Submitted:
01 May 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- 1)
- Threshold Arbitrariness: Choosing a threshold to binarize the network is subjective. A high threshold may fracture the network, losing weak but biologically significant connections, while a low threshold may introduce noise.
- 2)
- Scale Sensitivity: Brain networks exhibit multiscale organization. Fixed-scale graph metrics (e.g., path length, clustering coefficient) fail to capture the hierarchical topology of neural degeneration.
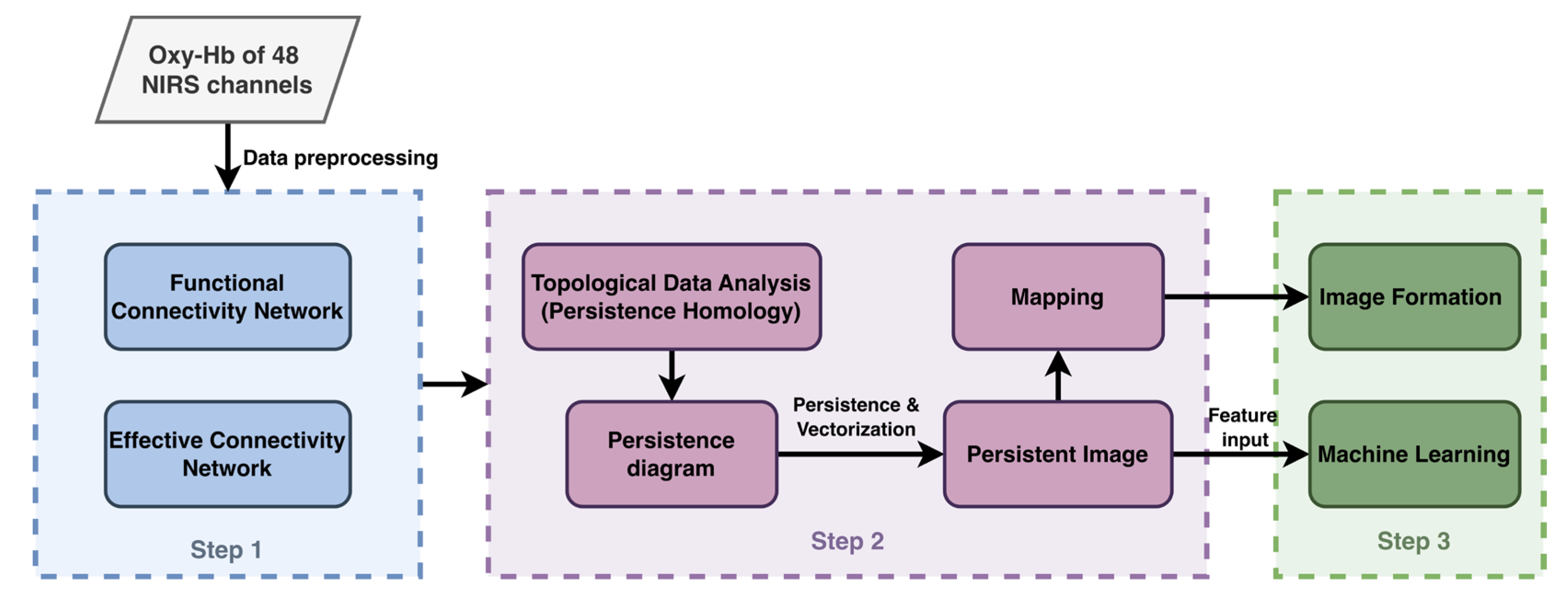
- Transforms raw fNIRS time-series into Effective Connectivity networks using Granger Causality, capturing the directionality of information flow.
- Extracts topological invariants (Betti numbers) via Persistent Homology, identifying stable structures (clusters, loops and higher-order voids) that persist across scales.
- Vectorizes these abstract topological features into Persistence Images (PIs), creating a stable, fixed-dimensional input for machine learning classifiers.
2. Literature Review
