Submitted:
15 January 2023
Posted:
20 January 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data preparation
2.1.1. Data acquisition of herbs from the online database
2.2. Hyperlipidemia-associated targets prediction
2.3. Protein-protein interaction (PPI) network construction
2.4. Gene ontology (GO) terms and KEGG pathway enrichment analysis
2.5. Chemicals and Antibodies
2.6. Preparation of Samples
2.7. Cell culture and treatment
2.8. Cell viability assay
2.9. Western blot analysis
2.10. Quantitative real-time polymerase chain reaction
2.11. Oil Red O staining
2.12. Statistical analysis
3. Results
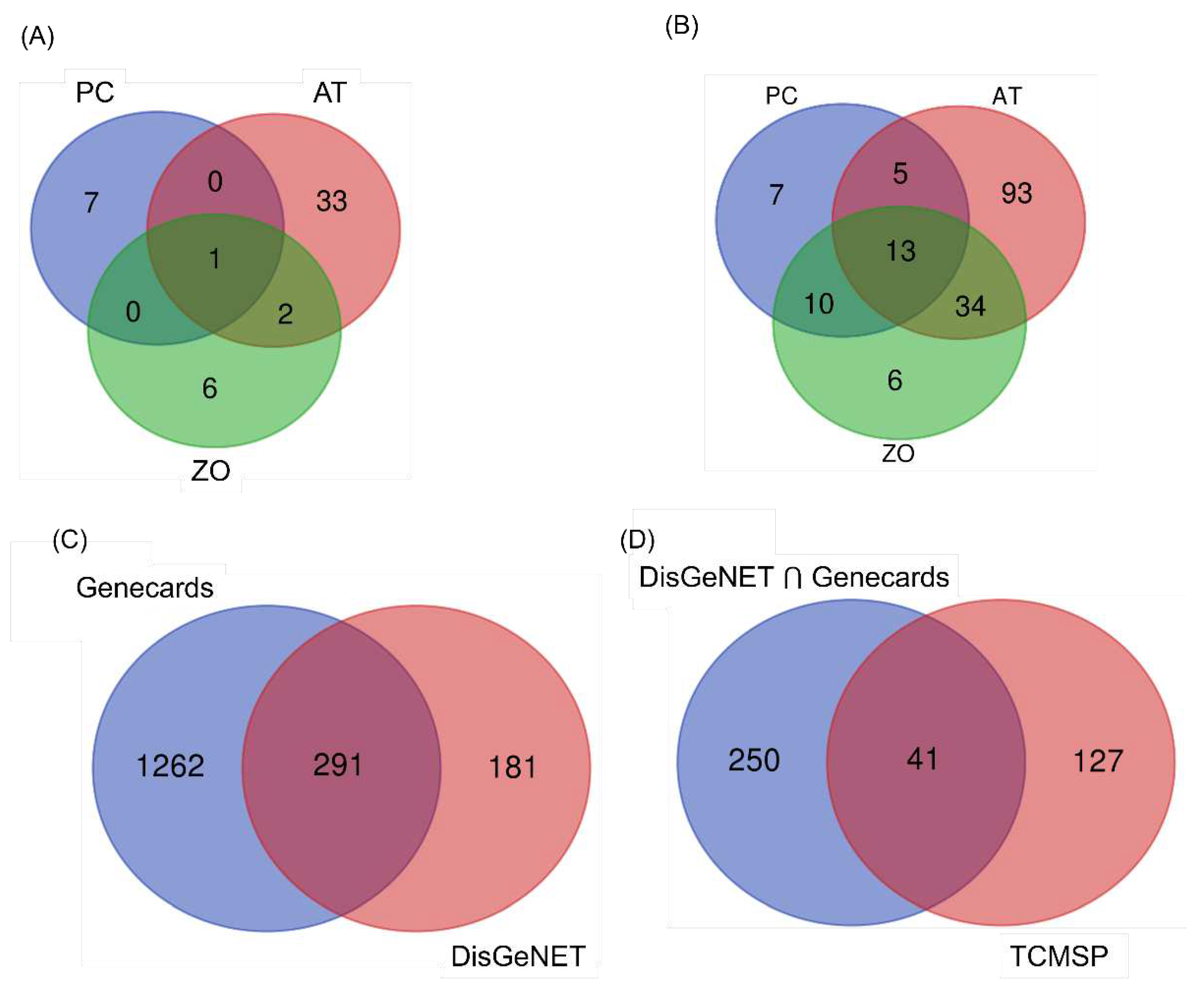
3.1. Selection of potential compounds from AT, PC, and ZO
