Submitted:
27 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
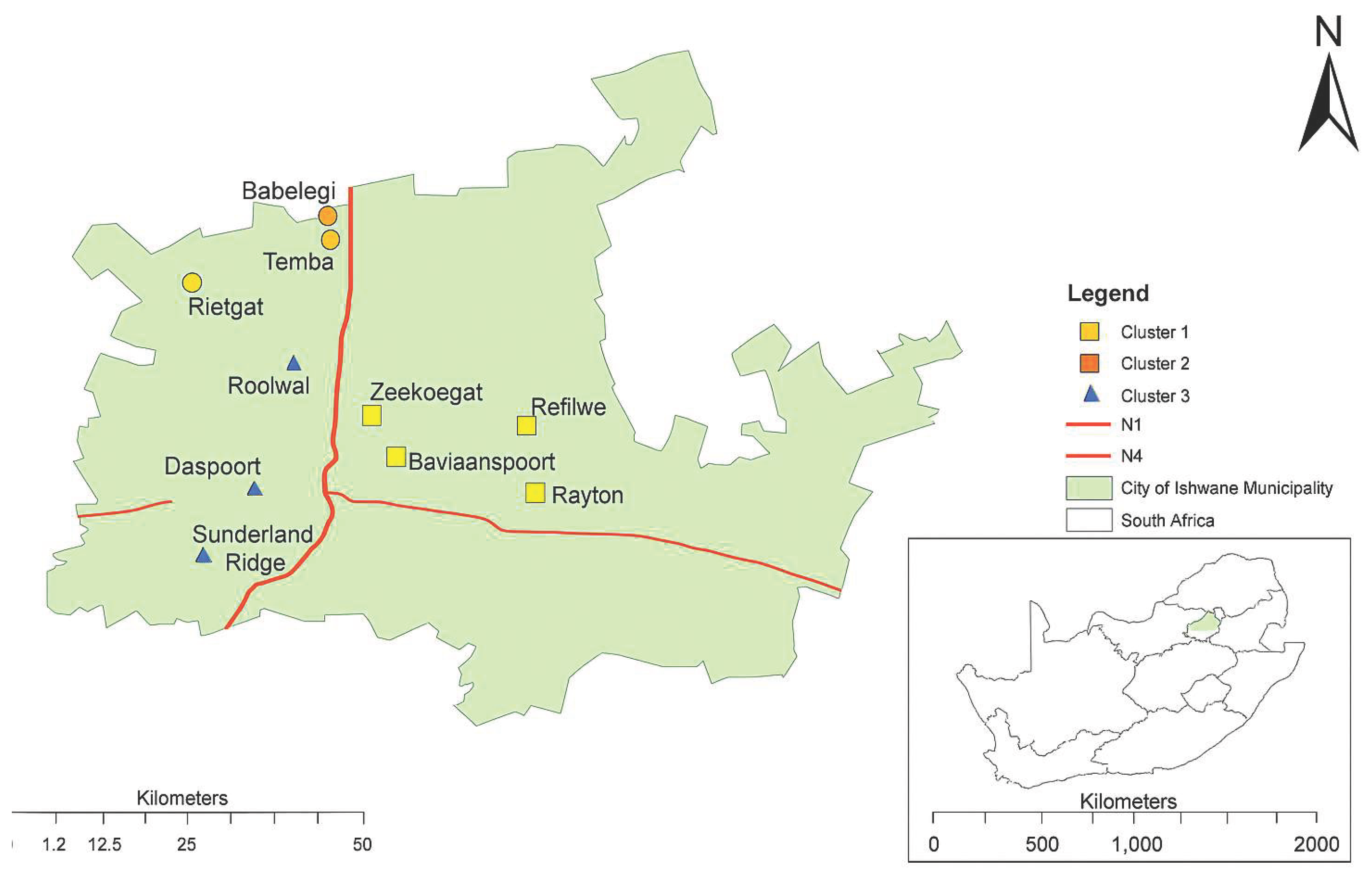
2.1. Study Area and Sample Collection
2.2. Identification of V. cholerae
2.3. Genus-Specific PCR
2.4. Identification of Non-O1/Non-O139 V. cholerae
2.5. Repetitive Extragenic Palindromic PCR
2.6. Whole Genome Sequencing