2.1. Machine Learning in AD Diagnosis
2.2. fNIRS in Cognitive Neuroscience
2.3. Topological Data Analysis (TDA) in Medicine
3. Materials & Methodology
3.1. Participants and Experimental Protocol
3.1.1. Dataset Balancing via Augmentation
3.1.2. Hyperparameter Optimization
3.1.3. Evaluation Metrics
3.2. fNIRS Data Acquisition and Detailed Preprocessing
- 1)
- Optical Density Conversion
- 2)
- Motion Artifact Correction (Moving Standard Deviation):
- 3)
- Band-pass Filtering:
- 4)
- Modified Beer-Lambert Law (MBLL):
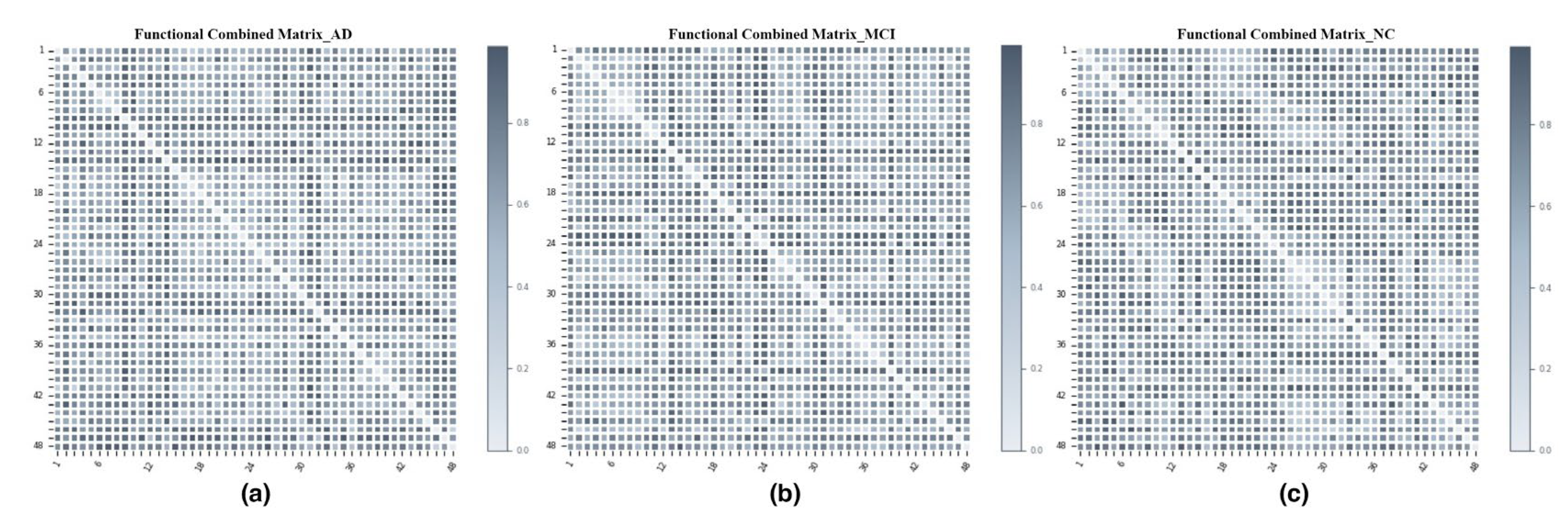
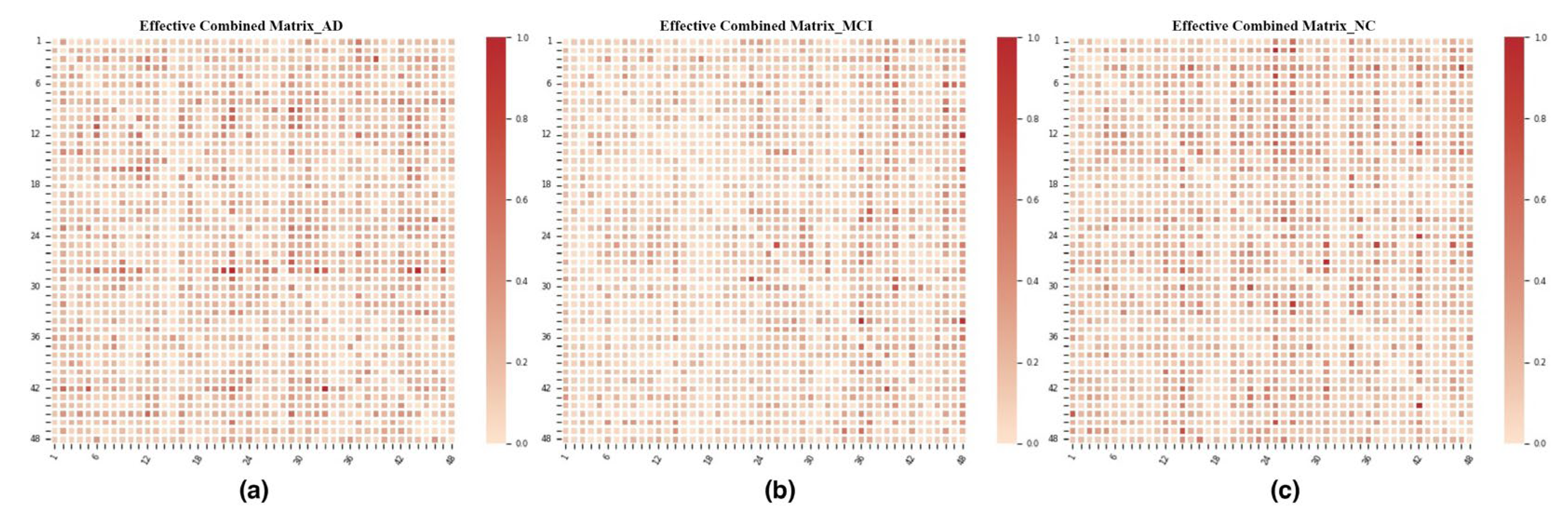
3.3. Metric Space Construction: Connectivity Networks
3.4. Topological Data Analysis: Persistent Homology
3.4.1. Mathematical Foundations: From Simplices to Homology Groups
3.4.2. The Vietoris-Rips Filtration
- 1)
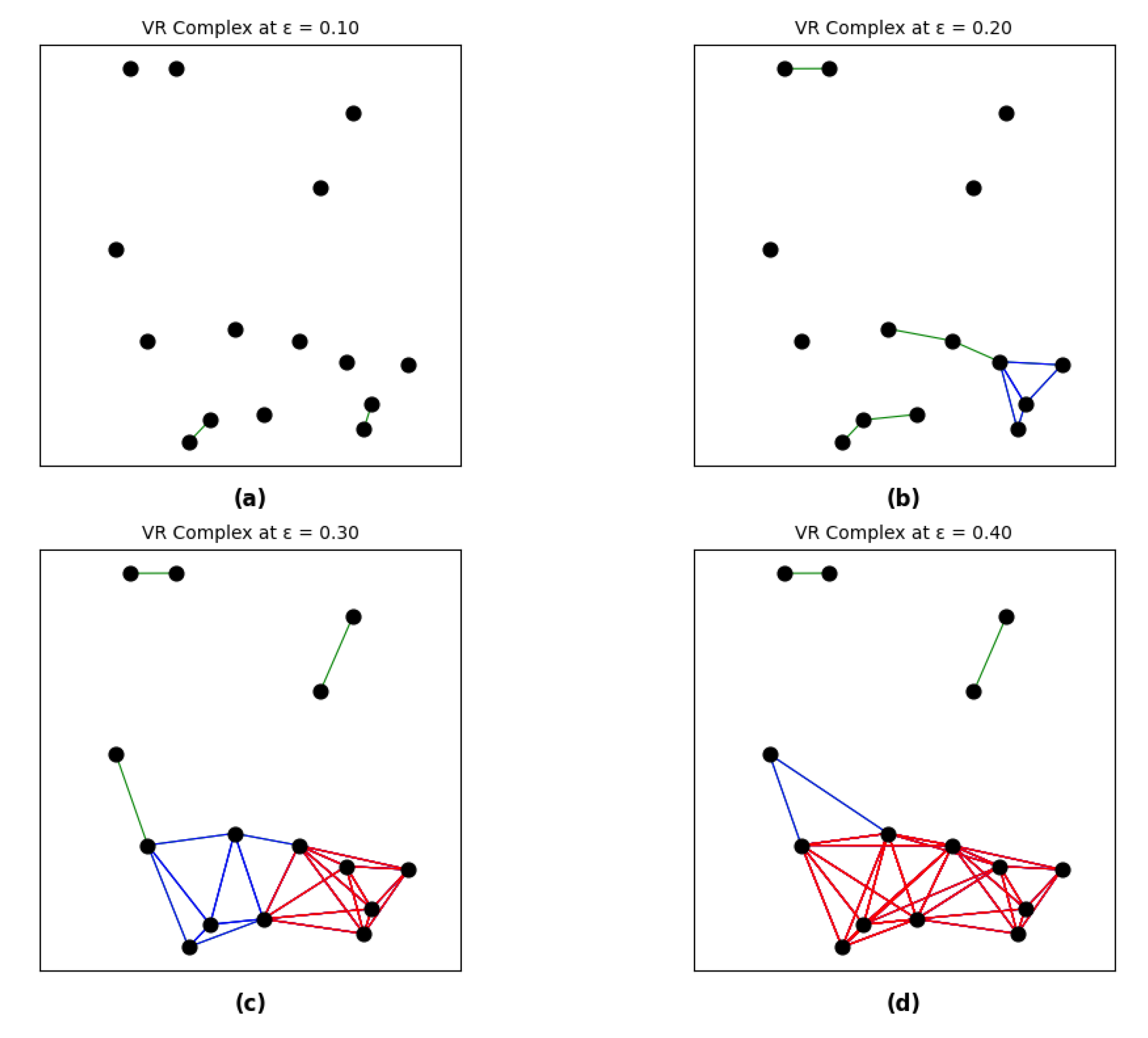
- At (or low , e.g., = 0.10 in Figure 4a): The network starts as a cloud of disconnected points. In our study, this corresponds to and, where no functional integration has yet occurred.
