Submitted:
23 April 2026
Posted:
24 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
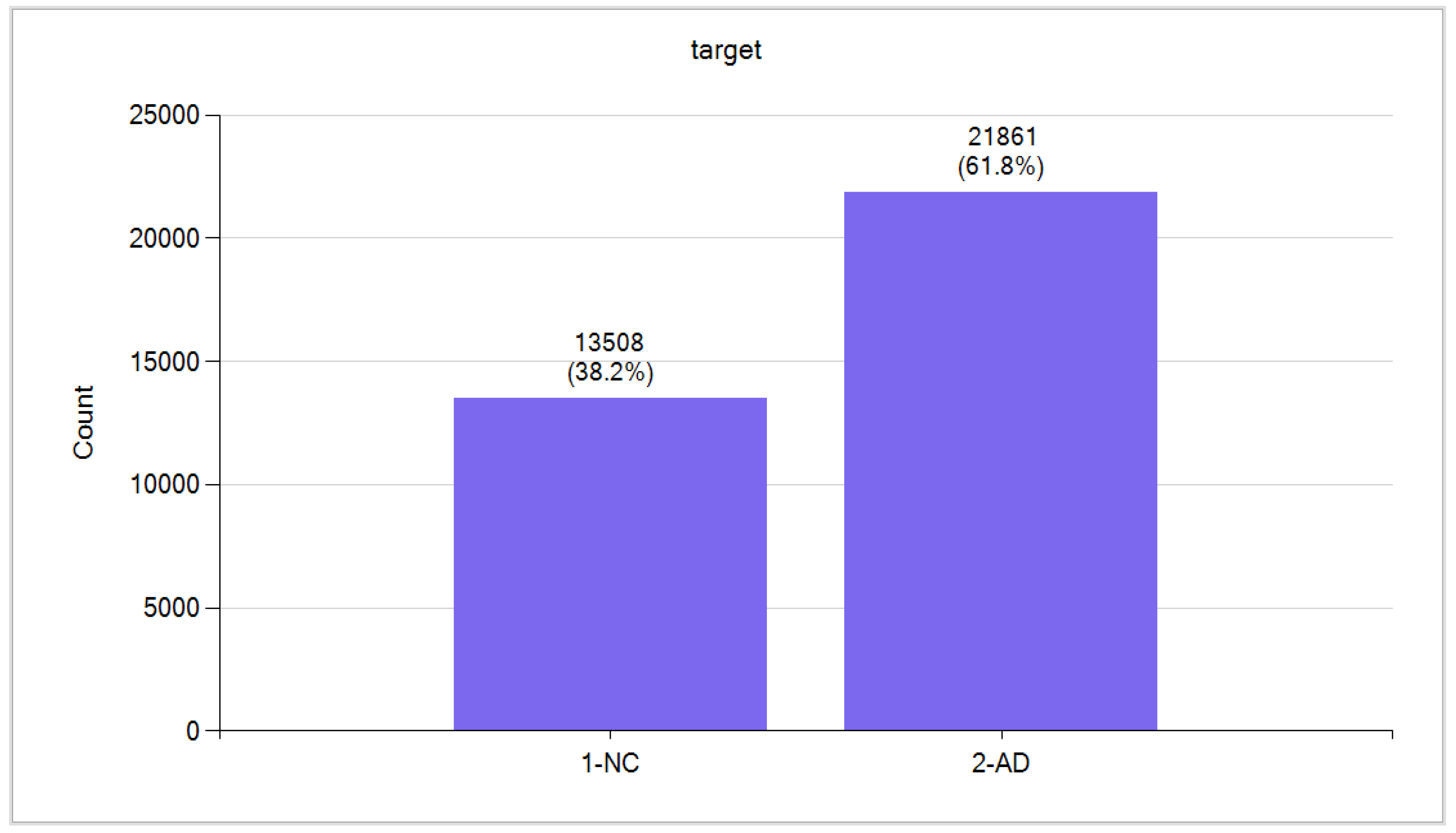
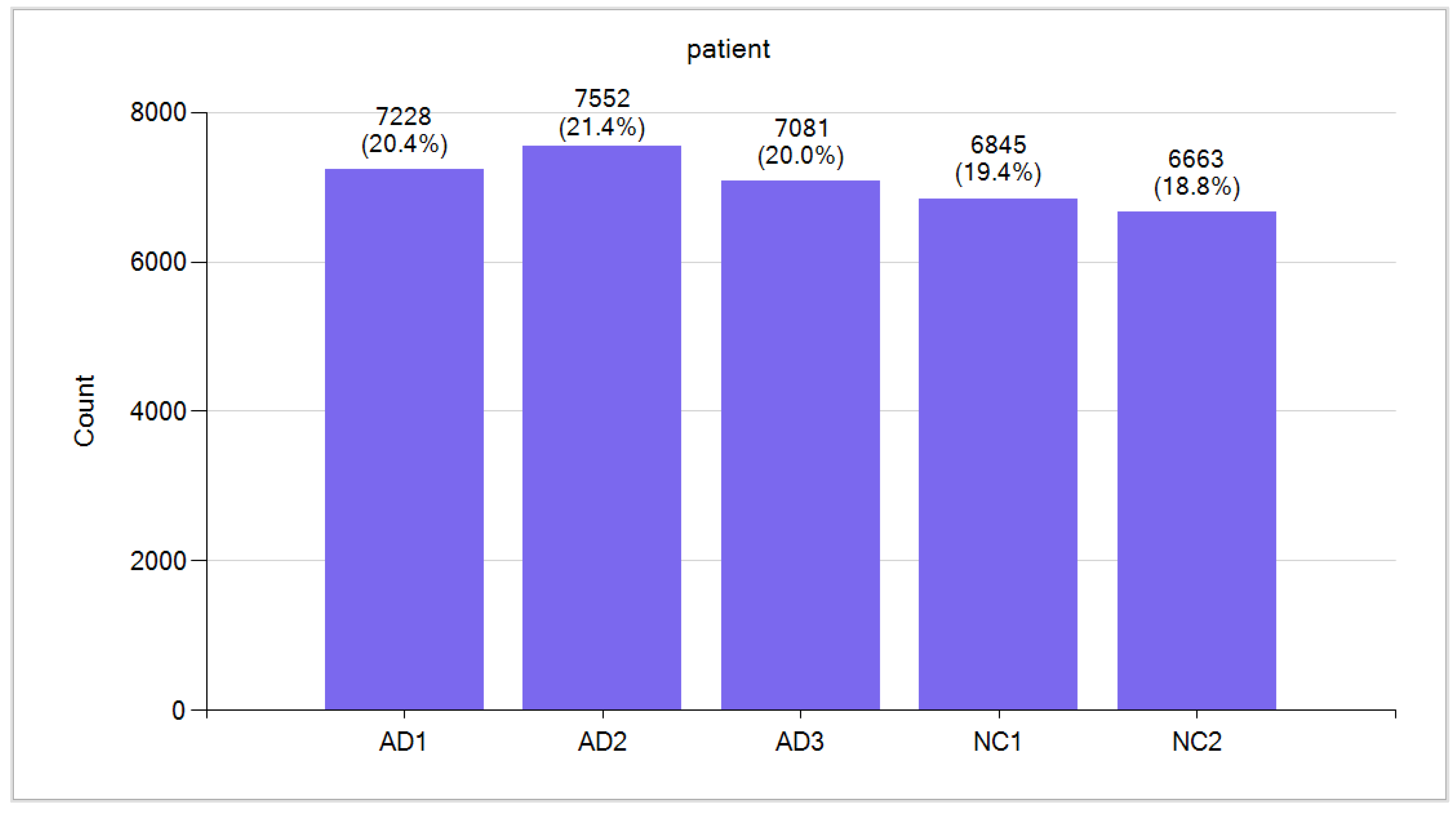
