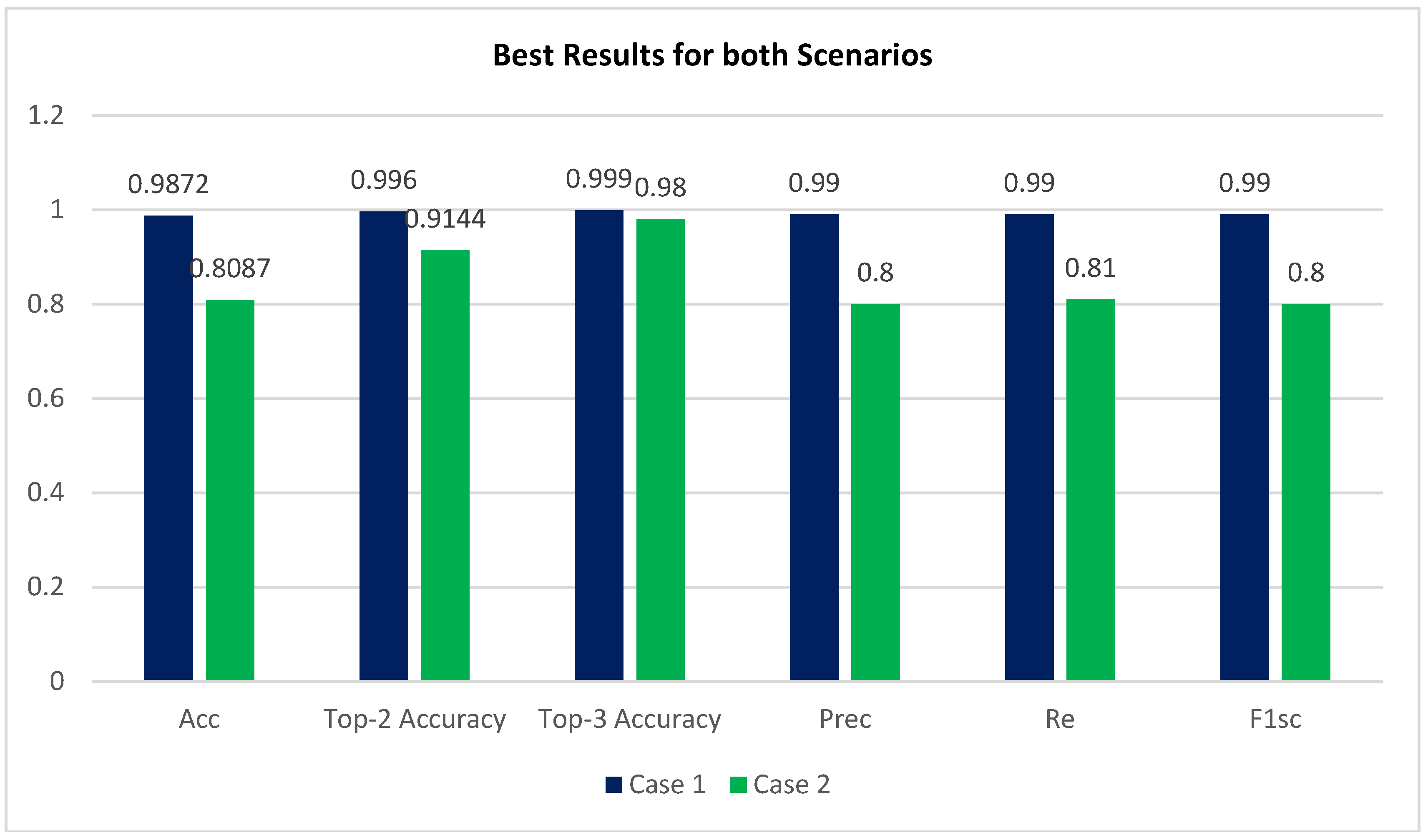
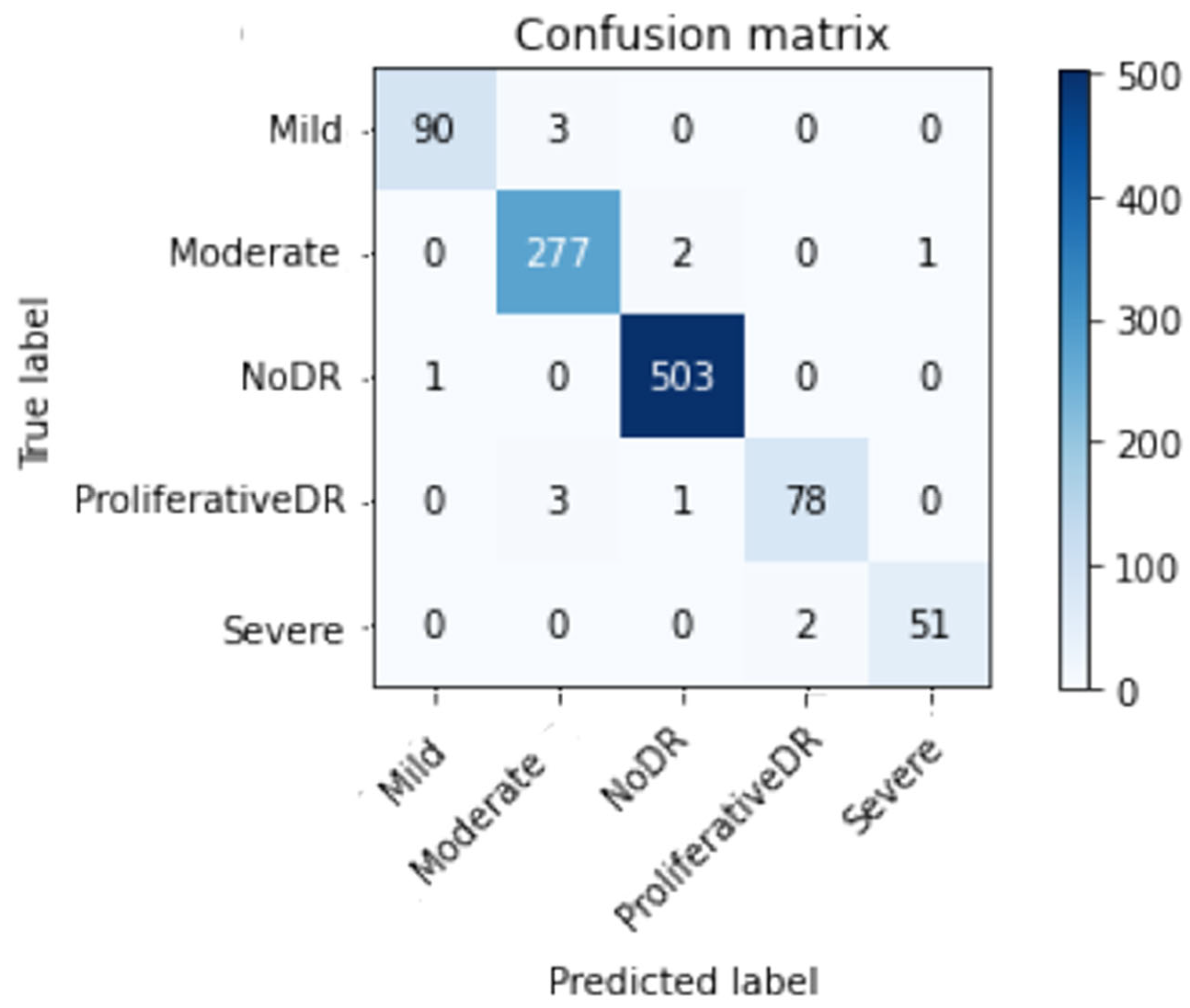
- Accuracy (Acc), Confusion matrix (CM), precision (Prec), recall (Re), top n accuracy, and the F1-score (F1sc) are the indicators used in a comprehensive comparative study to determine the viability of the proposed system.
- Pre-trained networks trained on the APTOS data set are fine-tuned with the use of an Inception-V3 weight-tuning algorithm.