- 2)
- As increases (e.g., = 0.20 to 0.30 in Figure 4b and c): Edges begin to form between channels with strong causal links. This process leads to the merging of connected components (death of H0 features) and the formation of cyclic pathways or loops (birth of H1 features), representing the emergence of local information processing clusters.
- 3)
- At sufficiently large (e.g., = 0.40 in Figure 4d): Most loops become "filled" by triangles or higher-order simplices (death of H1), and subsequent voids become "filled" by tetrahedrons or higher-order simplices (death of H2). In this stage, the network transitions into a single giant component, reflecting a state of global integration.
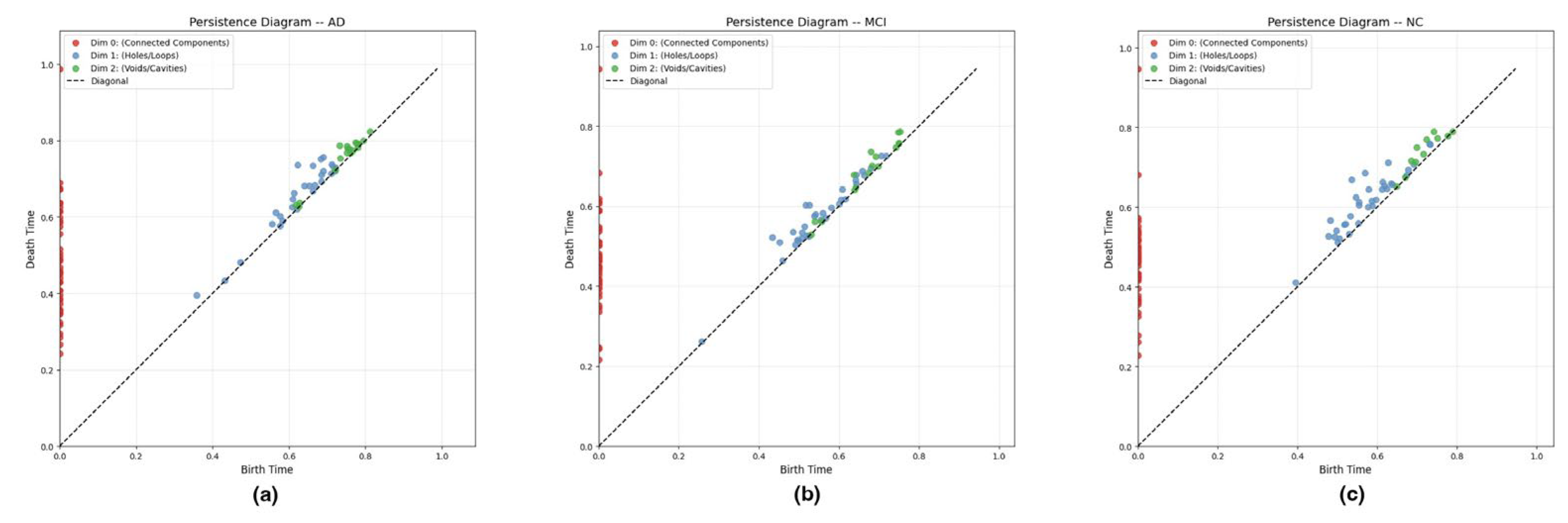
3.4.3. Persistence Diagrams and Stability Theory
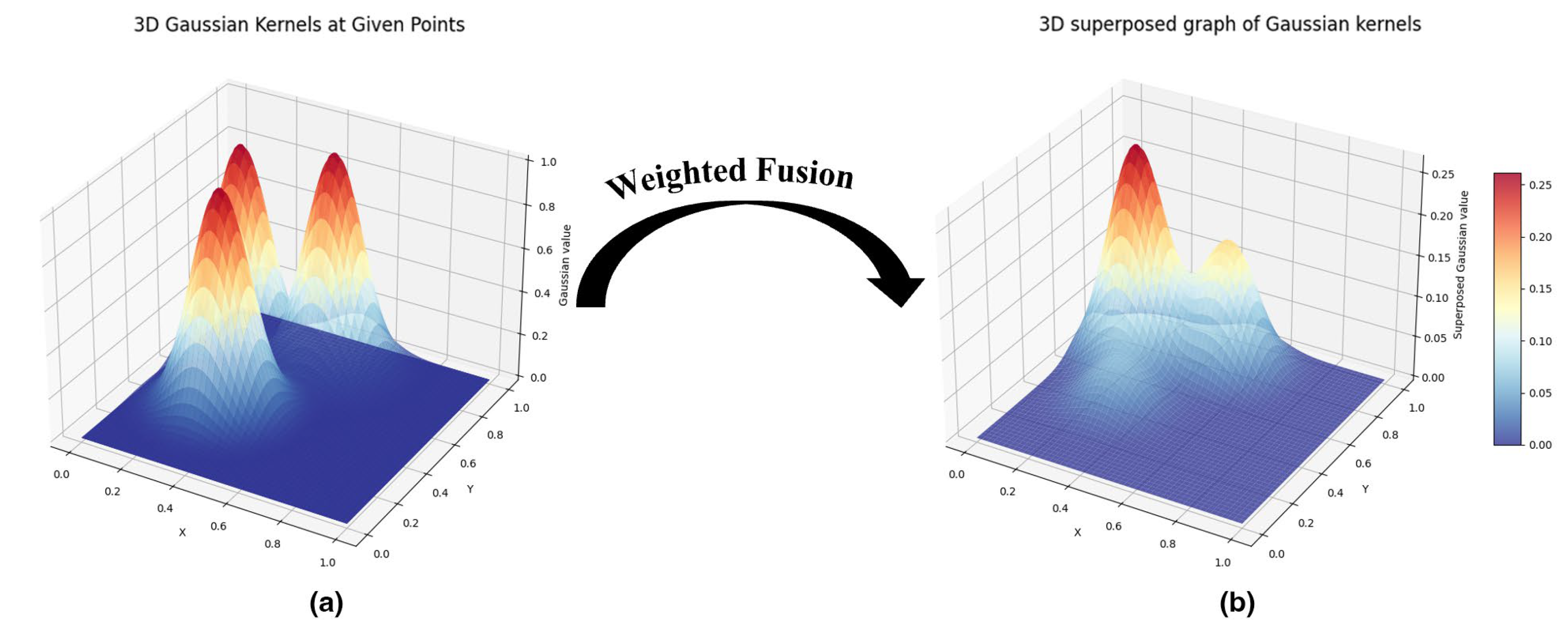
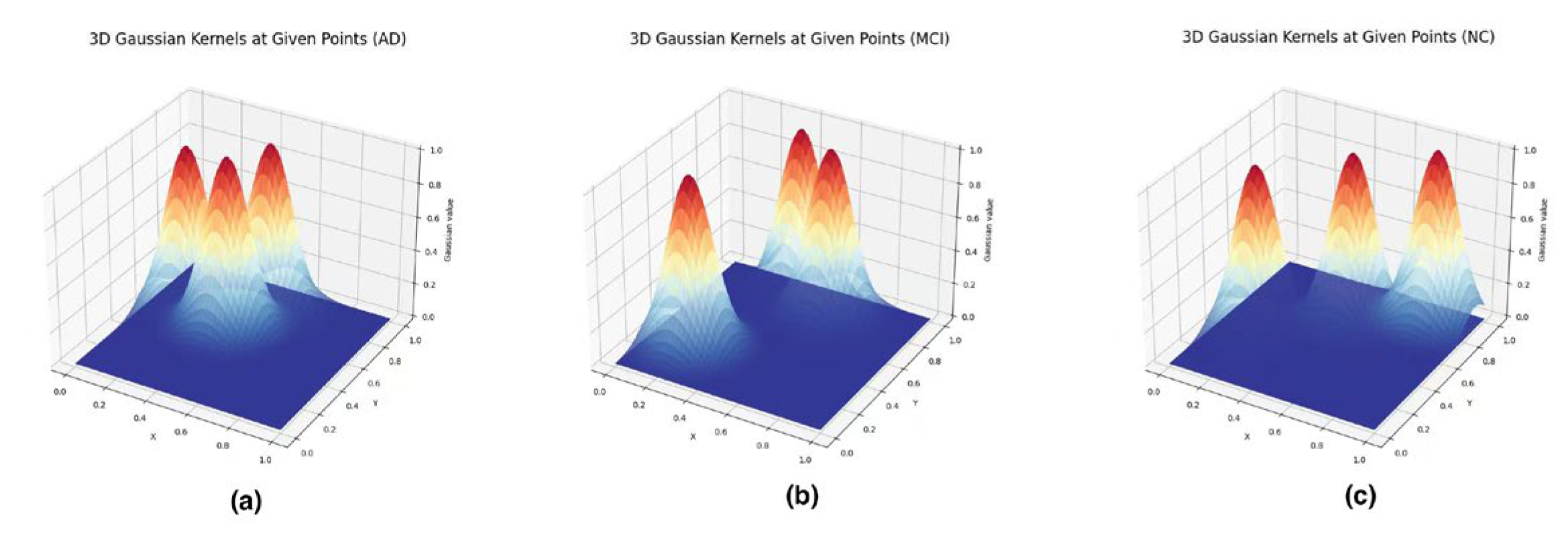
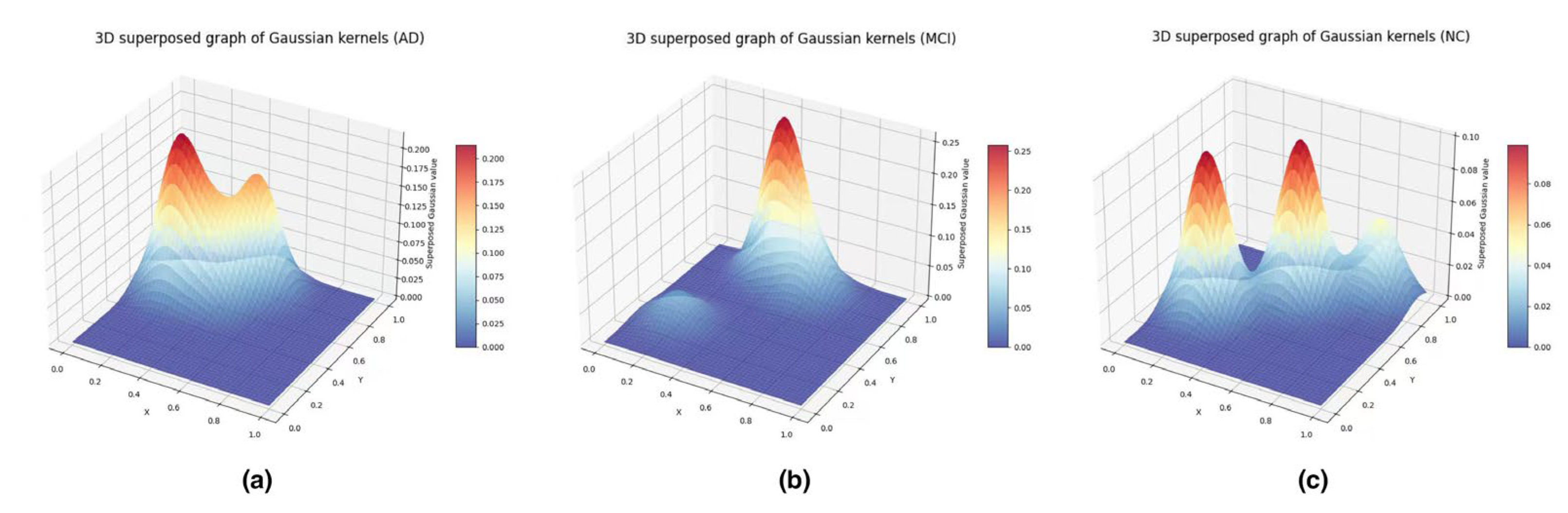
3.4.4. Feature Vectorization: Persistence Images
3.4.5. Machine Learning Classification Framework
- 1)
- Base Classifiers: The ensemble aggregates predictions from three robust algorithms:
- 2)
- Ensemble Strategy: We utilized a Soft Voting mechanism. For a given input sample x, each base classifier outputs a probability distribution over the classes . The final prediction is the class with the highest average probability:
4. Results
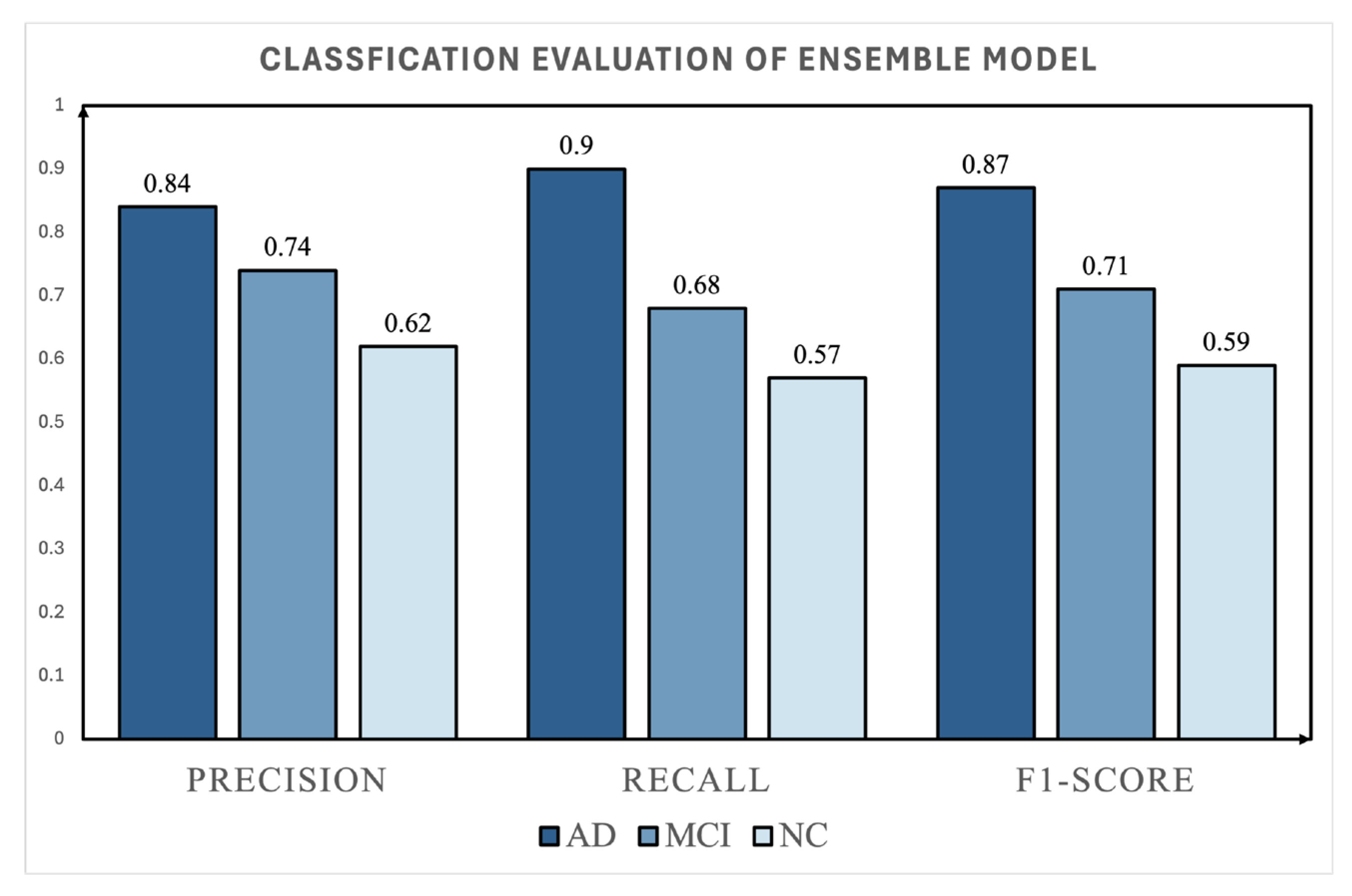
4.1. Ensemble Model Performance
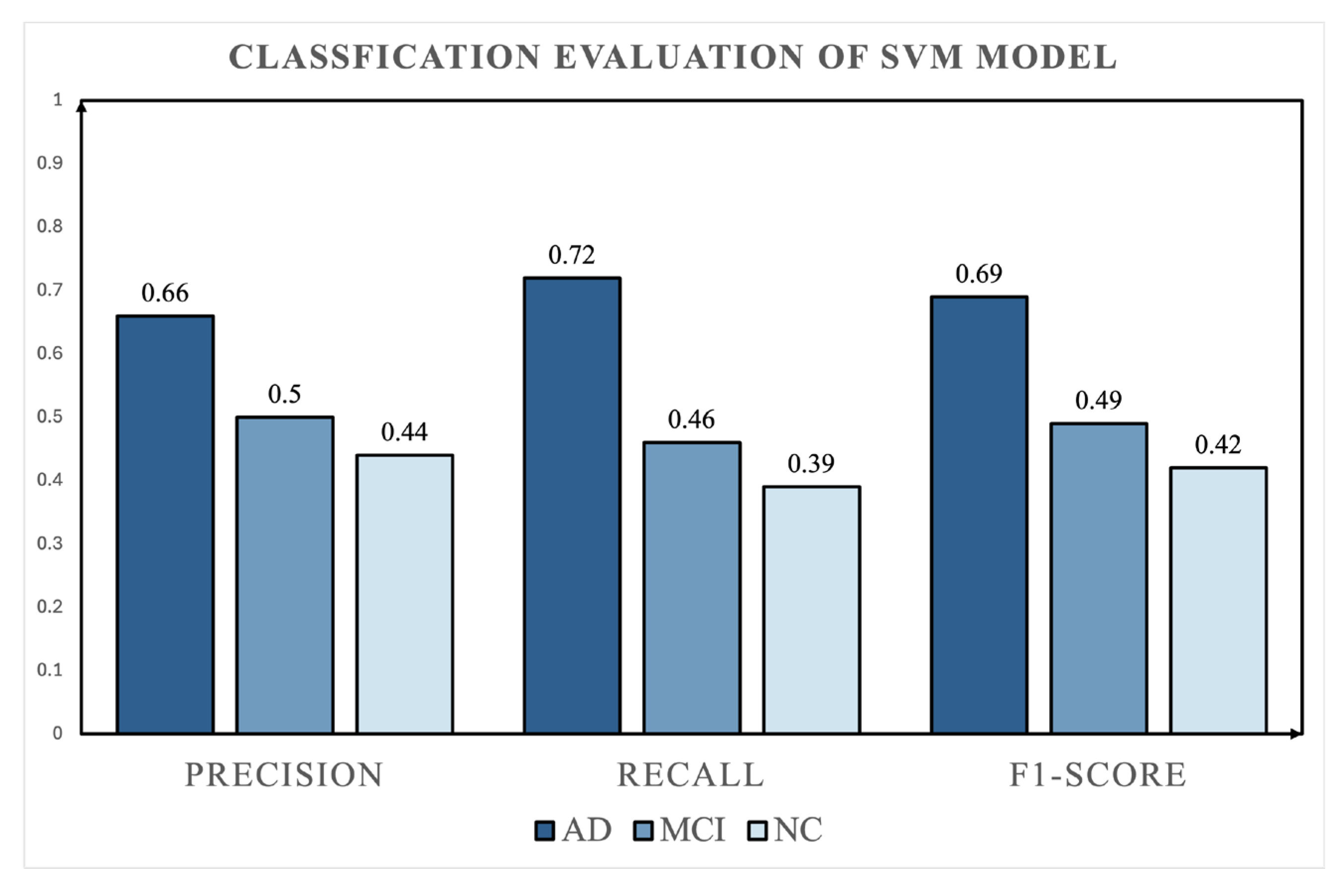
4.2. Baseline Comparison
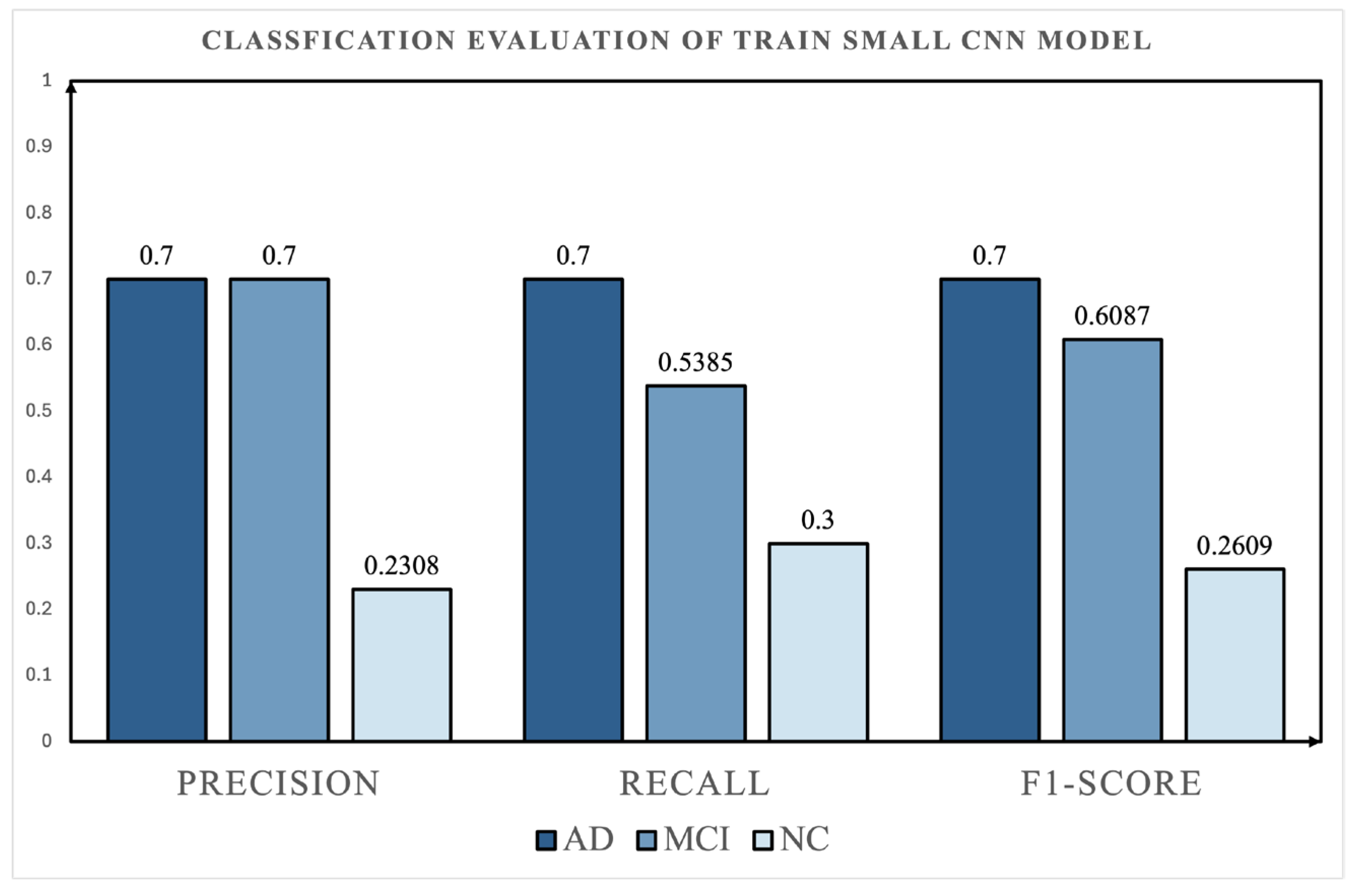
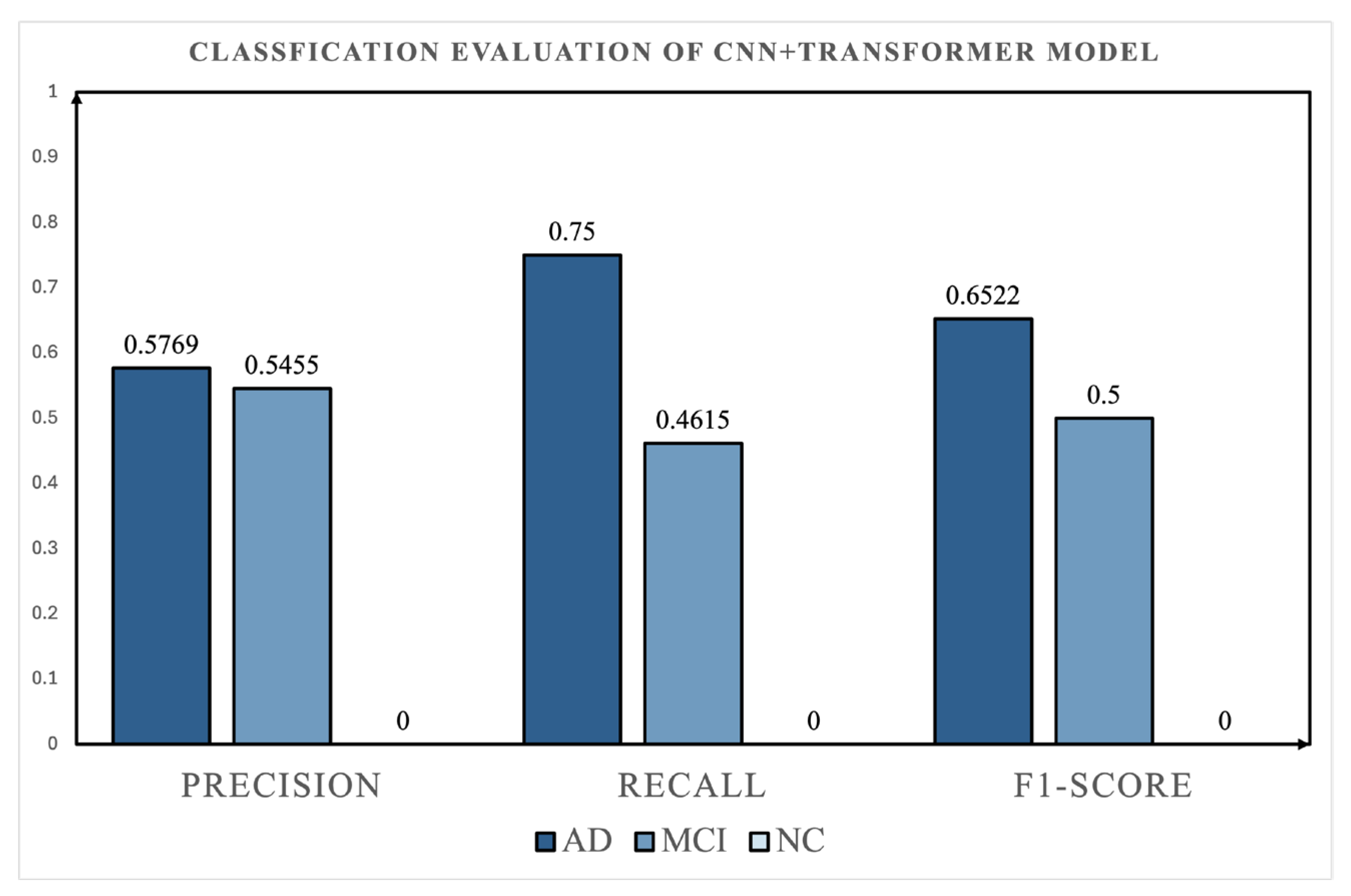
4.3. Comparison with Deep Learning Baselines