3.2. Target prediction
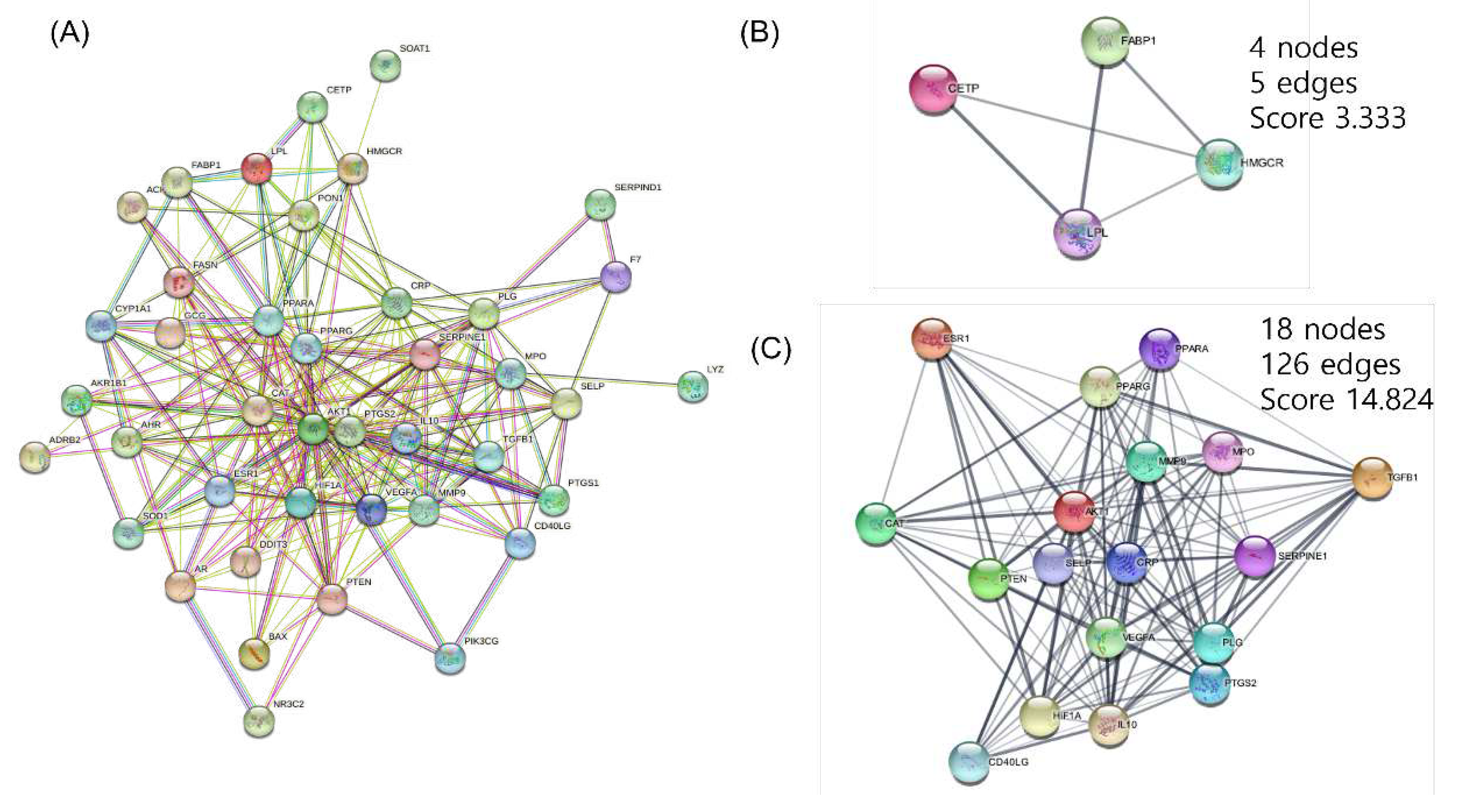
3.3. PPI networks construction and analysis
| Degree | stress | Betweenness centrality | |
|---|---|---|---|
| AKT1 | 31 | 894 | 0.131579 |
| PPARG | 27 | 682 | 0.092949 |
| PTGS2 | 26 | 494 | 0.049889 |
| CAT | 26 | 484 | 0.049086 |
| VEGFA | 25 | 454 | 0.042859 |
| PPARA | 23 | 448 | 0.049087 |
| CRP | 23 | 542 | 0.066034 |
| IL10 | 22 | 238 | 0.018832 |
| SERPINE1 | 21 | 476 | 0.049813 |
| MMP9 | 19 | 164 | 0.011888 |
| HIF1A | 19 | 178 | 0.012385 |
| PLG | 18 | 472 | 0.057540 |
| ESR1 | 17 | 154 | 0.010555 |
| PTEN | 16 | 228 | 0.023134 |
| MPO | 15 | 314 | 0.053684 |
| TGFB1 | 14 | 26 | 0.001131 |
| SELP | 12 | 98 | 0.007016 |
