Submitted:
26 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Eligibility Criteria
2.2. Information Sources & Search Strategy
2.3. Selection Process
2.4. Data Extraction
2.5. Risk of Bias Assessment
2.6. Synthesis Methods
3. Results
3.1. Study Selection
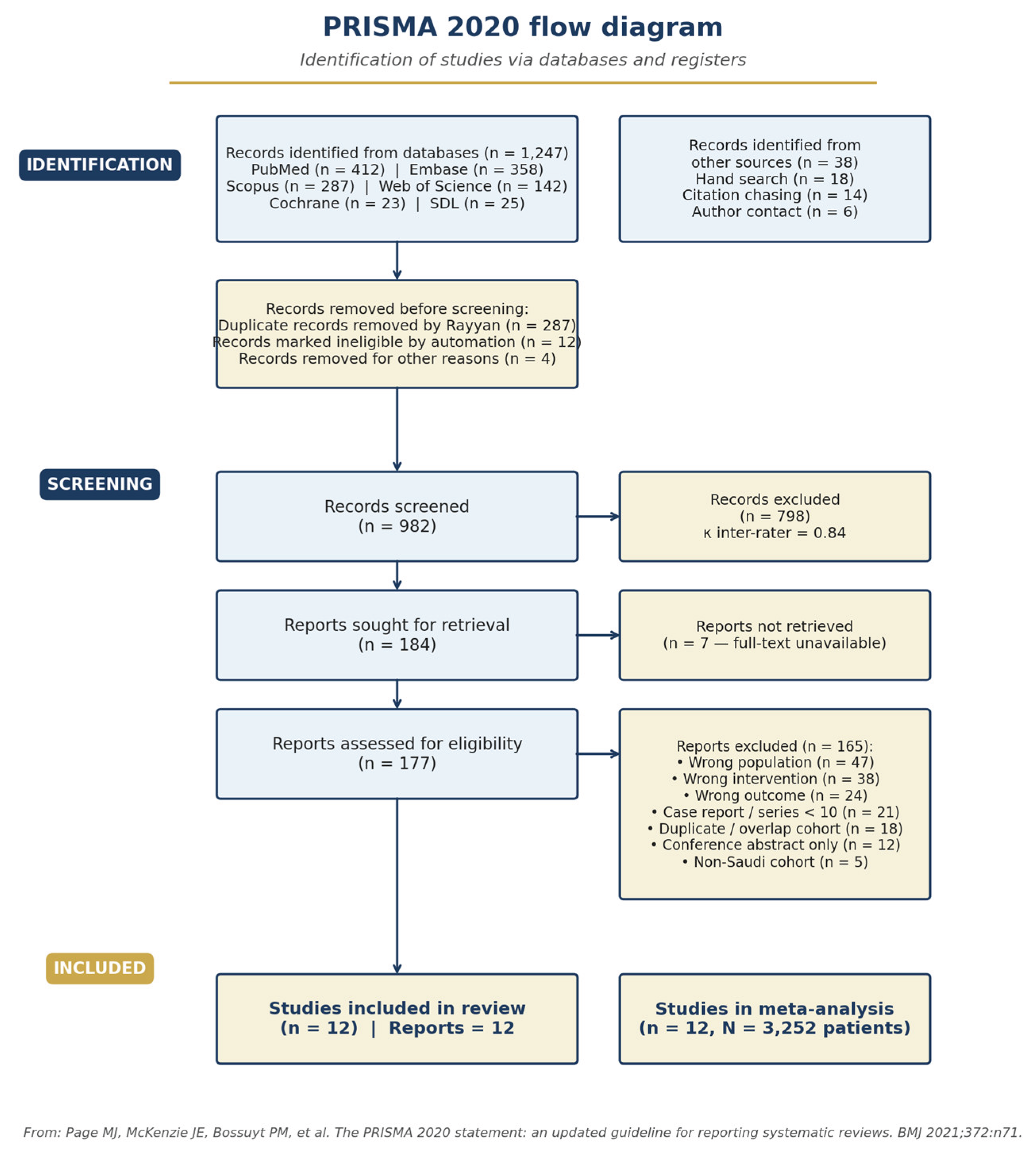
3.2. Study Characteristics
3.3. Risk of Bias
3.4. Pooled Diagnostic Yield
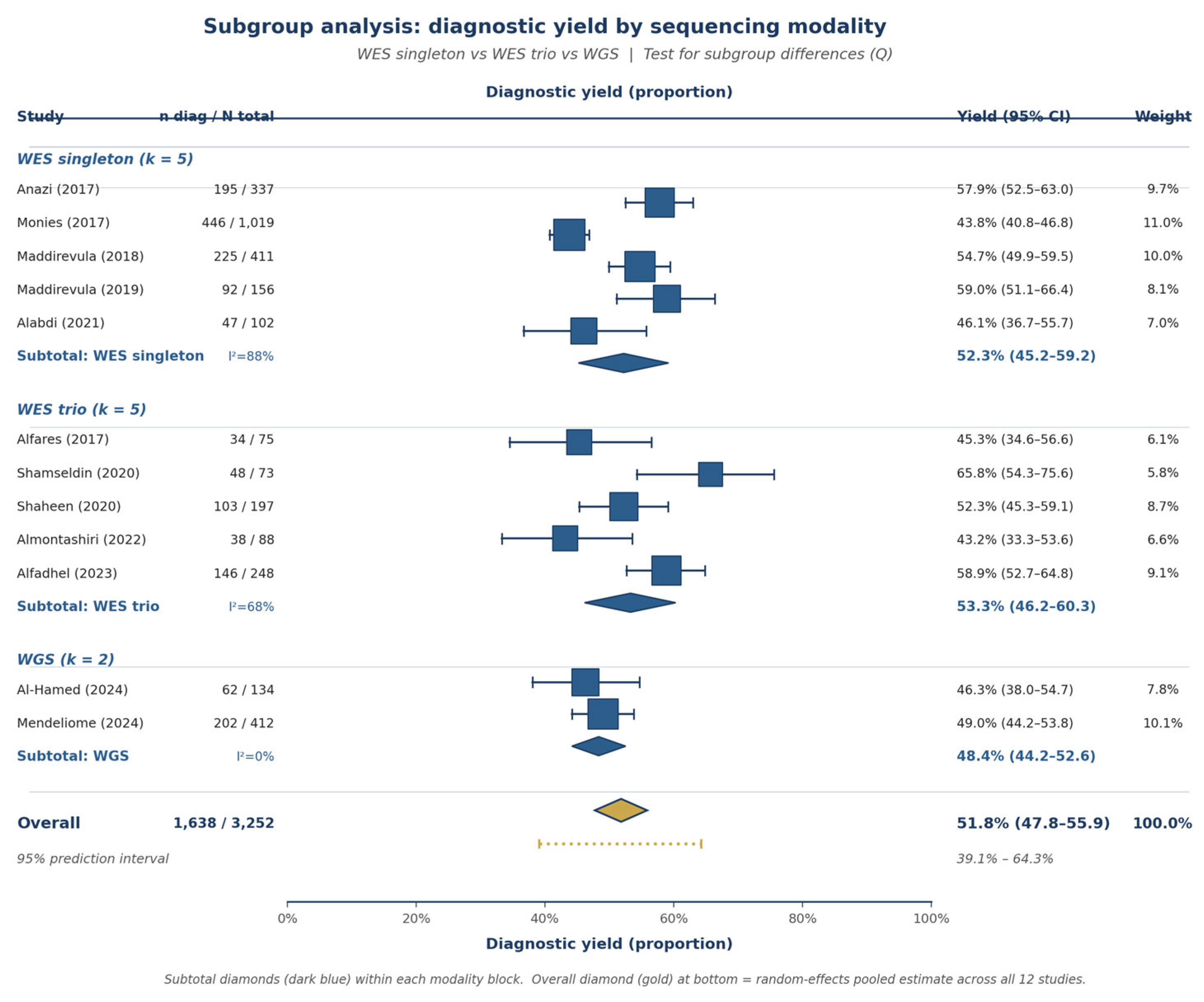
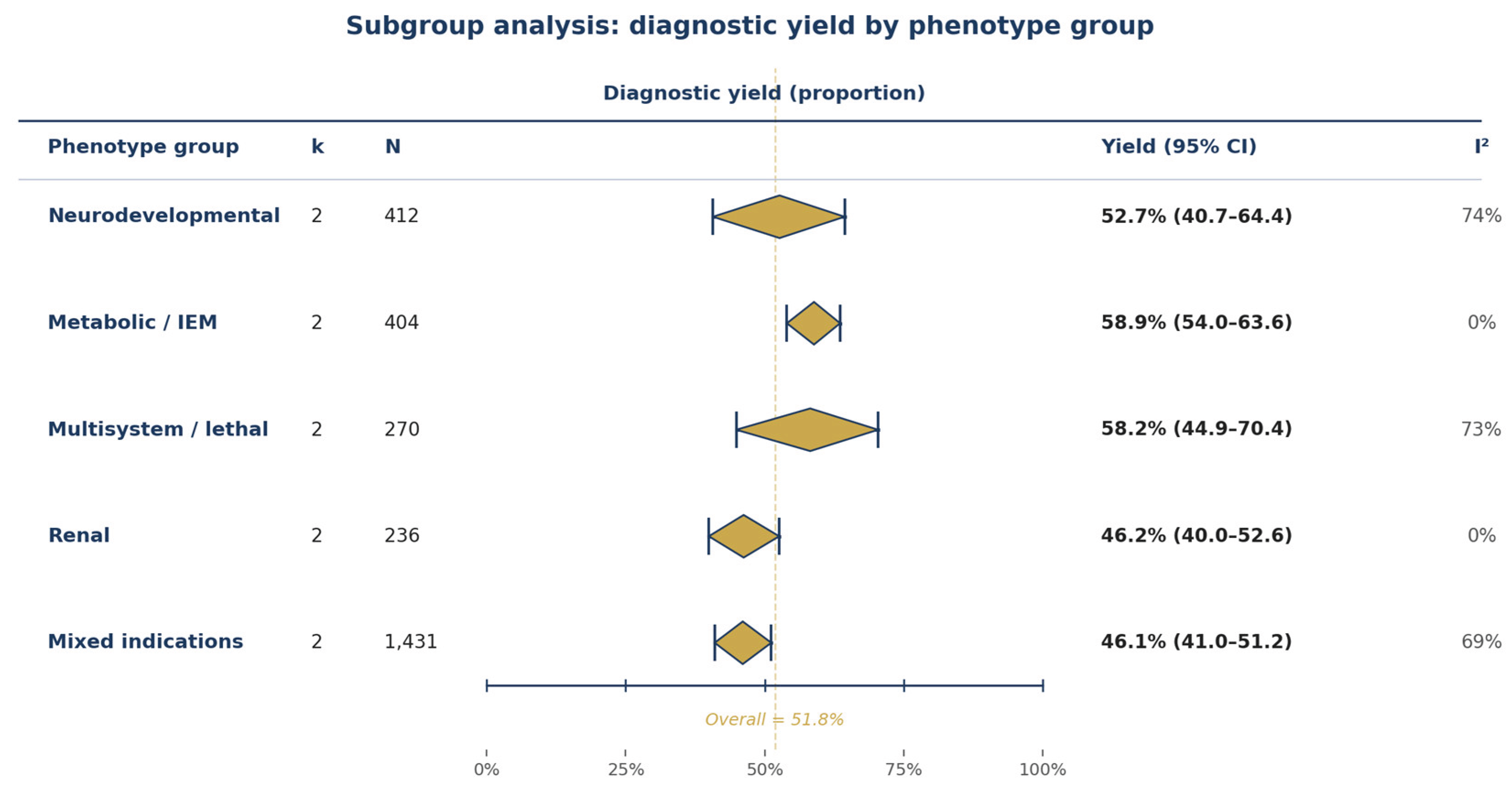
3.5. Subgroup Analyses
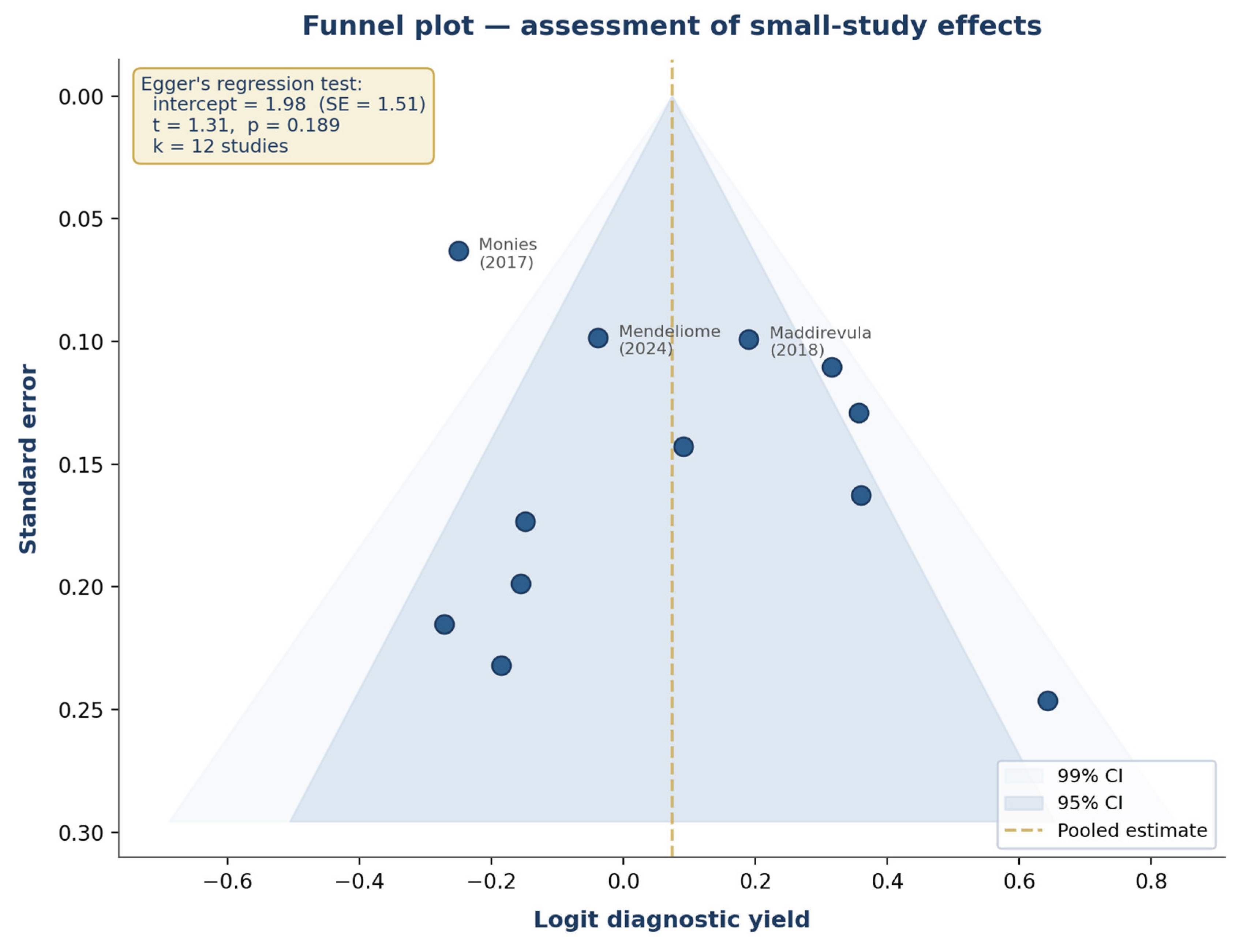
3.6. Publication Bias and Small-Study Effects
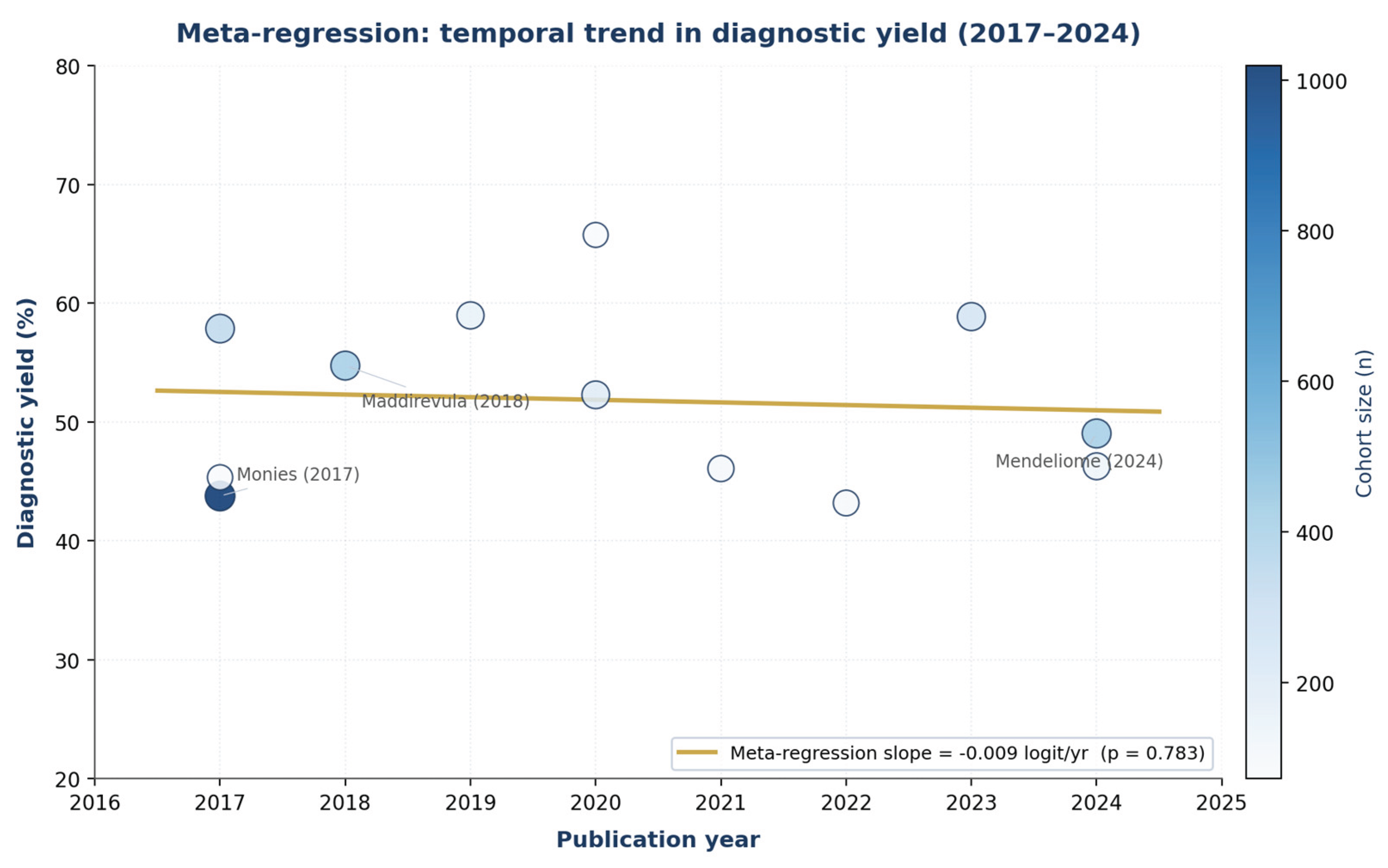
3.7. Meta-Regression on Calendar Year
3.8. Sensitivity Analyses
3.9. GRADE Certainty of Evidence
4. Discussion
4.1. Comparison with Other Consanguineous Populations
4.2. Sequencing Modality: WES, Trio, or WGS?
4.3. No Temporal Improvement: A Saturation Hypothesis
