Submitted:
24 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
DNA Extraction and Microbiome Sequencing
Statistical Analysis
Results
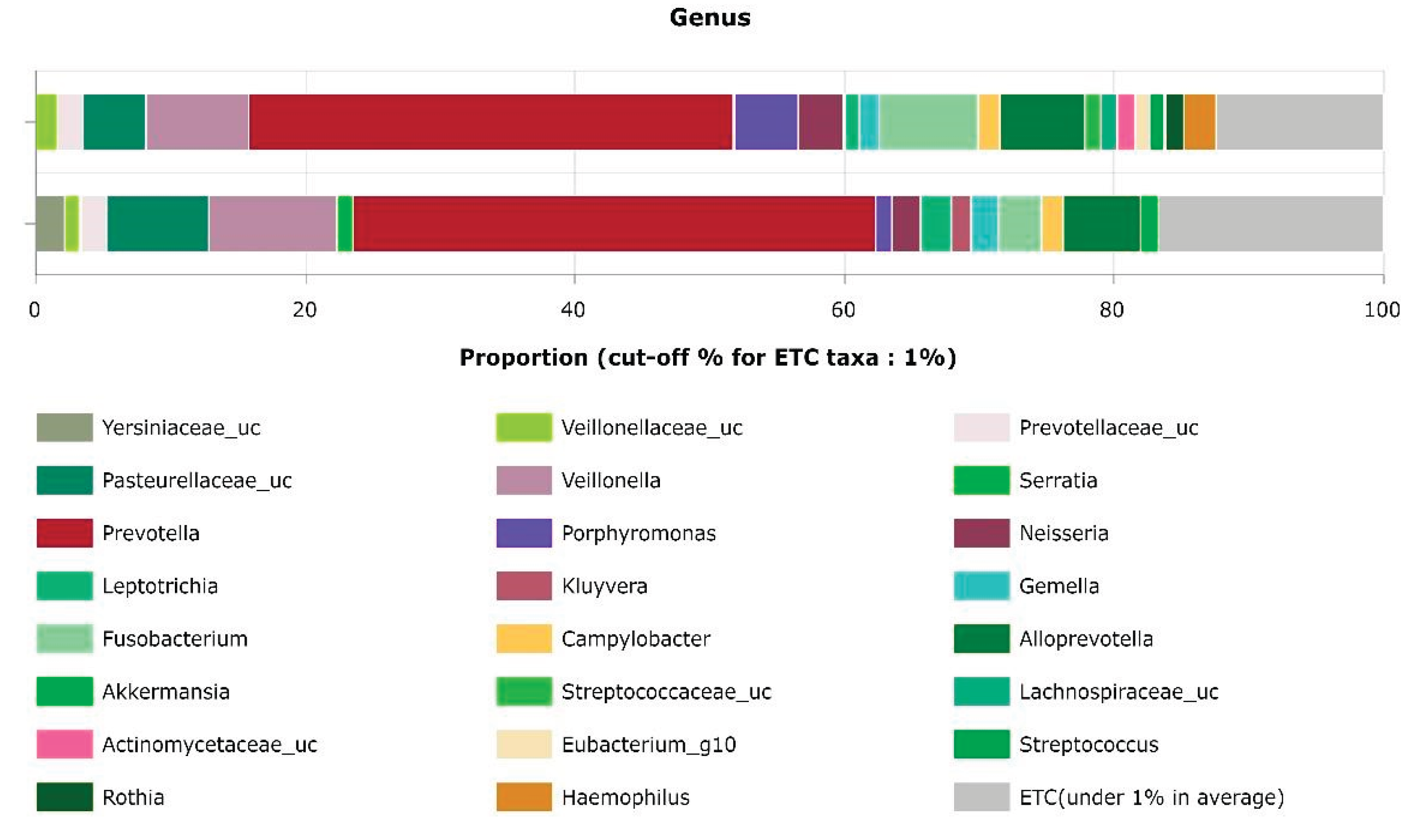
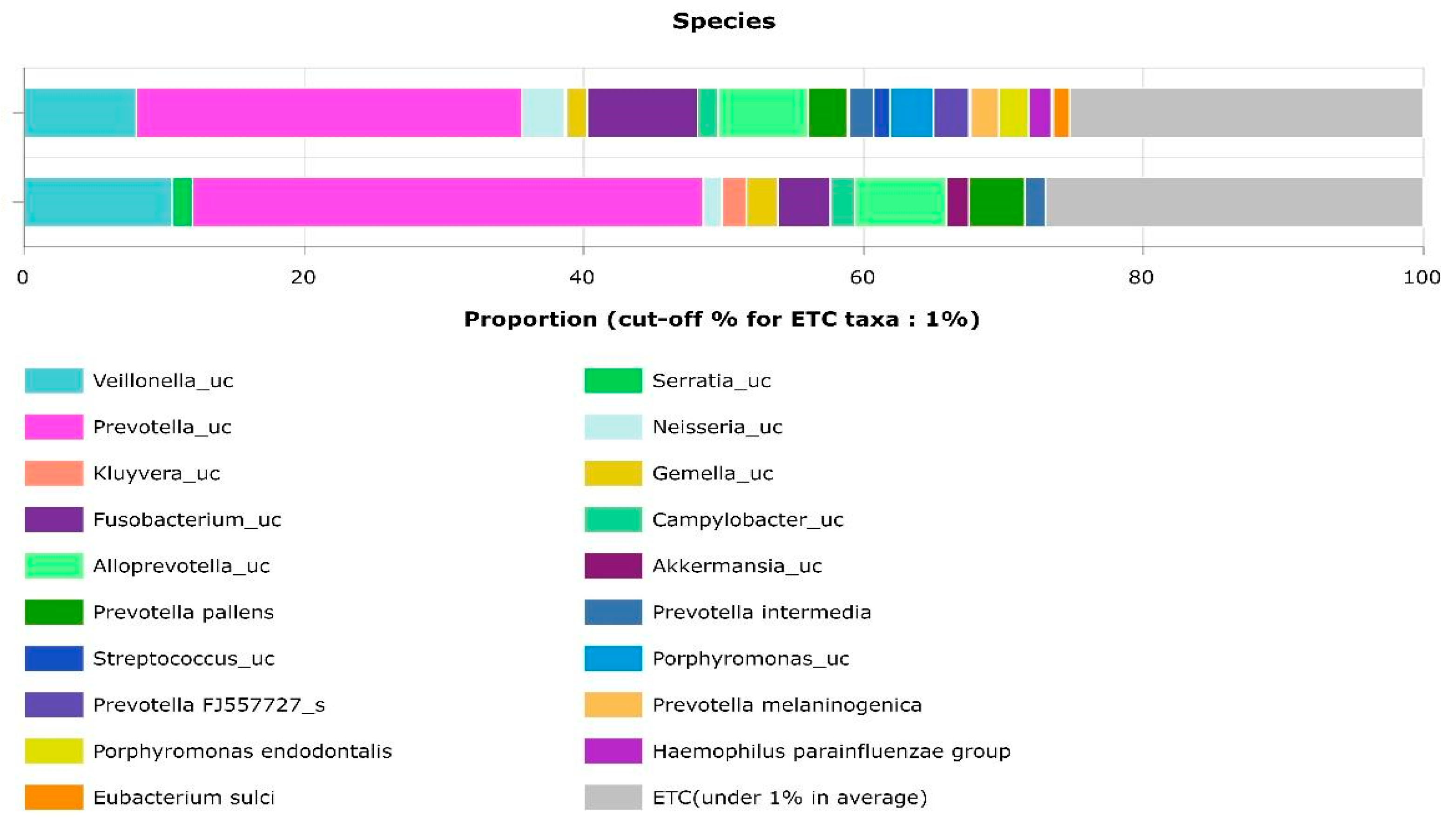
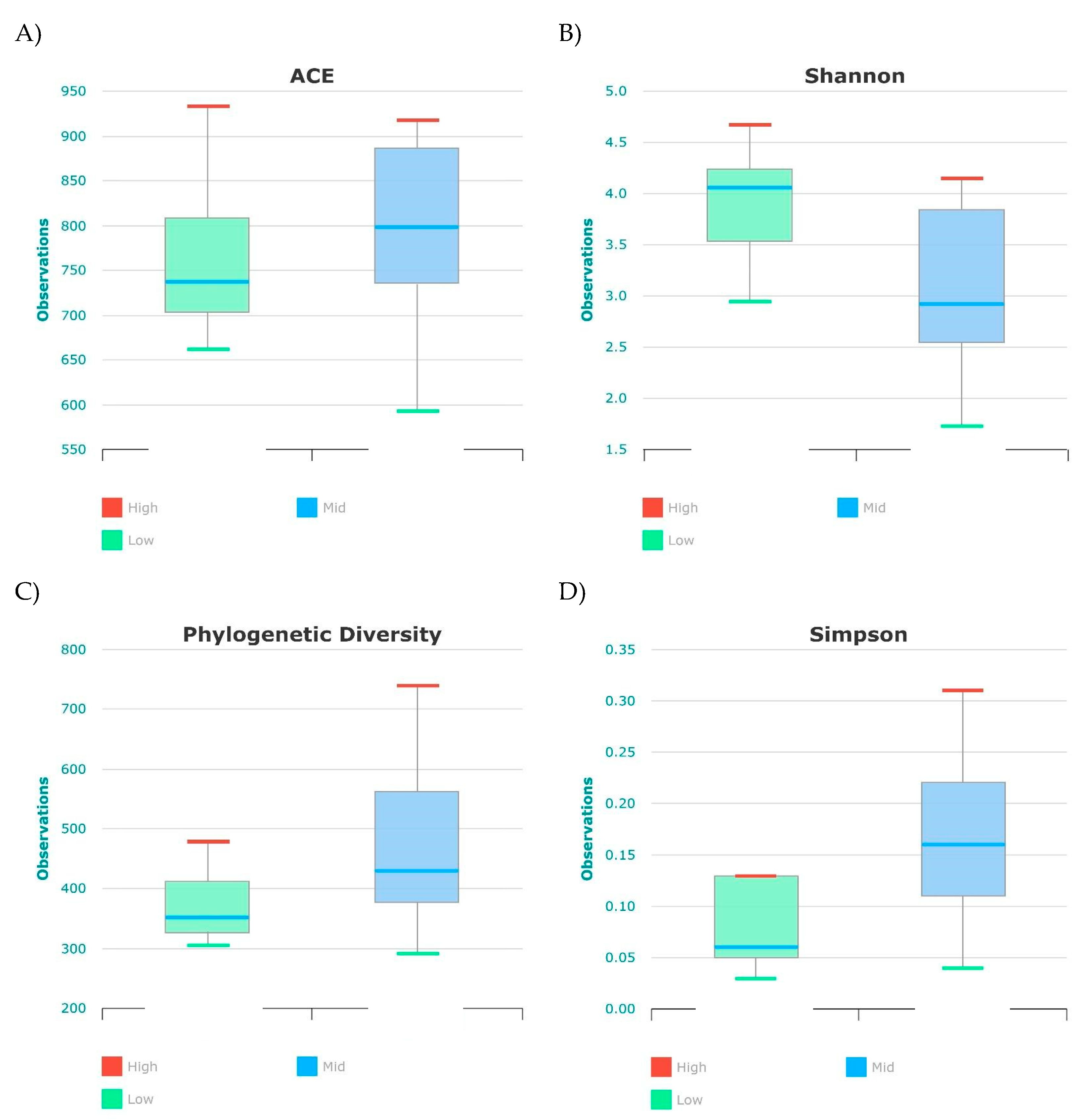
Upper Airway Microbiome
Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chong, B.; Jayabaskaran, J.; Jauhari, S.M.; Chan, S.P.; Goh, R.; Kueh, M.T.W.; Li, H.; Chin, Y.H.; Kong, G.; Anand, V.V.; et al. Global Burden of Cardiovascular Diseases: Projections from 2025 to 2050. Eur. J. Prev. Cardiol. 2024. [CrossRef]
- Mensah, G.A.; Habtegiorgis Abate, Y.; Abbasian, M.; Abd-Allah, F.; Abdollahi, A.; Abdollahi, M.; Morad Abdulah, D.; Abdullahi, A.; Abebe, A.M.; Abedi, A.; et al. Global Burden of Cardiovascular Diseases and Risks, 1990-2022. J. Am. Coll. Cardiol. 2023, 82, 2350–2473. [CrossRef]
- Roth, G.A.; Mensah, G.A.; Johnson, C.O.; Addolorato, G.; Ammirati, E.; Baddour, L.M.; Barengo, N.C.; Beaton, A.; Benjamin, E.J.; Benziger, C.P.; et al. Global Burden of Cardiovascular Diseases and Risk Factors, 1990-2019: Update From the GBD 2019 Study. J. Am. Coll. Cardiol. 2020, 76, 2982–3021.
- Morgan, A.D.; Zakeri, R.; Quint, J.K. Defining the Relationship between COPD and CVD: What Are the Implications for Clinical Practice? Ther. Adv. Respir. Dis. 2018, 12, 1753465817750524. [CrossRef]
- Garcia-Nuñez, M.; Millares, L.; Pomares, X.; Ferrari, R.; Pérez-Brocal, V.; Gallego, M.; Espasa, M.; Moya, A.; Monsó, E. Severity-Related Changes of Bronchial Microbiome in Chronic Obstructive Pulmonary Disease. J. Clin. Microbiol. 2014, 52, 4217–4223. [CrossRef]