- By adopting a varied training procedure backed by various permutations of training strategies, the general reliability of the suggested method is enhanced, and overfitting is avoided (e.g., learning rate, data augmentation, batch size, and validation patience).
- The APTOS dataset was used during both the training and evaluation phases of the model’s development. By employing stringent 80:20 hold-out validation, the model achieved a remarkable 98.71% accuracy of classification using enhancement techniques and 80.87% without using enhancement techniques.
2. Related Work
3. Research Methodology
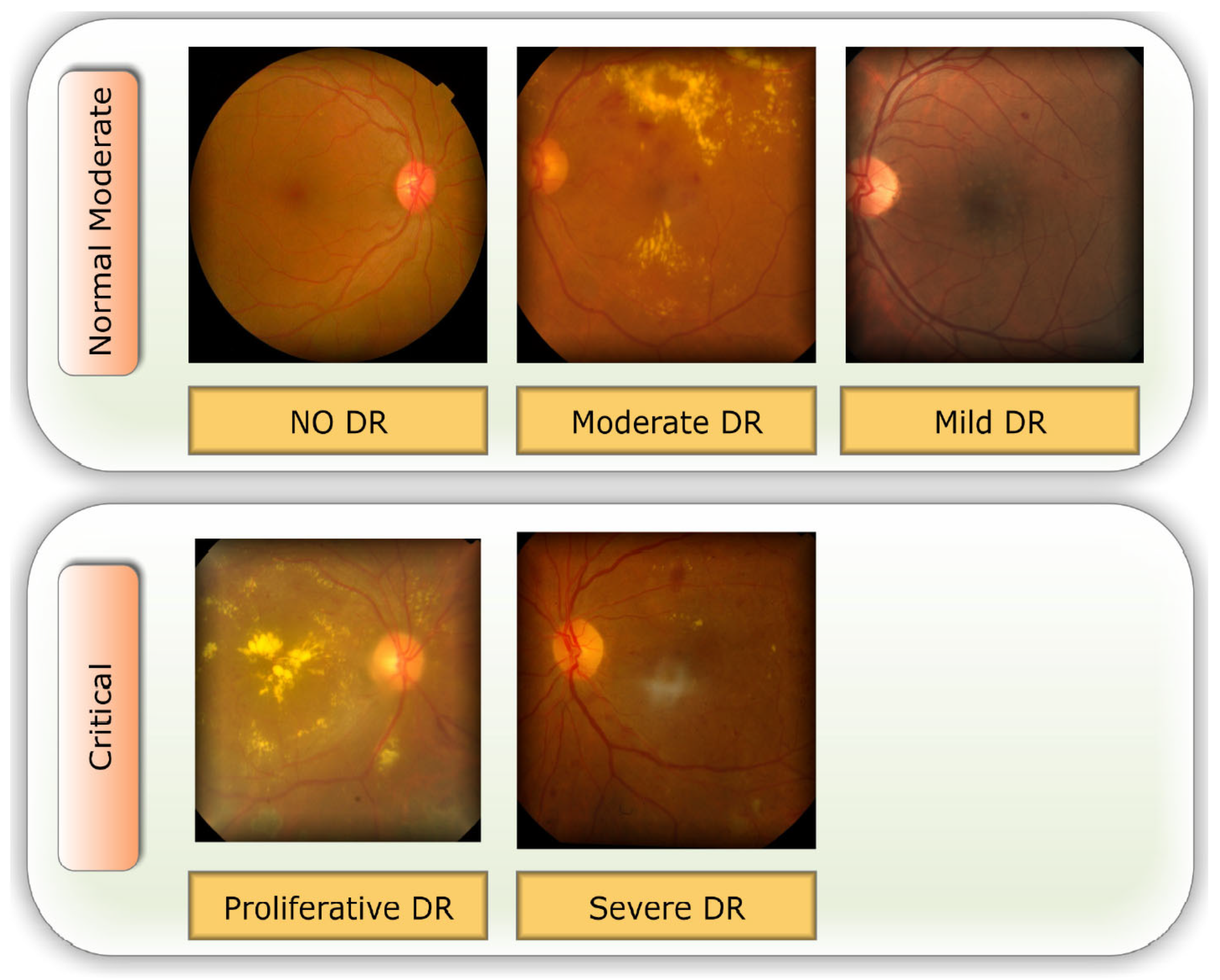
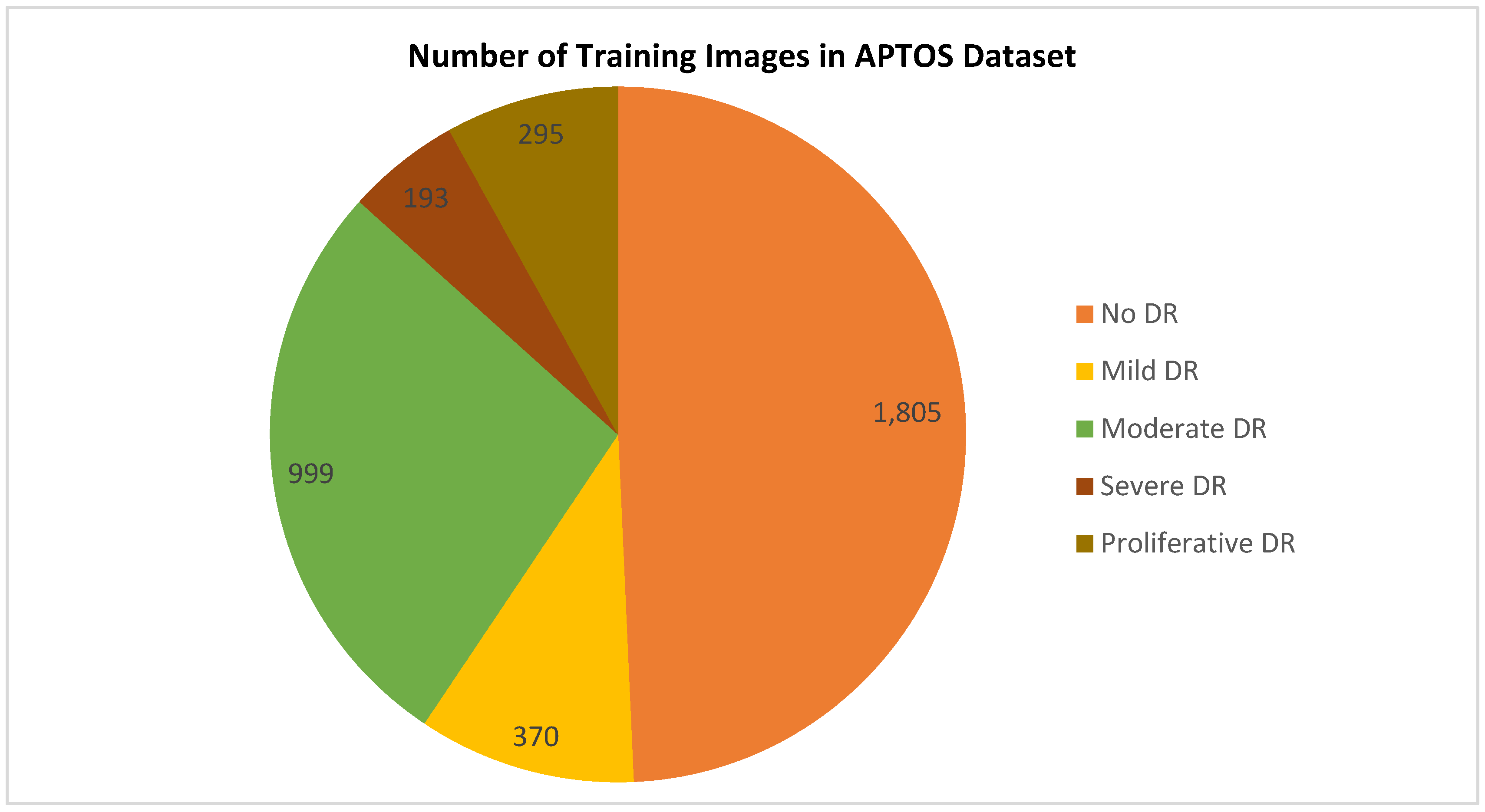
3.1. Data set Description
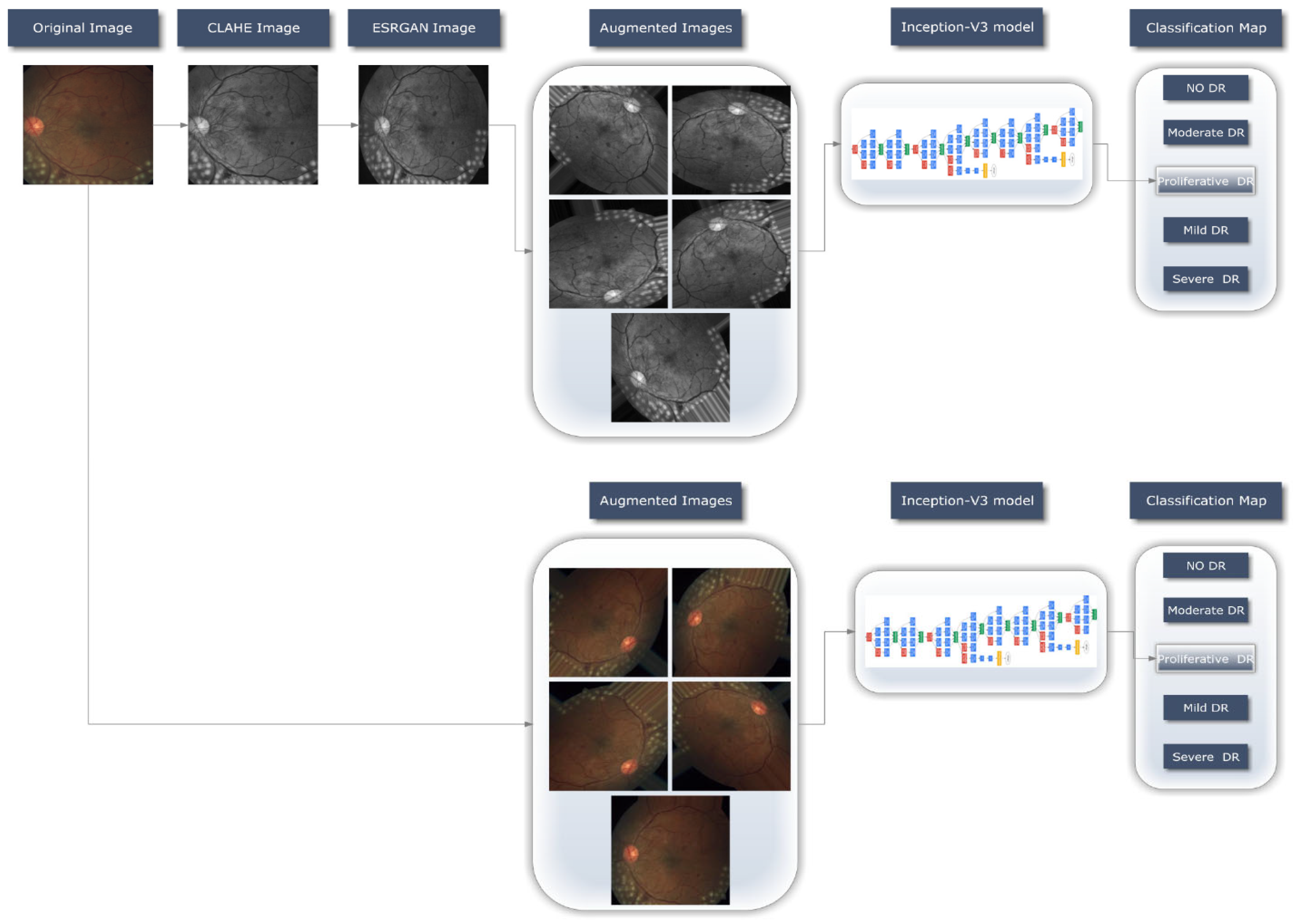
3.2. Proposed Methodology
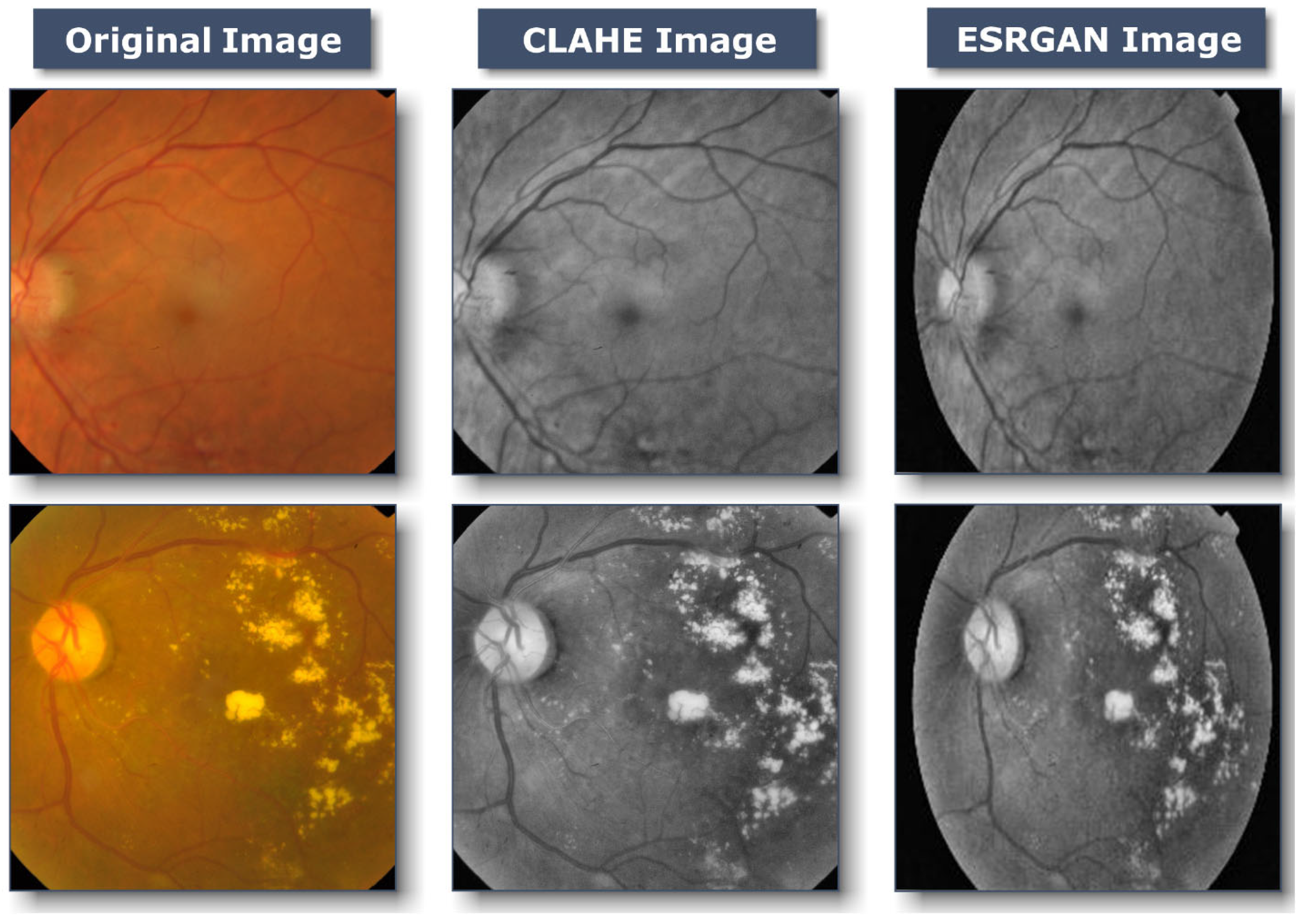
3.2.1. Preprocessing Using CLAHE and ESRGAN
- CLAHE
- Resize each picture to 224*224*3 pixels
- ESRGAN
- Normalization
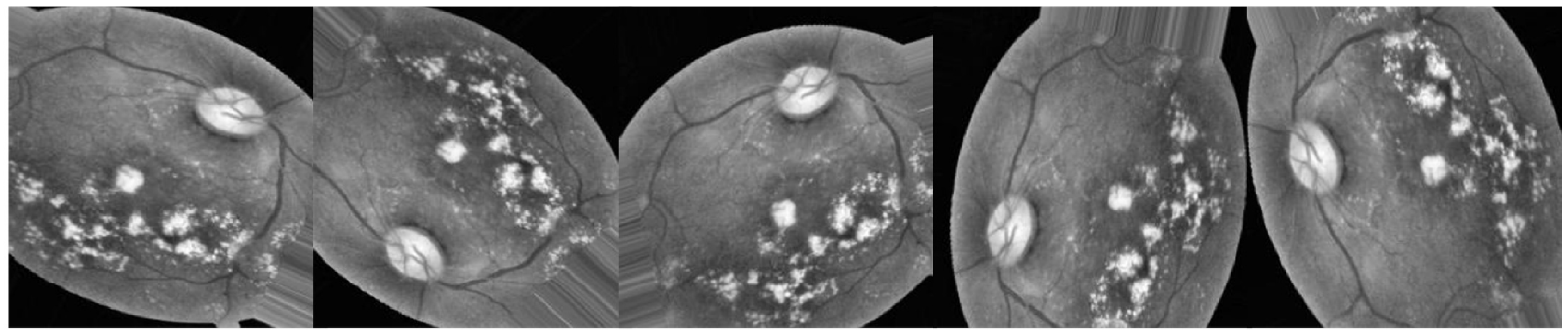
3.2.3. Data Augmentation
3.2.4. Learning Model (Inception-V3)
4. Experimental Results
4.1. Instruction and Setup of Inception-V3
4.2. Evaluative Parameters
Performance lof Inception-V3 Model Outcomes:
Evaluation Considering a Variety of Other Methodologies
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Atwany, M.Z., A.H. Sahyoun, and M. Yaqub, Deep learning techniques for diabetic retinopathy classification: A survey. IEEE Access, 2022. [CrossRef]
- Amin, J., M. Sharif, and M. Yasmin, A review on recent developments for detection of diabetic retinopathy. Scientifica, 2016. 2016. [CrossRef]
- Kharroubi, A.T. and H.M. Darwish, Diabetes mellitus: The epidemic of the century. World journal of diabetes, 2015. 6(6): p. 850. [CrossRef]
- Alex, S.A., Jhanjhi, N.Z., Humayun, M., Ibrahim, A.O. and Abulfaraj, A.W., 2022. Deep LSTM Model for Diabetes Prediction with Class Balancing by SMOTE. Electronics, 11(17), p.2737. [CrossRef]
- Taylor, R. and D. Batey, Handbook of retinal screening in diabetes: diagnosis and management. 2012: John Wiley & Sons.