| FASN | 12 | 104 | 0.011726 |
| PON1 | 11 | 100 | 0.012281 |
| LPL | 11 | 72 | 0.006200 |
| CD40LG | 11 | 42 | 0.004880 |
| AR | 11 | 94 | 0.010259 |
| GCG | 11 | 42 | 0.005491 |
| AHR | 11 | 30 | 0.001772 |
| SOD1 | 10 | 12 | 0.000844 |
| HMGCR | 9 | 256 | 0.051766 |
| DDIT3 | 9 | 26 | 0.001189 |
| CYP1A1 | 9 | 50 | 0.005523 |
| BAX | 8 | 8 | 0.000513 |
| PTGS1 | 8 | 4 | 0.000256 |
| AKR1B1 | 7 | 2 | 0.000056 |
| FABP1 | 7 | 18 | 0.001893 |
| CETP | 6 | 6 | 0.000569 |
| F7 | 5 | 32 | 0.005104 |
| PIK3CG | 4 | 2 | 0.000214 |
| ACHE | 4 | 6 | 0.000341 |
| ADRB2 | 3 | 0 | 0.000000 |
| NR3C2 | 3 | 8 | 0.000383 |
| SERPIND1 | 2 | 0 | 0.000000 |
| LYZ | 1 | 0 | 0.000000 |
| SOAT1 | 1 | 0 | 0.000000 |
