Submitted:
30 April 2026
Posted:
01 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- 1.
- We design a lightweight diagnostic framework based on ELM that uses readily collected demographic and clinical features, aiming to enable practical, scalable screening for OSA and SDB in resource-constrained settings.
- 2.
- We systematically integrate eleven metaheuristic algorithms covering evolutionary, math-based, physics-based, and swarm-based families to optimize ELM hidden-layer weights and biases under a common objective and protocol.
- 3.
- We perform an extensive empirical study on two real datasets, benchmarking all metaheuristic-optimized ELM variants against standard ML baselines in terms of accuracy, F1-score, ROC-AUC, generalization gap, convergence behaviour, and computational time.
- 4.
- We show that metaheuristic-optimized ELM models achieve consistent improvements over plain ELM and several baselines on the OSA dataset and provide moderate but meaningful gains in discrimination on the more imbalanced SDB dataset.
2. Related Works
2.1. OSA Detection Using Demographic and Clinical Data
2.2. Machine Learning and Deep Learning Models in OSA
2.3. Metaheuristic-Based Optimization of ELM
3. Materials and Methods
3.1. Dataset Description
3.1.1. Obstructive Sleep Apnea (OSA) Dataset
3.1.2. Sleep-Disordered Breathing (SDB) Detection Dataset
| Dataset | No. samples | No. features | Negative samples | Positive samples | Positive rate (%) |
|---|---|---|---|---|---|
| OSA | 274 | 31 | 149 | 125 | 45.6 |
| SDB | 500 | 10 | 119 | 381 | 76.2 |
3.1.3. Summary and Feature Characteristics
3.2. Data Processing
3.2.1. OSA Dataset Preprocessing
| Feature | Type | In OSA | In SDB | Description |
|---|---|---|---|---|
| Race | Categorical | Yes | No | Race / ethnic group category. |
| Age | Numeric (continuous) | Yes | Yes | Patient age in years. |
| Sex | Categorical (binary) | Yes | Yes | Patient sex. |
| BMI* | Numeric / categorical | Yes | Yes | Body mass index or BMI category. |
| Epworth | Numeric (ordinal) | Yes | No | Epworth Sleepiness Scale total score. |
| Wast | Numeric (continuous) | Yes | No | Waist circumference (cm). |
| Hip | Numeric (continuous) | Yes | No | Hip circumference (cm). |
| RDI | Numeric (continuous) | Yes | No | Respiratory Disturbance Index per hour. |
| Neck | Numeric (continuous) | Yes | No | Neck circumference (cm). |
| M.Friedman | Ordinal categorical | Yes | No | Friedman tongue position grade (1–4). |
| Co-morbid | Categorical | Yes | No | Presence or count of comorbidities. |
| Snoring | Categorical (binary) | Yes | Yes | Snoring indicator. |
| Daytime sleepiness | Categorical (binary) | Yes | No | Self-reported daytime sleepiness. |
| DM | Categorical (binary) | Yes | No | Diabetes mellitus status. |
| HTN | Categorical (binary) | Yes | No | Hypertension status. |
| CAD | Categorical (binary) | Yes | No | Coronary artery disease status. |