- Lee, J.J.; Kim, S.H.; Lee, M.J.; Kim, B.K.; Song, W.J.; Park, H.W.; Cho, S.H.; Hong, S.J.; Chang, Y.S.; Kim, B.S. Different Upper Airway Microbiome and Their Functional Genes Associated with Asthma in Young Adults and Elderly Individuals. Allergy 2019, 74, 709–719. [CrossRef]
- Koren, O.; Spor, A.; Felin, J.; Fåk, F.; Stombaugh, J.; Tremaroli, V.; Behre, C.J.; Knight, R.; Fagerberg, B.; Ley, R.E.; et al. Human Oral, Gut, and Plaque Microbiota in Patients with Atherosclerosis. Proc. Natl. Acad. Sci. U. S. A. 2011, 108 Suppl 1, 4592–4598. [CrossRef]
- Nazari, Z.; Abiri, R.; Mohajerani, H.R. Comparison of the Oral Microbiota of Patients with Atherosclerosis and Healthy Controls by Denaturing Gradient Gel Electrophoresis. Cell. Mol. Biol. 2024, 70, 201–206. [CrossRef]
- Fåk, F.; Tremaroli, V.; Bergström, G.; Bäckhed, F. Oral Microbiota in Patients with Atherosclerosis. Atherosclerosis 2015, 243, 573–578. [CrossRef]
- Cai, Y.Y.; Huang, F.Q.; Lao, X.; Lu, Y.; Gao, X.; Alolga, R.N.; Yin, K.; Zhou, X.; Wang, Y.; Liu, B.; et al. Integrated Metagenomics Identifies a Crucial Role for Trimethylamine-Producing Lachnoclostridium in Promoting Atherosclerosis. NPJ Biofilms Microbiomes 2022, 8. [CrossRef]
- Ramirez, J.A. Isolation of Chlamydia Pneumoniae from the Coronary Artery of a Patient with Coronary Atherosclerosis. The Chlamydia Pneumoniae/Atherosclerosis Study Group. Ann. Intern. Med. 1996, 125, 979–982. [CrossRef]
- Stark, B.A.; DeCleene, N.K.; Desai, E.C.; Hsu, J.M.; Johnson, C.O.; Lara-Castor, L.; LeGrand, K.E.; A, P.B.; Aalipour, M.A.; Aalruz, H.; et al. Global, Regional, and National Burden of Cardiovascular Diseases and Risk Factors in 204 Countries and Territories, 1990-2023. J. Am. Coll. Cardiol. 2025, 86, 2167–2243. [CrossRef]
