Submitted:
20 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
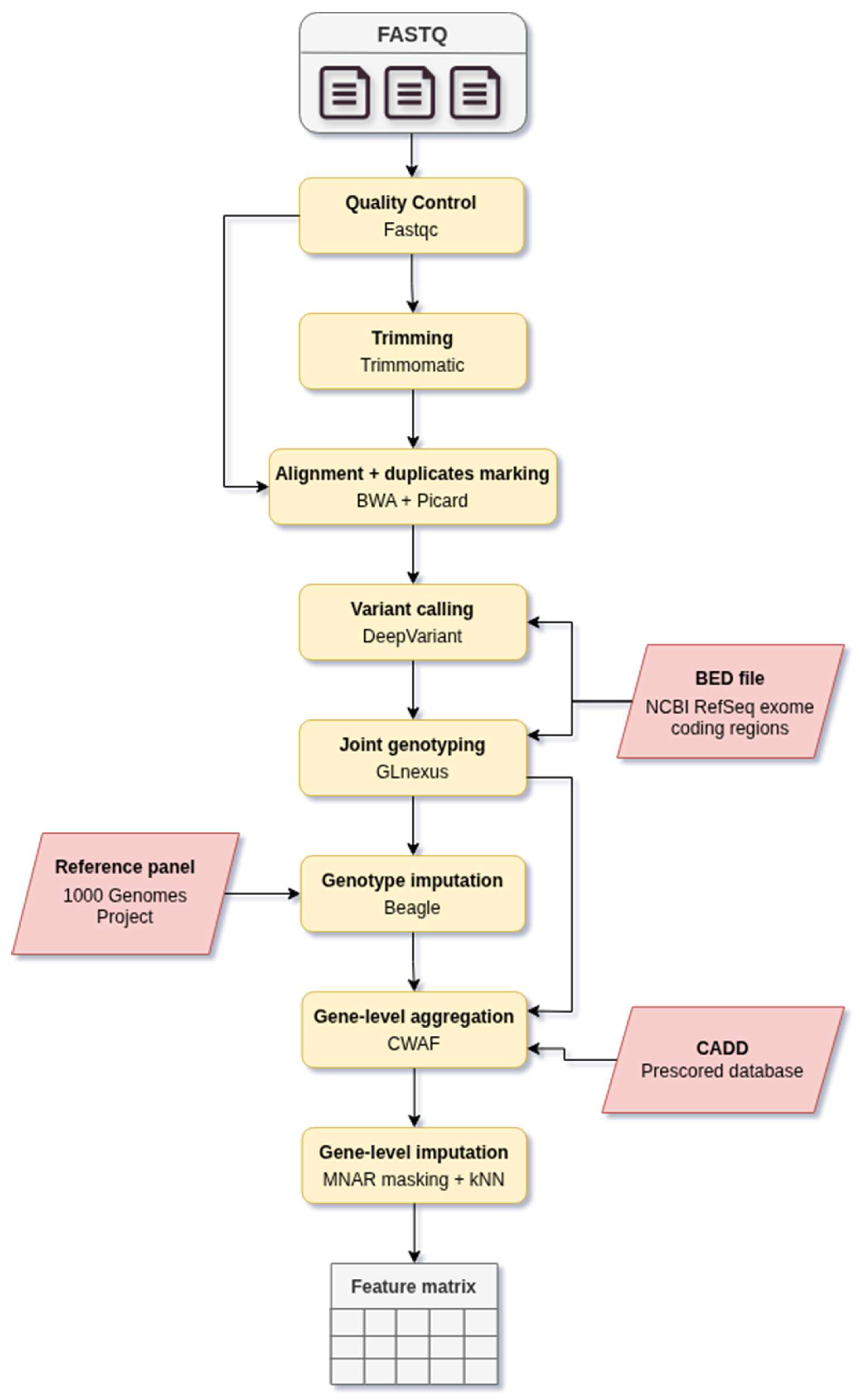
2.1. Pipeline Overview
2.2. Reads Quality Control and Mapping
2.3. Variant Calling and Joint Genotyping
2.4. Genotype Imputation
2.5. Variants Post-Processing and Filtering
2.7. Gene-Level Aggregation
2.8. Gene-Level Feature Imputation
2.9. Benchmark Study
2.9.1. Samples and Data
2.9.2. Benchmark Analysis
3. Results
3.1. Variant-Level Processing
3.2. Gene-Level Processing
3.2.1. Variant Aggregation and Genotype Imputation Performance