| CVA | Categorical (binary) | Yes | No | History of cerebrovascular accident. |
| TST | Numeric (continuous) | Yes | No | Total sleep time. |
| Sleep Effic | Numeric (continuous) | Yes | No | Sleep efficiency (%). |
| REM AHI | Numeric (continuous) | Yes | No | Apnea–Hypopnea Index in REM sleep. |
| NREM AHI | Numeric (continuous) | Yes | No | Apnea–Hypopnea Index in NREM sleep. |
| Supine AHI | Numeric (continuous) | Yes | No | Apnea–Hypopnea Index in supine position. |
| Apnea Index | Numeric (continuous) | Yes | No | Number of apnea events per hour. |
| Hypopnea Index | Numeric (continuous) | Yes | No | Number of hypopnea events per hour. |
| Berlin Q | Categorical | Yes | No | Berlin questionnaire risk category. |
| Arousal index | Numeric (continuous) | Yes | No | Number of arousals per hour. |
| Awakening Index | Numeric (continuous) | Yes | No | Number of awakenings per hour. |
| PLM Index | Numeric (continuous) | Yes | No | Periodic limb movement index per hour. |
| Mins.SaO2 | Numeric (continuous) | Yes | No | Minutes below oxygen saturation threshold. |
| Mins.SaO2Desats | Numeric (continuous) | Yes | No | Minutes with oxygen desaturation events. |
| Lowest Sa02 | Numeric (continuous) | Yes | No | Lowest oxygen saturation recorded. |
| Oxygen_Saturation | Numeric (continuous) | No | Yes | Average oxygen saturation during recording. |
| AHI | Numeric (continuous) | No | Yes | Overall Apnea–Hypopnea Index. |
| ECG_Heart_Rate | Numeric (continuous) | No | Yes | Heart rate derived from ECG. |
| SpO2 | Numeric (continuous) | No | Yes | Average peripheral oxygen saturation (%). |
| Nasal_Airflow | Numeric (continuous) | No | Yes | Normalized nasal airflow signal. |
| Chest_Movement | Numeric (continuous) | No | Yes | Normalized chest movement signal. |
3.2.2. SDB Dataset Preprocessing
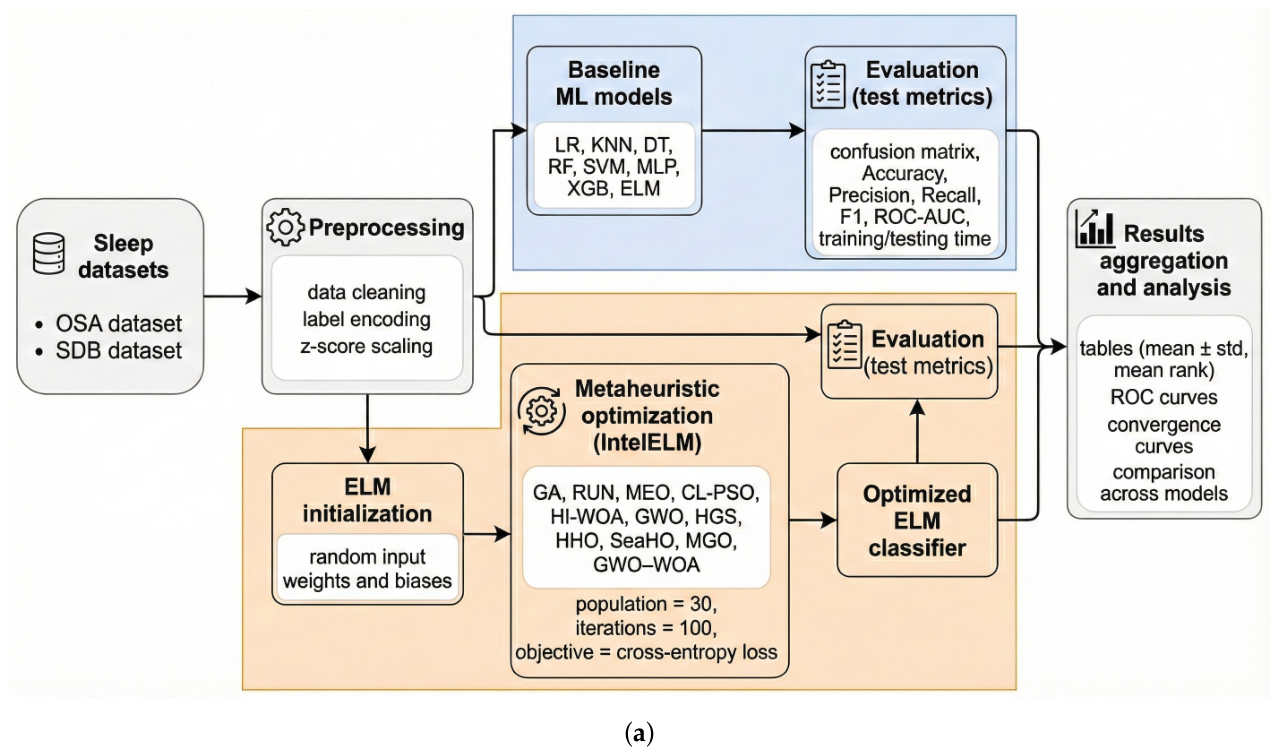
3.3. Proposed Optimized-ELM Framework
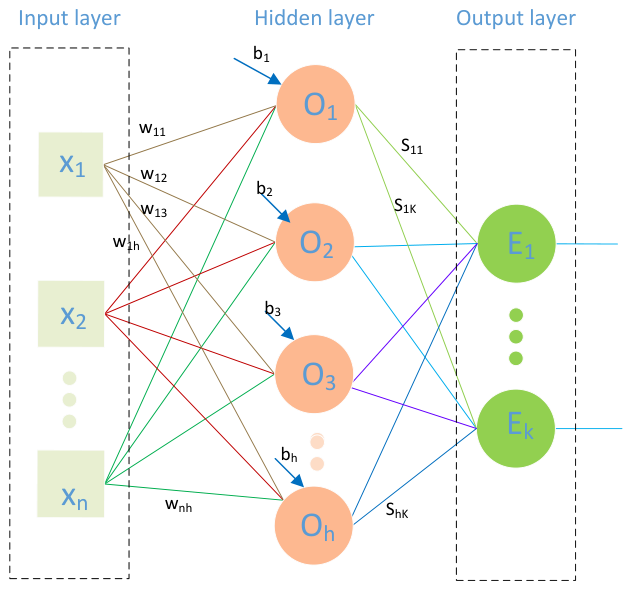
3.3.1. Basic ELM Classifier and Mathematical Formulation
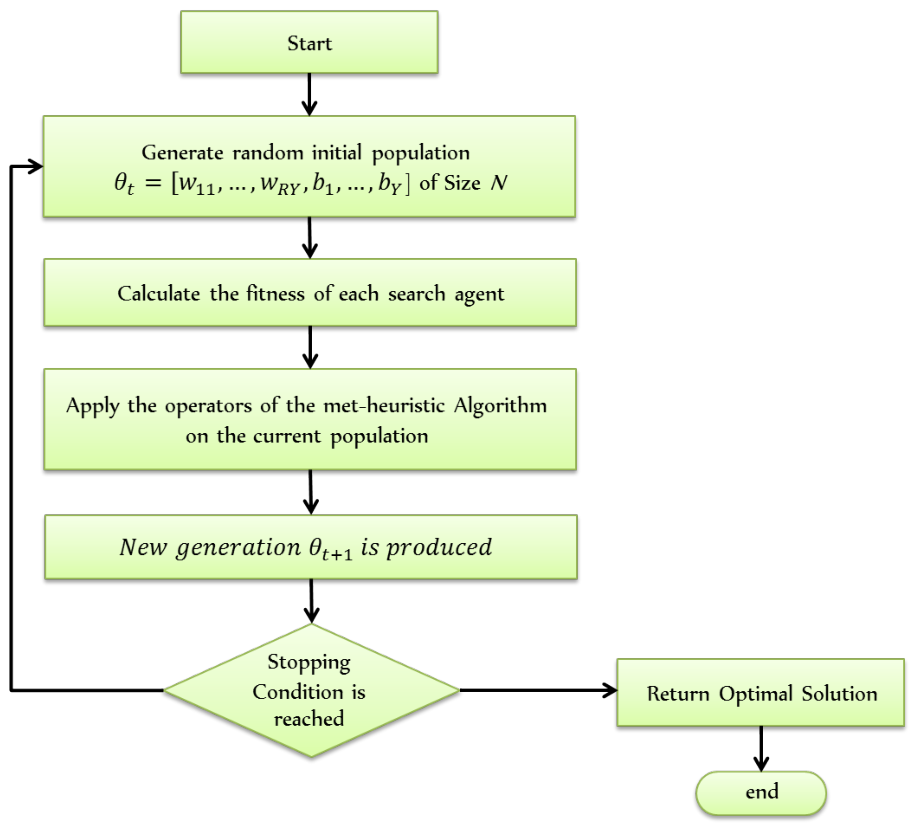
3.3.2. Optimization Methodology
Solution Encoding
Objective Function
3.3.3. Integration of Metaheuristics with ELM
3.4. Experimental Setup and Evaluation Protocol
3.4.1. Data Splitting and Repeated Experiments
3.4.2. Baseline Classifiers and Hyperparameters
3.4.3. Metaheuristic-Optimized ELM Configuration
3.4.4. Evaluation Measures
Confusion matrix for binary diagnosis
- Positive class: patient with obstructive sleep apnea or sleep-disordered breathing (OSA/SDB present).
- Negative class: patient without OSA/SDB (no sleep apnea).
- True Positive (TP): correctly predicted OSA/SDB cases.
- True Negative (TN): correctly predicted non-OSA/non-SDB cases.
- False Positive (FP): predicted OSA/SDB, but the patient is actually non-OSA/non-SDB.
- False Negative (FN): predicted non-OSA/non-SDB, but the patient actually has OSA/SDB.
