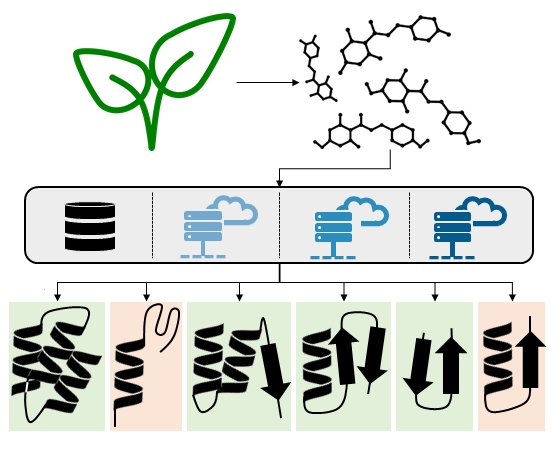
Natural products comprise a rich reservoir for innovative drug leads and are a constant source of bioactive compounds. To find pharmacological targets for new or already known natural products using modern computer-aided methods is a current endeavor in drug discovery. Nature’s treasures, however, could be used more effectively. Yet, reliable pipelines for large scale target prediction of natural products are still rare. We have developed an in silico workflow consisting of four independent, stand-alone target prediction tools and evaluated its performance on dihydrochalcones (DHCs) – a well-known class of natural products. Thereby, we revealed four previously unreported protein targets for DHCs, namely 5-lipoxygenase, cyclooxygenase-1, 17β- hydroxysteroid dehydrogenase 3, and aldo-keto reductase 1C3. Moreover, we provide a thorough strategy on how to perform computational target prediction and guidance on using the respective tools.