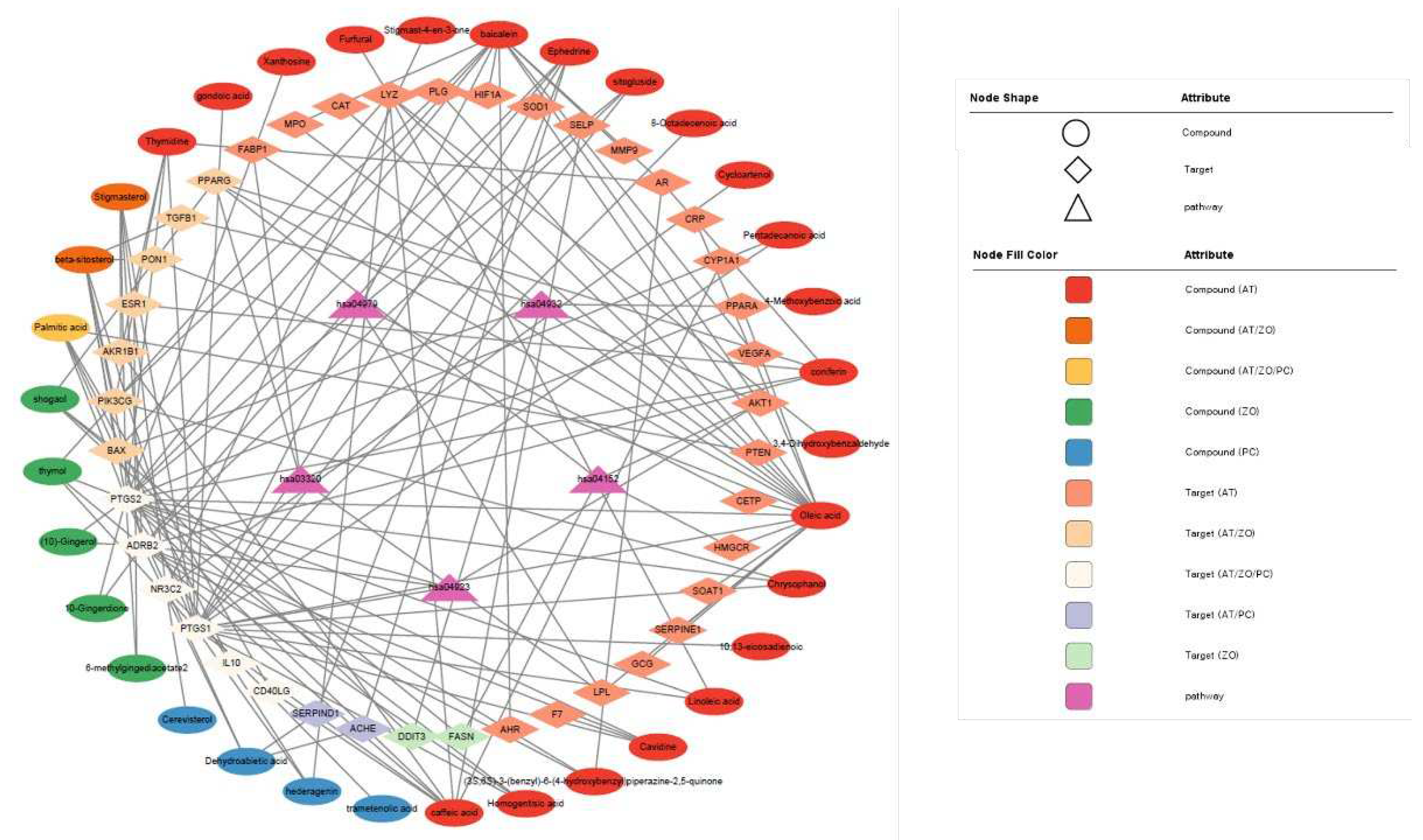
3.4. CTP visualization of Cytoscape
| KEGG Pathway | |||
| Entry | Pathway | FDR | Genes |
| hsa04923 | Regulation of lipolysis in adipocytes | 0.00039 | AKT1, PTGS1, PTGS2, ADRB2 |
| hsa04932 | Non-alcoholic fatty liver disease | 0.00054 | AKT1, BAX, DDIT3, PPARA, TGFB1 |
| hsa03320 | PPAR signaling pathway | 0.00062 | FABP1, LPL, PPARA, PPARG |
| hsa04152 | AMPK signaling pathway | 0.0019 | AKT1, FASN, HMGCR, PPARG |
| hsa04979 | Cholesterol metabolism | 0.0023 | CETP, LPL, SOAT1 |
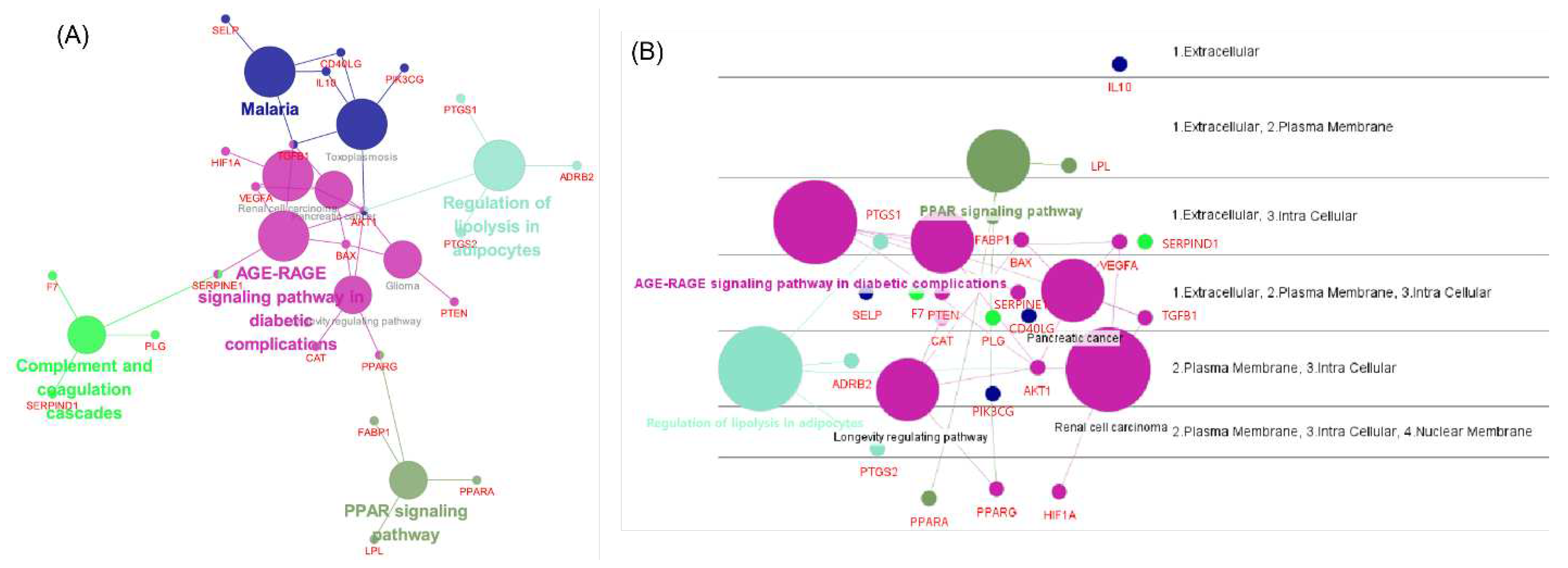
3.5. Network analysis using ClueGO, CluePedia
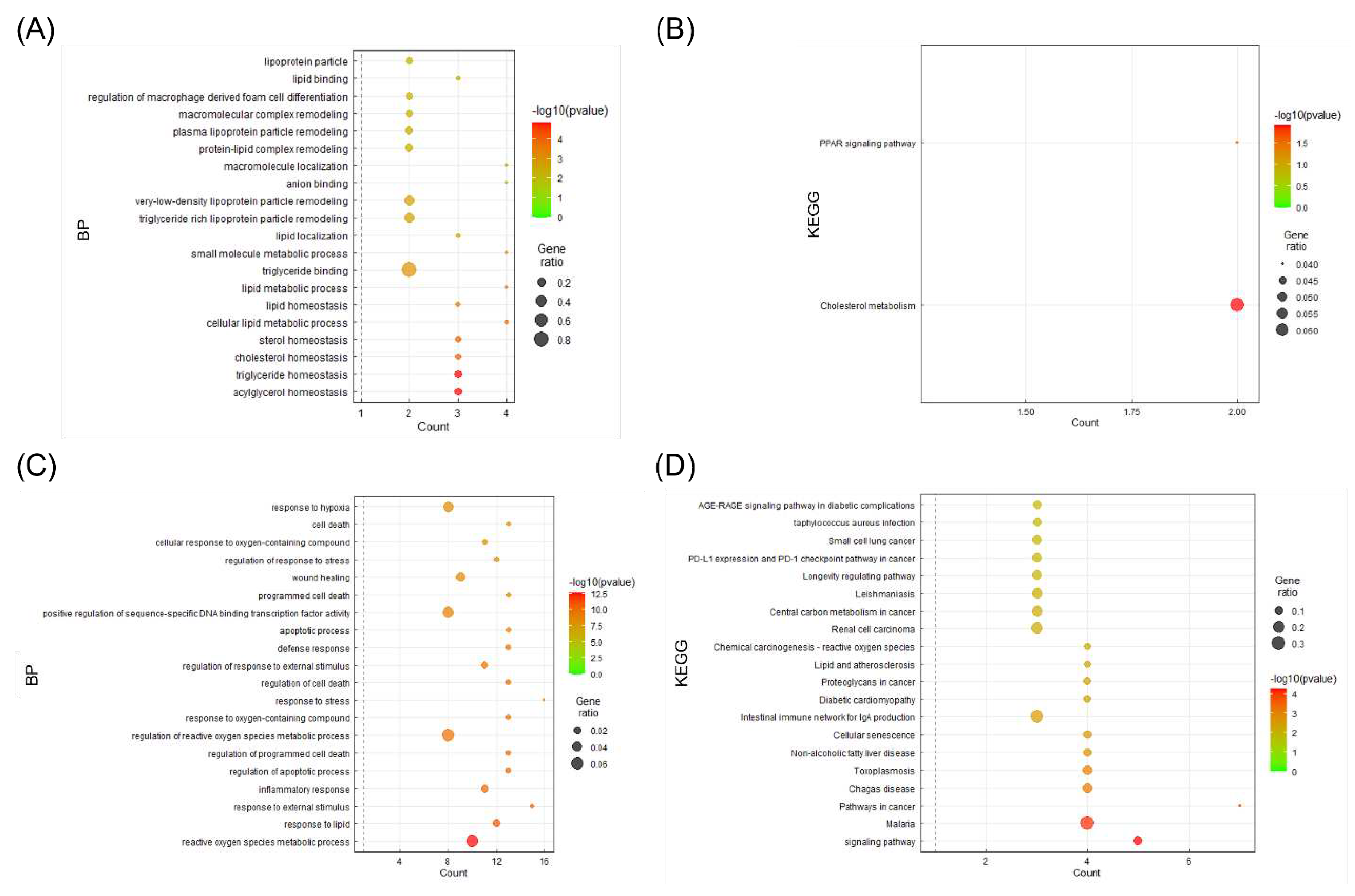
3.6. BP and KEGG enrichment analysis