Classification Quality Metrics
Computational Time
3.4.5. Environment and Tools
4. Experimental Results
4.1. Results and Analysis of Baseline Models
4.1.1. Results on OSA Dataset
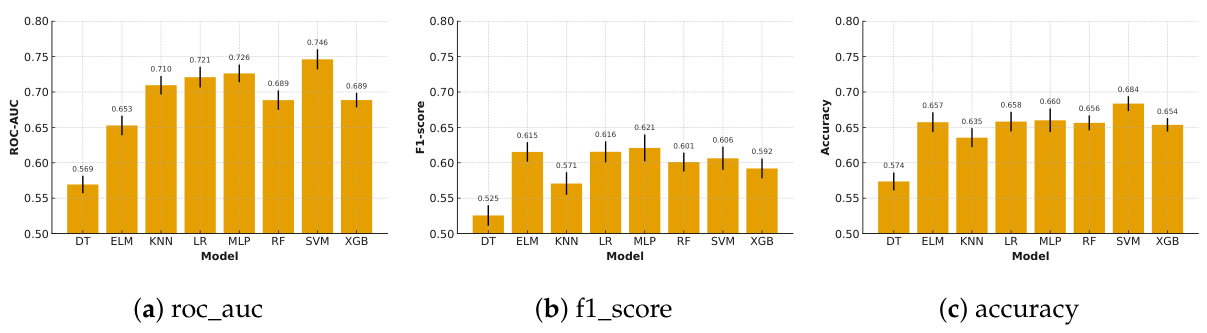
Testing Performance
| model | Measure | accuracy | precision | recall | f1 | roc_auc | mean rank |
|---|---|---|---|---|---|---|---|
| DT | Avg | 0.5736 | 0.5345 | 0.5220 | 0.5255 | 0.5693 | 8.0 |
| Std | 0.0564 | 0.0684 | 0.0794 | 0.0651 | 0.0552 | ||
| ELM | Avg | 0.6573 | 0.6332 | 0.6020 | 0.6153 | 0.6527 | 4.4 |
| Std | 0.0628 | 0.0751 | 0.0680 | 0.0620 | 0.0616 | ||
| KNN | Avg | 0.6355 | 0.6175 | 0.5340 | 0.5707 | 0.7096 | 6.4 |
| Std | 0.0603 | 0.0865 | 0.0749 | 0.0713 | 0.0584 | ||
| LR | Avg | 0.6582 | 0.6331 | 0.6040 | 0.6155 | 0.7209 | 3.2 |
| Std | 0.0615 | 0.0720 | 0.0850 | 0.0664 | 0.0653 | ||
| MLP | Avg | 0.6600 | 0.6334 | 0.6160 | 0.6210 | 0.7263 | 2.0 |
| Std | 0.0749 | 0.0943 | 0.1025 | 0.0847 | 0.0561 | ||
| RF | Avg | 0.6564 | 0.6423 | 0.5740 | 0.6010 | 0.6885 | 4.4 |
| Std | 0.0472 | 0.0785 | 0.0902 | 0.0593 | 0.0611 | ||
| SVM | Avg | 0.6836 | 0.6973 | 0.5440 | 0.6063 | 0.7462 | 2.6 |
| Std | 0.0470 | 0.0758 | 0.1017 | 0.0736 | 0.0633 | ||
| XGB | Avg | 0.6536 | 0.6393 | 0.5600 | 0.5920 | 0.6886 | 5.0 |
| Std | 0.0423 | 0.0609 | 0.0954 | 0.0624 | 0.0460 |
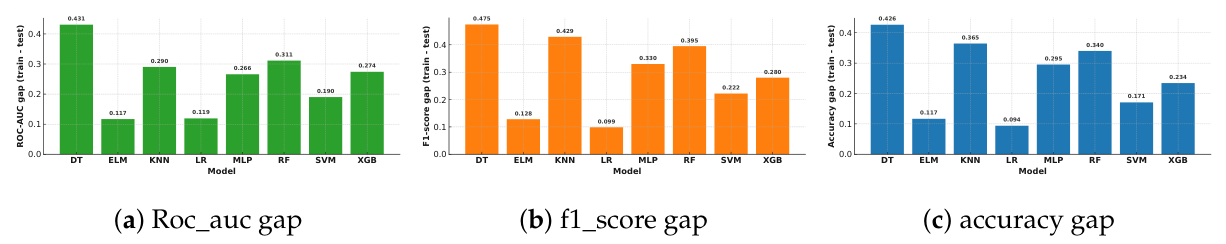
Training Performance and Overfitting
| Model | accuracy | f1-score | roc_auc | ||||||
|---|---|---|---|---|---|---|---|---|---|
| train | test | train | test | train | test | ||||
| DT | 1.0000 | 0.5736 | 0.4264 | 1.0000 | 0.5255 | 0.4745 | 1.0000 | 0.5693 | 0.4307 |
| ELM | 0.7740 | 0.6573 | 0.1167 | 0.7435 | 0.6153 | 0.1282 | 0.7696 | 0.6527 | 0.1169 |
| KNN | 1.0000 | 0.6355 | 0.3645 | 1.0000 | 0.5707 | 0.4293 | 1.0000 | 0.7096 | 0.2904 |
| LR | 0.7521 | 0.6582 | 0.0939 | 0.7142 | 0.6155 | 0.0987 | 0.8402 | 0.7209 | 0.1193 |
| MLP | 0.9553 | 0.6600 | 0.2953 | 0.9509 | 0.6210 | 0.3299 | 0.9923 | 0.7263 | 0.2660 |
| RF | 0.9966 | 0.6564 | 0.3402 | 0.9962 | 0.6010 | 0.3952 | 0.9999 | 0.6885 | 0.3114 |
| SVM | 0.8543 | 0.6836 | 0.1707 | 0.8282 | 0.6063 | 0.2219 | 0.9364 | 0.7462 | 0.1902 |
| XGB | 0.8879 | 0.6536 | 0.2343 | 0.8719 | 0.5920 | 0.2799 | 0.9627 | 0.6886 | 0.2741 |
Training and Testing Time
4.1.2. Results on SDB Dataset
4.2. Results of Metaheuristic-Optimized ELM
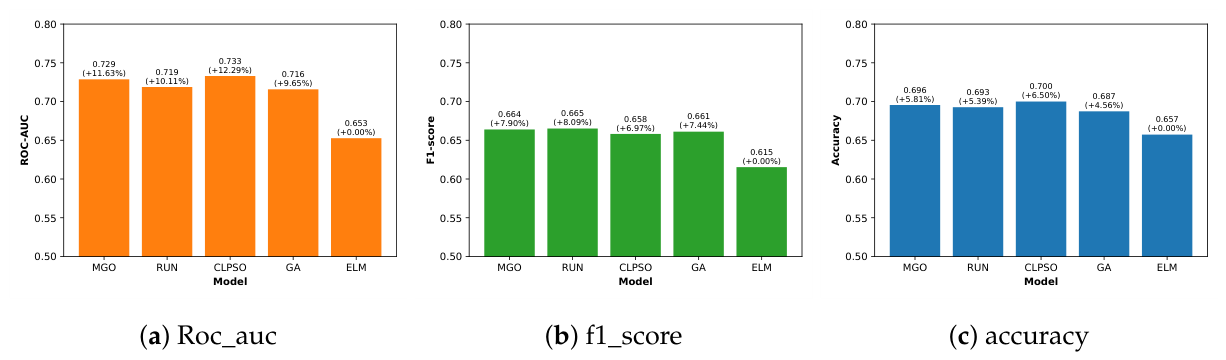
4.2.1. Results on OSA Dataset
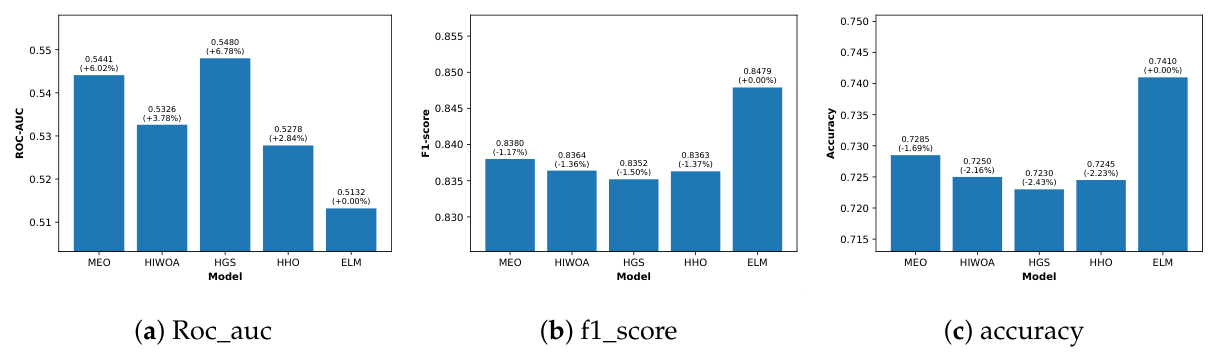
4.2.2. Results on SDB Dataset
4.2.3. Computational Cost of Optimization
4.3. Discussion and Limitations
4.3.1. Limitations of the Study
- The experiments rely on two datasets from a specific clinical context, each with a relatively small sample size. Accordingly, the generality of the conclusions to other populations or acquisition protocols is not guaranteed.