- Rojas, Y.G.; Duque, C.A.T.; Figueredo, M. del C.; Hernández, F.; Cardona, C.C.; Lasalvia, P.; Rosselli, D. Estimación de La Prevalencia de EPOC En Colombia a Partir Del Registro Individual de Prestaciones de Servicios de Salud (RIPS). Revista Colombiana de Neumología 2019, 31, 5–15. [CrossRef]
- Caballero, A.; Torres-Duque, C.A.; Jaramillo, C.; Bolívar, F.; Sanabria, F.; Osorio, P.; Orduz, C.; Guevara, D.P.; Maldonado, D. Prevalence of COPD in Five Colombian Cities Situated at Low, Medium, and High Altitude (PREPOCOL Study). Chest 2008, 133, 343–349. [CrossRef]
- Ospina, J.; Garcia-Morales, O.M.; Gaviria, M.C. Beyond Reported Rates: Detection-Adjusted COPD Prevalence and Underdiagnosis Patterns in Colombia. Int. J. Chron. Obstruct. Pulmon. Dis. 2026, 21, 1–13. [CrossRef]
- Hou, D.; Liu, Z.; Li, X.; Shen, P.; Li, W.; Zhang, M.; Cheang, I.F.; Lin, H.; Zhan, S.; Sun, F.; et al. Impacts of COPD Exacerbation History on Mortality and Severe Cardiovascular Events among Patients with COPD in China: A Retrospective Cohort Study. Respir. Res. 2025, 26, 252. [CrossRef]
- Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43(11):5721–32. [CrossRef]
- Al Bataineh MT, Hamoudi RA, Dash NR, Ramakrishnan RK, Almasalmeh MA, Sharif HA, et al. Altered respiratory microbiota composition and functionality associated with asthma early in life. BMC Infect Dis. 2020;20(1):697. [CrossRef]
- Zaura E, Keijser BJ, Huse SM, Crielaard W. Defining the healthy "core microbiome" of oral microbial communities. BMC Microbiol. 2009;9:259. [CrossRef]
- Li B, Xia Y, Hu B. Infection and atherosclerosis: TLR-dependent pathways. Cell Mol Life Sci. 2020;77(14):2751–69. [CrossRef]