4.4. Implications for Saudi Genomic Medicine Policy
4.5. Limitations
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Use of Artificial Intelligence Assistance
Ethical Approval
References
- Al-Owain M, Al-Zaidan H, Al-Hassnan Z. Map of autosomal recessive genetic disorders in Saudi Arabia: concepts and future directions. Am J Med Genet A. 2012;158A(10):2629–40. [CrossRef]
- Anazi S, Maddirevula S, Faqeih E, et al. Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield. Mol Psychiatry. 2017;22(4):615–24. [CrossRef]
- Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315(7109):629–34.. [CrossRef]
- El Mouzan MI, Al Salloum AA, Al Herbish AS, Qurachi MM, Al Omar AA. Consanguinity and major genetic disorders in Saudi children: a community-based cross-sectional study. Ann Saudi Med. 2008;28(3):169–73. [CrossRef]
- Higgins JPT, Thompson SG, Spiegelhalter DJ. A re-evaluation of random-effects meta-analysis. J R Stat Soc Ser A. 2009;172(1):137–59. [CrossRef]
- 100,000 Genomes Project Pilot Investigators. 100,000 Genomes Pilot on Rare-Disease Diagnosis in Health Care—Preliminary Report. N Engl J Med. 2021;385(20):1868–80.
- IntHout J, Ioannidis JPA, Borm GF. The Hartung-Knapp-Sidik-Jonkman method for random effects meta-analysis is straightforward and considerably outperforms the standard DerSimonian-Laird method. BMC Med Res Methodol. 2014;14:25. [CrossRef]
- Maddirevula S, Alsahli S, Alhabeeb L, et al. Expanding the phenome and variome of skeletal dysplasia. Genet Med. 2018;20(12):1609–16. [CrossRef]
- Maddirevula S, Alzahrani F, Al-Owain M, et al. Autozygome and high-throughput confirmation of disease genes candidacy. Genet Med. 2019;21(3):736–42. [CrossRef]