Data
Data Analysis
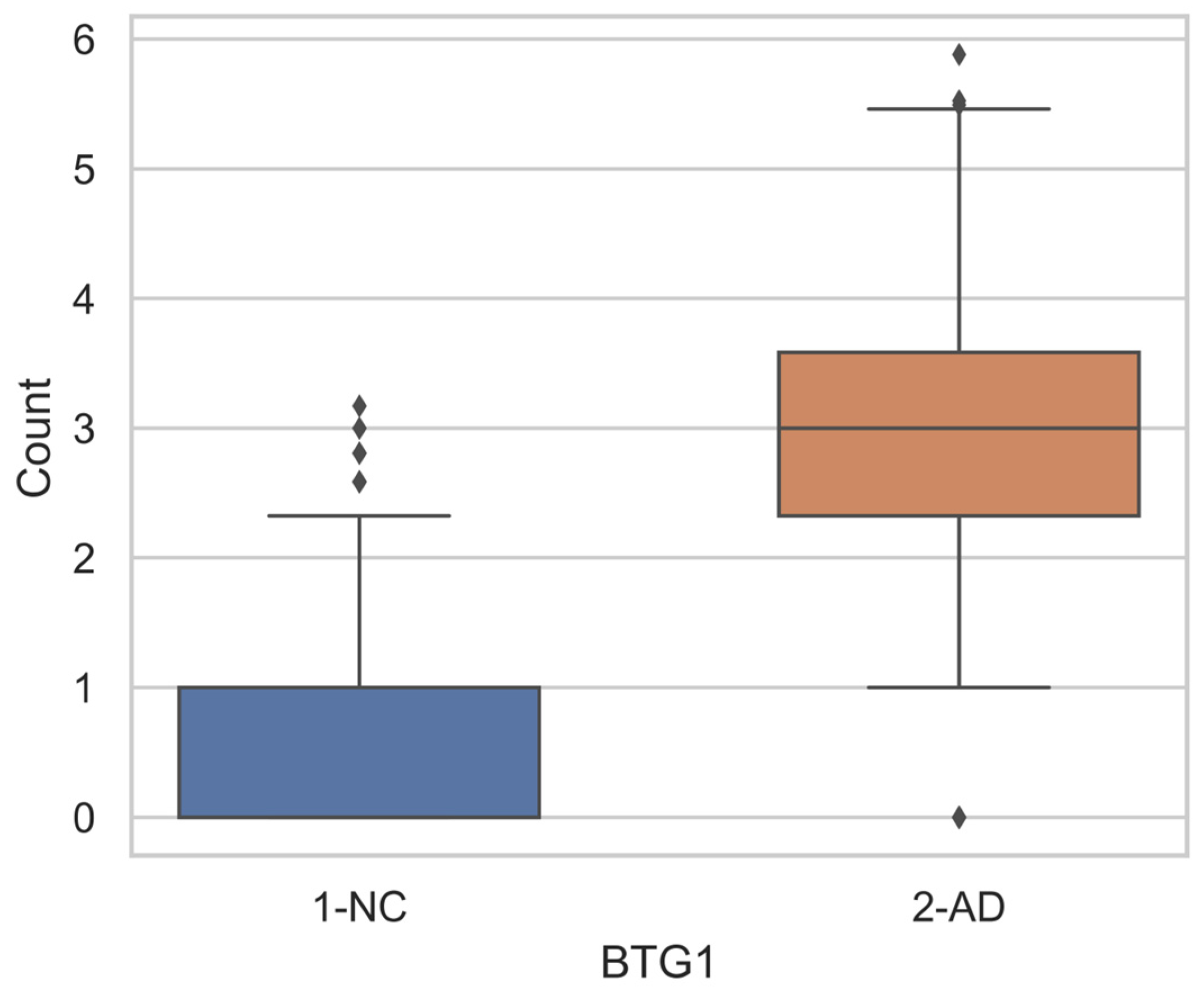
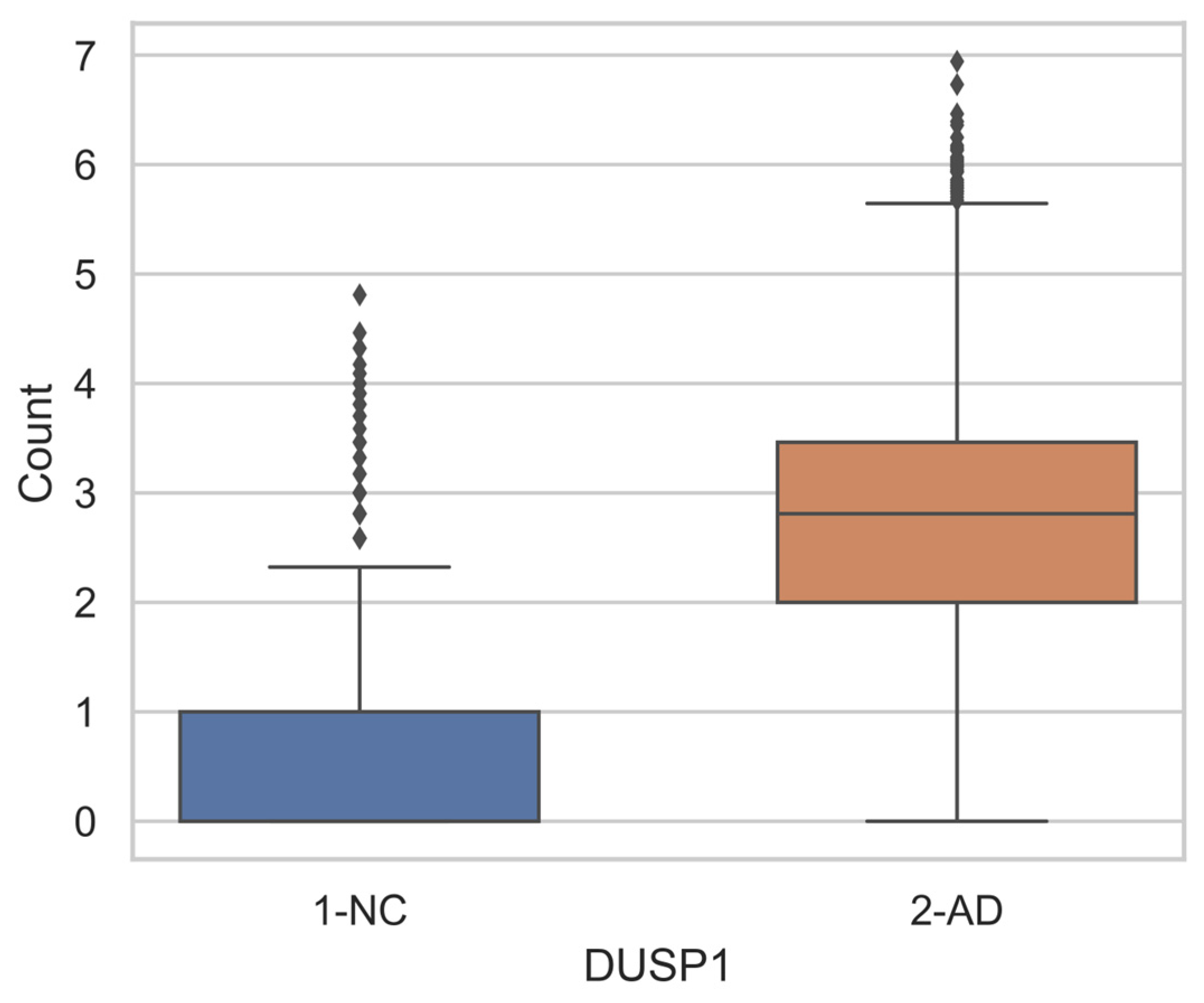
Differential Gene Expression
Pathway Analysis
Gene Ontology Analysis
Exploratory Classification of AD Using Peripheral-Blood Single-cell Transcriptomics
Discussion
Conclusion
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Xu H, Jia J. Single-Cell RNA Sequencing of Peripheral Blood Reveals Immune Cell Signatures in Alzheimer's Disease. Front Immunol. 2021 Aug 9;12:645666.