- Kurita-Ochiai T, Yamamoto M. Periodontal pathogens and atherosclerosis: implications of inflammation and oxidative modification of LDL. Biomed Res Int. 2014;2014:595981. [CrossRef]
- Schenkein HA, Papapanou PN, Genco R, Sanz M. Mechanisms underlying the association between periodontitis and atherosclerotic disease. Periodontol 2000. 2020;83(1):90–106. [CrossRef]
- Velsko IM, Chukkapalli SS, Rivera-Kweh MF, Zheng D, Aukhil I, Lucas AR, et al. Periodontal pathogens invade gingiva and aortic adventitia and elicit inflammasome activation in αvβ6 integrin-deficient mice. Infect Immun. 2015;83(12):4582–93. [CrossRef]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–14. [CrossRef]
- Dickson RP, Erb-Downward JR, Martinez FJ, Huffnagle GB. The microbiome and the respiratory tract. Annu Rev Physiol. 2016;78:481–504. [CrossRef]
- Guan WJ, Yuan JJ, Li HM, Gao YH, Huang Y, Chen CL, et al. Proteobacteria community compositions correlate with bronchiectasis severity. Int J Tuberc Lung Dis. 2018;22(9):1095–105. [CrossRef]
- Wang Z, Bafadhel M, Haldar K, Spivak A, Mayhew D, Miller BE, et al. Lung microbiome dynamics during COPD exacerbations. Am J Respir Crit Care Med. 2016;194(8):916–27. [CrossRef]
- Jie Z, Xia H, Zhong SL, Feng Q, Li S, Liang S, et al. The gut microbiome in atherosclerotic cardiovascular disease. Nat Commun. 2017;8(1):845. [CrossRef]
- Jia W, Li H, Zhao L, Nicholson JK. Gut microbiota: a potential new territory for drug targeting. Nat Rev Drug Discov. 2008;7(2):123–9. [CrossRef]
- Zhu Q, Gao R, Wu W, Qin H. The role of gut microbiota in atherosclerosis and related diseases. Front Pharmacol. 2018;9:454. [CrossRef]