5. Discussion
5.1. Comparative Advantage of Topology-Based Representation
5.2. Ablation Discussion
- 1)
- Connectivity Metric:
- 2)
- PI Resolution:
5.3. Limitations and Future Work
6. Conclusions
- 1)
- Topological Biomarkers: The Persistence Images revealed that AD brains exhibit a "topological simplification"—a loss of high-persistence loops and an increase in fragmented components. This aligns with the disconnection hypothesis.
- 2)
- Methodological Robustness: Our pipeline avoids the critical pitfall of arbitrary thresholding in network neuroscience. The use of Granger Causality further enhanced the model by incorporating directional coupling.
- 3)
- Clinical Potential: With an accuracy of nearly 77% using only 48 fNIRS channels, this approach offers a low-cost, portable screening tool that could be deployed in community clinics, unlike MRI or PET.
- 4)
- Comparative Validation with Deep Learning Baselines: Supplementary experiments using a Small CNN and a CNN+Transformer model showed that direct temporal modeling did not outperform the proposed topology-based ensemble framework under the current setting. In particular, the CNN+Transformer model exhibited poorer class balance and weak NC recognition, further highlighting the robustness of the proposed TDA-based representation.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nawaz, A.; Anwar, S.M.; Liaqat, R.; Iqbal, J.; Bagci, U.; Majid, M. Deep Convolutional Neural Network Based Classification of Alzheimer’s Disease Using MRI Data. 2020 IEEE 23rd International Multitopic Conference (INMIC), Bahawalpur, Pakistan, 2020; pp. 1–6. [Google Scholar] [CrossRef]