- Piacentini S, Polimanti R, Squitti R, Ventriglia M, Cassetta E, Vernieri F, Rossini PM, Manfellotto D, Fuciarelli M. GSTM1 null genotype as risk factor for late-onset Alzheimer's disease in Italian patients. J Neurol Sci. 2012 Jun 15;317(1-2):137-40.
- Haure-Mirande JV, Audrain M, Ehrlich ME, Gandy S. Microglial TYROBP/DAP12 in Alzheimer's disease: Transduction of physiological and pathological signals across TREM2. Mol Neurodegener. 2022 Aug 24;17(1):55.
- Loera-Valencia R, Piras A, Ismail MAM, Manchanda S, Eyjolfsdottir H, Saido TC, Johansson J, Eriksdotter M, Winblad B, Nilsson P. Targeting Alzheimer's disease with gene and cell therapies. J Intern Med. 2018 Jul;284(1):2-36.
- Nazarian A, Yashin AI, Kulminski AM. Summary-Based Methylome-Wide Association Analyses Suggest Potential Genetically Driven Epigenetic Heterogeneity of Alzheimer's Disease. J Clin Med. 2020 May 15;9(5):1489.
- Daneshafrooz N, Bagherzadeh Cham M, Majidi M, Panahi B. Identification of potentially functional modules and diagnostic genes related to amyotrophic lateral sclerosis based on the WGCNA and LASSO algorithms. Sci Rep. 2022 Nov 22;12(1):20144.
- Ali M, Garcia P, Lunkes LP, Sciortino A, Thomas M, Heurtaux T, Grzyb K, Halder R, Coowar D, Skupin A, Buée L, Blum D, Buttini M, Glaab E. Single cell transcriptome analysis of the THY-Tau22 mouse model of Alzheimer's disease reveals sex-dependent dysregulations. Cell Death Discov. 2024 Mar 7;10(1):119.
- Perobelli S, Alessandrini F, Zoccatelli G, Nicolis E, Beltramello A, Assael BM, Cipolli M. Diffuse alterations in grey and white matter associated with cognitive impairment in Shwachman-Diamond syndrome: evidence from a multimodal approach. Neuroimage Clin. 2015 Feb 27;7:721-31.
- Boora GK, Kulkarni AA, Kanwar R, Beyerlein P, Qin R, Banck MS, Ruddy KJ, Pleticha J, Lynch CA, Behrens RJ, Züchner S, Loprinzi CL, Beutler AS. Association of the Charcot-Marie-Tooth disease gene ARHGEF10 with paclitaxel induced peripheral neuropathy in NCCTG N08CA (Alliance). J Neurol Sci. 2015 Oct 15;357(1-2):35-40.
- Farioli-Vecchioli S, Micheli L, Saraulli D, Ceccarelli M, Cannas S, Scardigli R, Leonardi L, Cinà I, Costanzi M, Ciotti MT, Moreira P, Rouault JP, Cestari V, Tirone F. Btg1 is Required to Maintain the Pool of Stem and Progenitor Cells of the Dentate Gyrus and Subventricular Zone. Front Neurosci. 2012 Aug 30;6:124.
- Arango-Lievano M, Peguet C, Catteau M, Parmentier ML, Wu S, Chao MV, Ginsberg SD, Jeanneteau F. Deletion of Neurotrophin Signaling through the Glucocorticoid Receptor Pathway Causes Tau Neuropathology. Sci Rep. 2016 Nov 16;6:37231.
- Qi C, Liu F, Zhang W, Han Y, Zhang N, Liu Q, Li H. Alzheimer's disease alters the transcriptomic profile of natural killer cells at single-cell resolution. Front Immunol. 2022 Nov 2;13:1004885.
- Bobkova NV, Lyabin DN, Medvinskaya NI, Samokhin AN, Nekrasov PV, Nesterova IV, Aleksandrova IY, Tatarnikova OG, Bobylev AG, Vikhlyantsev IM, Kukharsky MS, Ustyugov AA, Polyakov DN, Eliseeva IA, Kretov DA, Guryanov SG, Ovchinnikov LP. The Y-Box Binding Protein 1 Suppresses Alzheimer's Disease Progression in Two Animal Models. PLoS One. 2015 Sep 22;10(9):e0138867.