- Liu Z, Liu H-Y, Zhou H, Zhan Q, Lai W, Zeng Q, et al. Moderate-intensity exercise affects gut microbiome composition and influences cardiac function in myocardial infarction mice. Front Microbiol. 2017;8:1687. [CrossRef]
- Gebrayel P, Nicco C, Al Khodor S, Bilinski J, Caselli E, Comelli EM, et al. Microbiota medicine: towards clinical revolution. J Transl Med. 2022;20(1):111. [CrossRef]
- Lynch SV, Pedersen O. The human intestinal microbiome in health and disease. N Engl J Med. 2016;375(24):2369–79. [CrossRef]
- Lee JT, Kim CM, Ramakrishnan V. Microbiome and disease in the upper airway. Curr Opin Allergy Clin Immunol. 2019;19(1):1–6. [CrossRef]
- Yuan Y, Wang C, Wang G, Guo X, Jiang S, Zuo X, et al. Airway microbiome and serum metabolomics analysis identify differential candidate biomarkers in allergic rhinitis. Front Immunol. 2022;12:771136. [CrossRef]
- Park SC, Park I-H, Lee JS, Park SM, Kang SH, Hong S-M, et al. Microbiome of unilateral chronic rhinosinusitis: a controlled paired analysis. Int J Environ Res Public Health. 2021;18(18):9878. [CrossRef]
- Stearns JC, Davidson CJ, McKeon S, Whelan FJ, Fontes ME, Schryvers AB, et al. Culture and molecular-based profiles show shifts in bacterial communities of the upper respiratory tract that occur with age. ISME J. 2015;9(5):1246–59. [CrossRef]
- Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011;9(4):244–53. [CrossRef]
- Xu Y, Huang Y, Shen Z, Shi L. The nasal microbiome predicting bronchopulmonary dysplasia in preterm infants. Sci Rep. 2022;12(1):7727. [CrossRef]
- Bianco MR, Ralli M, Modica DM, Amata M, Poma S, Mattina G, et al. The role of probiotics in chronic rhinosinusitis treatment: an update of the current literature. Healthcare (Basel). 2021;9(12):1715. [CrossRef]