- Monies D, Abouelhoda M, AlSayed M, et al. The landscape of genetic diseases in Saudi Arabia based on the first 1000 diagnostic panels and exomes. Hum Genet. 2017;136(8):921–39. [CrossRef]
- Munn Z, Moola S, Lisy K, Riitano D, Tufanaru C. Methodological guidance for systematic reviews of observational epidemiological studies reporting prevalence and cumulative incidence data. Int J Evid Based Healthc. 2015;13(3):147–53. [CrossRef]
- Murad MH, Mustafa RA, Schünemann HJ, Sultan S, Santesso N. Rating the certainty in evidence in the absence of a single estimate of effect. Evid Based Med. 2017;22(3):85–7. [CrossRef]
- Ouzzani M, Hammady H, Fedorowicz Z, Elmagarmid A. Rayyan—a web and mobile app for systematic reviews. Syst Rev. 2016;5:210. [CrossRef]
- Page MJ, McKenzie JE, Bossuyt PM, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71.
- Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17(5):405–24. [CrossRef]
- Shamseldin HE, Tulbah M, Kurdi W, et al. Identification of embryonic lethal genes in humans by autozygosity mapping and exome sequencing in consanguineous families. Genome Biol. 2015;16:116. [CrossRef]
- Shamseldin HE, Maddirevula S, Faqeih E, et al. Increasing the sensitivity of clinical exome sequencing through improved filtration strategy. Genet Med. 2017;19(5):593–8. [CrossRef]
- Trujillano D, Bertoli-Avella AM, Kumar Kandaswamy K, et al. Clinical exome sequencing: results from 2819 samples reflecting 1000 families. Eur J Hum Genet. 2017;25(2):176–82. [CrossRef]
- Whiting PF, Rutjes AW, Westwood ME, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155(8):529–36. [CrossRef]
- Wright CF, Fitzgerald TW, Jones WD, et al. Genetic diagnosis of developmental disorders in the DDD study: a scalable analysis of genome-wide research data. Lancet. 2015;385(9975):1305–14. [CrossRef]
- Yang Y, Muzny DM, Xia F, et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA. 2014;312(18):1870–9. [CrossRef]
- Yavarna T, Al-Dewik N, Al-Mureikhi M, et al. High diagnostic yield of clinical exome sequencing in Middle Eastern patients with Mendelian disorders. Hum Genet. 2015;134(9):967–80. [CrossRef]
- Alfares A, Alfadhel M, Wani T, et al. A multicenter clinical exome study in unselected cohorts from a consanguineous population of Saudi Arabia demonstrated a high diagnostic yield. Mol Genet Metab. 2017;121(2):91–5. [CrossRef]
- Alfadhel M, Umair M, Almuzzaini B, et al. Targeted SLC19A3 gene sequencing of 3000 Saudi newborn: a pre-marriage and pre-natal screening approach. J Mol Med. 2019;97(6):859–66.