- Alyoubi, W.L., W.M. Shalash, and M.F. Abulkhair, Diabetic retinopathy detection through deep learning techniques: A review. Informatics in Medicine Unlocked, 2020. 20: p. 100377. [CrossRef]
- Dubow, M., et al., Classification of human retinal microaneurysms using adaptive optics scanning light ophthalmoscope fluorescein angiography. Investigative ophthalmology & visual science, 2014. 55(3): p. 1299-1309. [CrossRef]
- Mazhar, K., et al., Severity of diabetic retinopathy and health-related quality of life: the Los Angeles Latino Eye Study. Ophthalmology, 2011. 118(4): p. 649-655. [CrossRef]
- Willis, J.R., et al., Vision-related functional burden of diabetic retinopathy across severity levels in the United States. JAMA ophthalmology, 2017. 135(9): p. 926-932. [CrossRef]
- Vora, P. and S. Shrestha, Detecting diabetic retinopathy using embedded computer vision. Applied Sciences, 2020. 10(20): p. 7274. [CrossRef]
- Humayun, Mamoona, Muhammad Ibrahim Khalil, Saleh Naif Almuayqil, and Noor Zaman Jhanjhi. "Framework for Detecting Breast Cancer Risk Presence Using Deep Learning." Electronics 12, no. 2 (2023): 403. [CrossRef]
- Szegedy, C., et al. Inception-v4, inception-resnet and the impact of residual connections on learning. in Thirty-first AAAI conference on artificial intelligence. 2017.
- Xia, X., C. Xu, and B. Nan. Inception-v3 for flower classification. in 2017 2nd international conference on image, vision and computing (ICIVC). 2017. IEEE.
- APTOS 2019 Blindness Detection. 2019, Kaggle.: Kaggle.
- Pizer, S.M., et al., Adaptive histogram equalization and its variations. Computer vision, graphics, and image processing, 1987. 39(3): p. 355-368. [CrossRef]
- Ledig, C., et al. Photo-realistic single image super-resolution using a generative adversarial network. in Proceedings of the IEEE conference on computer vision and pattern recognition. 2017.
- Al-Antary, M.T. and Y. Arafa, Multi-scale attention network for diabetic retinopathy classification. IEEE Access, 2021. 9: p. 54190-54200. [CrossRef]
- Gargeya, R. and T. Leng, Automated identification of diabetic retinopathy using deep learning. Ophthalmology, 2017. 124(7): p. 962-969. [CrossRef]
- Pak, A., et al., Comparative analysis of deep learning methods of detection of diabetic retinopathy. Cogent Engineering, 2020. 7(1): p. 1805144. [CrossRef]
- Kazakh-British, N.P., A. Pak, and D. Abdullina. Automatic detection of blood vessels and classification in retinal images for diabetic retinopathy diagnosis with application of convolution neural network. in Proceedings of the 2018 international conference on sensors, signal and image processing. 2018.
- Macsik, P., et al., Local Binary CNN for Diabetic Retinopathy Classification on Fundus Images. Acta Polytechnica Hungarica, 2022. 19(7). [CrossRef]
- Khalifa, N.E.M., et al., Deep transfer learning models for medical diabetic retinopathy detection. Acta Informatica Medica, 2019. 27(5): p. 327. [CrossRef]
- Hemanth, D.J., O. Deperlioglu, and U. Kose, An enhanced diabetic retinopathy detection and classification approach using deep convolutional neural network. Neural Computing and Applications, 2020. 32(3): p. 707-721. [CrossRef]
- Maqsood, S., R. Damaševičius, and R. Maskeliūnas, Hemorrhage detection based on 3D CNN deep learning framework and feature fusion for evaluating retinal abnormality in diabetic patients. Sensors, 2021. 21(11): p. 3865. [CrossRef]
- Das, S., et al., Deep learning architecture based on segmented fundus image features for classification of diabetic retinopathy. Biomedical Signal Processing and Control, 2021. 68: p. 102600. [CrossRef]
- Saranya, P., et al. Red Lesion Detection in Color Fundus Images for Diabetic Retinopathy Detection. in Proceedings of International Conference on Deep Learning, Computing and Intelligence. 2022. Springer.
