Submitted:
24 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
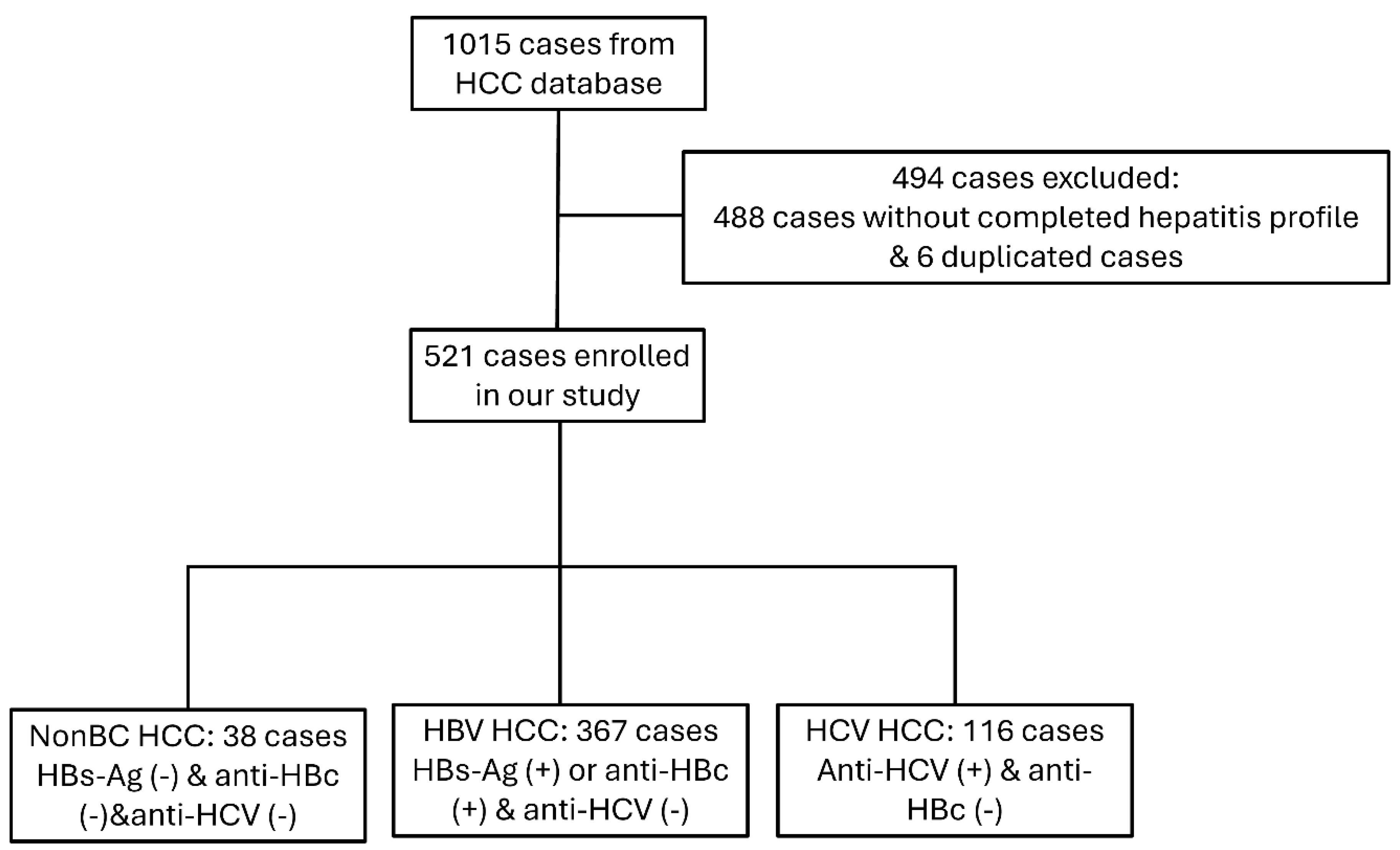
2.1. Subjects
2.2. Study Design
2.3. Statistical Analysis
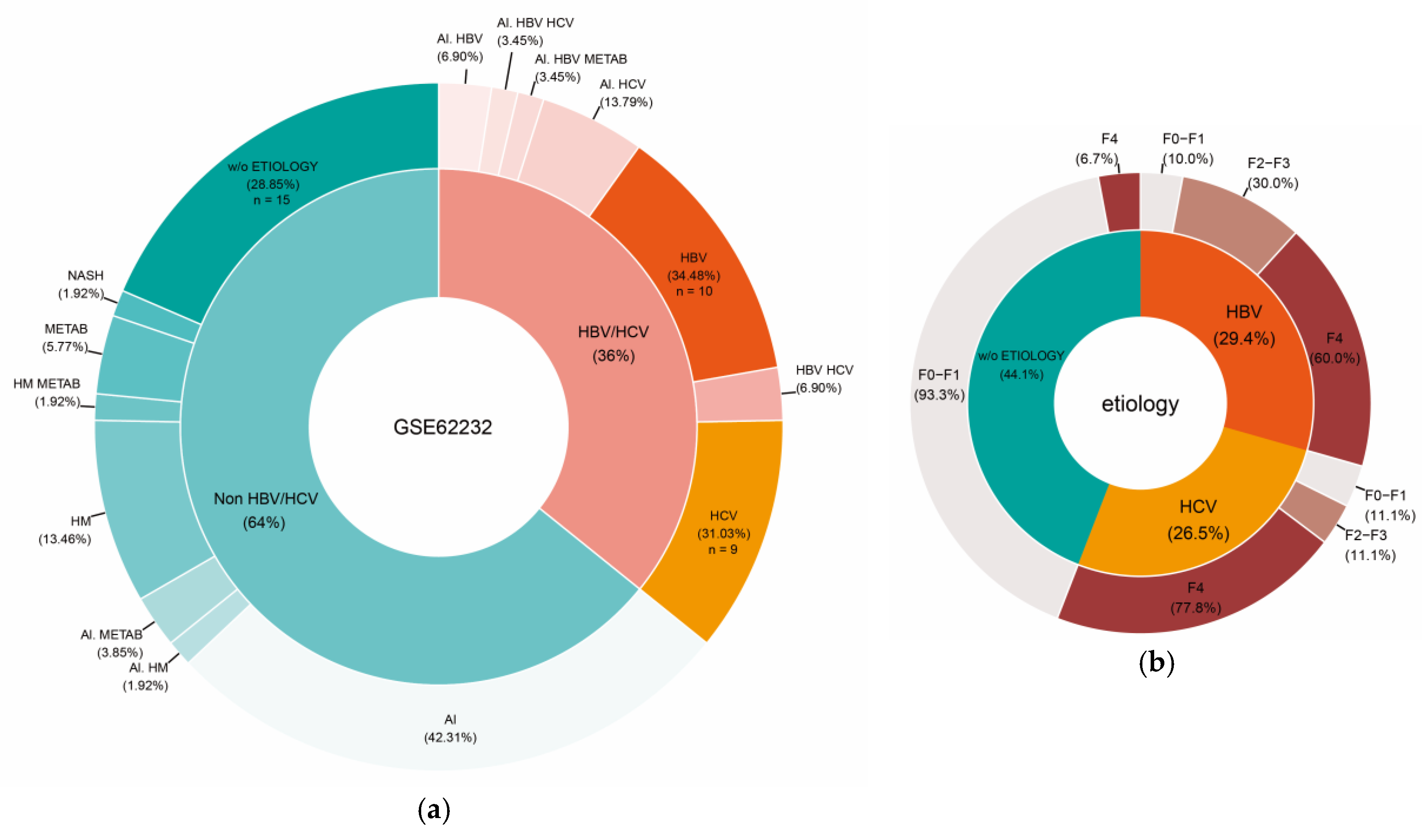
2.4. Gene Expression and Cell-Type Profiling
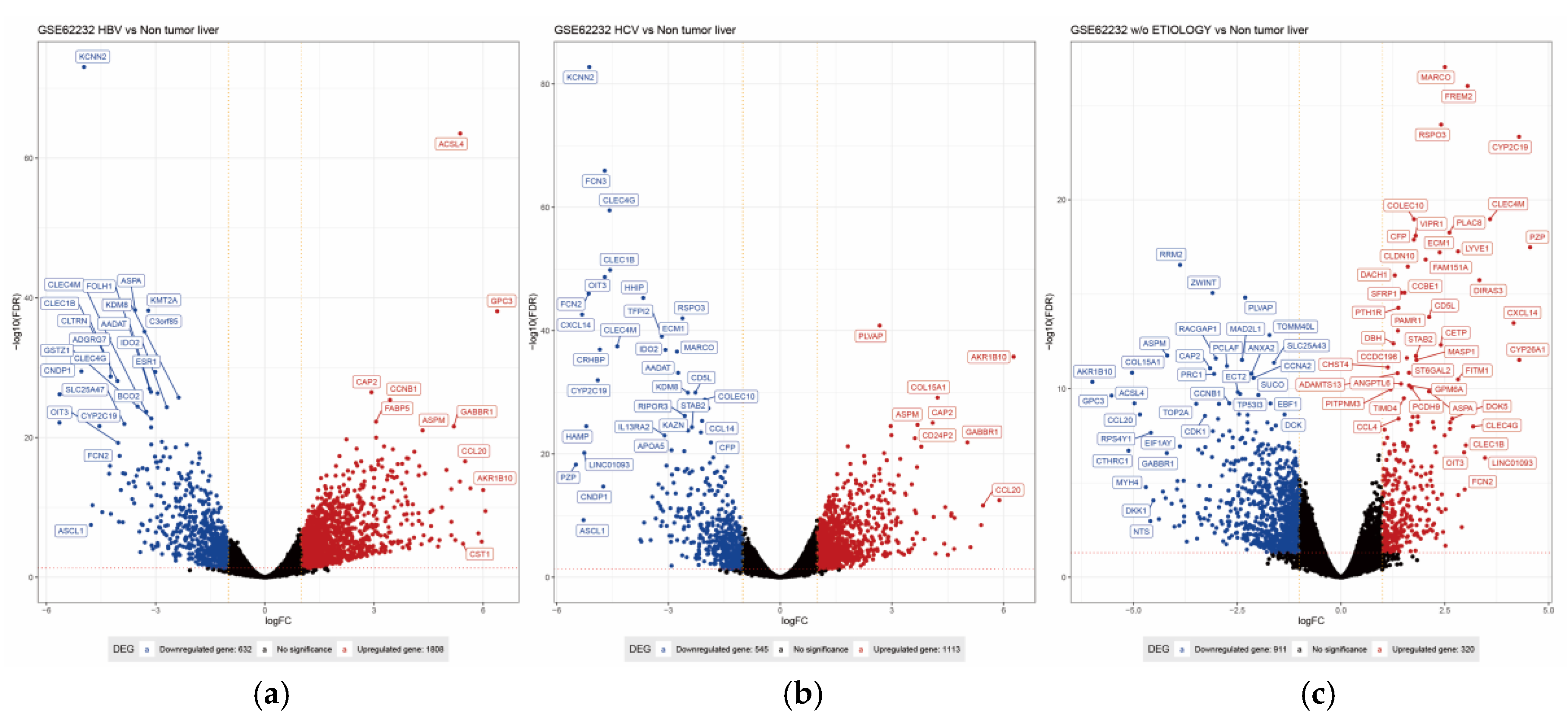
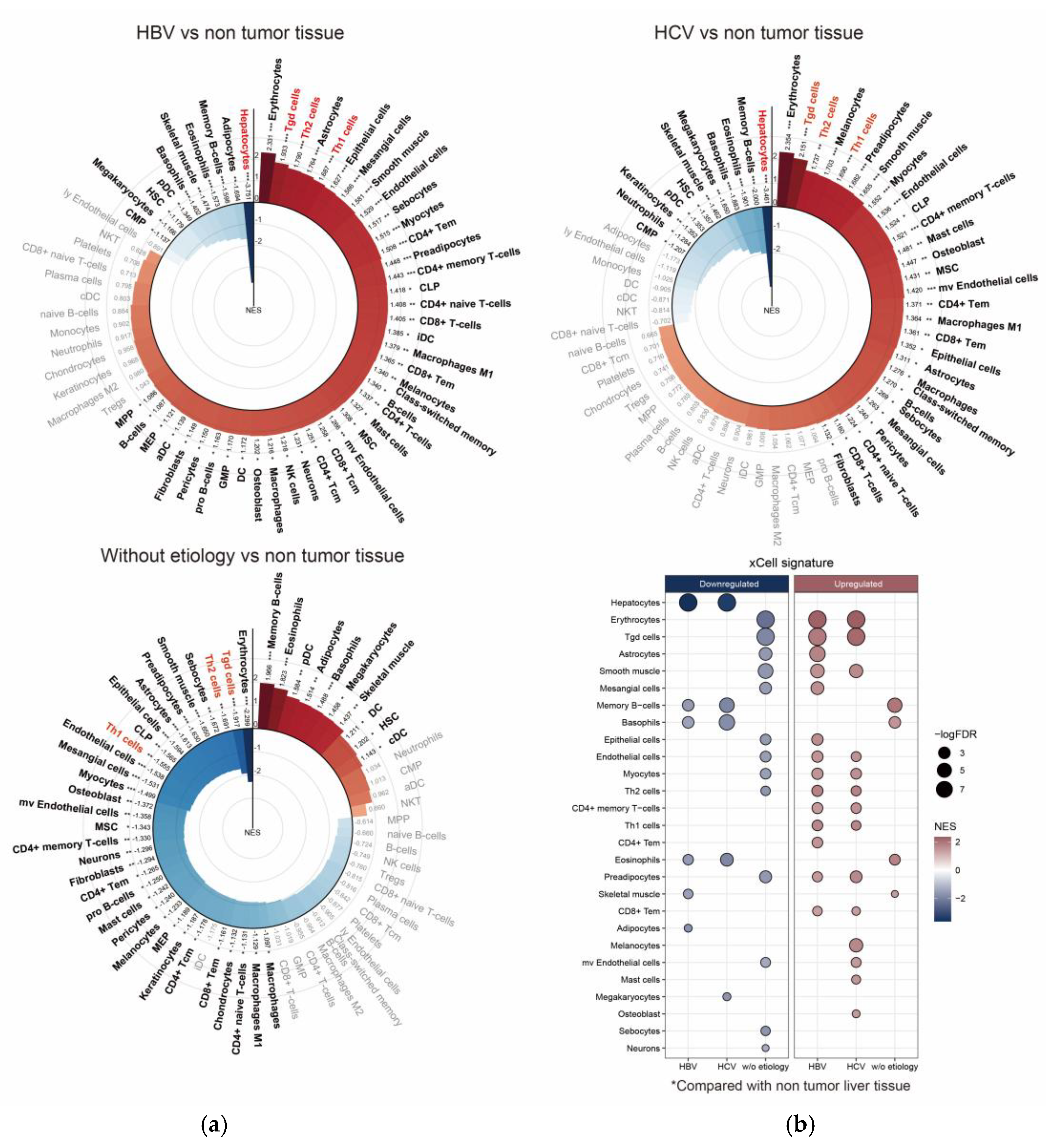
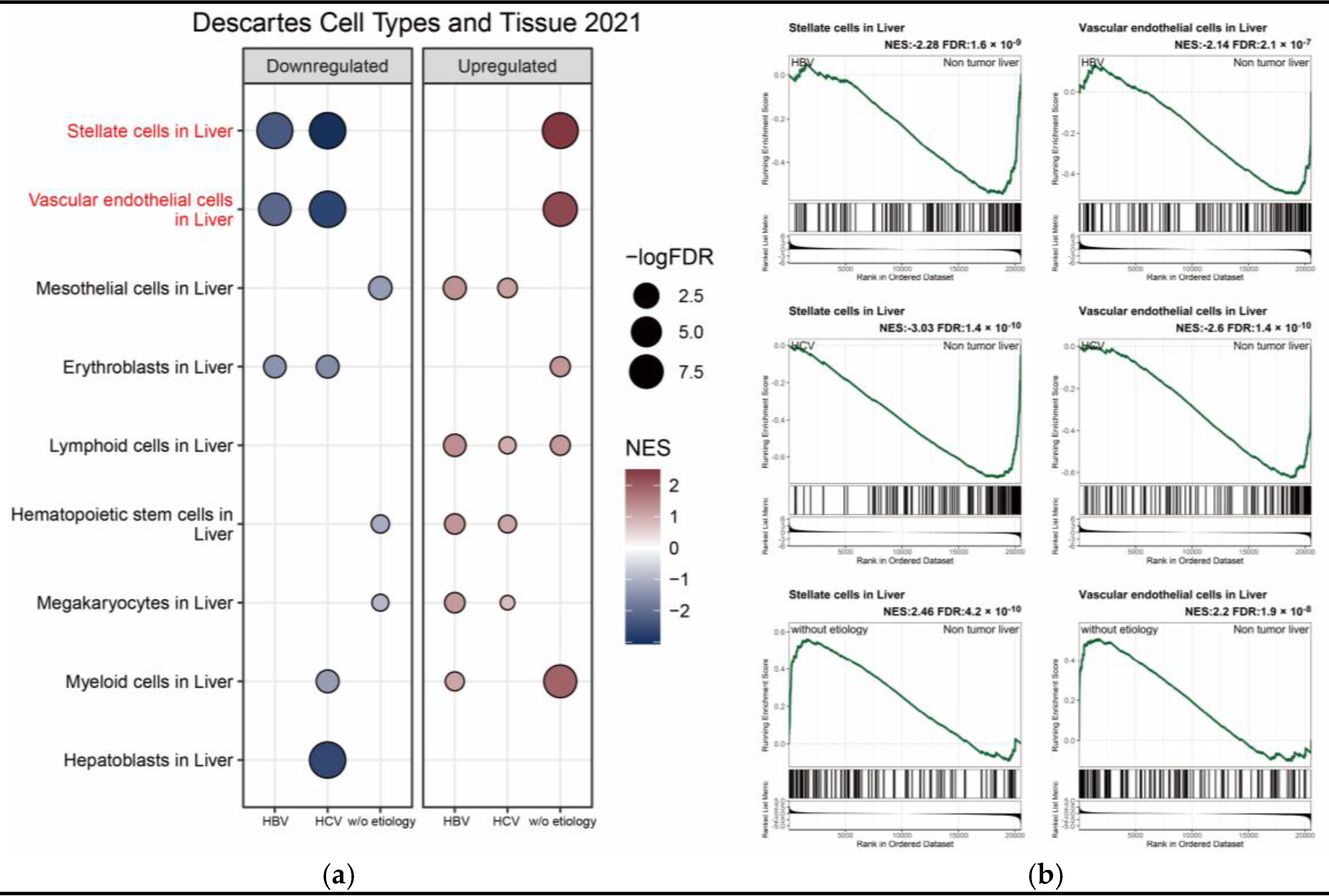
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AFP | Alpha-fetoprotein |
| ALT | Alanine aminotransferase |
| AST | Aspartate aminotransferase |
| BMI | Body mass index |
| CD | Cluster of differentiation |
| CMP | Common myeloid progenitors |
| CPL | Common lymphoid progenitors |
| DC | Dendritic cells |
| DEG | Differential Expression Gene |
| FDR | False discovery rate |
| GEO | Gene Expression Omnibus |
| GMP | Granulocyte myeloid progenitors |
| GSEA | Gene Set Enrichment Analysis |
| HBV | Hepatitis B virus |
| HCC | Hepatocellular carcinoma |
| HCV | Hepatitis C virus |
| HSC | Hematopoietic stem cells |
| INR | International normalized ratio |
| MELD-Na | Model for End-Stage Liver Disease (with Sodium) |