- The SDB dataset is highly imbalanced, and although appropriate metrics are used, residual bias toward the majority class may still affect the reported performance.
- The analysis focuses on global performance metrics. However, model interpretability and feature importance, which are essential for clinical adoption, were not explored in this study.
- The metaheuristic algorithms, the ELM architecture, and the investigated ML models were configured using reasonable but fixed hyperparameter settings. More exhaustive hyperparameter tuning could further improve performance or change the relative rankings of the methods.
- Metaheuristic optimization introduces a non-negligible offline training cost, which may limit its use when frequent re-training is required or when computational resources are very constrained
5. Conclusion and Future Work
Acknowledgments
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| model | Measure | accuracy | precision | recall | f1 | roc_auc | mean rank |
|---|---|---|---|---|---|---|---|
| DT | Avg | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | ||
| ELM | Avg | 0.7740 | 0.7708 | 0.7195 | 0.7435 | 0.7696 | 7.2 |
| Std | 0.0218 | 0.0239 | 0.0454 | 0.0293 | 0.0230 | ||
| KNN | Avg | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | ||
| LR | Avg | 0.7521 | 0.7536 | 0.6790 | 0.7142 | 0.8402 | 7.8 |
| Std | 0.0133 | 0.0151 | 0.0229 | 0.0173 | 0.0108 | ||
| MLP | Avg | 0.9553 | 0.9516 | 0.9505 | 0.9509 | 0.9923 | 4.0 |
| Std | 0.0100 | 0.0089 | 0.0224 | 0.0115 | 0.0024 | ||
| RF | Avg | 0.9966 | 0.9975 | 0.9950 | 0.9962 | 0.9999 | 3.0 |
| Std | 0.0036 | 0.0044 | 0.0061 | 0.0039 | 0.0002 | ||
| SVM | Avg | 0.8543 | 0.8959 | 0.7710 | 0.8282 | 0.9364 | 6.0 |
| Std | 0.0164 | 0.0172 | 0.0377 | 0.0224 | 0.0072 | ||
| XGB | Avg | 0.8879 | 0.9112 | 0.8365 | 0.8719 | 0.9627 | 5.0 |
| Std | 0.0152 | 0.0224 | 0.0268 | 0.0178 | 0.0057 |
| model | Measure | accuracy | precision | recall | f1 | roc_auc | mean rank |
|---|---|---|---|---|---|---|---|
| DT | Avg | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | ||
| ELM | Avg | 0.7795 | 0.7860 | 0.9770 | 0.8711 | 0.5612 | 7.0 |
| Std | 0.0101 | 0.0089 | 0.0088 | 0.0053 | 0.0226 | ||
| KNN | Avg | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0000 | 1.0 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | ||
| LR | Avg | 0.7621 | 0.7624 | 0.9995 | 0.8650 | 0.6098 | 6.0 |
| Std | 0.0009 | 0.0002 | 0.0012 | 0.0006 | 0.0136 | ||
| MLP | Avg | 0.8208 | 0.8172 | 0.9856 | 0.8935 | 0.8698 | 5.0 |
| Std | 0.0090 | 0.0098 | 0.0065 | 0.0046 | 0.0124 | ||
| RF | Avg | 0.9965 | 0.9958 | 0.9997 | 0.9977 | 1.0000 | 1.0 |
| Std | 0.0024 | 0.0028 | 0.0015 | 0.0015 | 0.0001 | ||
| SVM | Avg | 0.7656 | 0.7649 | 1.0000 | 0.8668 | 0.5389 | 8.0 |
| Std | 0.0034 | 0.0026 | 0.0000 | 0.0017 | 0.4601 | ||
| XGB | Avg | 0.7626 | 0.7626 | 1.0000 | 0.8653 | 0.9324 | 4.0 |
| Std | 0.0006 | 0.0004 | 0.0000 | 0.0003 | 0.0106 |
References
- Lee, Y.C.; Lu, C.T.; Chuang, L.P.; Lee, L.A.; Fang, T.J.; Cheng, W.N.; Li, H.Y. Pharmacotherapy for obstructive sleep apnea - A systematic review and meta-analysis of randomized controlled trials. Sleep Med. Rev. 2023, 70, 101809. [Google Scholar] [CrossRef] [PubMed]
- Meyer, E.J.; Wittert, G.A. Approach the Patient With Obstructive Sleep Apnea and Obesity. J. Clin. Endocrinol. Metab. 2024, 109, e1267–e1279. Available online: https://academic.oup.com/jcem/article-pdf/109/3/e1267/56680639/dgad572.pdf. [CrossRef]
- Piriyajitakonkij, M.; Warin, P.; Lakhan, P.; Leelaarporn, P.; Kumchaiseemak, N.; Suwajanakorn, S.; Pianpanit, T.; Niparnan, N.; Mukhopadhyay, S.C.; Wilaiprasitporn, T. SleepPoseNet: Multi-View Learning for Sleep Postural Transition Recognition Using UWB. IEEE J. Biomed. Health Inform. 2020, 1–1. [Google Scholar] [CrossRef] [PubMed]
- Kaditis, A.G.; Alonso Alvarez, M.L.; Boudewyns, A.; Alexopoulos, E.I.; Ersu, R.; Joosten, K.; Larramona, H.; Miano, S.; Narang, I.; Trang, H.; et al. Obstructive sleep disordered breathing in 2- to 18-year-old children: diagnosis and management. Eur. Respir. J. 2016, 47, 69–94. [Google Scholar] [CrossRef]
- Banluesombatkul, N.; Ouppaphan, P.; Leelaarporn, P.; Lakhan, P.; Chaitusaney, B.; Jaimchariya, N.; Chuangsuwanich, E.; Chen, W.; Phan, H.; Dilokthanakul, N.; et al. MetaSleepLearner: A Pilot Study on Fast Adaptation of Bio-signals-Based Sleep Stage Classifier to New Individual Subject Using Meta-Learning. IEEE J. Biomed. Health Inform. 2020, 1–1. [Google Scholar] [CrossRef]
- American Sleep Apnea Association. Available online: https://www.sleepapnea.org/learn/sleep-apnea-information-clinicians/ (accessed on 2019-04-22).