- Thomas, N.M. and S. Albert Jerome, Grading and Classification of Retinal Images for Detecting Diabetic Retinopathy Using Convolutional Neural Network, in Advances in Electrical and Computer Technologies. 2022, Springer. p. 607-614.
- Crane, A. and M. Dastjerdi, Effect of Simulated Cataract on the Accuracy of an Artificial Intelligence Algorithm in Detecting Diabetic Retinopathy in Color Fundus Photos. Investigative Ophthalmology & Visual Science, 2022. 63(7): p. 2100–F0089-2100–F0089.
- Majumder, S. and N. Kehtarnavaz, Multitasking deep learning model for detection of five stages of diabetic retinopathy. IEEE Access, 2021. 9: p. 123220-123230. [CrossRef]
- Deshpande, A. and J. Pardhi, Automated detection of Diabetic Retinopathy using VGG-16 architecture. International Research Journal of Engineering and Technology, 2021. 8(03).
- Yadav, S., P. Awasthi, and S. Pathak, RETINA IMAGE AND DIABETIC RETINOPATHY: A DEEP LEARNING BASED APPROACH.
- Kobat, S.G., et al., Automated Diabetic Retinopathy Detection Using Horizontal and Vertical Patch Division-Based Pre-Trained DenseNET with Digital Fundus Images. Diagnostics, 2022. 12(8): p. 1975. [CrossRef]
- Reza, A.M., Realization of the contrast limited adaptive histogram equalization (CLAHE) for real-time image enhancement. Journal of VLSI signal processing systems for signal, image and video technology, 2004. 38(1): p. 35-44. [CrossRef]
- Jolicoeur-Martineau, A. The relativistic discriminator: a key element missing from standard GAN. arXiv 2018, arXiv:1807.00734. [Google Scholar]
- Humayun, M. and Alsayat, A., 2022. Prediction Model for Coronavirus Pandemic Using Deep Learning. Comput. Syst. Sci. Eng., 40(3), pp.947-961. [CrossRef]
- Hamza, M., Tehsin, S., Humayun, M., Almufareh, M.F. and Alfayad, M., 2022. A Comprehensive Review of Face Morph Generation and Detection of Fraudulent Identities. Applied Sciences, 12(24), p.12545. [CrossRef]
- Krizhevsky, A., I. Sutskever, and G.E. Hinton, Imagenet classification with deep convolutional neural networks. Advances in neural information processing systems, 2012. 25.
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Serre, T., et al., Robust object recognition with cortex-like mechanisms. IEEE transactions on pattern analysis and machine intelligence, 2007. 29(3): p. 411-426. [CrossRef]
- Lin, M.; Chen, Q.; Yan, S. Network in network. arXiv 2013, arXiv:1312.4400. [Google Scholar]
- Szegedy, C., et al. Going deeper with convolutions. in Proceedings of the IEEE conference on computer vision and pattern recognition. 2015.
- Arora, S., et al. Provable bounds for learning some deep representations. in International conference on machine learning. 2014. PMLR.
- Maqsood, Z. and M.K. Gupta, Automatic Detection of Diabetic Retinopathy on the Edge, in Cyber Security, Privacy and Networking. 2022, Springer. p. 129-139.
- Lahmar, C. and A. Idri, Deep hybrid architectures for diabetic retinopathy classification. Computer Methods in Biomechanics and Biomedical Engineering: Imaging & Visualization, 2022: p. 1-19. [CrossRef]
- Oulhadj, M., et al., Diabetic retinopathy prediction based on deep learning and deformable registration. Multimedia Tools and Applications, 2022: p. 1-19. [CrossRef]
- Gangwar, A.K. and V. Ravi, Diabetic retinopathy detection using transfer learning and deep learning, in Evolution in Computational Intelligence. 2021, Springer. p. 679-689.
- Lahmar, C. and A. Idri, On the value of deep learning for diagnosing diabetic retinopathy. Health and Technology, 2022. 12(1): p. 89-105. [CrossRef]
- Canayaz, M., Classification of diabetic retinopathy with feature selection over deep features using nature-inspired wrapper methods. Applied Soft Computing, 2022. 128: p. 109462. [CrossRef]
- Escorcia-Gutierrez, J., et al. Analysis of Pre-trained Convolutional Neural Network Models in Diabetic Retinopathy Detection Through Retinal Fundus Images. in International Conference on Computer Information Systems and Industrial Management. 2022. Springer.
- Salluri, D.K., V. Sistla, and V.K.K. Kolli, HRUNET: Hybrid Residual U-Net for automatic severity prediction of Diabetic Retinopathy. Computer Methods in Biomechanics and Biomedical Engineering: Imaging & Visualization, 2022: p. 1-12. [CrossRef]
- Yadav, S. and P. Awasthi, DIABETIC RETINOPATHY DETECTION USING DEEP LEARNING AND INCEPTION-V3 MODEL.
