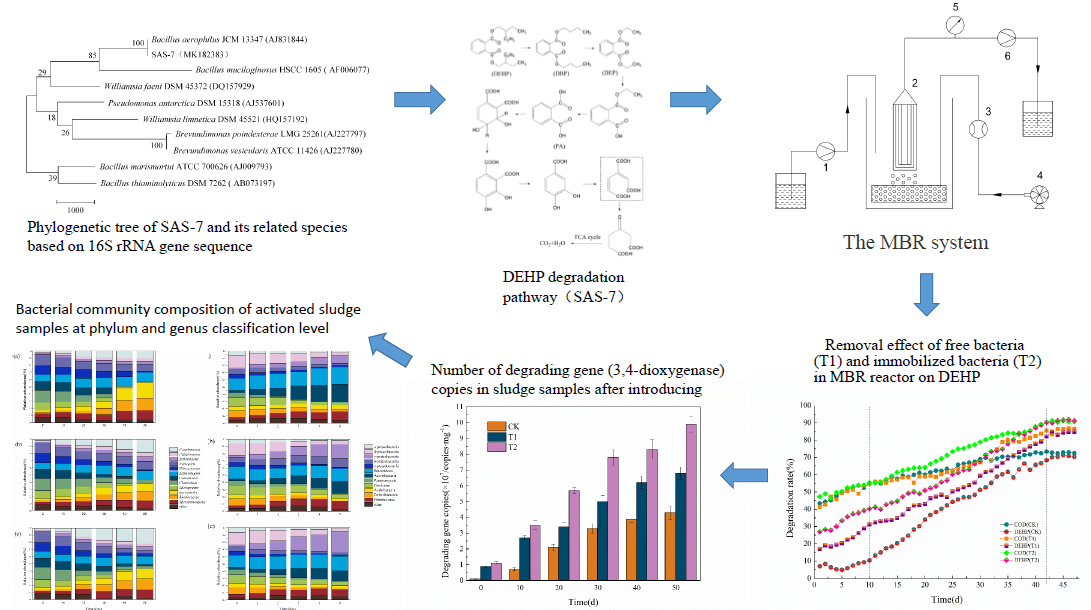
A bacterial strain that could effectively degrade DEHP was isolated from the activated sludge and identified as Bacillus sp. by DNA sequencing. The biochemical degradation pathway of DEHP was further analyzed by GC-MS, and the results showed that DEHP was first decomposed into phthalates (DBP). Diuretic sylycol (DEP) was then generated, and phthalates (PA) were generated by a continuous de-ehelateization reaction. Phthalic acid (PA) was oxidized, dehydrogenated, and decarboxylated into protocatechins. Protocatechins enter the TCA cycle through orthotopic ring opening. To enhance DEHP degradation, sodium alginate and calcium chloride were used as embedding and cross-linking materials, and the strain was immobilized. The immobilization conditions were optimized via an orthogonal experiment, and the results showed that the optimal immobilization conditions were SA mass fraction of 4%, CaCl2 mass fraction of 5%, ratio of bacteria to SA of 1:1, and the crosslinking time of 6 hours. The immobilized bacteria agent was further applied to MBR systems. The results showed that the removal rate of DEHP (5mg/L) in the system by immobilized bacteria was 91.9%, which is significantly higher than that of free bacteria. The 3, 4-dioxygenase gene and microbial community dynamics were analyzed by q-PCR and Illumina Miseq sequencing. The q-PCR results showed that the number of copies of 3, 4-dioxygenase gene in the immobilized system was significantly higher than that of free bacteria. Illumina Miseq sequencing results showed that Micromonospora, Rhodococcus, Bacteroides and Pseudomonas were the dominant generas in the MBR system. The analysis of bacterial community structure indicated that immobilization technology had a positive impact on the system stability. The results implied that this immobilized technique had potential applications in DEHP wastewater treatment.