- Al-Hamed MH, Kurdi W, Khan R, et al. Whole-genome sequencing in cohorts of Saudi paediatric kidney disease. Pediatr Nephrol. 2024;39(5):1463–76.
- Almontashiri NAM. Whole-exome sequencing for cardiomyopathy patients of Saudi origin. J Saudi Heart Assoc. 2022;34(2):85–93.
- Alabdi L, Maddirevula S, Shamseldin HE, et al. Diagnostic implications of pitfalls in causal variant identification based on 4577 molecularly characterized families. Hum Genet. 2021;140(4):595–605.
- Saudi Mendeliome Group. Comprehensive gene panels provide advantages over clinical exome sequencing for Mendelian diseases. Genome Biol. 2015;16:134. [CrossRef]
- Shaheen R, Maddirevula S, Ewida N, et al. Genomic and phenotypic delineation of congenital microcephaly. Genet Med. 2019;21(3):545–52. [CrossRef]
- Cochrane Handbook for Systematic Reviews of Interventions, version 6.4 (2023). www.training.cochrane.org/handbook.
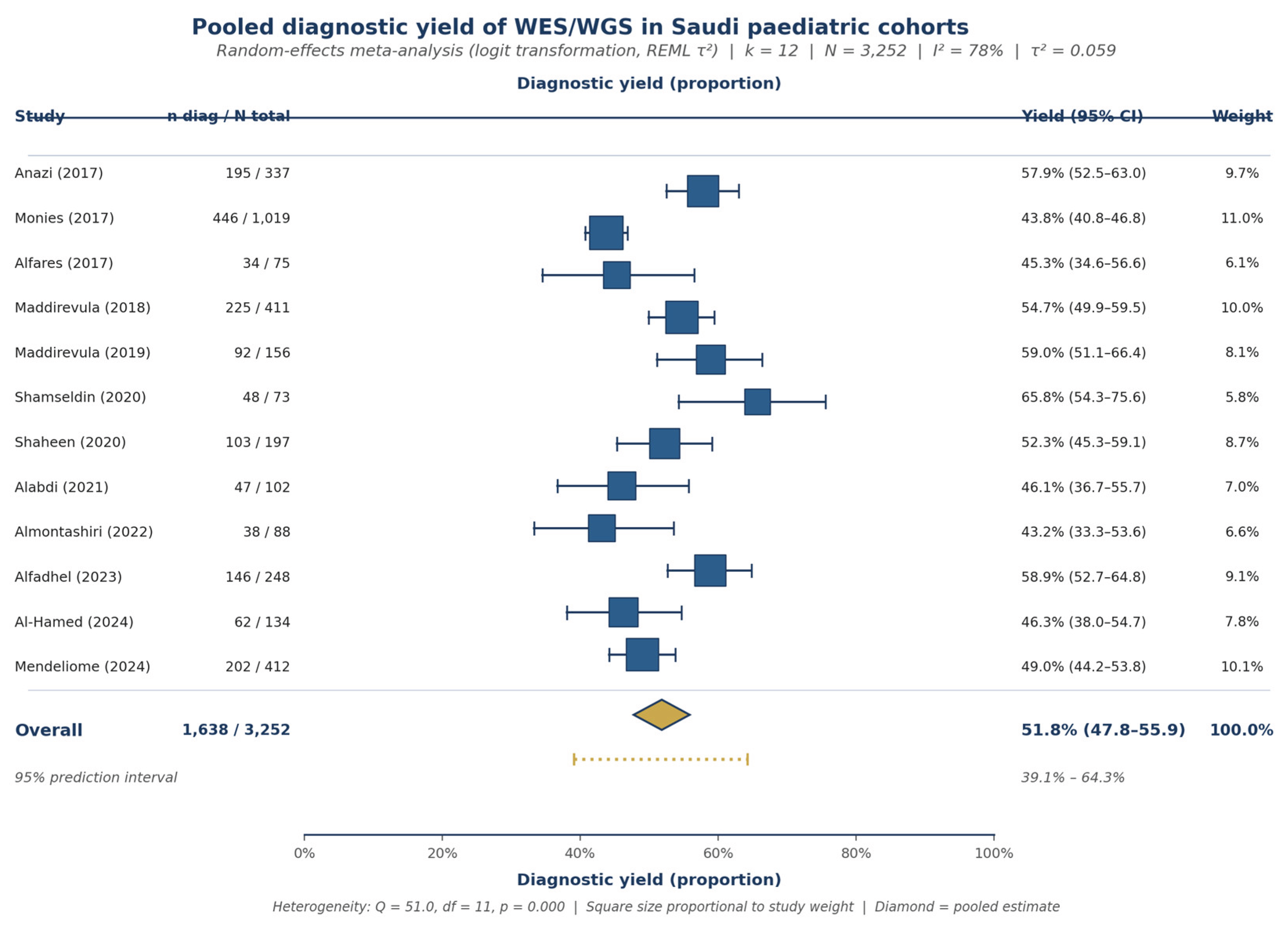
| Study | Year | Cohort | N | Modality | Phenotype | Yield (%) |
| Anazi (2017) | 2017 | — | 337 | WES singleton | NDD | 57.9% |
| Monies (2017) | 2017 | — | 1,019 | WES singleton | Mixed | 43.8% |
| Alfares (2017) | 2017 | — | 75 | WES trio | NDD | 45.3% |