2.6. Bioinformatics Analysis
2.6. Phylogenetic Analysis
3. Results
3.1. Phenotypic and Molecular Identification of NOVC Isolates
3.2. Whole Genome Characteristics of NOVC Isolates
3.3. Genomic Characteristics of NOVC Isolates
3.4. Mobile Genetic Elements
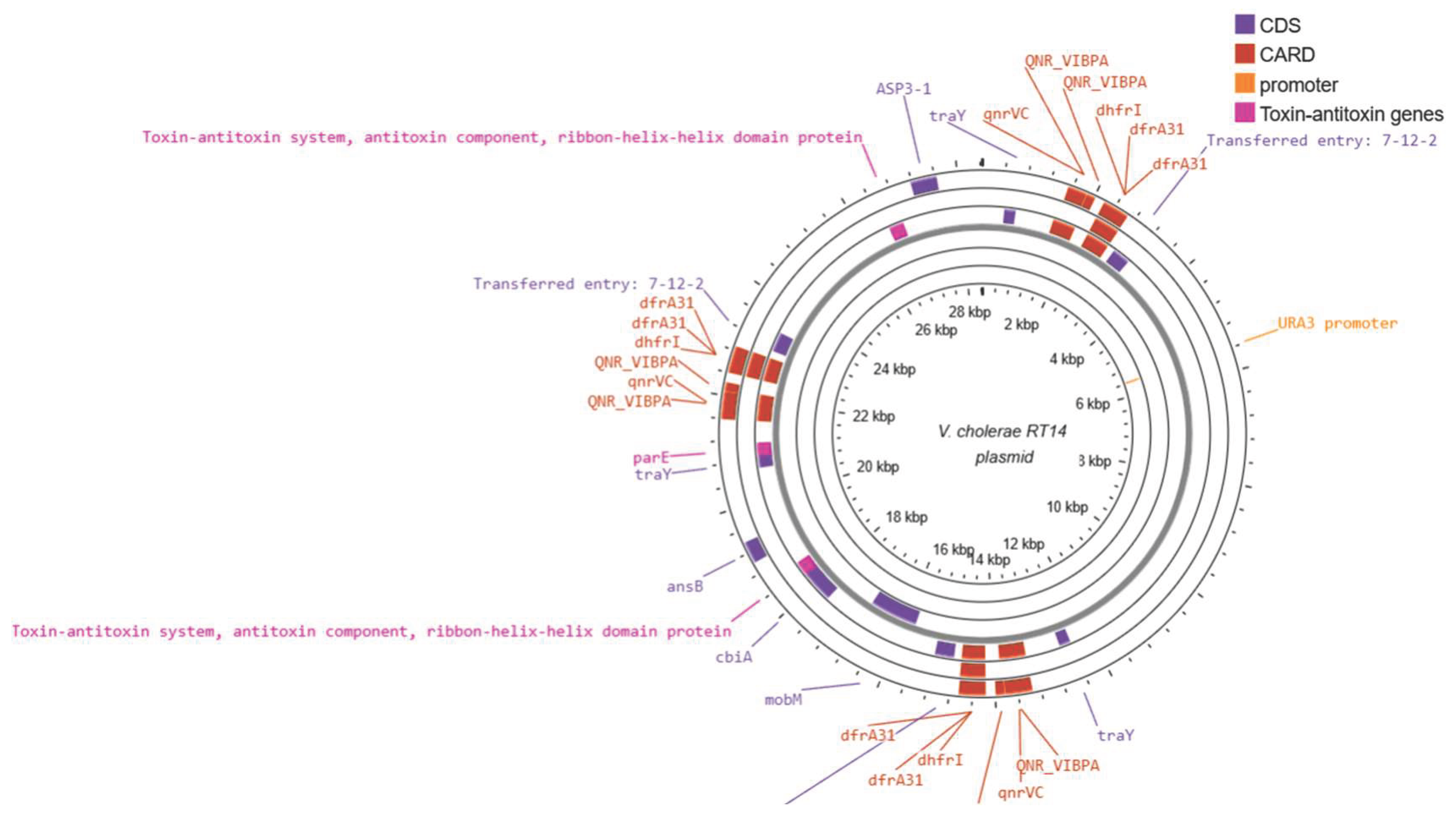
3.5. Plasmids
3.6. Antibiotic Resistance Profiles of NOVC Isolates