| MEP | Megakaryocyte–erythroid progenitor cells |
| MPP | Multipotent progenitors |
| MSC | Mesenchymal stem cells |
| NAFLD | Nonalcoholic fatty liver disease |
| NASH | Nonalcoholic steatohepatitis |
| NBNC | Non-HBV and non-HCV |
| NES | Normalized enrichment scores |
| NK cells | Natural killer cells |
| PLT | Platelet |
| S.E.M | Standard error of the mean |
| Tgd cell | Gamma delta T cell |
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R. L.; Torre, L. A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2018, 68(6), 394–424. [Google Scholar] [CrossRef] [PubMed]
- El-Serag, H. B.; Rudolph, K. L. Hepatocellular Carcinoma: Epidemiology and Molecular Carcinogenesis. Gastroenterology 2007, 132(7), 2557–2576. [Google Scholar] [CrossRef] [PubMed]
- KOGA, K. Clinical Aspects of Cryptogenic Hepatocellular Carcinoma. Kurume Med. J. 1998, 45(1), 105–111. [Google Scholar] [CrossRef] [PubMed]
- Fattovich, G.; Giustina, G.; Realdi, G.; Corrocher, R.; Schalm, S. W. Long-Term Outcome of Hepatitis B E Antigen-Positive Patients with Compensated Cirrhosis Treated with Interferon Alfa. Hepatology 1997, 26(5), 1338–1342. [Google Scholar] [CrossRef] [PubMed]