- AASM. American Academy of Sleep Medicine: Economic burden of undiagnosed sleep apnea in U.S. is nearly $150B per year, 2023. Accessed: Aug. 8, 2016.
- Kuo, N.Y.; Tsai, H.J.; Tsai, S.J.; Yang, A.C. Efficient Screening in Obstructive Sleep Apnea Using Sequential Machine Learning Models, Questionnaires, and Pulse Oximetry Signals: Mixed Methods Study. J. Med. Internet Res. 2024, 26, e51615. [Google Scholar] [CrossRef]
- Sheta, A.; Turabieh, H.; Braik, M.; Surani, S.R. Diagnosis of Obstructive Sleep Apnea Using Logistic Regression and Artificial Neural Networks Models. In Proceedings of the Proceedings of the Future Technologies Conference (FTC) 2019, Cham; Arai, K., Bhatia, R., Kapoor, S., Eds.; 2020; pp. 766–784. [Google Scholar]
- Aiyer, I.; Shaik, L.; Sheta, A.; Surani, S. Review of Application of Machine Learning as a Screening Tool for Diagnosis of Obstructive Sleep Apnea. Medicina 2022, 58. [Google Scholar] [CrossRef]
- Surani, S.; Sheta, A.; Turabieh, H.; Park, J.; Mathur, S.; Katangur, A. Diagnosis of Sleep Apnea Using artificial Neural Network and binary Particle Swarm Optimization for Feature Selection. Chest 2019, 156, A136. [Google Scholar] [CrossRef]
- Markowska-Kaczmar, U.; Kosturek, M. Extreme learning machine versus classical feedforward network: Comparison from the usability perspective. Neural Comput. Appl. 2021, 33, 15121–15144. [Google Scholar] [CrossRef]
- Abu Al-Haija, Q.; Altamimi, S.; AlWadi, M. Analysis of Extreme Learning Machines (ELMs) for intelligent intrusion detection systems: A survey. Expert Syst. With Appl. 2024, 253, 124317. [Google Scholar] [CrossRef]
- Albadr, M.; Tiun, S.; Ayob, M.; Al-Dhief, F. Particle Swarm Optimization-Based Extreme Learning Machine for COVID-19 Detection. Cogn. Comput. 2022, 16, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Lu, S.; Wang, S.H.; Zhang, Y.D. A review on extreme learning machine. Multimed. Tools Appl. 2022, 81, 41611–41660. [Google Scholar] [CrossRef]
- Eshtay, M.; Faris, H.; Obeid, N. Metaheuristic-based extreme learning machines: a review of design formulations and applications. Int. J. Mach. Learn. Cybern. 2019, 10, 1543–1561. [Google Scholar] [CrossRef]
- Thaher, T.; Sheta, A.; Awad, M.; Aldasht, M. Enhanced variants of crow search algorithm boosted with cooperative based island model for global optimization. Expert Syst. With Appl. 2024, 238, 121712. [Google Scholar] [CrossRef]
- Qian, Y.; Dharmage, S.C.; Hamilton, G.S.; Lodge, C.J.; Lowe, A.J.; Zhang, J.; Bowatte, G.; Perret, J.L.; Senaratna, C.V. Longitudinal risk factors for obstructive sleep apnea: A systematic review. Sleep Med. Rev. 2023, 71, 101838. [Google Scholar] [CrossRef]
- Chang, J.L.; Goldberg, A.N.; Alt, J.A.; Mohammed, A.; Ashbrook, L.; Auckley, D.; Ayappa, I.; Bakhtiar, H.; Barrera, J.E.; Bartley, B.L.; et al. International Consensus Statement on Obstructive Sleep Apnea. Int. Forum Allergy Rhinol. 2023, 13, 1061–1482. Available online: https://onlinelibrary.wiley.com/doi/pdf/10.1002/alr.23079. [CrossRef]
- Duarte, M.; Pereira-Rodrigues, P.; Ferreira-Santos, D. The role of novel digital clinical tools in the screening or diagnosis of obstructive sleep apnea: systematic review. J. Med. Internet Res. 2023, 25, e47735. [Google Scholar] [CrossRef]
- Maniaci, A.; Riela, P.M.; Iannella, G.; Lechien, J.R.; La Mantia, I.; De Vincentiis, M.; Cammaroto, G.; Calvo-Henriquez, C.; Di Luca, M.; Chiesa Estomba, C.; et al. Machine learning identification of obstructive sleep apnea severity through the patient clinical features: a retrospective study. Life 2023, 13, 702. [Google Scholar] [CrossRef]
- Srivastava, G.; Chauhan, A.; Kargeti, N.; Pradhan, N.; Dhaka, V.S. ApneaNet: A hybrid 1DCNN-LSTM architecture for detection of Obstructive Sleep Apnea using digitized ECG signals. Biomed. Signal Process. Control 2023, 84, 104754. [Google Scholar] [CrossRef]
- Brennan, H.L.; Kirby, S.D. Barriers of artificial intelligence implementation in the diagnosis of obstructive sleep apnea. J. Otolaryngol.-Head. Neck Surg. 2022, 51, 16. [Google Scholar] [CrossRef]
- Yeh, E.; Wong, E.; Tsai, C.W.; Gu, W.; Chen, P.L.; Leung, L.; Wu, I.C.; Strohl, K.P.; Folz, R.J.; Yar, W.; et al. Detection of obstructive sleep apnea using Belun Sleep Platform wearable with neural network-based algorithm and its combined use with STOP-Bang questionnaire. PLoS ONE 2021, 16, e0258040. [Google Scholar] [CrossRef]
- Shi, E.; Zhang, Y.; Cao, Z.; Ma, L.; Yuan, Y.; Niu, X.; Su, Y.; Xie, Y.; Chen, X.; Xing, L.; et al. Application and interpretation of machine learning models in predicting the risk of severe obstructive sleep apnea in adults. BMC Med. Inform. Decis. Mak. 2023, 23. [Google Scholar] [CrossRef]