3.7. Virulence-Associated Genes of NOVC Isolates
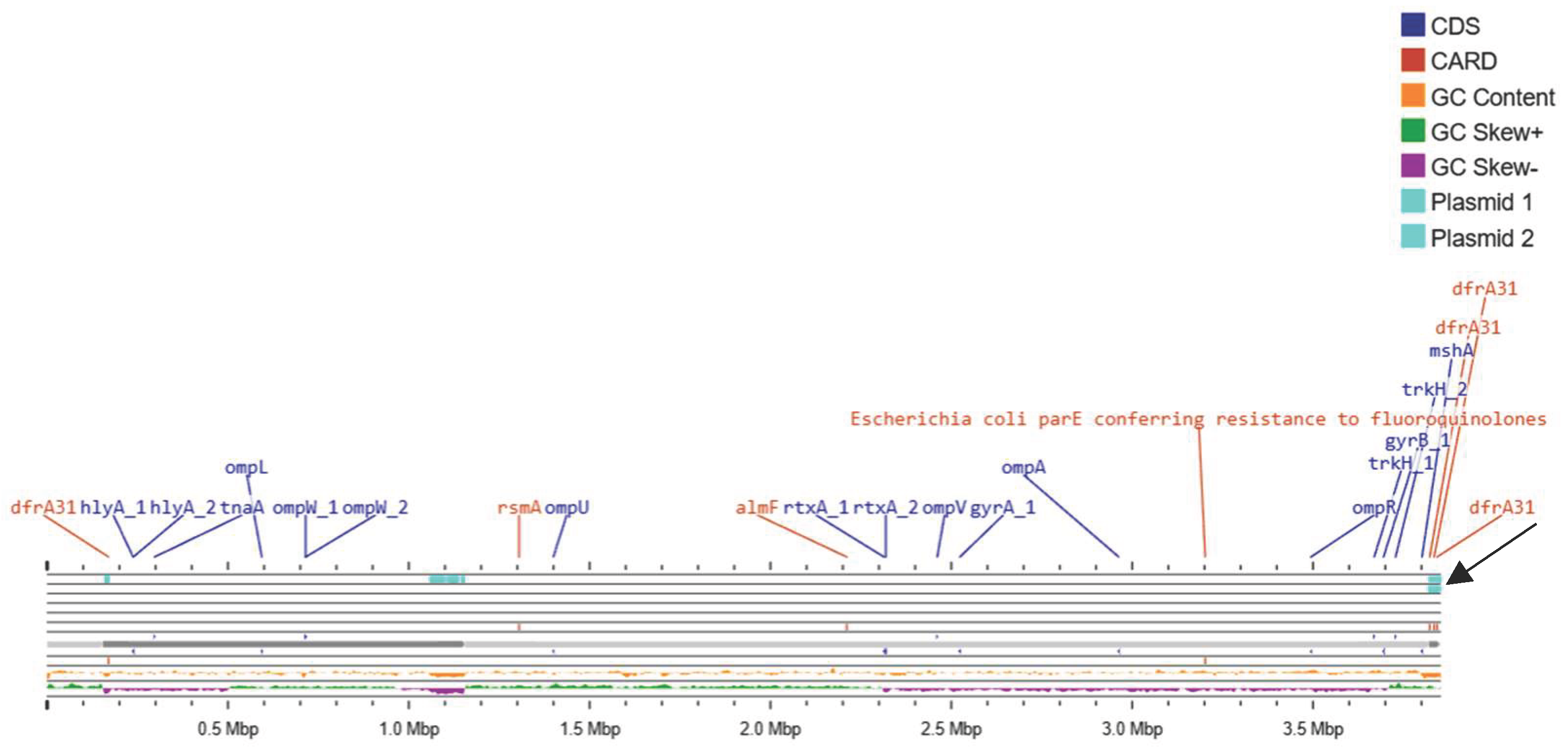
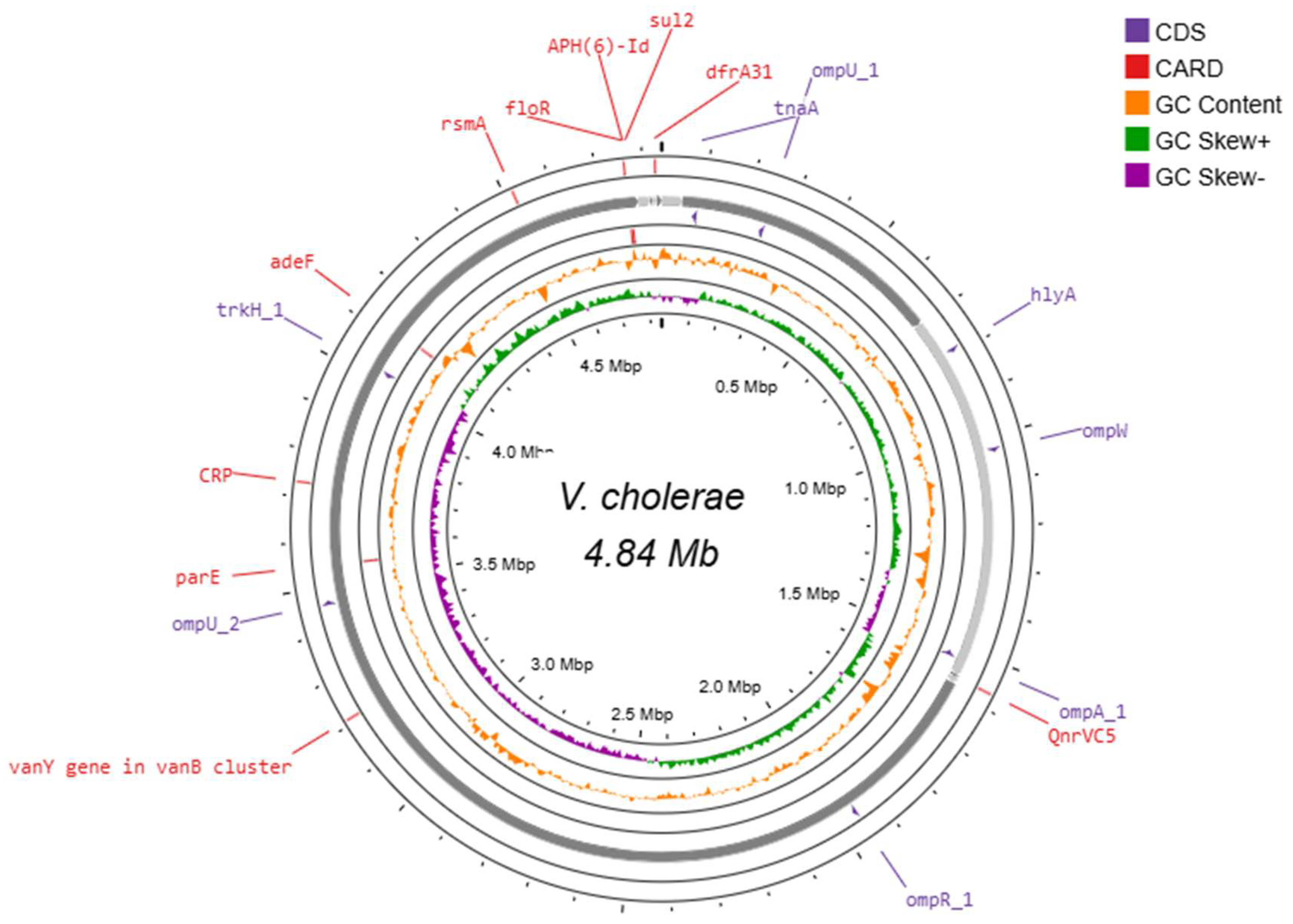
3.8. Circular Representation of a Whole Genome Map of NOVC Isolates
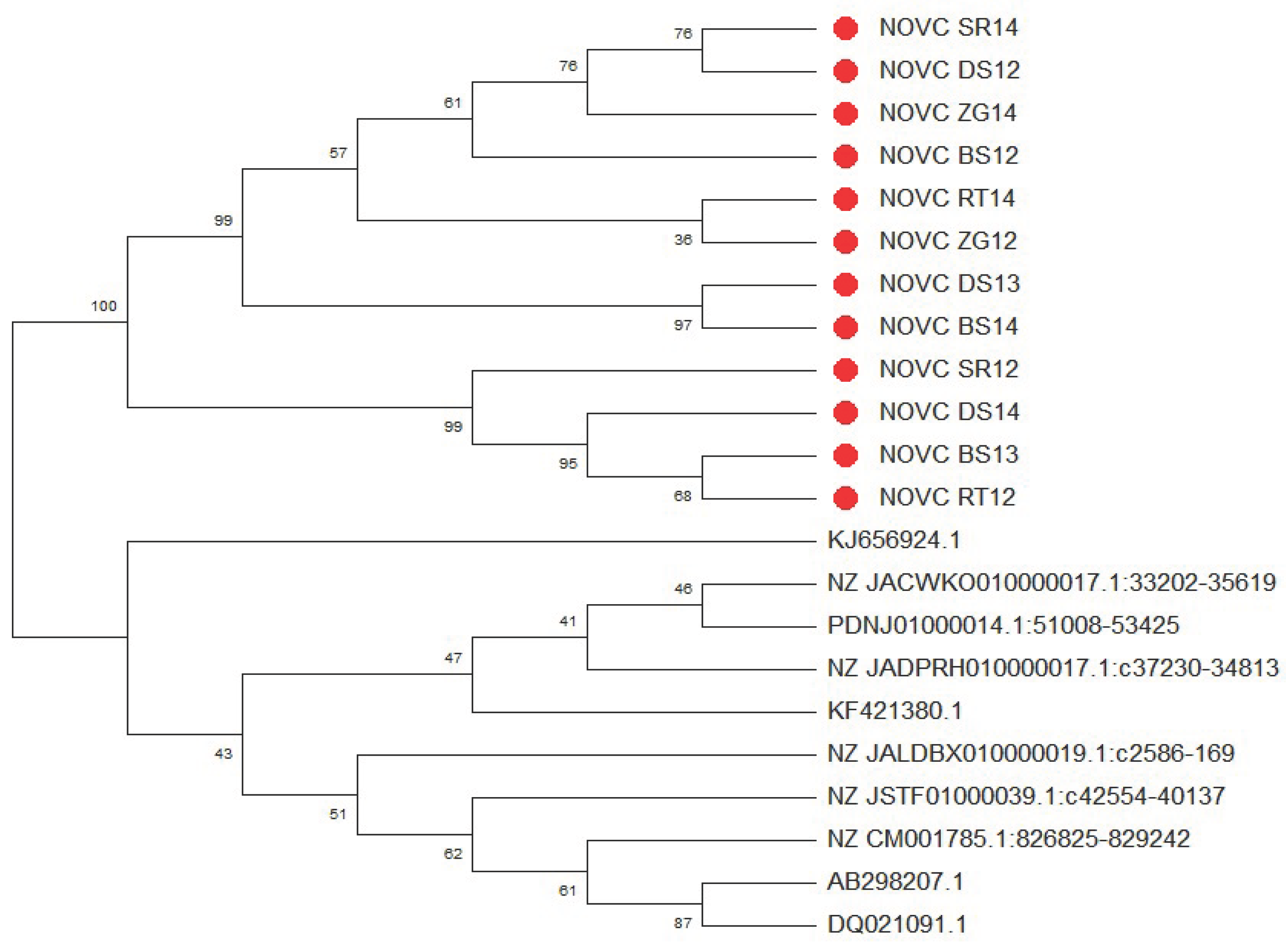
3.9. Phylogenetic Analysis of the gyrB Gene in NOVC Isolates
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Montero, D.A.; Vidal, R.M.; Velasco, J.; George, S.; Lucero, Y.; Gómez, L.A.; et al. Vibrio cholerae, classification, pathogenesis, immune response, and trends in vaccine development. Front Med [Internet]. 5 May 2023, 10. Available online: https://www.frontiersin.org/journals/medicine/articles/10.3389/fmed.2023.1155751/full.