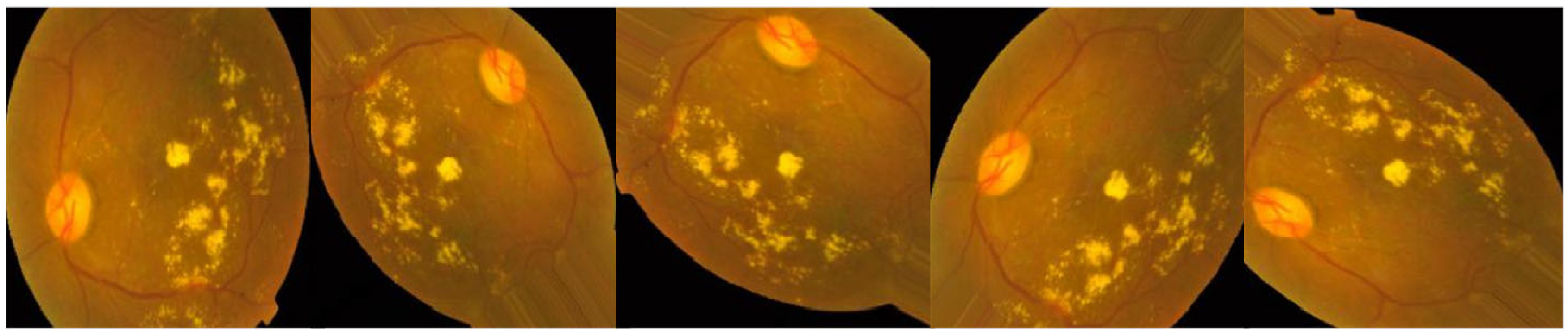
| Reference | Year | Technique | Classes | Dataset |
| [17] | 2021 | multi-scale attention network (MSA-Net) | 5 | APTOS |
| Eyepacs | ||||
| [21] | 2022 | local binary convolutional neural network (LBCNN) | 2 | APTOS |
| [26] | 2022 | support vector machine (SVM) | 2 | APTOS |
| IDRiD | ||||
| [27] | 2022 | CNN | 2 | APTOS |
| [28] | 2022 | Inception-ResNet-v2 | 5 | APTOS |
| [29] | 2021 | Squeeze Excitation Densely Connected deep CNN |
5 | APTOS |
| EyePACS | ||||
| [30] | 2021 | VGG-16 | 5 | APTOS |
| [31] | 2022 | VGG16 | 2 | APTOS |
| DenseNet121 | ||||
| [32] | 2022 | DenseNet201 | 5 | APTOS |
| 3 | New Dataset |
| Class Index | DR Level | # Images |
| 0 | No | 1,805 |
| 1 | Mild | 370 |
| 2 | Moderate | 999 |
| 3 | Severe | 193 |
| 4 | Proliferate | 295 |
| Acc | Prec | Re | F1sc | Top-2 Accuracy | Top-3 Accuracy |
|---|---|---|---|---|---|
| 0.9872 | 0.99 | 0.99 | 0.99 | 0.996 | 0.999 |
| Acc | Prec | Re | F1sc | Top-2 Accuracy | Top-3 Accuracy |
|---|---|---|---|---|---|
| 0.8087 | 0.80 | 0.81 | 0.80 | 0.9144 | 0.9800 |
| Prec | Re | F1sc | Total images | |
|---|---|---|---|---|
| Mild DR | 0.99 | 0.97 | 0.98 | 93 |
| Moderate DR | 0.98 | 0.99 | 0.98 | 280 |
| No DR | 0.99 | 1.00 | 1.00 | 504 |
| Proliferative DR | 0.97 | 0.95 | 0.96 | 82 |
| Severe DR | 0.98 | 0.96 | 0.97 | 53 |
| Average | 0.99 | 0.99 | 0.99 | 1012 |
| Prec | Re | F1sc | Total images | |
|---|---|---|---|---|
| Mild DR | 0.58 | 0.62 | 0.60 | 93 |
| Moderate DR | 0.70 | 0.78 | 0.74 | 280 |
| No DR | 0.97 | 0.97 | 0.97 | 504 |
| Proliferative DR | 0.68 | 0.48 | 0.56 | 82 |
| Severe DR | 0.43 | 0.31 | 0.36 | 53 |
| Average | 0.80 | 0.81 | 0.80 | 1012 |
| Reference | Technique | Accuracy |
|---|---|---|
| [17] | MSA-Net | 84.6% |
| [21] | LBCNN | 97.41% |
| [26] | SVM | 94.5% |
| [27] | CNN | 95.3% |
| [28] | Inception-ResNet-v2 | 97.0%, |
| [30] | VGG-16 | 74.58% |
| [31] | VGG16 | 73.26% |
| DenseNet121 | 96.11% | |
| [32] | DenseNet201 | 93.85% |
| [43] | EfficientNet-B6 | 86.03% |
| [44] | SVM classifier and MobileNet_V2 for feature extraction | 88.80% |
| [45] | Densenet-121, Xception, Inception-v3, Resnet-50 | 85.28% |
| [46] | Inception-ResNet-v2 | 72.33% |
| [47] | MobileNet_V2 | 93.09% |
| [48] | EfficientNet and DenseNet | 96.32% |
| [49] | VGG16 | 96.86% |
| [50] | Hybrid Residual U-Net | 94% |
| [51] | Inception-v3 | 88.1% |
| Proposed Methodology | Inception-V3 ( without using CLAHE + ESRGAN) Case 2 | 80.87% |
| Inception-V3 (using CLAHE + ESRGAN) Case 1 | 98.7% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