- Cai, H.; Sheng, X.; Wu, G.; Hu, B.; Cheung, Y.-M.; Chen, J. Brain Network Classification for Accurate Detection of Alzheimer’s Disease via Manifold Harmonic Discriminant Analysis. IEEE Trans. Neural Netw. Learn. Syst. 2024, 35, 17266–17280. [Google Scholar] [CrossRef] [PubMed]
- Bhandarkar, A.; Naik, P.; Vakkund, K.; Junjappanavar, S.; Bakare, S.; Pattar, S. Deep Learning Based Computer Aided Diagnosis of Alzheimer’s Disease: A Snapshot of Last 5 Years, Gaps, and Future Directions. Artif. Intell. Rev. 2024, 57, 30. [Google Scholar] [CrossRef]
- Aisen, P.S.; Cummings, J.; Jack, C.R., Jr.; Morris, J.C.; Sperling, R.; Frölich, L.; Jones, R.W.; Dowsett, S.A.; Matthews, B.R.; Raskin, J.; Scheltens, P.; Dubois, B. On the Path to 2025: Understanding the Alzheimer’s Disease Continuum. Alzheimers Res. Ther. 2017, 9, 60. [Google Scholar] [CrossRef] [PubMed]
- Faux, N.G.; Rembach, A.; Wiley, J.; Ellis, K.A.; Ames, D.; Fowler, C.J.; Martins, R.N.; Pertile, K.K.; Rumble, R.L.; Trounson, B.; Masters, C.L.; Bush, A.I.; The AIBL Research Group. An Anemia of Alzheimer’s Disease. Mol. Psychiatry 2014, 19, 1227–1234. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Popuri, K.; Ding, G.W.; Balachandar, R.; Beg, M.F. Alzheimer’s Disease Neuroimaging Initiative. Multimodal and Multiscale Deep Neural Networks for the Early Diagnosis of Alzheimer’s Disease Using Structural MR and FDG-PET Images. Sci. Rep. 2018, 8, 5697. [Google Scholar] [CrossRef] [PubMed]