- Banluesombatkul, N.; Rakthanmanon, T.; Wilaiprasitporn, T. Single Channel ECG for Obstructive Sleep Apnea Severity Detection Using a Deep Learning Approach. In Proceedings of the TENCON 2018 - 2018 IEEE Region 10 Conference, 2018; pp. 2011–2016. [Google Scholar] [CrossRef]
- L., Z.; D., F.; R., U.; D., K. 0311 Automated Apnea and Hypopnea Event Detection Using Deep Learning. In Sleep; Copyright - Copyright © 2018 Sleep Research Society, 2018; Volume 41, pp. A119–A120. [Google Scholar]
- Faust, O.; Barika, R.; Shenfield, A.; Ciaccio, E.J.; Acharya, U.R. Accurate detection of sleep apnea with long short-term memory network based on RR interval signals. Knowl.-Based Syst. 2021, 212, 106591. [Google Scholar] [CrossRef]
- Haberfeld, C.; Sheta, A.; Hossain, M.S.; Turabieh, H.; Surani, S. SAS Mobile Application for Diagnosis of Obstructive Sleep Apnea Utilizing Machine Learning Models. In Proceedings of the 2020 11th IEEE Annual Ubiquitous Computing, Electronics Mobile Communication Conference (UEMCON), 2020; pp. 0522–0529. [Google Scholar] [CrossRef]
- Azimi, H.; Xi, P.; Bouchard, M.; Goubran, R.; Knoefel, F. Machine Learning-Based Automatic Detection of Central Sleep Apnea Events From a Pressure Sensitive Mat. IEEE Access 2020, 8, 173428–173439. [Google Scholar] [CrossRef]
- Huang, W.; Lee, P.; Liu, Y.; Lai, F. 0495 Prediction Of Obstructive Sleep Apnea Using Machine Learning Technique. Sleep 2018, 41, A186–A186. [Google Scholar] [CrossRef]
- K., T.; K., J.W.; L., K. Detection of sleep disordered breathing severity using acoustic biomarker and machine learning techniques. Biomed. Eng. OnLine 2018, 17, 16. [Google Scholar] [CrossRef]
- Alshaer, H.; Hummel, R.; Mendelson, M.; Marshal, T.; Bradley, T.D. Objective Relationship Between Sleep Apnea and Frequency of Snoring Assessed by Machine Learning. J. Clin. Sleep Med. 2019, 15, 463–470. [Google Scholar] [CrossRef]
- Bozkurt, F.; Uçar, M.K.; Bozkurt, M.R.; Bilgin, C. Detection of abnormal respiratory events with single channel ECG and hybrid machine learning model in patients with obstructive sleep apnea. IRBM 2020. [Google Scholar] [CrossRef]
- Sheta, A.; Turabieh, H.; Thaher, T.; Too, J.; Mafarja, M.; Hossain, M.S.; Surani, S.R. Diagnosis of obstructive sleep apnea from ECG signals using machine learning and deep learning classifiers. Appl. Sci. 2021, 11, 6622. [Google Scholar] [CrossRef]
- Ferreira-Santos, D.; Amorim, P.; Silva Martins, T.; Monteiro-Soares, M.; Pereira Rodrigues, P. Enabling early obstructive sleep apnea diagnosis with machine learning: systematic review. J. Med. Internet Res. 2022, 24, e39452. [Google Scholar] [CrossRef]
- Bahrami, M.; Forouzanfar, M. Sleep apnea detection from single-lead ECG: A comprehensive analysis of machine learning and deep learning algorithms. IEEE Trans. Instrum. Meas. 2022, 71, 1–11. [Google Scholar] [CrossRef]
- Kohzadi, Z.; Safdari, R.; Haghighi, K.S. Evaluation of the PSO metaheuristic algorithm in different types of sleep apnea diagnosis using RR intervals. J. Biomed. Phys. Eng. 2023, 13, 147. [Google Scholar] [CrossRef]
- Pouramirarsalani, S.; Maleki, S.E.; Rajebi, S.; Manaf, N.V.; Roohany, A. Diagnosis of sleep apnea by optimal fuzzy system based on respiratory signals. In Proceedings of the 2024 10th International Conference on Artificial Intelligence and Robotics (QICAR); IEEE, 2024; pp. 100–105. [Google Scholar]
- Sheta, A.; Thaher, T.; Surani, S.R.; Turabieh, H.; Braik, M.; Too, J.; Abu-El-Rub, N.; Mafarjah, M.; Chantar, H.; Subramanian, S. Diagnosis of Obstructive Sleep Apnea Using Feature Selection, Classification Methods, and Data Grouping Based Age, Sex, and Race. Diagnostics 2023, 13. [Google Scholar] [CrossRef]
- Sleep Disordered Breathing Detection. Kaggle dataset. Accessed: 2025-12-13.
- Golowich, N.; Rakhlin, A.; Shamir, O. Size-independent sample complexity of neural networks. Inf. Inference A J. IMA 2020, 9, 473–504. Available online: https://academic.oup.com/imaiai/article-pdf/9/2/473/33321322/iaz007.pdf. [CrossRef]
- Huang, G.B.; Zhu, Q.Y.; Siew, C.K. Extreme learning machine: a new learning scheme of feedforward neural networks. In Proceedings of the 2004 IEEE international joint conference on neural networks (IEEE Cat. No. 04CH37541); Ieee, 2004; Vol. 2, pp. 985–990. [Google Scholar]
- Eshtay, M.; Faris, H.; Obeid, N. Metaheuristic-based extreme learning machines: a review of design formulations and applications. Int. J. Mach. Learn. Cybern. 2019, 10, 1543–1561. [Google Scholar] [CrossRef]