- Ortega, D.R.; Kjær, A.; Briegel, A. The chemosensory systems of Vibrio cholerae. Mol. Microbiol. 2020, 114(3), 367–76. [Google Scholar] [CrossRef]
- Liang, K.Y.H.; Orata, F.D.; Islam, M.T.; Nasreen, T.; Alam, M.; Tarr, C.L.; et al. A Vibrio cholerae Core Genome Multilocus Sequence Typing Scheme To Facilitate the Epidemiological Study of Cholera. J. Bacteriol. 2020, 202(24). [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yang, C.; Sun, Z.; Zheng, W.; Zhang, W.; Yu, H.; et al. Genomic epidemiology of Vibrio cholerae reveals the regional and global spread of two epidemic non-toxigenic lineages. PLoS Neglected Trop. Dis. 2020, 14(2), e0008046. [Google Scholar] [CrossRef] [PubMed]
- Bhandari, M.; Jennison, A.V.; Rathnayake, I.U.; Huygens, F. Evolution, distribution and genetics of atypical Vibrio cholerae – A review. Infect. Genet. Evol. 2021, 89, 104726. [Google Scholar] [CrossRef]
- Singh, D.V.; Matte, M.H.; Matte, G.R.; Jiang, S.; Sabeena, F.; Shukla, B.N.; et al. Molecular Analysis of Vibrio choleraeO1, O139, non-O1, and non-O139 Strains: Clonal Relationships between Clinical and Environmental Isolates. Appl. Environ. Microbiol. 2001, 67(2), 910–21. [Google Scholar] [CrossRef]
- Fu, S.; Hao, J.; Jin, S.; Wu, K.; Wang, Y.; Ye, S.; et al. A Human Intestinal Infection Caused by a Novel Non-O1/O139 Vibrio cholerae Genotype and Its Dissemination Along the River. Front Public Health [Internet] 2019, 7. Available online: https://www.frontiersin.org/journals/public-health/articles/10.3389/fpubh.2019.00100/full. [CrossRef]
- Vezzulli, L.; Baker-Austin, C.; Kirschner, A.; Pruzzo, C.; Martinez-Urtaza, J. Global emergence of environmental non-O1/O139 Vibrio cholerae infections linked with climate change: a neglected research field? Environ. Microbiol. 2020, 22(10), 4342–55. [Google Scholar] [CrossRef]
- Li, X.; Wu, Y.; Sun, X.; Ma, J.; Li, X.; Liu, C.; et al. Non-O1/non-O139 Vibrio cholerae bacteraemia in mainland China from 2005 to 2019: clinical, epidemiological and genetic characteristics. Epidemiol. Infect. 2020, 148, e186. [Google Scholar] [CrossRef]
- le Roux, W.J.; Masoabi, D.; de Wet, C.M.E.; Venter, S.N. Evaluation of a rapid polymerase chain reaction based identification technique for Vibrio cholerae isolates. Water Sci. Technol. 2004, 50(1), 229–32. [Google Scholar] [CrossRef]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37(5), 540–6. [Google Scholar] [CrossRef] [PubMed]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75(12), 3491–500. [Google Scholar] [CrossRef]
- Jolley, K.A.; Maiden, M.C. BIGSdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinform. 2010, 11(1), 595. [Google Scholar] [CrossRef]
- Alcock, B.P.; Huynh, W.; Chalil, R.; Smith, K.W.; Raphenya, A.R.; Wlodarski, M.A.; et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2023, 51(D1), D690–9. [Google Scholar] [CrossRef]
- Brown, C.L.; Mullet, J.; Hindi, F.; Stoll, J.E.; Gupta, S.; Choi, M.; et al. mobileOG-db: a Manually Curated Database of Protein Families Mediating the Life Cycle of Bacterial Mobile Genetic Elements. Appl. Environ. Microbiol. 2022, 88(18), e00991-22. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4(4), 406–25. [Google Scholar]
- Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 1985, 125(1), 1–5. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. 2004, 101(30), 11030–5. [Google Scholar] [CrossRef]
- Street, R.; Nkambule, S.; Mahlangeni, N.; Mthethwa, M.; Blose, N.; Genthe, B.; et al. Wastewater and environmental surveillance for Vibrio cholerae: a scoping review. J. Water Health 2025, 23(6), 715–26. [Google Scholar] [CrossRef]
- Hounmanou, Y.M.; Mdegela, R.H.; Dougnon, T.V.; Mhongole, O.J.; Mayila, E.S.; Malakalinga, J.; Makingi, G.; Dalsgaard, A. Toxigenic Vibrio cholerae O1 in vegetables and fish raised in wastewater irrigated fields and stabilization ponds during a non-cholera outbreak period in Morogoro, Tanzania: an environmental health study. BMC Res. Notes 2016, 9(1), 466.6. [Google Scholar] [CrossRef] [PubMed]
- Baron, S.; Larvor, E.; Chevalier, S.; Jouy, E.; Kempf, I.; Granier, S.A.; et al. Antimicrobial Susceptibility among Urban Wastewater and Wild Shellfish Isolates of Non-O1/Non-O139 Vibrio cholerae from La Rance Estuary (Brittany, France). Front Microbiol [Internet]. 2017, 8. Available online: https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2017.01637/full.