- Pinti, P.; Tachtsidis, I.; Hamilton, A.; Hirsch, J.; Aichelburg, C.; Gilbert, S.; Burgess, P.W. The Present and Future Use of Functional Near-Infrared Spectroscopy (fNIRS) for Cognitive Neuroscience. Ann. N. Y. Acad. Sci. 2020, 1464, 5–29. [Google Scholar] [CrossRef] [PubMed]
- Xie, L.; Liu, Y.; Gao, Y.; Zhou, J. Functional Near-Infrared Spectroscopy in Neurodegenerative Disease: A Review. Front. Neurosci. 2024, 18, 1469903. [Google Scholar] [CrossRef] [PubMed]
- Pinti, P.; Tachtsidis, I.; Hamilton, A.; Hirsch, J.; Aichelburg, C.; Gilbert, S.; Burgess, P.W. The Present and Future Use of Functional Near-Infrared Spectroscopy (fNIRS) for Cognitive Neuroscience. Ann. N. Y. Acad. Sci. 2020, 1464, 5–29. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, I.; Gómez, A.; Carmona, M.; Huesa, G.; Porta, S.; Riera-Codina, M.; Biagioli, M.; Gustincich, S.; Aso, E. Neuronal Hemoglobin Is Reduced in Alzheimer’s Disease, Argyrophilic Grain Disease, Parkinson’s Disease, and Dementia with Lewy Bodies. J. Alzheimers Dis. 2011, 23, 537–550. [Google Scholar] [CrossRef] [PubMed]
- Ali, D.; Asaad, A.; Jimenez, M.-J.; Nanda, V.; Paluzo-Hidalgo, E.; Soriano-Trigueros, M. A Survey of Vectorization Methods in Topological Data Analysis. IEEE Trans. Pattern Anal. Mach. Intell. 2023, 45, 14069–14080. [Google Scholar] [CrossRef] [PubMed]
- Bhandarkar, A.; Naik, P.; Vakkund, K.; Junjappanavar, S.; Bakare, S.; Pattar, S. Deep Learning Based Computer Aided Diagnosis of Alzheimer’s Disease: A Snapshot of Last 5 Years, Gaps, and Future Directions. Artif. Intell. Rev. 2024, 57, 30. [Google Scholar] [CrossRef]