- Van Thieu, N.; Houssein, E.H.; Oliva, D.; Hung, N.D. IntelELM: A python framework for intelligent metaheuristic-based extreme learning machine. Neurocomputing 2025, 618, 129062. [Google Scholar] [CrossRef]
- Nguyen, T.; Hoang, B.; Nguyen, G.; Nguyen, B.M. A new workload prediction model using extreme learning machine and enhanced tug of war optimization. Procedia Comput. Sci. 2020, 170, 362–369. [Google Scholar] [CrossRef]
- Whitley, D. A genetic algorithm tutorial. Stat. Comput. 1994, 4, 65–85. [Google Scholar] [CrossRef]
- Ahmadianfar, I.; Heidari, A.A.; Gandomi, A.H.; Chu, X.; Chen, H. RUN beyond the metaphor: An efficient optimization algorithm based on Runge Kutta method. Expert Syst. With Appl. 2021, 181, 115079. [Google Scholar] [CrossRef]
- Gupta, S.; Deep, K.; Mirjalili, S. An efficient equilibrium optimizer with mutation strategy for numerical optimization. Appl. Soft Comput. 2020, 96, 106542. [Google Scholar] [CrossRef]
- Liang, J.; Qin, A.; Suganthan, P.; Baskar, S. Comprehensive learning particle swarm optimizer for global optimization of multimodal functions. IEEE Trans. Evol. Comput. 2006, 10, 281–295. [Google Scholar] [CrossRef]
- Tang, C.; Sun, W.; Wu, W.; Xue, M. A hybrid improved whale optimization algorithm. In Proceedings of the 2019 IEEE 15th International Conference on Control and Automation (ICCA), 2019; pp. 362–367. [Google Scholar] [CrossRef]
- Lou, L.; Xia, W.; Sun, Z.; Quan, S.; Yin, S.; Gao, Z.; Lin, C. COVID-19 mortality prediction using ensemble learning and grey wolf optimization. PeerJ Comput. Sci. 2023, 9, e1209. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Chen, H.; Heidari, A.A.; Gandomi, A.H. Hunger games search: Visions, conception, implementation, deep analysis, perspectives, and towards performance shifts. Expert Syst. With Appl. 2021, 177, 114864. [Google Scholar] [CrossRef]
- Heidari, A.A.; Mirjalili, S.; Faris, H.; Aljarah, I.; Mafarja, M.; Chen, H. Harris hawks optimization: Algorithm and applications. Future Gener. Comput. Syst. 2019, 97, 849–872. [Google Scholar] [CrossRef]
- Zhao, S.; Zhang, T.; Ma, S.; Wang, M. Sea-horse optimizer: a novel nature-inspired meta-heuristic for global optimization problems. Appl. Intell. 2022, 53, 11833–11860. [Google Scholar] [CrossRef]
- Abdollahzadeh, B.; Gharehchopogh, F.S.; Khodadadi, N.; Mirjalili, S. Mountain Gazelle Optimizer: A new Nature-inspired Metaheuristic Algorithm for Global Optimization Problems. Adv. Eng. Softw. 2022, 174, 103282. [Google Scholar] [CrossRef]
- Obadina, O.O.; Thaha, M.A.; Althoefer, K.; Shaheed, M.H. Dynamic characterization of a master–slave robotic manipulator using a hybrid grey wolf–whale optimization algorithm. J. Vib. Control 2022, 28, 1992–2003. [Google Scholar] [CrossRef]

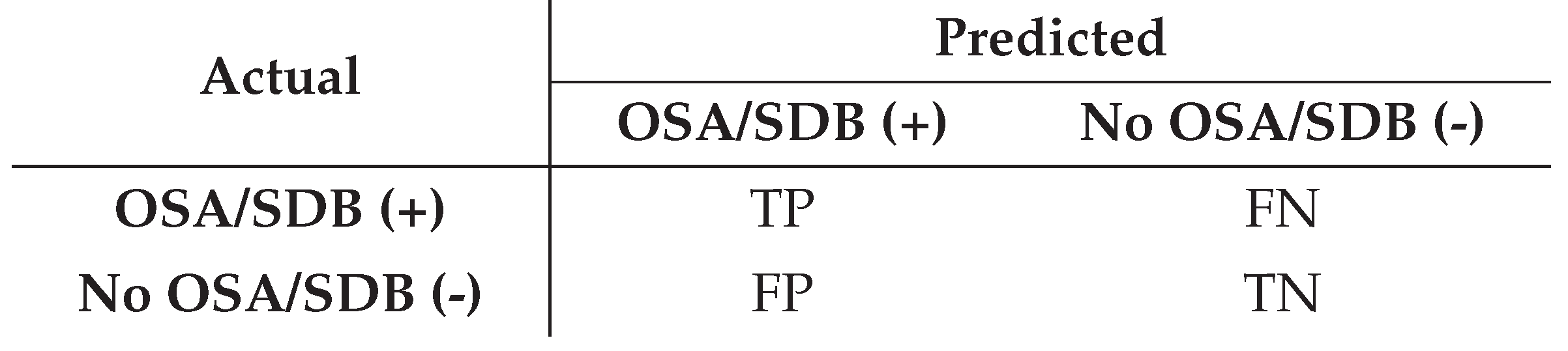
| Model | Key hyperparameters |
|---|---|
| ELM | layer_sizes = (50); act_name = relu |
| LR (LogReg) | max_iter = 200 |
| RF | n_estimators = 20 |
| SVM (RBF) | kernel = rbf; probability = True |
| XGB | n_estimators = 20; max_depth = 4; learning_rate = 0.05; |
| subsample = 0.8; colsample_bytree = 0.8; | |
| eval_metric = logloss | |
| MLP | hidden_layer_sizes = (100,); activation = relu; solver = adam; |
| max_iter = 200 | |
| KNN | n_neighbors = 5; weights = distance; metric = minkowski |
| DT | max_depth = None |
| model | Measure | train_time | test_time | mean rank |
|---|---|---|---|---|
| DT | Avg | 0.0063 | 0.0005 | 2.0 |
| Std | 0.0019 | 0.0003 | ||
| ELM | Avg | 0.0046 | 0.0001 | 1.0 |
| Std | 0.0054 | 0.0001 | ||
| KNN | Avg | 0.1295 | 0.0051 | 6.5 |
| Std | 0.5372 | 0.0013 | ||
| LR | Avg | 0.0076 | 0.0006 | 3.0 |
| Std | 0.0037 | 0.0004 | ||
| MLP | Avg | 0.3751 | 0.0009 | 6.0 |
| Std | 0.0939 | 0.0004 | ||
| RF | Avg | 0.0622 | 0.0066 | 6.5 |
| Std | 0.0218 | 0.0027 | ||
| SVM | Avg | 0.0507 | 0.0070 | 6.5 |
| Std | 0.0178 | 0.0033 | ||
| XGB | Avg | 0.0286 | 0.0014 | 4.5 |
| Std | 0.0065 | 0.0006 |
| model | Measure | accuracy | precision | recall | f1 | roc_auc | mean rank |
|---|---|---|---|---|---|---|---|
| DT | Avg | 0.6345 | 0.7695 | 0.7408 | 0.7542 | 0.5194 | 6.4 |
| Std | 0.0467 | 0.0258 | 0.0544 | 0.0369 | 0.0521 | ||
| ELM | Avg | 0.7410 | 0.7650 | 0.9513 | 0.8479 | 0.5132 | 4.8 |
| Std | 0.0295 | 0.0116 | 0.0355 | 0.0194 | 0.0301 | ||
| KNN | Avg | 0.7190 | 0.7629 | 0.9145 | 0.8317 | 0.5426 | 5.6 |
| Std | 0.0279 | 0.0142 | 0.0275 | 0.0176 | 0.0508 | ||
| LR | Avg | 0.7590 | 0.7598 | 0.9987 | 0.8630 | 0.5380 | 4.2 |