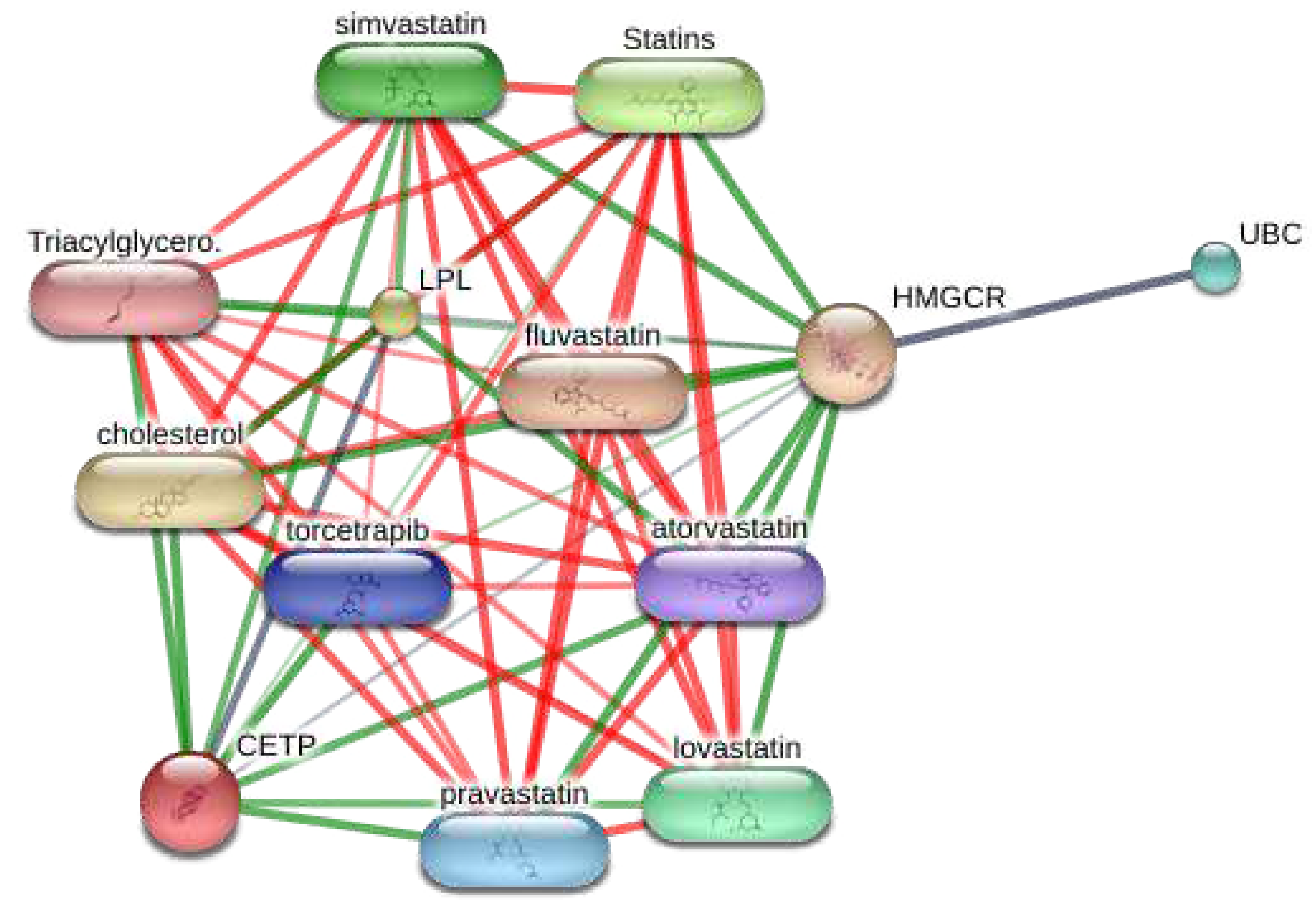
3.7. Visualization of the target-chemical interaction using STITCH
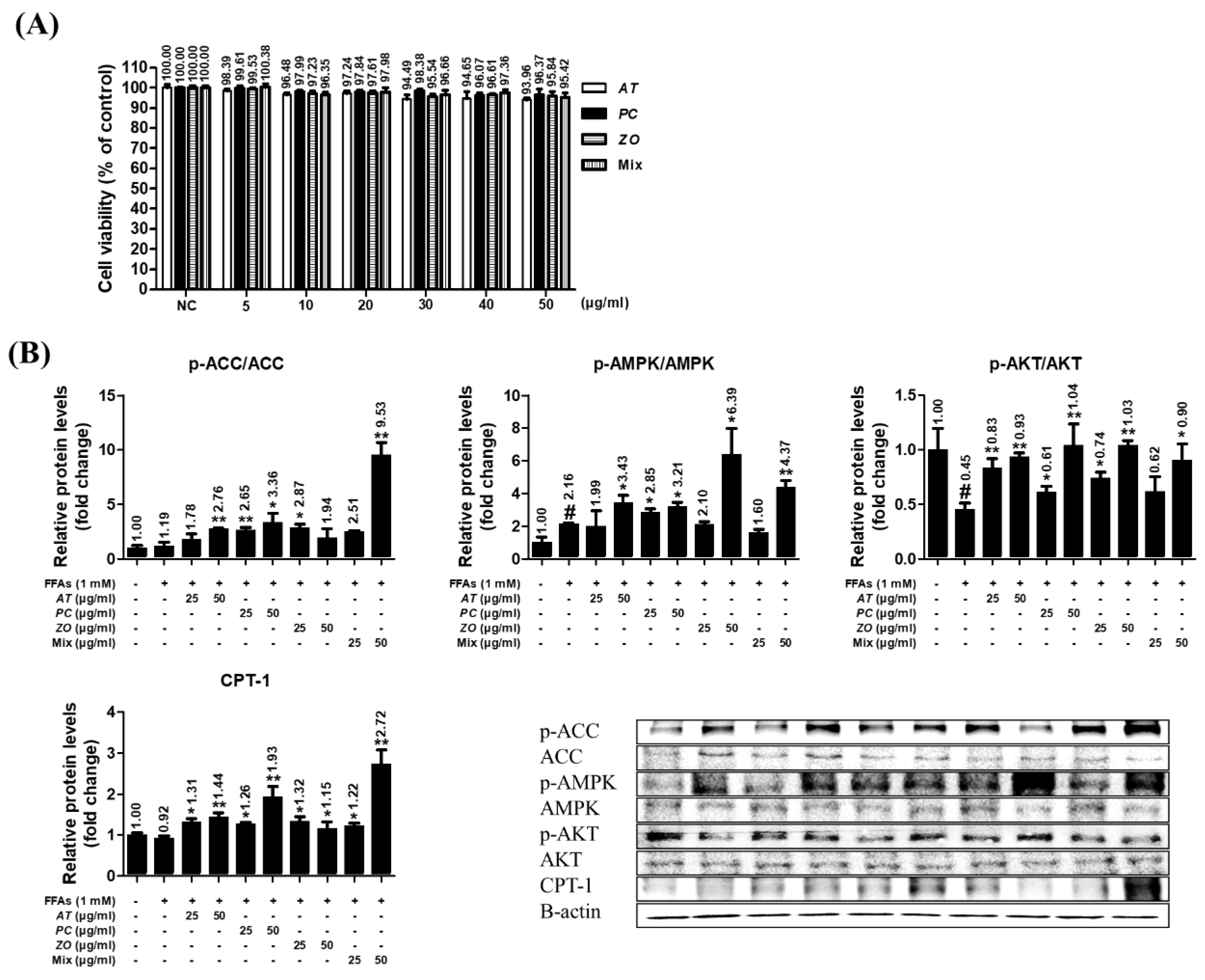
3.8. AT, PC, ZO, and mixed extract (MIX) improved the energy metabolism-related proteins in the hepatic steatosis model.
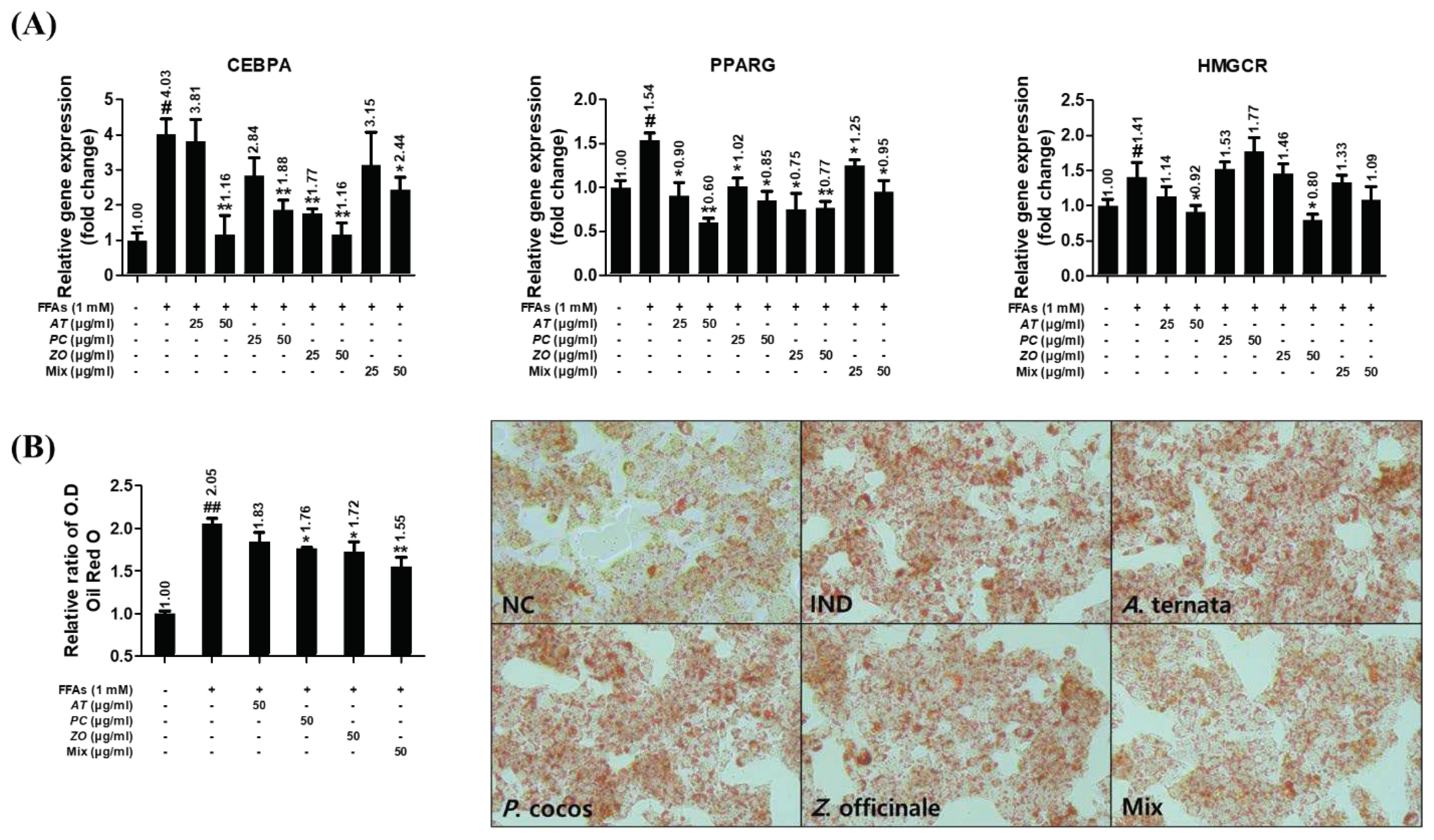
3.9. AT, PC, and ZO regulated the expression of genes related to lipogenesis and reduced FFAs-induced intracellular lipid accumulation.
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- He, N.; Ye, H. Exercise and hyperlipidemia. Physical Exercise for Human Health 2020, 79-90.
- Organization, W.H. The world health report 2002: reducing risks, promoting healthy life; World Health Organization: 2002.