- Hu, K.-O.; Tong, M. The Long-Term Clinical Outcomes of Patients with HCV-Related Compensated Cirrhosis and History of Parenteral Exposure. Gastroenterology 1998, 114, A1258–A1259. [Google Scholar] [CrossRef]
- Serfaty, L.; Aumaître, H.; Chazouillères, O.; Bonnand, A.-M.; Rosmorduc, O.; Poupon, R. E.; Poupon, R. Determinants of Outcome of Compensated Hepatitis c Virus-Related Cirrhosis. Hepatology 1998, 27(5), 1435–1440. [Google Scholar] [CrossRef] [PubMed]
- Gawrieh, S.; Dakhoul, L.; Miller, E.; Scanga, A.; deLemos, A.; Kettler, C.; Burney, H.; Liu, H.; Abu-Sbeih, H.; Chalasani, N.; et al. Characteristics, Aetiologies and Trends of Hepatocellular Carcinoma in Patients without Cirrhosis: A United States Multicentre Study. Aliment. Pharmacol. Ther. 2019, 50(7), 809–821. [Google Scholar] [CrossRef] [PubMed]
- Leung, C. Characteristics of Hepatocellular Carcinoma in Cirrhotic and Non-Cirrhotic Non-Alcoholic Fatty Liver Disease. World J. Gastroenterol. 2015, 21(4), 1189. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-H.; Lin, J.