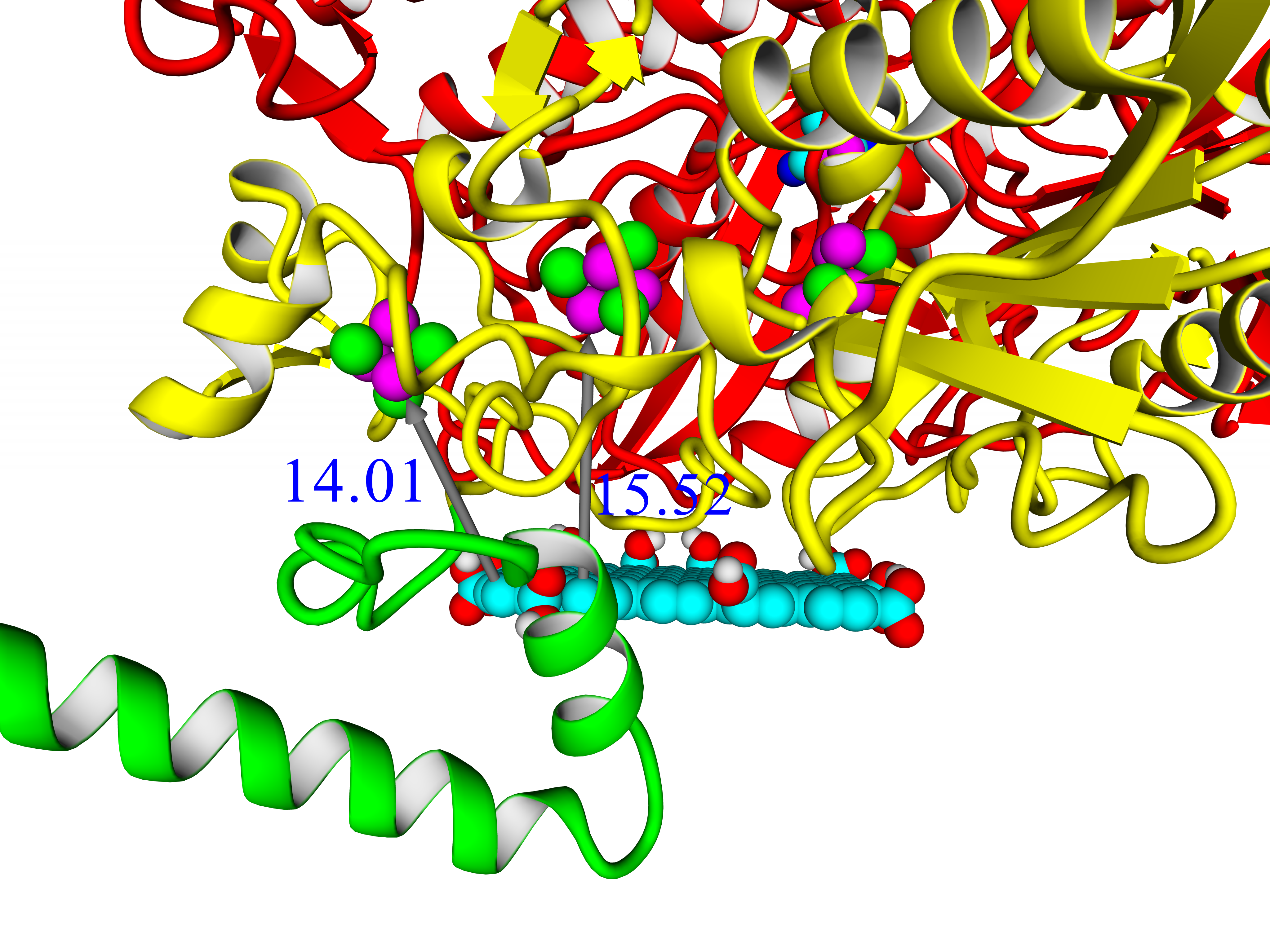
Three-dimensional structure of six closely related hydrogenases from purple bacteria has been modeled by combining template-based and ab initio modeling approach. The results lead to conclusion that there should be 4Fe3S-cluster in the structure of these enzymes. Thus, these hydrogenases could drive attention for exploring their oxygen tolerance and practical applicability in hydrogen fuel cells. Analysis of 4Fe3S-cluster's microenvironment showed intragroup heterogeneity. Possible function of the C-terminal part of the small subunit in membrane binding has been discussed. Comparison of the built models with existing hydrogenases of the same subgroup (membrane-bound oxygen-tolerant hydrogenases) has been carried out. Analysis of intramolecular interactions in the large subunits showed statistically reliable differences in number of hydrophobic interactions and number of ionic interactions. Molecular tunnels were mapped in the models and compared with structures from PDB. Protein-protein docking showed that these enzymes could exchange electrons in oligomeric state, which is important for oxygen-tolerant hydrogenases. Molecular docking with model electrode compounds showed mostly the same results as with hydrogenases from E.coli, H. marinus, R. eutropha, S. enterica; some interesting results were shown in case of HupSL from Rba. sphaeroides and Rvx. gelatinosus.