- Uelze, L.; Grützke, J.; Borowiak, M.; Hammerl, J.A.; Juraschek, K.; Deneke, C.; et al. Typing methods based on whole genome sequencing data. One Health Outlook 2020, 2(1), 3. [Google Scholar] [CrossRef]
- Octavia, S.; Salim, A.; Kurniawan, J.; Lam, C.; Leung, Q.; Ahsan, S.; et al. Population Structure and Evolution of Non-O1/Non-O139 Vibrio cholerae by Multilocus Sequence Typing. PLoS ONE 2013, 8(6), e65342. [Google Scholar] [CrossRef]
- Arteaga, M.; Velasco, J.; Rodriguez, S.; Vidal, M.; Arellano, C.; Silva, F.; Carreño, L.J.; Vidal, R.; Montero, D.A. Genomic characterization of the non-O1/non-O139 Vibrio cholerae strain that caused a gastroenteritis outbreak in Santiago, Chile, 2018. Microb. Genom. 2020, 6(3), e000340. [Google Scholar] [CrossRef]
- Luo, Y.; Ye, J.; Jin, D.; Ding, G.; Zhang, Z.; Mei, L.; et al. Molecular analysis of non-O1/non-O139 Vibrio choleraeisolated from hospitalised patients in China. BMC Microbiol. 2013, 13(1), 52. [Google Scholar] [CrossRef]
- Mkhize, L.; Marimani, M.; Duze, S.T. Characterization of Vibrio cholerae from the Jukskei River in Johannesburg, South Africa. Lett. Appl. Microbiol. 2025, 78(3), ovaf036. [Google Scholar] [CrossRef]
- Iyer, L.; Vadivelu, J.; Puthucheary, S.D. Detection of virulence associated genes, haemolysin and protease amongst Vibrio cholerae isolated in Malaysia. Epidemiol. Infect. 2000, 125(1), 27–34. [Google Scholar] [CrossRef]
- Rajpara, N.; Vinothkumar, K.; Mohanty, P.; Singh, A.K.; Singh, R.; Sinha, R.; et al. Synergistic Effect of Various Virulence Factors Leading to High Toxicity of Environmental V. cholerae Non-O1/ Non-O139 Isolates Lacking ctx Gene: Comparative Study with Clinical Strains. PLoS ONE 2013, 8(9), e76200. [Google Scholar] [CrossRef]
- Yang, W.; Li, P.; Lei, S.; Yu, Y.; Liu, S.; You, C. Case report: Characterization and bioinformatics analysis of non-O1/O139 Vibrio cholerae strain isolated from a choledochoduodenal fistula patient with septicemia. Front Med. [Internet] 2024, 11. Available online: https://www.frontiersin.org/journals/medicine/articles/10.3389/fmed.2024.1458143/full.
- Dutta, D.; Kaushik, A.; Kumar, D.; Bag, S. Foodborne Pathogenic Vibrios: Antimicrobial Resistance. Front Microbiol [Internet]. 30 June 2021, 12. Available online: https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2021.638331/full.
- Tang, J.; Li, S.; Zhang, M.; Li, F.; Tang, Y.; Yang, F. Whole Genome Analysis of a Non-O1, Non-O139 Vibrio cholerae Detected from Human Blood in China. Infect. Drug. Resist. 2023, 16, 5453–61. [Google Scholar] [CrossRef] [PubMed]
- Lepuschitz, S.; Baron, S.; Larvor, E.; Granier, S.A.; Pretzer, C.; Mach, R.L.; et al. Phenotypic and Genotypic Antimicrobial Resistance Traits of Vibrio cholerae Non-O1/Non-O139 Isolated From a Large Austrian Lake Frequently Associated With Cases of Human Infection. Front Microbiol [Internet]. 8 Nov 2019, 10. Available online: https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2019.02600/full.