- Ebanks B, Ingram TL, Chakrabarti L. ATP synthase and Alzheimer's disease: putting a spin on the mitochondrial hypothesis. Aging (Albany NY). 2020 Aug 27;12(16):16647-16662.
- Jiao L, Liu Y, Yu XY, Pan X, Zhang Y, Tu J, Song YH, Li Y. Ribosome biogenesis in disease: new players and therapeutic targets. Signal Transduct Target Ther. 2023 Jan 9;8(1):15.
- Lehmann DJ, Barnardo MC, Fuggle S, Quiroga I, Sutherland A, Warden DR, Barnetson L, Horton R, Beck S, Smith AD. Replication of the association of HLA-B7 with Alzheimer's disease: a role for homozygosity? J Neuroinflammation. 2006 Dec 18;3:33.
- Astarita G, Jung KM, Berchtold NC, Nguyen VQ, Gillen DL, Head E, Cotman CW, Piomelli D. Deficient liver biosynthesis of docosahexaenoic acid correlates with cognitive impairment in Alzheimer's disease. PLoS One. 2010 Sep 8;5(9):e12538.
- Lemon N, Canepa E, Ilies MA, Fossati S. Carbonic Anhydrases as Potential Targets Against Neurovascular Unit Dysfunction in Alzheimer's Disease and Stroke. Front Aging Neurosci. 2021 Nov 16;13:772278.
- Choi HJ, Jeong YJ, Kim J, Hoe HS. EGFR is a potential dual molecular target for cancer and Alzheimer's disease. Front Pharmacol. 2023 Aug 2;14:1238639.
- Tang, X., Tena, J., Di Lucente, J. et al. Transcriptomic and glycomic analyses highlight pathway-specific glycosylation alterations unique to Alzheimer’s disease. Sci Rep 13, 7816 (2023).
- Ling Y, Hu L, Chen J, Zhao M, Dai X. The mechanism of mitochondrial metabolic gene PMAIP1 involved in Alzheimer's disease process based on bioinformatics analysis and experimental validation. Clinics (Sao Paulo). 2024 Apr 30;79:100373.
- Du S, Zheng H. Role of FoxO transcription factors in aging and age-related metabolic and neurodegenerative diseases. Cell Biosci. 2021 Nov 2;11(1):188.
- Raffaele I, Silvestro S, Mazzon E. MicroRNAs and MAPKs: Evidence of These Molecular Interactions in Alzheimer's Disease. Int J Mol Sci. 2023 Mar 1;24(5):4736.
- Kapoor A, Nation DA. Role of Notch signaling in neurovascular aging and Alzheimer's disease. Semin Cell Dev Biol. 2021 Aug;116:90-97.
- Li J, Xu M, Zhou H, Ma J, Potter H. Alzheimer presenilins in the nuclear membrane, interphase kinetochores, and centrosomes suggest a role in chromosome segregation. Cell. 1997 Sep 5;90(5):917-27.
- Potter H. Cell Cycle and Chromosome Segregation Defects in Alzheimer's Disease. In: Madame Curie Bioscience Database [Internet]. Austin (TX): Landes Bioscience; 2000-2013.
- Talman V, Pascale A, Jäntti M, Amadio M, Tuominen RK. Protein Kinase C Activation as a Potential Therapeutic Strategy in Alzheimer's Disease: Is there a Role for Embryonic Lethal Abnormal Vision-like Proteins? Basic Clin Pharmacol Toxicol. 2016 Aug;119(2):149-60.
- Asghari K, Niknam Z, Mohammadpour-Asl S, Chodari L. Cellular junction dynamics and Alzheimer's disease: a comprehensive review. Mol Biol Rep. 2024 Feb 1;51(1):273.
- Oladele JO, Oyeleke OM, Oladele OT, Olaniyan M. Neuroprotective mechanism of Vernonia amygdalina in a rat model of neurodegenerative diseases. Toxicol Rep. 2020 Sep 14;7:1223-1232.