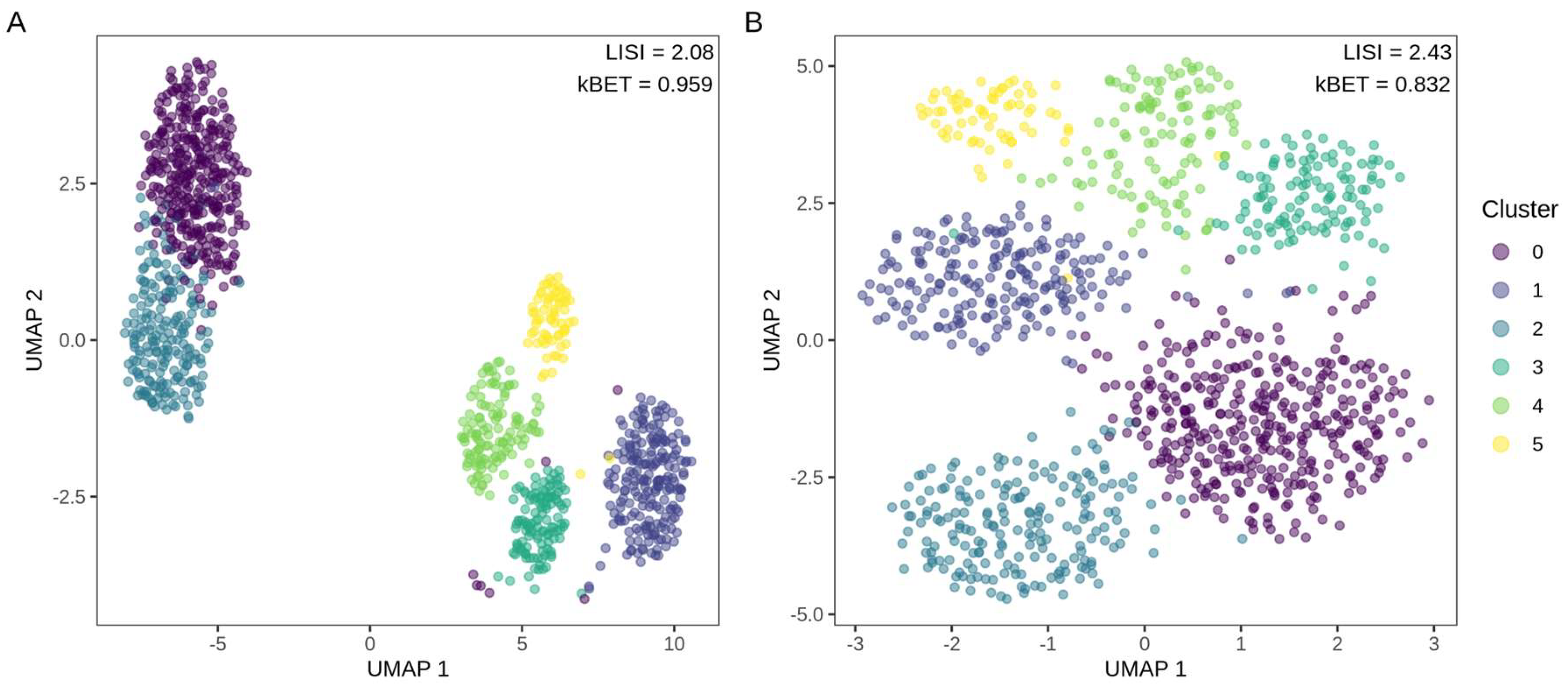
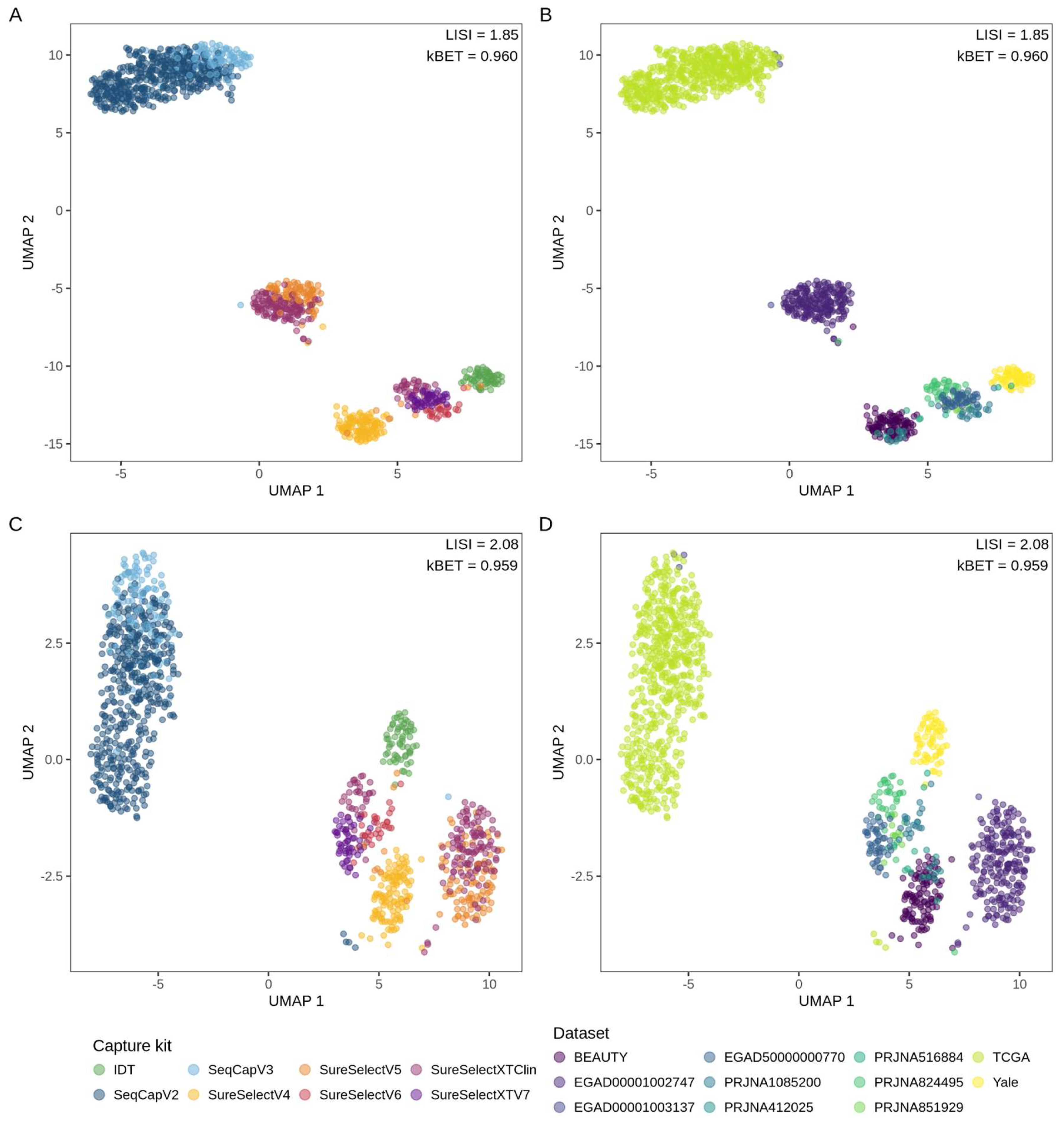
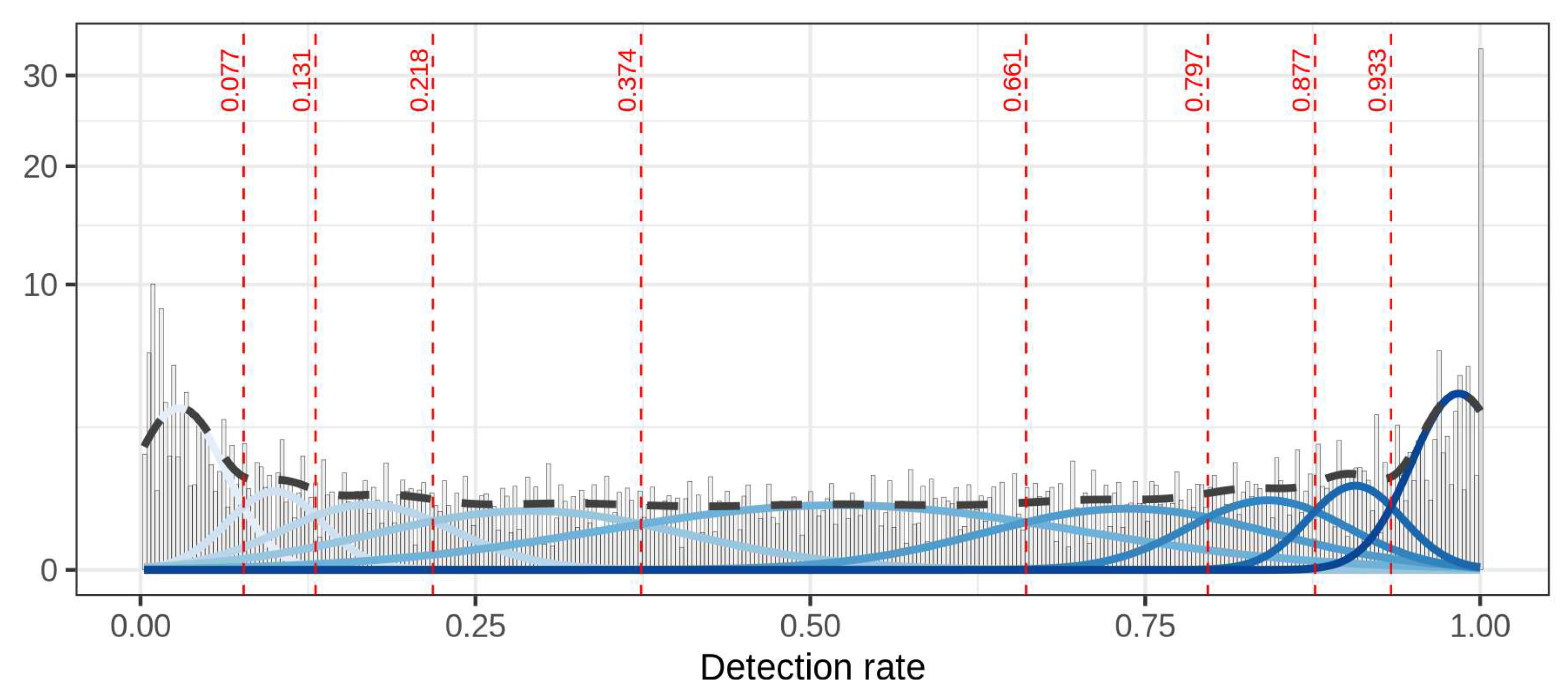
3.2.2. Sample Clustering
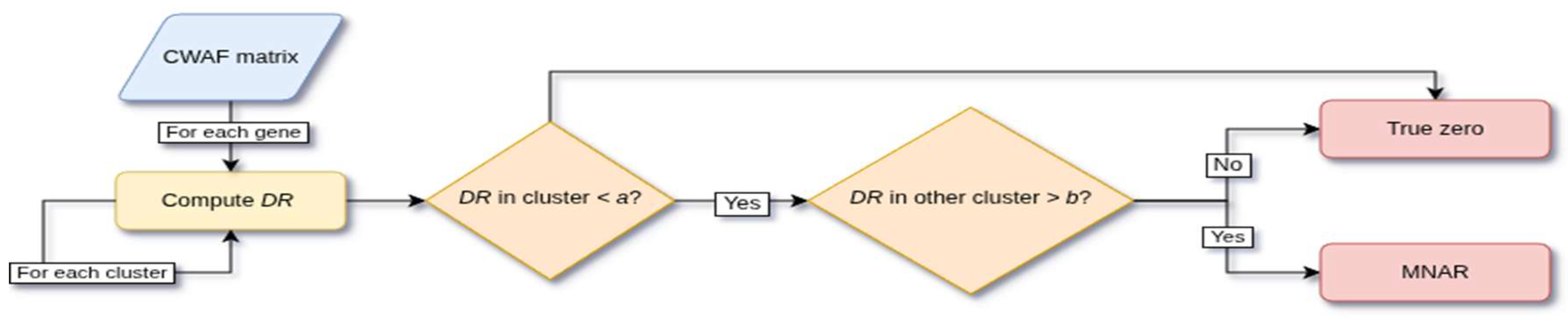
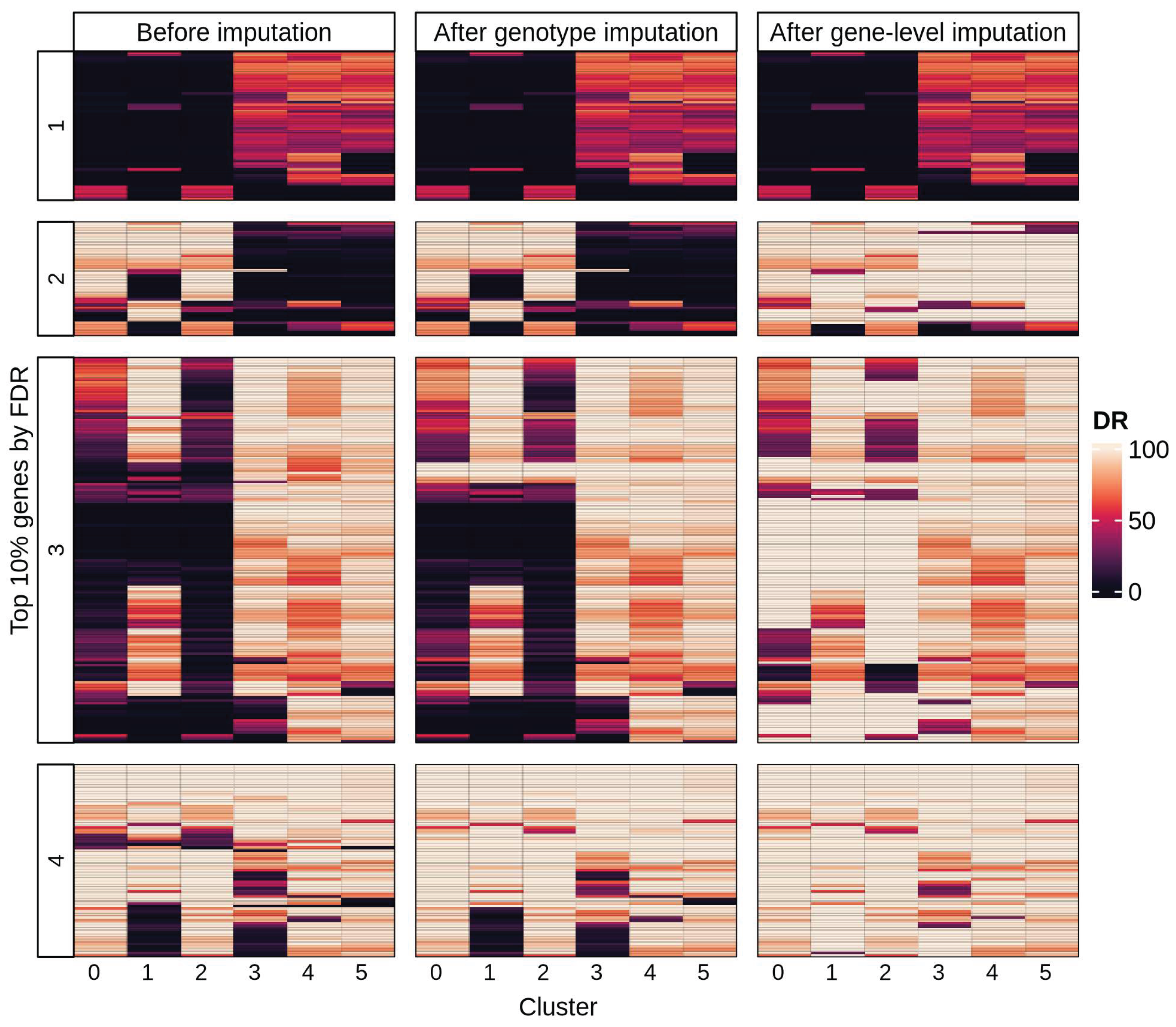
3.2.3. MNAR Masking and Gene-Level Imputation
3.2.4. Gene Detection
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 1kGP | The 1000 Genomes Project |
| AF | Allele fraction |
| CCDS | Consensus coding sequence |
| CWAF | CADD-weighted allele fraction |
| DR | Gene detection rate |