- Mevada, V.; Patel, R.; Dudhagara, P.; Chaudhari, R.; Vohra, M.; Khan, V.; JH Shyu, D.; Chen, Y.Y.; Zala, D. Whole Genome Sequencing and Pan-Genomic Analysis of Multidrug-Resistant Vibrio cholerae VC01 Isolated from a Clinical Sample. Microorganisms 2023, 11(8), 2030. [Google Scholar] [CrossRef] [PubMed]
- Nohejl, T.; Valcek, A.; Papousek, I.; Palkovicova, J.; Wailan, A.M.; Pratova, H.; et al. Genomic analysis of qnr-harbouring IncX plasmids and their transferability within different hosts under induced stress. BMC Microbiol. 2022, 22(1), 136. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.T.; Tang, H.J.; Chao, C.M.; Lai, C.C. Clinical Manifestations of Non-O1 Vibrio cholerae Infections. PLoS ONE 2015, 10(1), e0116904. [Google Scholar] [CrossRef]
- MacLean, R.C.; San Millan, A. Microbial Evolution: Towards Resolving the Plasmid Paradox. Curr. Biol. 2015, 25(17), R764–7. [Google Scholar] [CrossRef]
- Carattoli, A. Plasmids and the spread of resistance. Int. J. Med. Microbiol. 2013, 303(6), 298–304. [Google Scholar] [CrossRef]
- Ceccarelli, D.; Garriss, G.; Choi, S.Y.; Hasan, N.A.; Stepanauskas, R.; Pop, M.; et al. Characterization of Two Cryptic Plasmids Isolated in Haiti from Clinical Vibrio cholerae Non-O1/Non-O139. Front Microbiol. 2017, 8, 2283. [Google Scholar] [CrossRef] [PubMed]
- Praszkier, J.; Pittard, A.J. Control of replication in I-complex plasmids. Plasmid 2005, 53(2), 97–112. [Google Scholar] [CrossRef] [PubMed]
- Zatyka, M.; Thomas, C.M. Control of genes for conjugative transfer of plasmids and other mobile elements. FEMS Microbiol. Rev. 1998, 21(4), 291–319. [Google Scholar] [CrossRef]
- Rawlings, D.E.; Tietze, E. Comparative biology of IncQ and IncQ-like plasmids. Microbiol. Mol. Biol. Rev. 2001, 65(4), 481–96. [Google Scholar] [CrossRef]
- Piergentili, P.; Castellani-Pastoris, M.; Fellini, R.D.; Farisano, G.; Bonello, C.; Rigoli, e; et al. Transmission of Non O Group 1 Vibrio cholerae by Raw Oyster Consumption. Int. J. Epidemiol. 1984, 13(3), 340–3. [Google Scholar] [CrossRef]
- Vollmer, W.; von Rechenberg, M.; Höltje, J.V. Demonstration of Molecular Interactions between the Murein Polymerase PBP1B, the Lytic Transglycosylase MltA, and the Scaffolding Protein MipA of Escherichia coli *. J. Biol. Chem. 1999, 274(10), 6726–34. [Google Scholar] [CrossRef] [PubMed]
- Francia, M.V.; Varsaki, A.; Garcillán-Barcia, M.P.; Latorre, A.; Drainas, C.; de la Cruz, F. A classification scheme for mobilization regions of bacterial plasmids. FEMS Microbiol. Rev. 2004, 28(1), 79–100. [Google Scholar] [CrossRef]
- Hays, J.P.; Eadie, K.; Verduin, C.M.; Verbrugh, H.; van, Belkum A. A novel plasmid (pEMCJH03) isolated from moraxella catarrhalis possibly useful as a cloning and expression vector within this species. Plasmid 2005, 53(3), 263–8. [Google Scholar] [CrossRef]
- Perwez, T.; Meyer, R. MobB protein stimulates nicking at the R1162 origin of transfer by increasing the proportion of complexed plasmid DNA. J. Bacteriol. 1996, 178(19), 5762–7. [Google Scholar] [CrossRef]
- Polyvalent Proteins, a Pervasive Theme in the Intergenomic Biological Conflicts of Bacteriophages and Conjugative Elements | Journal of Bacteriology [Internet]. 4 Nov 2025. Available online: https://journals.asm.org/doi/full/10.1128/jb.00245-17.
- Guzmán-Herrador, D.L.; Llosa, M. The secret life of conjugative relaxases. Plasmid 2019, 104, 102415. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Orejas, R.; Espinosa, M.; Yeo, C.C. The Importance of the Expendable: Toxin–Antitoxin Genes in Plasmids and Chromosomes. Front Microbiol [Internet]. 4 Aug 2017, 8. Available online: https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2017.01479/full.