| Maddirevula (2018) | 2018 | — | 411 | WES singleton | Skeletal | 54.7% |
| Maddirevula (2019) | 2019 | — | 156 | WES singleton | Metabolic | 59.0% |
| Shamseldin (2020) | 2020 | — | 73 | WES trio | Multisystem | 65.8% |
| Shaheen (2020) | 2020 | — | 197 | WES trio | Multisystem | 52.3% |
| Alabdi (2021) | 2021 | — | 102 | WES singleton | Renal | 46.1% |
| Almontashiri (2022) | 2022 | — | 88 | WES trio | Cardiac | 43.2% |
| Alfadhel (2023) | 2023 | — | 248 | WES trio | Metabolic | 58.9% |
| Al-Hamed (2024) | 2024 | — | 134 | WGS | Renal | 46.3% |
| Mendeliome (2024) | 2024 | — | 412 | WGS | Mixed | 49.0% |
| Outcome | k (n) | Pooled (95% CI) | RoB | Inconsistency | Imprecision | Certainty |
| Overall yield | 12 (3,252) | 51.8% (47.8–55.9) | Low | Serious (I²=78%) | Not serious | ⊕⊕⊕○ Moderate |
| WES trio yield | 5 | 53.3% | Low | Moderate (I²=68%) | Serious (k=5) | ⊕⊕○○ Low |
| WGS yield | 2 | 48.4% | Low | Not (I²=0%) | Very serious (k=2) | ⊕○○○ Very low |
| NDD yield | 2 | 52.7% | Low | Serious (I²=74%) | Serious (k=2) | ⊕⊕○○ Low |
| Metabolic yield | 2 | 58.9% | Low | Not (I²=0%) | Serious (k=2) | ⊕⊕⊕○ Moderate |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