- Henares D, Brotons P, de Sevilla MF, Fernandez-Lopez A, Hernandez-Bou S, Perez-Argüello A, et al. Differential nasopharyngeal microbiota composition in children according to respiratory health status. Microb Genom. 2021;7(10):000661. [CrossRef]
- Charlson ES, Chen J, Custers-Allen R, Bittinger K, Li H, Sinha R, et al. Disordered microbial communities in the upper respiratory tract of cigarette smokers. PLoS One. 2010;5(12):e15216. [CrossRef]
- de Steenhuijsen Piters WAA, Huijskens EGW, Wyllie AL, Biesbroek G, van den Bergh MR, Veenhoven RH, et al. Dysbiosis of upper respiratory tract microbiota in elderly pneumonia patients. ISME J. 2016;10(1):97–108. [CrossRef]
- Ubags ND, Marsland BJ. Mechanistic insight into the function of the microbiome in lung diseases. Eur Respir J. 2017;50(3):170–89. [CrossRef]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–14. [CrossRef]
- Dickson RP, Erb-Downward JR, Martinez FJ, Huffnagle GB. The microbiome and the respiratory tract. Annu Rev Physiol. 2016;78:481–504. [CrossRef]
- Abusleme L, Dupuy AK, Dutzan N, Silva N, Burleson JA, Strausbaugh LD, et al. The subgingival microbiome in health and periodontitis and its relationship with community biomass and inflammation. ISME J. 2013;7(5):1016–25. [CrossRef]
- Abreu A, Milke-García M, Argüello-Arévalo G, Calderón-de la Barca A, Carmona-Sánchez R, Consuelo-Sánchez A, et al. Fibra dietaria y microbiota, revisión narrativa de un grupo de expertos de la Asociación Mexicana de Gastroenterología. Rev Gastroenterol Mex. 2021;86(3):287–304. [CrossRef]
- Li W, Ma Z. FBA ecological guild: trio of firmicutes-bacteroidetes alliance against actinobacteria in human oral microbiome. Sci Rep. 2020;10(1):287. [CrossRef]