| Std | 0.0031 | 0.0007 | 0.0040 | 0.0020 | 0.0527 | ||
| MLP | Avg | 0.7460 | 0.7697 | 0.9500 | 0.8502 | 0.5430 | 3.2 |
| Std | 0.0237 | 0.0094 | 0.0327 | 0.0160 | 0.0582 | ||
| RF | Avg | 0.7300 | 0.7625 | 0.9368 | 0.8405 | 0.5329 | 5.6 |
| Std | 0.0192 | 0.0104 | 0.0288 | 0.0130 | 0.0532 | ||
| SVM | Avg | 0.7600 | 0.7600 | 1.0000 | 0.8636 | 0.4855 | 3.4 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0568 | ||
| XGB | Avg | 0.7600 | 0.7600 | 1.0000 | 0.8636 | 0.5804 | 2.0 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0511 |
| model | Measure | accuracy | precision | recall | f1 | roc_auc | mean rank |
|---|---|---|---|---|---|---|---|
| CLPSO | Avg | 0.7000 | 0.6798 | 0.6420 | 0.6582 | 0.7329 | 3.0 |
| Std | 0.0423 | 0.0457 | 0.0904 | 0.0602 | 0.0382 | ||
| GA | Avg | 0.6873 | 0.6541 | 0.6740 | 0.6611 | 0.7157 | 4.0 |
| Std | 0.0415 | 0.0513 | 0.0771 | 0.0462 | 0.0408 | ||
| GWO | Avg | 0.6818 | 0.6500 | 0.6560 | 0.6515 | 0.7164 | 5.8 |
| Std | 0.0492 | 0.0561 | 0.0716 | 0.0552 | 0.0519 | ||
| GWOWOA | Avg | 0.6827 | 0.6573 | 0.6460 | 0.6483 | 0.7223 | 4.8 |
| Std | 0.0406 | 0.0556 | 0.0737 | 0.0446 | 0.0381 | ||
| HGS | Avg | 0.6818 | 0.6533 | 0.6480 | 0.6468 | 0.7074 | 6.6 |
| Std | 0.0499 | 0.0561 | 0.0968 | 0.0650 | 0.0485 | ||
| HHO | Avg | 0.6745 | 0.6464 | 0.6300 | 0.6343 | 0.7066 | 9.6 |
| Std | 0.0627 | 0.0687 | 0.1153 | 0.0812 | 0.0505 | ||
| HIWOA | Avg | 0.6573 | 0.6304 | 0.5960 | 0.6112 | 0.6839 | 11.6 |
| Std | 0.0597 | 0.0705 | 0.0860 | 0.0733 | 0.0550 | ||
| MEO | Avg | 0.6755 | 0.6506 | 0.6220 | 0.6333 | 0.7214 | 8.0 |
| Std | 0.0418 | 0.0528 | 0.0846 | 0.0589 | 0.0559 | ||
| MGO | Avg | 0.6955 | 0.6689 | 0.6640 | 0.6639 | 0.7286 | 2.2 |
| Std | 0.0368 | 0.0497 | 0.0679 | 0.0419 | 0.0478 | ||
| RUN | Avg | 0.6927 | 0.6590 | 0.6740 | 0.6651 | 0.7187 | 2.6 |
| Std | 0.0504 | 0.0547 | 0.0771 | 0.0581 | 0.0539 | ||
| SeaHO | Avg | 0.6791 | 0.6486 | 0.6440 | 0.6445 | 0.7111 | 8.0 |
| Std | 0.0614 | 0.0726 | 0.0917 | 0.0744 | 0.0594 | ||
| ELM | Avg | 0.6573 | 0.6332 | 0.6020 | 0.6153 | 0.6527 | 11.2 |
| Std | 0.0628 | 0.0751 | 0.0680 | 0.0620 | 0.0616 |
| model | Measure | accuracy | precision | recall | f1 | roc_auc | mean rank |
|---|---|---|---|---|---|---|---|
| CLPSO | Avg | 0.7175 | 0.7579 | 0.9230 | 0.8323 | 0.4985 | 8.6 |
| Std | 0.0183 | 0.0077 | 0.0215 | 0.0121 | 0.0516 | ||
| GA | Avg | 0.7140 | 0.7649 | 0.9007 | 0.8271 | 0.5300 | 8.2 |
| Std | 0.0190 | 0.0095 | 0.0251 | 0.0130 | 0.0571 | ||
| GWO | Avg | 0.7120 | 0.7673 | 0.8914 | 0.8246 | 0.5454 | 7.2 |
| Std | 0.0253 | 0.0134 | 0.0280 | 0.0168 | 0.0518 | ||
| GWOWOA | Avg | 0.7180 | 0.7681 | 0.9013 | 0.8293 | 0.5404 | 6.0 |
| Std | 0.0164 | 0.0106 | 0.0184 | 0.0104 | 0.0506 | ||
| HGS | Avg | 0.7230 | 0.7624 | 0.9237 | 0.8352 | 0.5480 | 5.0 |
| Std | 0.0187 | 0.0118 | 0.0207 | 0.0116 | 0.0493 | ||
| HHO | Avg | 0.7245 | 0.7620 | 0.9270 | 0.8363 | 0.5278 | 5.6 |
| Std | 0.0295 | 0.0143 | 0.0306 | 0.0189 | 0.0722 | ||
| HIWOA | Avg | 0.7250 | 0.7631 | 0.9257 | 0.8364 | 0.5326 | 4.6 |
| Std | 0.0164 | 0.0097 | 0.0231 | 0.0109 | 0.0447 | ||
| MEO | Avg | 0.7285 | 0.7666 | 0.9243 | 0.8380 | 0.5441 | 3.0 |
| Std | 0.0278 | 0.0152 | 0.0276 | 0.0174 | 0.0647 | ||
| MGO | Avg | 0.6935 | 0.7545 | 0.8842 | 0.8139 | 0.5094 | 11.6 |
| Std | 0.0303 | 0.0112 | 0.0404 | 0.0216 | 0.0667 | ||
| RUN | Avg | 0.7165 | 0.7635 | 0.9086 | 0.8296 | 0.5215 | 8.0 |
| Std | 0.0320 | 0.0174 | 0.0294 | 0.0201 | 0.0803 | ||
| SeaHO | Avg | 0.7205 | 0.7648 | 0.9132 | 0.8323 | 0.5450 | 5.6 |
| Std | 0.0190 | 0.0092 | 0.0223 | 0.0126 | 0.0329 | ||
| ELM | Avg | 0.7410 | 0.7650 | 0.9513 | 0.8479 | 0.5132 | 3.4 |
| Std | 0.0295 | 0.0116 | 0.0355 | 0.0194 | 0.0301 |
| model | Measure | OSA | SDB |
|---|---|---|---|
| CLPSO | Avg | 56.7140 | 30.8554 |
| Std | 1.4979 | 3.4613 | |
| GA | Avg | 12.2153 | 13.8607 |
| Std | 0.3092 | 3.4478 | |
| GWO | Avg | 11.4065 | 10.5191 |
| Std | 0.2875 | 0.3267 | |
| GWOWOA | Avg | 11.4524 | 14.6931 |
| Std | 0.5361 | 2.7607 | |
| HGS | Avg | 10.6527 | 10.5610 |
| Std | 0.5867 | 0.9460 | |
| HHO | Avg | 18.6361 | 18.7242 |
| Std | 0.7000 | 1.3378 | |
| HIWOA | Avg | 12.9272 | 12.9110 |
| Std | 0.8009 | 0.7845 | |
| MEO | Avg | 17.5411 | 22.7018 |
| Std | 0.5319 | 4.3979 | |
| MGO | Avg | 40.1385 | 46.5233 |
| Std | 1.1942 | 4.0455 | |
| RUN | Avg | 20.2531 | 24.6024 |
| Std | 0.7883 | 9.1322 | |
| SeaHO | Avg | 17.4395 | 17.7751 |
| Std | 0.5797 | 2.9496 | |
| ELM | Avg | 0.0046 | 0.0911 |
| Std | 0.0054 | 0.2988 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