| FDR | False discovery rate |
| FFPE | Formalin-fixed paraffin-embedded |
| kBET | k-nearest neighbor Batch Effect Test |
| kNN | K-nearest neighbors |
| LD | Linear dichroism |
| LISI | Local inverse Simpson’s index |
| MNAR | Missing-not-at-random |
| PARC | Phenotyping by accelerated refined community-partitioning |
| SNV | Single nucleotide variation |
| TCGA | The Cancer Genome Atlas |
| UMAP | Uniform manifold approximation and projection |
| WES | Whole-exome sequencing |
| WGS | Whole-genome sequencing |
Appendix A
Appendix A.1
| Database | Version |
|---|---|
| RefGene | hg38 2020-08-17 |
| ExAC | v0.3 |
| gnomAD exomes | v4.1 |
| 1000 Genomes | 2015-08 |
| dbSNP | build 150 |
| ClinVar | 2024-12-1 |
| dbNSFP | v4.7a |
References
- Mardis, E. R. DNA sequencing technologies: 2006-2016. Nat. Protoc. 2017, vol. 12(no. 2), 213–218. [Google Scholar] [CrossRef]
- Auton, A. , A global reference for human genetic variation. Nature 2015, vol. 526(no. 7571), 68–74. [Google Scholar] [CrossRef]
- Weinstein, J. N. , The Cancer Genome Atlas Pan-Cancer Analysis Project. Nat. Genet. 2013, vol. 45(no. 10), 1113–1120. [Google Scholar] [CrossRef]
- Sudlow, C. , UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015, vol. 12(no. 3), e1001779. [Google Scholar] [CrossRef]
- Buckley, A. R. , Pan-cancer analysis reveals technical artifacts in TCGA germline variant calls. BMC Genomics 2017, vol. 18(no. 1), 458. [Google Scholar] [CrossRef]
- Next-generation data filtering in the genomics era | Nature Reviews Genetics. Available online: https://www.nature.com/articles/s41576-024-00738-6 (accessed on Mar. 19 2026).
