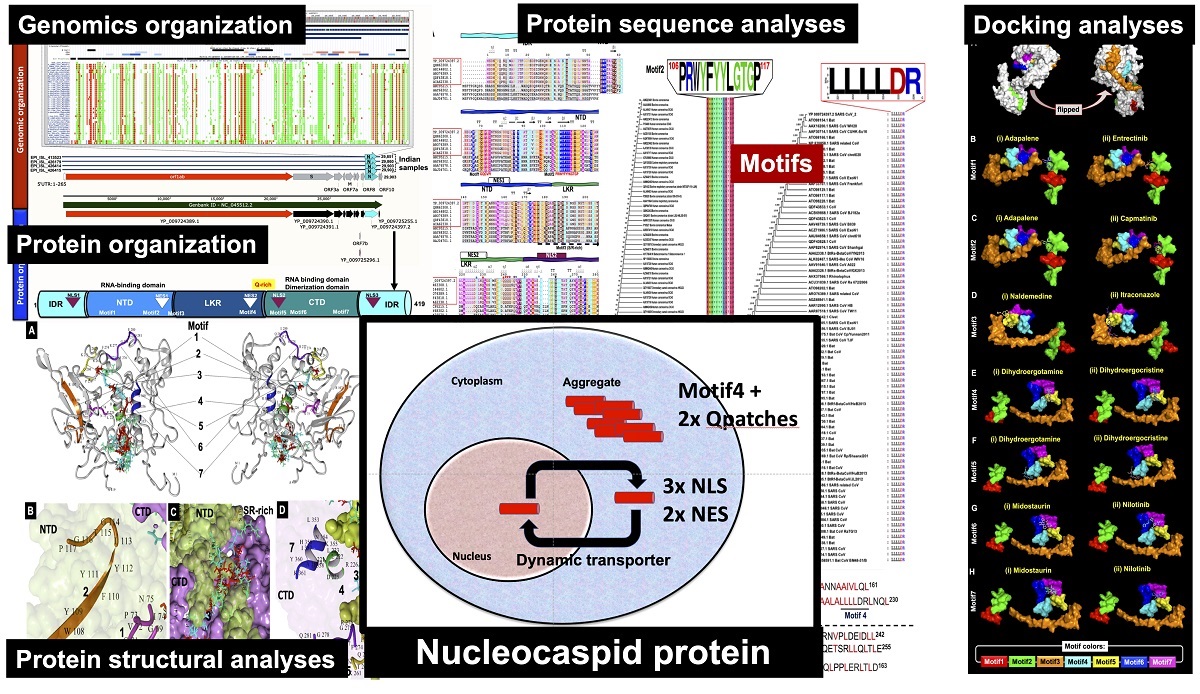
Severe acute respiratory syndrome novel coronavirus 2 (SARS-CoV-2) has caused the global pandemic as COVID-19, which is the most notorious global public health crisis in the last 100 years. SARS-CoV-2 is composed of four structural proteins and several non-structured proteins. The multi-facet nucleocapsid (N) protein is the major component of structural proteins of CoVs, However, there are no dedicated genomic, sequences and structural analyses focusing on potential roles of N protein. Hence, there is an urgent requirement of a detailed study on N protein of SARS-CoV-2. Herein, we are presenting a comprehensive study on N protein from SARS-CoV-2. We have identified seven motifs conserved in the three major domains namely N-terminal domain, linker regions and the C-terminal domains. Out of seven motifs, six motifs are conserved across different members of coronaviridae, while motif4 is specific for SARS CoVs with potential amyloidogenic properties. Additionally, we report this protein has large patches of disordered regions flanking with these seven motifs. These motifs are hubs of epitopes with 67 experimentally verified epitopes from related viruses. We report the presence of three nuclear localization signals (NLS1-NLS3 mapped to 36-41, 256-26, and 363-389 residues, respectively) and two nuclear export signals (NES1-NLS2 from 151-161 and 217-230 residues, respectively) in the N protein of SARS-CoV-2. These deciphered two Q-patches as Q-patch1 and Q-patch2, mapped in the regions of 266-306, and 361-418 residues, which potentially help in the aggregation of the viral proteins along with 219LALLLLDR226 patch. Additionally, we have identified 14 antiviral drugs potentially binding to seven motifs of N-proteins using docking-based drug discovery methods.