- Song, D.-x.; Jiang, J.-g. Hypolipidemic components from medicine food homology species used in China: pharmacological and health effects. Archives of Medical Research 2017, 48, 569-581. [CrossRef]
- Jain, K.S.; Kathiravan, M.; Somani, R.S.; Shishoo, C.J. The biology and chemistry of hyperlipidemia. Bioorganic & medicinal chemistry 2007, 15, 4674-4699. [CrossRef]
- Ruixing, Y.; Jinzhen, W.; Weixiong, L.; Yuming, C.; Dezhai, Y.; Shangling, P. The environmental and genetic evidence for the association of hyperlipidemia and hypertension. Journal of hypertension 2009, 27, 251-258. [CrossRef]
- Naukkarinen, J.; Ehnholm, C.; Peltonen, L. Genetics of familial combined hyperlipidemia. Current opinion in lipidology 2006, 17, 285-290. [CrossRef]
- Lee, J.; Hoang, T.; Lee, S.; Kim, J. Association Between Dietary Patterns and Dyslipidemia in Korean Women. Frontiers in Nutrition 2021, 8. [CrossRef]
- Bandyopadhyay, D.; Qureshi, A.; Ghosh, S.; Ashish, K.; Heise, L.R.; Hajra, A.; Ghosh, R.K. Safety and efficacy of extremely low LDL-cholesterol levels and its prospects in hyperlipidemia management. Journal of lipids 2018, 2018. [CrossRef]
- Shattat, G.F. A review article on hyperlipidemia: types, treatments and new drug targets. Biomedical and Pharmacology Journal 2015, 7, 399-409. [CrossRef]
- Thompson, P.D.; Panza, G.; Zaleski, A.; Taylor, B. Statin-associated side effects. Journal of the American College of Cardiology 2016, 67, 2395-2410.
- Brault, M.; Ray, J.; Gomez, Y.-H.; Mantzoros, C.S.; Daskalopoulou, S.S. Statin treatment and new-onset diabetes: a review of proposed mechanisms. Metabolism 2014, 63, 735-745. [CrossRef]
- Yu, Q.; Chen, Y.; Xu, C.-B. Statins and new-onset diabetes mellitus: LDL receptor may provide a key link. Frontiers in Pharmacology 2017, 8, 372. [CrossRef]
- Zhang, R.; Zhu, X.; Bai, H.; Ning, K. Network pharmacology databases for traditional Chinese medicine: review and assessment. Frontiers in pharmacology 2019, 10, 123. [CrossRef]
- Zhao, S.; Li, S. Network-based relating pharmacological and genomic spaces for drug target identification. PloS one 2010, 5, e11764. [CrossRef]
- Banerjee, S.; Bhattacharjee, P.; Kar, A.; Mukherjee, P.K. LC–MS/MS analysis and network pharmacology of Trigonella foenum-graecum–A plant from Ayurveda against hyperlipidemia and hyperglycemia with combination synergy. Phytomedicine 2019, 60, 152944. [CrossRef]
- Huang, Y.; Guo, S.; Yang, J.; Tang, Y.; Zhu, X.; Ren, S. An Objective Diagnosis Model with Integrated Metabolic and Immunity Parameters for Phlegm-Dampness Constitution. Evidence-Based Complementary and Alternative Medicine 2022, 2022. [CrossRef]
- Jiang, Y.-H.; Zhang, P.; Tao, Y.; Liu, Y.; Cao, G.; Zhou, L.; Yang, C.-H. Banxia Baizhu Tianma decoction attenuates obesity-related hypertension. Journal of Ethnopharmacology 2021, 266, 113453. [CrossRef]
- Lee, A.Y.; Park, W.; Kang, T.-W.; Cha, M.H.; Chun, J.M. Network pharmacology-based prediction of active compounds and molecular targets in Yijin-Tang acting on hyperlipidaemia and atherosclerosis. Journal of Ethnopharmacology 2018, 221, 151-159. [CrossRef]
- Lee, S.M.; Lee, J.; Kang, E.; Kim, H.-L.; Hwang, G.-S.; Jung, J. Lipidomic analysis reveals therapeutic effects of Yijin-Tang on high-fat/high-cholesterol diet-induced obese mice. Phytomedicine 2020, 74, 152936. [CrossRef]
- Zhang, Y.; Wang, Z.; Zhang, Y.; Tong, H.; Zhang, Y.; Lu, T. Potential mechanisms for traditional Chinese medicine in treating airway mucus hypersecretion associated with coronavirus disease 2019. Frontiers in Molecular Biosciences 2020, 7, 577285.
- Liu, N.; Huo, G.; Zhang, L.; Zhang, X. Effect of Zingiber OfficinaleRosc on lipid peroxidation in hyperlipidemia rats. Wei sheng yan jiu= Journal of Hygiene Research 2003, 32, 22-23.
- Chen, H.; Chen, L.; Tang, D.-D.; Chen, D.-Q.; Miao, H.; Zhao, Y.-Y.; Ma, S.-C. Metabolomics reveals hyperlipidemic biomarkers and antihyperlipidemic effect of Poria cocos. Current Metabolomics 2016, 4, 104-115. [CrossRef]
- Kim, Y.-J.; Shin, Y.-O.; Ha, Y.-W.; Lee, S.; Oh, J.-K.; Kim, Y.S. Anti-obesity effect of Pinellia ternata extract in Zucker rats. Biological and Pharmaceutical Bulletin 2006, 29, 1278-1281. [CrossRef]
- Nemes, K.; Åberg, F. Interpreting lipoproteins in nonalcoholic fatty liver disease. Current opinion in lipidology 2017, 28, 355-360. [CrossRef]
- Hydes, T.; Alam, U.; Cuthbertson, D.J. The impact of macronutrient intake on non-alcoholic fatty liver disease (NAFLD): too much fat, too much carbohydrate, or just too many calories? Frontiers in nutrition 2021, 8, 640557. [CrossRef]
- Chen, M.; Guo, W.-L.; Li, Q.-Y.; Xu, J.-X.; Cao, Y.-J.; Liu, B.; Yu, X.-D.; Rao, P.-F.; Ni, L.; Lv, X.-C. The protective mechanism of Lactobacillus plantarum FZU3013 against non-alcoholic fatty liver associated with hyperlipidemia in mice fed a high-fat diet. Food & function 2020, 11, 3316-3331. [CrossRef]
- Deprince, A.; Haas, J.T.; Staels, B. Dysregulated lipid metabolism links NAFLD to cardiovascular disease. Molecular metabolism 2020, 42, 101092. [CrossRef]
- Świderska, E.; Strycharz, J.; Wróblewski, A.; Szemraj, J.; Drzewoski, J.; Śliwińska, A. Role of PI3K/AKT pathway in insulin-mediated glucose uptake. Blood Glucose Levels 2018, 1, 1-18.