- Oddo S. The ubiquitin-proteasome system in Alzheimer's disease. J Cell Mol Med. 2008 Apr;12(2):363-73.
- Bouzid, H., Belk, J.A., Jan, M. et al. Clonal hematopoiesis is associated with protection from Alzheimer’s disease. Nat Med 29, 1662–1670 (2023).
| Downregulated | Upregulated | ||
| Gene Symbol | Gene Name | Gene Symbol | Gene Name |
| GSTM1 | Glutathione S-Transferase Mu 1 | BTG1 | BTG Anti-Proliferation Factor 1 |
| MYADM | Myeloid Associated Diff. Marker | DUSP1 | Dual Specificity Phosphatase 1 |
| MTRNR2L2 | MT-RNR2 Like 2 (Pseudogene) | PNRC1 | Proline Rich Nuclear Receptor Coactivator 1 |
| TYROBP | Transmembrane Immune Signaling Adaptor TYROBP | ZFP36L2 | ZFP36 Ring Finger Protein Like 2 |
| KLF6 | KLF Transcription Factor 6 | RPS2 | Ribosomal Protein S2 |
| CFAP20 | Cilia And Flagella Associated Protein | YBX1 | Y-Box Binding Protein 1 |
| YPEL5 | Yippee Like 5 | ATP5F1D | ATP Synthase F1 Subunit Delta |
| TSC22D3 | TSC22 Domain Family Member 3 | NOP53 | NOP53 Ribosome Biogenesis Factor |
| SBDS | SBDS Ribosome Maturation Factor | HLA-C | Major Histocompatibility Complex, Class I, C |
| ARHGEF10 | Rho Guanine Nucleotide Exchange Factor 10 | TRIR | Telomerase RNA Component Interacting RNase |
| Downregulated | Upregulated |
| Biosynthesis of DHA-derived sulfido conjugates | FOXO-mediated transcription of cell cycle genes |
| Biosynthesis of maresin conjugates in tissue regeneration (MCTR) | RMTs methylate histone arginines |
| Reversible hydration of carbon dioxide | RAF-independent MAPK1/3 activation |
| Inhibition of Signaling by Overexpressed EGFR | FOXO-mediated transcription |
| Signaling by Overexpressed Wild-Type EGFR in Cancer | PTK6 Regulates Proteins Involved in RNA Processing |
| Activation, myristolyation of BID and translocation to mitochondria | Noncanonical activation of NOTCH3 |
| Defective B3GALTL causes Peters-plus syndrome (PpS) | Amplification of signal from the kinetochores |
| O-glycosylation of TSR domain-containing proteins | Amplification of signal from unattached kinetochores via a MAD2 |
| Activation of NOXA and translocation to mitochondria | Mitotic Spindle Checkpoint |
| Assembly of Viral Components at the Budding Site | EML4 and NUDC in mitotic spindle formation |
| Downregulated | Upregulated |
| (BP) negative regulation of protein kinase C signaling | (BP) positive regulation of endothelial cell differentiation |
| (BP) cell-cell junction maintenance | (BP) positive regulation of fibroblast apoptotic process |
| (BP) membrane raft organization | (BP) positive regulation of myoblast differentiation |
| (BP) nitrobenzene metabolic process | (BP) positive regulation of ubiquitin-protein transferase activity |
| (BP) positive regulation of feeding behavior | (BP) negative regulation of mitotic cell cycle |
| (BP) negative regulation of activation-induced cell death of T cells | (BP) definitive hemopoiesis |
| (BP) positive regulation of translational initiation in response to stress | (BP) somatic stem cell division |
| (BP) positive regulation of endoplasmic reticulum stress-induced eIF2 alpha dephosphorylation | (BP) negative regulation of mitotic cell cycle phase transition |
| (BP) positive regulation of peptidyl-serine dephosphorylation | (BP) negative regulation of stem cell differentiation |
| (BP) cellular detoxification of nitrogen compound | (BP) negative regulation of fat cell differentiation |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