-W.; Wu, L.-C.; Liu, C.-H.; Lai, M.-S. National Antiviral Treatment Program and the Incidence of Hepatocellular Carcinoma and Associated Mortality in Taiwan. Med. Care 2013, 51(10), 908–913. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.-Y.; Lee, Y.-H.; Liu, P.-H.; Hsia, C.-Y.; Huang, Y.-H.; Lin, H.-C.; Chiou, Y.-Y.; Lee, F.-Y.; Huo, T.-I. Decrypting Cryptogenic Hepatocellular Carcinoma: Clinical Manifestations, Prognostic Factors and Long-Term Survival by Propensity Score Model. PLoS ONE 2014, 9(2), e89373. [Google Scholar] [CrossRef] [PubMed]
- Nagaoki, Y.; Hyogo, Hideyuki; Ando, Y.; Kosaka, Y.; Uchikawa, Shinsuke; Nishida, Y.; Teraoka, Y.; Morio, K.; Fujino, H.; Ono, A.; et al. Increasing Incidence of Non-HBV- and Non-HCV-Related Hepatocellular Carcinoma: Single-Institution 20-Year Study. BMC Gastroenterol. 2021, 21(1). [Google Scholar] [CrossRef] [PubMed]
- Harouaka, D.; Engle, R. E.; Wollenberg, K.; Diaz, G.; Tice, A. B.; Zamboni, F.; Govindarajan, S.; Alter, H.; Kleiner, D. E.; Farci, P. Diminished Viral Replication and Compartmentalization of Hepatitis c Virus in Hepatocellular Carcinoma Tissue. Proc. Natl. Acad. Sci. 2016, 113(5), 1375–1380. [Google Scholar] [CrossRef] [PubMed]