- Ferreira, D.; Castro, R.; Machado, M.; Evangelista, T.; Silvestre, A.; Costa, A.; Coutinho, J.; Carepa, F.; Cortez-Pinto, H.; Rodrigues, C. Apoptosis and insulin resistance in liver and peripheral tissues of morbidly obese patients is associated with different stages of non-alcoholic fatty liver disease. Diabetologia 2011, 54, 1788-1798. [CrossRef]
- Niu, Y.; Li, S.; Na, L.; Feng, R.; Liu, L.; Li, Y.; Sun, C. Mangiferin decreases plasma free fatty acids through promoting its catabolism in liver by activation of AMPK. PLoS one 2012, 7, e30782. [CrossRef]
- Lee, W.H.; Kim, S.G. AMPK-dependent metabolic regulation by PPAR agonists. PPAR research 2010, 2010. [CrossRef]
- Liu, S.; Jing, F.; Yu, C.; Gao, L.; Qin, Y.; Zhao, J. AICAR-induced activation of AMPK inhibits TSH/SREBP-2/HMGCR pathway in liver. PLoS One 2015, 10, e0124951.
- Vecchione, G.; Grasselli, E.; Compalati, A.D.; Ragazzoni, M.; Cortese, K.; Gallo, G.; Voci, A.; Vergani, L. Ethanol and fatty acids impair lipid homeostasis in an in vitro model of hepatic steatosis. Food and Chemical Toxicology 2016, 90, 84-94. [CrossRef]
- Ru, J.; Li, P.; Wang, J.; Zhou, W.; Li, B.; Huang, C.; Li, P.; Guo, Z.; Tao, W.; Yang, Y. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. Journal of cheminformatics 2014, 6, 1-6.
- Xu, X.; Zhang, W.; Huang, C.; Li, Y.; Yu, H.; Wang, Y.; Duan, J.; Ling, Y. A novel chemometric method for the prediction of human oral bioavailability. International journal of molecular sciences 2012, 13, 6964-6982. [CrossRef]
- Tao, W.; Xu, X.; Wang, X.; Li, B.; Wang, Y.; Li, Y.; Yang, L. Network pharmacology-based prediction of the active ingredients and potential targets of Chinese herbal Radix Curcumae formula for application to cardiovascular disease. Journal of ethnopharmacology 2013, 145, 1-10. [CrossRef]
- Yue, S.-J.; Liu, J.; Feng, W.-W.; Zhang, F.-L.; Chen, J.-X.; Xin, L.-T.; Peng, C.; Guan, H.-S.; Wang, C.-Y.; Yan, D. System pharmacology-based dissection of the synergistic mechanism of Huangqi and Huanglian for diabetes mellitus. Frontiers in Pharmacology 2017, 8, 694. [CrossRef]
- Safran, M.; Dalah, I.; Alexander, J.; Rosen, N.; Iny Stein, T.; Shmoish, M.; Nativ, N.; Bahir, I.; Doniger, T.; Krug, H. GeneCards Version 3: the human gene integrator. Database 2010, 2010. [CrossRef]
- Piñero, J.; Queralt-Rosinach, N.; Bravo, A.; Deu-Pons, J.; Bauer-Mehren, A.; Baron, M.; Sanz, F.; Furlong, L.I. DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database 2015, 2015. [CrossRef]
- Bardou, P.; Mariette, J.; Escudié, F.; Djemiel, C.; Klopp, C. jvenn: an interactive Venn diagram viewer. BMC bioinformatics 2014, 15, 1-7. [CrossRef]
- Cline, M.S.; Smoot, M.; Cerami, E.; Kuchinsky, A.; Landys, N.; Workman, C.; Christmas, R.; Avila-Campilo, I.; Creech, M.; Gross, B. Integration of biological networks and gene expression data using Cytoscape. Nature protocols 2007, 2, 2366-2382. [CrossRef]
- Bindea, G.; Galon, J.; Mlecnik, B. CluePedia Cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics 2013, 29, 661-663. [CrossRef]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.-H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091-1093. [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic acids research 2019, 47, D607-D613.
- Szklarczyk, D.; Santos, A.; Von Mering, C.; Jensen, L.J.; Bork, P.; Kuhn, M. STITCH 5: augmenting protein–chemical interaction networks with tissue and affinity data. Nucleic acids research 2016, 44, D380-D384.
- Dennis, G.; Sherman, B.T.; Hosack, D.A.; Yang, J.; Gao, W.; Lane, H.C.; Lempicki, R.A. DAVID: database for annotation, visualization, and integrated discovery. Genome biology 2003, 4, 1-11.
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal statistical society: series B (Methodological) 1995, 57, 289-300. [CrossRef]
- Bonnot, T.; Gillard, M.B.; Nagel, D.H. A simple protocol for informative visualization of enriched gene ontology terms. Bio-protocol 2019, e3429-e3429. [CrossRef]
- Lim, D.-W.; Kim, H.; Lee, S.-J.; Yu, G.-R.; Kim, J.-E.; Park, W.-H. Jwa Kum Whan attenuates nonalcoholic fatty liver disease by modulating glucose metabolism and the insulin signaling pathway. Evidence-Based Complementary and Alternative Medicine 2019, 2019. [CrossRef]
- Yu, G.R.; Lim, D.W.; Karunarathne, W.A.H.M.; Kim, G.Y.; Kim, H.; Kim, J.E.; Park, W.H. A non-polar fraction of Saponaria officinalis L. acted as a TLR4/MD2 complex antagonist and inhibited TLR4/MyD88 signaling in vitro and in vivo. The FASEB Journal 2022, 36, e22387.
- Yu, G.-R.; Lee, S.-J.; Kim, D.-H.; Lim, D.-W.; Kim, H.; Park, W.-H.; Kim, J.-E. Literature-based drug repurposing in traditional Chinese medicine: Reduced inflammatory M1 macrophage polarization by Jisil Haebaek Gyeji-Tang alleviates cardiovascular disease in vitro and ex vivo. Evidence-Based Complementary and Alternative Medicine 2020, 2020. [CrossRef]
- Lin, Y.; Ren, N.; Li, S.; Chen, M.; Pu, P. Novel anti-obesity effect of scutellarein and potential underlying mechanism of actions. Biomedicine & Pharmacotherapy 2019, 117, 109042. [CrossRef]
- Yao, H.-R.; Liu, J.; Plumeri, D.; Cao, Y.-B.; He, T.; Lin, L.; Li, Y.; Jiang, Y.-Y.; Li, J.; Shang, J. Lipotoxicity in HepG2 cells triggered by free fatty acids. American journal of translational research 2011, 3, 284.