- Bateman, A.; Coggill, P.; Finn, R.D. DUFs: families in search of function. Acta Cryst. F. 2010, 66(10), 1148–52. [Google Scholar] [CrossRef]
- Goodacre, N.F.; Gerloff, D.L.; Uetz, P. Protein domains of unknown function are essential in bacteria. MBio 2014, 5(1), 10–128. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Orejas, R.; Espinosa, M.; Yeo, C.C. The importance of the expendable: toxin–antitoxin genes in plasmids and chromosomes. Front. Microbiol. 2017, 8, 1479. [Google Scholar] [CrossRef]
- Harrison, E.; Brockhurst, M.A. Plasmid-mediated horizontal gene transfer is a coevolutionary process. Trends Microbiol. 2012, 20(6), 262–7. [Google Scholar] [CrossRef]
- Korch, C.; Hagblom, P.; Ohman, H.; Göransson, M.; Normark, S. Cryptic plasmid of Neisseria gonorrhoeae: complete nucleotide sequence and genetic organization. J. Bacteriol. 1985, 163(2), 430–8. [Google Scholar] [CrossRef] [PubMed]
- Hasan, N.A.; Grim, C.J.; Haley, B.J.; Chun, J.; Alam, M.; Taviani, E.; et al. Comparative genomics of clinical and environmental Vibrio mimicus. Proc. Natl. Acad. Sci. 2010, 107(49), 21134–9. [Google Scholar] [CrossRef]
- Srikhanta, Y.N.; Atack, J.M.; Beacham, I.R.; Jennings, M.P. Distinct physiological roles for the two l-asparaginase isozymes of Escherichia coli. Biochem. Biophys. Res. Commun. 2013, 436(3), 362–5. [Google Scholar] [CrossRef]
| Isolate ID | Taxon | Genome size |
|---|---|---|
| BS12 | V. cholerae | 3.87 Mb |
| BS13 | V. cholerae | 4.25 Mb |
| BS14 | V. cholerae | 3.73 Mb |
| DS12 | Vibrio spp. | 4.81 Mb |
| DS13 | Vibrio spp. | 4.84 Mb |
| DS14 | V. cholerae | 3.95 Mb |
| ZG12 | V. cholerae | 4.13 Mb |
| ZG14 | V. cholerae | 4.92 Mb |
| RT12 | V. cholerae | 3.95 Mb |
| RT14 | V. cholerae | 3.86 Mb |
| SR12 | V. cholerae | 3.84 Mb |
| SR14 | V. cholerae | 3.86 Mb |
| Isolate id | G+C content (%) | Genes | CDS | tRNA | tmRNA | rRNA | sORF | CRISPR |
|---|---|---|---|---|---|---|---|---|
| BS12 | 48% | 6,459 | 6,328 | 96 | 1 | 33 | 6,800 | 1 |
| BS13 | 47% | 7,152 | 7,031 | 90 | 1 | 29 | 7,273 | 1 |
| BS14 | 48% | 7,474 | 7,365 | 88 | 0 | 20 | 6,180 | 1 |
| DS13 | 50% | 5,320 | 5,179 | 108 | 1 | 31 | 9,591 | 1 |
| DS12 | 50% | 4,919 | 4,787 | 105 | 1 | 25 | 9,283 | 1 |
| DS14 | 48% | 4,289 | 4,157 | 102 | 1 | 28 | 5,983 | 1 |
| ZG12 | 47% | 3,761 | 3,624 | 100 | 1 | 32 | 5,221 | 1 |
| ZG14 | 48% | 8,446 | 8,302 | 109 | 1 | 34 | 8,727 | 0 |
| RT12 | 48% | 3,562 | 3,430 | 102 | 1 | 29 | 5,385 | 0 |
| RT14 | 47% | 5,154 | 5,022 | 100 | 1 | 29 | 6,297 | 2 |
| SR12 | 47% | 3,564 | 3,434 | 101 | 1 | 27 | 5,221 | 1 |
| SR14 | 44% | 4,527 | 4,397 | 98 | 1 | 27 | 4,672 | 4 |
| Isolate ID | P | IE | RRR | STD | T | Total |
|---|---|---|---|---|---|---|
| BS12 | 69 | 31 | 118 | 24 | 29 | 271 |
| BS13 | 41 | 112 | 119 | 24 | 30 | 326 |
| BS14 | 74 | 29 | 122 | 19 | 22 | 266 |
| DS12 | 64 | 45 | 79 | 11 | 23 | 222 |
| DS13 | 53 | 111 | 88 | 12 | 30 | 294 |
| DS14 | 33 | 45 | 77 | 47 | 21 | 223 |
| ZG12 | 30 | 65 | 69 | 26 | 15 | 205 |
| ZG14 | 127 | 54 | 125 | 25 | 37 | 378 |
| RT12 | 27 | 29 | 70 | 21 | 12 | 159 |
| RT14 | 44 | 43 | 100 | 11 | 28 | 226 |
| SR12 | 29 | 26 | 66 | 15 | 16 | 152 |
| SR14 | 32 | 152 | 99 | 44 | 51 | 378 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