- Ringelhan, M.; O’Connor, T.; Protzer, U.; Heikenwalder, M. The Direct and Indirect Roles of HBV in Liver Cancer: Prospective Markers for HCC Screening and Potential Therapeutic Targets. J. Pathol. 2014, 235(2), 355–367. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Zhang, J.; Xu, Y.; Wang, J.; Zhao, H.; Lei, J.; Zhou, Y.; Chen, Y.; Wu, L.; Zhou, M.; et al. OIT3 Mediates Macrophage Polarization and Facilitates Hepatocellular Carcinoma Progression. Cancer Immunol. Immunother. 2022, 71(11), 2677–2689. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.-N.; Ma, Y.; Luo, X.; Ma, Z.; Ma, L.-N.; Ding, X.-C. AKR1B10 Expression Characteristics in Hepatocellular Carcinoma and Its Correlation with Clinicopathological Features and Immune Microenvironment. Sci. Rep. 2024, 14(1). [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Jing, Y.; Xu, W.; Shi, X.; Zhang, W.; Chen, P.; Cao, X.; Han, X.; Duan, X.; Ren, J. The C-Type Lectin COLEC10 Is Predominantly Produced by Hepatic Stellate Cells and Involved in the Pathogenesis of Liver Fibrosis. Cell. Death Dis. 2023, 14(11). [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Shuai, C.; Gao, S.; Jiang, J.; Luan, J.; Lv, X. Chemokine CXCL14 Acts as a Potential Genetic Target for Liver Fibrosis. Int. Immunopharmacol. 2020, 89, 107067. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Ma, N.; Luo, H.; Chen, S.; Yu, F. Downregulated Long Non-Coding RNA LINC01093 in Liver Fibrosis Promotes Hepatocyte Apoptosis via Increasing Ubiquitination of SIRT1. J. Biochem. 2020, 168(3), 315–315. [Google Scholar] [CrossRef] [PubMed]
- Berumen, J.; Baglieri, J.; Kisseleva, T.; Mekeel, K. Liver Fibrosis: Pathophysiology and Clinical Implications. WIREs Mech. Dis. 2020, 13(1). [Google Scholar] [CrossRef] [PubMed]
| NBNC, N=38 | HBV, N=367 | HCV, N=116 | P value | ||
|---|---|---|---|---|---|
| NBNC versus HBV | NBNC versus HCV | ||||
| Age(years) | 64.45(41-91) | 62.88(27-93) | 68.62(40-97) | 0.7170 | 0.1432 |
| Sex(Male/Female) | 24/14 | 279/88 | 67/49 | 0.1141 | 0.5756 |
| BMI | 24.86(19.65-32.19) | 24.67(16.16-65.31) | 24.02(15.81-33.73) | 0.9687 | 0.5954 |