- Long, F.; Yang, H.; Xu, Y.; Hao, H.; Li, P. A strategy for the identification of combinatorial bioactive compounds contributing to the holistic effect of herbal medicines. Scientific reports 2015, 5, 1-11. [CrossRef]
- Efferth, T.; Koch, E. Complex interactions between phytochemicals. The multi-target therapeutic concept of phytotherapy. Current drug targets 2011, 12, 122-132. [CrossRef]
- Lahlou, M. Screening of natural products for drug discovery. Expert opinion on drug discovery 2007, 2, 697-705. [CrossRef]
- Jiao, X.; Jin, X.; Ma, Y.; Yang, Y.; Li, J.; Liang, L.; Liu, R.; Li, Z. A comprehensive application: molecular docking and network pharmacology for the prediction of bioactive constituents and elucidation of mechanisms of action in component-based Chinese medicine. Computational Biology and Chemistry 2021, 90, 107402. [CrossRef]
- Poornima, P.; Kumar, J.D.; Zhao, Q.; Blunder, M.; Efferth, T. Network pharmacology of cancer: From understanding of complex interactomes to the design of multi-target specific therapeutics from nature. Pharmacological Research 2016, 111, 290-302. [CrossRef]
- Xiao, P.-t.; Liu, S.-y.; Kuang, Y.-j.; Jiang, Z.-m.; Lin, Y.; Xie, Z.-s.; Liu, E.-H. Network pharmacology analysis and experimental validation to explore the mechanism of sea buckthorn flavonoids on hyperlipidemia. Journal of Ethnopharmacology 2021, 264, 113380. [CrossRef]
- Ye, J.; Li, L.; Hu, Z. Exploring the molecular mechanism of action of Yinchen Wuling powder for the treatment of hyperlipidemia, using network pharmacology, molecular docking, and molecular dynamics simulation. BioMed research international 2021, 2021. [CrossRef]
- Duncan, R.E.; Ahmadian, M.; Jaworski, K.; Sarkadi-Nagy, E.; Sul, H.S. Regulation of lipolysis in adipocytes. Annual review of nutrition 2007, 27, 79. [CrossRef]
- Kersten, S. Peroxisome proliferator activated receptors and lipoprotein metabolism. PPAR research 2008, 2008. [CrossRef]
- Grundy, S.M.; Vega, G.L. Fibric acids: effects on lipids and lipoprotein metabolism. The American journal of medicine 1987, 83, 9-20. [CrossRef]
- Ginsberg, H.N. Effects of statins on triglyceride metabolism. The American journal of cardiology 1998, 81, 32B-35B. [CrossRef]
- Bozkurt, B.; Aguilar, D.; Deswal, A.; Dunbar, S.B.; Francis, G.S.; Horwich, T.; Jessup, M.; Kosiborod, M.; Pritchett, A.M.; Ramasubbu, K. Contributory risk and management of comorbidities of hypertension, obesity, diabetes mellitus, hyperlipidemia, and metabolic syndrome in chronic heart failure: a scientific statement from the American Heart Association. Circulation 2016, 134, e535-e578. [CrossRef]
- Ul-Haq, Z.; Mackay, D.F.; Fenwick, E.; Pell, J.P. Impact of metabolic comorbidity on the association between body mass index and health-related quality of life: a Scotland-wide cross-sectional study of 5,608 participants. BMC Public Health 2012, 12, 1-7. [CrossRef]
| Compound List | ||||
| Herb | Name | CID | OB | DL |
| Zingiber officinale (ZO) |
(10)-Gingerol | 168115 | 19.14 | 0.28 |
| 10-Gingerdione | 5317591 | 21.42 | 0.29 | |
| 6-methylgingediacetate2 | 53179662 | 48.73 | 0.32 | |
| shogaol | 5281794 | 31.00 | 0.14 | |
| poriferast-5-en-3beta-ol | 457801 | 36.91 | 0.75 | |
| thymol | 6989 | 41.47 | 0.03 | |
| Poria cocos (PC) |
Cerevisterol | 10181133 | 37.96 | 0.77 |
| Dehydroabietic acid | 94391 | 14.93 | 0.28 | |
| (2R)-2-[(3S,5R,10S,13R,14R,16R,17R)-3,16-dihydroxy-4,4,10,13,14-pentamethyl-2,3,5,6,12,15,16,17-octahydro-1H-cyclopenta[a]phenanthren-17-yl]-6-methylhept-5-enoic acid | 10743008 | 30.93 | 0.81 | |
| Ergosterol peroxide | 5351516 | 40.36 | 0.81 | |
| ergosta-7,22E-dien-3beta-ol | 5283628 | 43.51 | 0.72 | |
| hederagenin | 73299 | 22.42 | 0.74 | |
| trametenolic acid | 12309443 | 38.71 | 0.80 | |
| Arum ternata (AT) | (3S,6S)-3-(benzyl)-6-(4-hydroxybenzyl)piperazine-2,5-quinone | 11438306 | 46.89 | 0.27 |
| 10,13-eicosadienoic | 549062 | 39.99 | 0.20 | |
| 3,4-Dihydroxybenzaldehyde | 8768 | 38.35 | 0.03 | |
| 4-Methoxybenzoic acid | 7478 | 29.69 | 0.03 | |
| 8-Octadecenoic acid | 5282758 | 33.13 | 0.14 | |
| Baicalein | 5281605 | 33.52 | 0.21 | |
| Xanthosine | 64959 | 44.72 | 0.21 | |
| caffeic acid | 689043 | 25.76 | 0.05 | |
| oct-1-ene | 8125 | 39.25 | 0.01 | |
| Baicalin | 64982 | 40.12 | 0.75 | |
| Choline | 305 | 0.47 | 0.01 | |
| 9-oxononanoic acid | 75704 | 19.60 | 0.03 | |
| docosanoic acid | 8125 | 15.69 | 0.26 | |
| 9-Heptadecanol | 136435 | 14.24 | 0.09 | |
| Palmitic acid | 985 | 19.30 | 0.10 | |
| Baicalein | 5281605 | 33.52 | 0.21 | |
| Hydroquinone | 785 | 29.26 | 0.02 | |
| Anethole | 637563 | 32.49 | 0.03 | |
| Adenine | 190 | 62.81 | 0.03 | |
| Cavidine | 193148 | 35.64 | 0.81 | |
| Chrysophanol | 10208 | 18.64 | 0.21 | |
| Coniferin | 5280372 | 10.28 | 0.27 | |
| Cycloartenol | 92110 | 38.69 | 0.78 | |
| Ephedrine | 9294 | 43.35 | 0.03 | |
| Furfural | 7362 | 34.35 | 0.01 | |
| gondoic acid | 5282768 | 30.70 | 0.20 | |
| Homogentisic acid | 780 | 92.44 | 0.04 | |
| Linoleic acid | 5280450 | 41.90 | 0.14 | |
| Oleic acid | 445639 | 33.13 | 0.14 | |
| Pentadecanoic acid | 13849 | 20.18 | 0.08 | |
| Sitogluside | 5742590 | 20.63 | 0.62 | |
| Stigmast-4-en-3-one | 5484202 | 36.08 | 0.76 | |
| Thymidine | 5789 | 11.34 | 0.11 | |
| AT∩ZO | beta-sitosterol | 222284 | 15.00 | 0.81 |
| Stigmasterol | 5280794 | 43.83 | 0.76 | |
| AT∩ZO∩PC | Palmitic acid | 985 | 19.30 | 0.10 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




