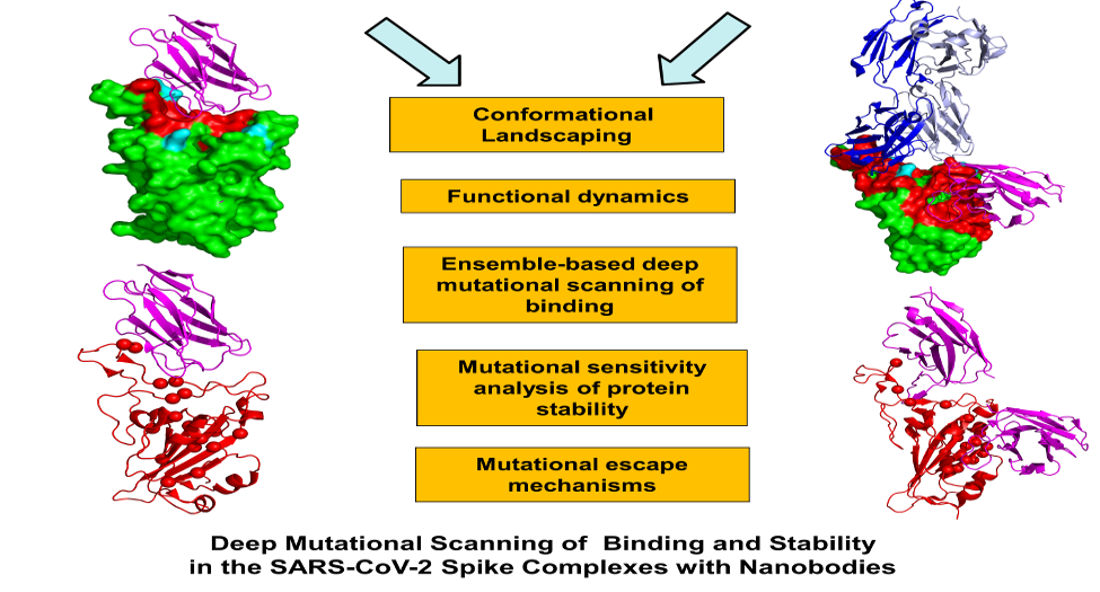
Structural and biochemical studies have recently revealed a range of rationally engineered nanobodies with efficient neutralizing capacity against SARS-CoV-2 virus and resilience against mutational escape. In this work, we combined atomistic simulations and conformational dynamics analysis with the ensemble-based mutational profiling of binding interactions for a diverse panel of SARS-CoV-2 spike complexes with nanobodies. Using this computational toolkit, we identified dynamic signatures and binding affinity fingerprints for the SARS-CoV-2 spike protein complexes with nanobodies Nb6 and Nb20, VHH E, a pair combination VHH E+U, a biparatopic nanobody VHH VE, and a combination of CC12.3 antibody and VHH V/W nanobodies. Through ensemble-based deep mutational profiling of stability and binding affinities, we identify critical hotspots and characterize molecular mechanisms of SARS-CoV-2 spike protein binding with single ultra-potent nanobodies, nanobody cocktails and biparatopic nanobodies. By quantifying dynamic and energetic determinants of the SARS-CoV-2 S binding with nanobodies, we also examine the effects of circulating variants and escaping mutations. We found that mutational escape mechanisms may be controlled through structurally and energetically adaptable binding hotspots located in the host receptor-accessible binding epitope that are dynamically coupled to the stability centers in the distant epitope targeted by VHH U/V/W nanobodies. The results of this study suggested a mechanism in which through cooperative dynamic changes, nanobody combinations and biparatopic nanobody can modulate the global protein response and induce the increased resilience to common escape mutants.