| Alcohol (Alcohol/Non-alcohol) | 8/30 | 91/276 | 24/92 | 0.6093 | 0.9618 |
| Creatinine | 1.019(0.5000-2.200) | 1.252(12.60-0.2000) | 1.356(0.4000-14.90) | 0.6014 | 0.4146 |
| Total bilirubin | 1.072(0.3000-4.400) | 2.014(0.2000-30.00) | 2.538(0.3000-42.40) | 0.4172 | 0.1736 |
| INR | 1.081(0.9000-1.800) | 1.093(0.7000-2.700) | 1.074(0.9-1.700) | 0.9288 | 0.9756 |
| AFP | 1685(0.000-21700) | 3522(0.000-42500) | 961.2(0.000-37500) | 0.5019 | 0.9142 |
| ALT | 41.63(4.000-199.0) | 56.19(4.000-505.0) | 56.49(6.000-469.0) | 0.4001 | 0.4520 |
| PLT | 209.3(53.00-484.0) | 199.0(30.00-698.0) | 164.5(38.00-524.0) | 0.8137 | 0.0415 |
| Fib-4 index | 3.191(0.2400-16.39) | 4.212(0.5300-34.67) | 6.077(0.8200-38.93) | 0.3804 | 0.0019 |
| MELD-Na score | 10.95(6.400-20.64) | 12.08(6.400-44.25) | 12.14(6.400-39.39) | 0.5782 | 0.6048 |
| Child Pugh score (A/B/C) | 20/8/1 | 224/61/26 | 62/22/7 | 0.4336 | 0.7051 |
| HCC stage (I/II/III/IV) | 14/6/8/8 | 136/40/131/56 | 44/28/30/13 | 0.2989 | 0.3569 |
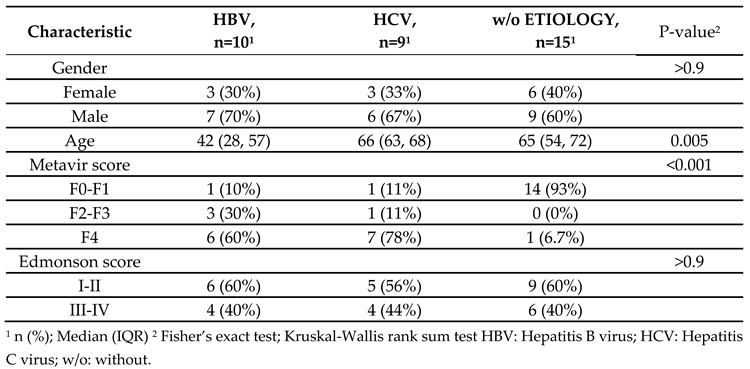 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