- Tsay J-CJ, Wu BG, Sulaiman I, Gershner K, Schluger R, Li Y, et al. Lower airway dysbiosis affects lung cancer progression. Cancer Discov. 2021;11(2):293–307. [CrossRef]
- Huang YJ, Nariya S, Harris JM, Lynch SV, Choy DF, Arron JR, et al. The airway microbiome in patients with severe asthma: associations with disease features and severity. J Allergy Clin Immunol. 2015;136(4):874–84. [CrossRef]
| Variable | Controls (n=50) |
Cases (n=50) |
|---|---|---|
| Age, y.o. mean ± SD | 44.9 ± 16.2 | 59.7 ± 13.4 |
| Sex, % (n) | ||
| Women | 34.2 (17) | 46.4 (23) |
| Men | 65.8 (33) | 53.6 (27) |
| Economic activity, % (n) | ||
| Housewives | 20.8 (10) | 31.0 (16) |
| Retired | 10.8 (5) | 8.3 (4) |
| Unemployed | 13.1 (7) | 11.9 (6) |
| Employee | 51.9 (26) | 47.0 (24) |
| Unspecified | 3.4 (2) | 1.8 (1) |
| Monthly income, USD median [IQR] | 325 [150; 600] | 250 [125; 381.3] |
| Smoking status, % (n) | ||
| Never | 81.2 (41) | 71.4 (36) |
| Former | 14.5 (7) | 22.0 (11) |
| Current | 4.3 (2) | 6.5 (3) |
| Pack-years median [IQR] | 4.35 [2.3; 10.6] | 8.6 [2.7; 17.9] |
| Pulmonary function (Pre-bronchodilator), mean ± SD | ||
| FVC (L) | 3.22 ± 0.67 | 2.85 ± 0.65 |
| FEV₁ (L) | 2.51 ± 0.54 | 2.05 ± 0.48 |
| FEV₁/FVC | 78.36 ± 6.65 | 72.7 ± 7.89 |
| Pulmonary function (Post-bronchodilator), mean ± SD | ||
| CVF (L) | 3.18 ± 0.63 | 2.74 ± 0.58 |
| VEF₁ (L) | 2.59 ± 0.54 | 2.11 ± 0.49 |
| FEV₁/FVC | 81.82 ± 6.22 | 76.85 ± 6.20 |
| Carotid Intima-Media Thickness, mean ± SD | 0.52 ± 0.19 | 0.91 ± 0.33 |
| Framingham 10-year risk, mean ± SD | 18.78 ± 20.22 | 42.37 ± 22.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