- Suk, H.-I.; Shen, D. Deep Learning-Based Feature Representation for AD/MCI Classification. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2013; Lecture Notes in Computer Science; Mori, K., Sakuma, I., Sato, Y., Barillot, C., Navab, N., Eds.; Springer: Berlin, Heidelberg, 2013; Vol. 8150, pp. 583–590. [Google Scholar] [CrossRef]
- Klöppel, S.; Stonnington, C.M.; Chu, C.; Draganski, B.; Scahill, R.I.; Rohrer, J.D.; Fox, N.C.; Jack, C.R.; Ashburner, J.; Frackowiak, R.S.J. Automatic Classification of MR Scans in Alzheimer’s Disease. Brain 2008, 131, 681–689. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liu, S.; Cai, W.; Pujol, S.; Kikinis, R.; Feng, D. Early Diagnosis of Alzheimer’s Disease with Deep Learning. 2014 IEEE 11th International Symposium on Biomedical Imaging (ISBI), Beijing, China, 2014; pp. 1015–1018. [Google Scholar] [CrossRef]
- Korolev, S.; Safiullin, A.; Belyaev, M.; Dodonova, Y. Residual and Plain Convolutional Neural Networks for 3D Brain MRI Classification. 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), Melbourne, VIC, Australia, 2017; pp. 835–838. [Google Scholar] [CrossRef]
- Hazarika, R.A.; Kandar, D.; Maji, A.K. A Novel Machine Learning Based Technique for Classification of Early-Stage Alzheimer’s Disease Using Brain Images. Multimed. Tools Appl. 2024, 83, 24277–24299. [Google Scholar] [CrossRef]
- Suk, H.-I.; Lee, S.-W.; Shen, D. the Alzheimer’s Disease Neuroimaging Initiative. Hierarchical Feature Representation and Multimodal Fusion with Deep Learning for AD/MCI Diagnosis. NeuroImage 2014, 101, 569–582. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liu, S.; Cai, W.; Che, H.; Pujol, S.; Kikinis, R.; Feng, D.; Fulham, M.J.; ADNI. Multimodal Neuroimaging Feature Learning for Multiclass Diagnosis of Alzheimer’s Disease. IEEE Trans. Biomed. Eng. 2015, 62, 1132–1140. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Zhong, S.; Zhang, Y. Multi-Level Fusion Network for Mild Cognitive Impairment Identification Using Multi-Modal Neuroimages. Phys. Med. Biol. 2023, 68, 095018. [Google Scholar] [CrossRef] [PubMed]
- Rao, K.N.; Gandhi, B.R.; Rao, M.V.; Javvadi, S.; Vellela, S.S.; Basha, S.K. Prediction and Classification of Alzheimer’s Disease Using Machine Learning Techniques in 3D MR Images. 2023 International Conference on Sustainable Computing and Smart Systems (ICSCSS), Coimbatore, India, 2023; pp. 85–90. [Google Scholar] [CrossRef]
| Classes | Total volumes | Testing set size | Precision | Recall | F1-Score |
| AD | 325 | 65 | 0.84 | 0.90 | 0.87 |
| MCI | 185 | 37 | 0.74 | 0.68 | 0.71 |
| NC | 140 | 28 | 0.62 | 0.57 | 0.59 |
| Total | 650 | 130 | 0.77 | — | — |
| Classes | Total volumes | Testing set size | Precision | Recall | F1-Score |
| AD | 325 | 65 | 0.66 | 0.72 | 0.69 |
| MCI | 185 | 37 | 0.50 | 0.46 | 0.49 |
| NC | 140 | 28 | 0.44 | 0.39 | 0.42 |
| Total | 650 | 130 | 0.58 | — | — |
| Method | Classes | Precision | Recall | F1-Score |
| Ensemble Model | AD | 0.84 | 0.90 | 0.87 |
| MCI | 0.74 | 0.68 | 0.71 | |
| NC Total |
0.62 0.7692 |
0.57 — |
0.59 — |
|
| Small CNN | AD | 0.70 | 0.70 | 0.70 |
| MCI | 0.70 | 0.54 | 0.61 | |
| NC Total |
0.23 0.5436 |
0.30 — |
0.26 — |
|
| CNN+Transformer | AD | 0.58 | 0.75 | 0.65 |
| MCI | 0.55 | 0.46 | 0.50 | |
| NC Total |
0.00 0.3741 |
0.00 — |
0.00 — |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