- Majewski, J.; Schwartzentruber, J.; Lalonde, E.; Montpetit, A.; Jabado, N. What can exome sequencing do for you? J. Med. Genet. 2011, vol. 48(no. 9), 580–589. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Lakatos, E.; Bakir, I. A.; Curtius, K.; Graham, T. A.; Mustonen, V. The mutational signatures of formalin fixation on the human genome. Nat. Commun. 2022, vol. 13(no. 1), 4487. [Google Scholar] [CrossRef] [PubMed]
- Krehenwinkel, H.; Wolf, M.; Lim, J. Y.; Rominger, A. J.; Simison, W. B.; Gillespie, R. G. Estimating and mitigating amplification bias in qualitative and quantitative arthropod metabarcoding. Sci. Rep. 2017, vol. 7(no. 1), 17668. [Google Scholar] [CrossRef] [PubMed]
- Pan, B. , Assessing reproducibility of inherited variants detected with short-read whole genome sequencing. Genome Biol. 2022, vol. 23(no. 1), 2. [Google Scholar] [CrossRef]
- Wickland, D. P.; et al. Impact of variant-level batch effects on identification of genetic risk factors in large sequencing studies. PLoS ONE 2021, vol. 16(no. 4), e0249305. [Google Scholar] [CrossRef]
- Jarosz, L.; Dai, J.; Ochocki, M.; Merta, J.; Pusztai, L.; Marczyk, M. Kit-Specific and Other Non-Biological-Based Biases in Germline Whole-Exome Analysis at the Gene Level. presented at the 17th International Conference on Bioinformatics Models, Methods and Algorithms, Mar. 2026; pp. 487–496. Available online: https://www.scitepress.org/PublicationsDetail.aspx?ID=A1n7u6b9T84=&t=1 (accessed on Mar. 27 2026).
- Johnson, W. E.; Li, C.; Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 2007, vol. 8(no. 1), 118–127. [Google Scholar] [CrossRef]
- Ritchie, M. E. , limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, vol. 43(no. 7), e47–e47. [Google Scholar] [CrossRef]
- Korsunsky. , Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 2019, vol. 16(no. 12), 1289–1296. [Google Scholar] [CrossRef]
- Benjamini, Y.; Speed, T. P. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 2012, vol. 40(no. 10), e72. [Google Scholar] [CrossRef]
- Diossy, M. , Strand Orientation Bias Detector to determine the probability of FFPE sequencing artifacts. Brief. Bioinform. 2021, vol. 22(no. 6), bbab186. [Google Scholar] [CrossRef] [PubMed]
- Heo, D.; et al. DEEPOMICS FFPE, a deep neural network model, identifies DNA sequencing artifacts from formalin fixed paraffin embedded tissue with high accuracy. Sci. Rep. 2024, vol. 14(no. 1), 2559. [Google Scholar] [CrossRef] [PubMed]
- Tellaetxe-Abete, M.; Calvo, B.; Lawrie, C. Ideafix: a decision tree-based method for the refinement of variants in FFPE DNA sequencing data. NAR Genomics Bioinforma. 2021, vol. 3(no. 4), lqab092. [Google Scholar] [CrossRef]
- Ikegami, M. , MicroSEC filters sequence errors for formalin-fixed and paraffin-embedded samples. Commun. Biol. 2021, vol. 4(no. 1), 1396. [Google Scholar] [CrossRef] [PubMed]
- Poplin, R.; et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 2018, vol. 36(no. 10), 983–987. [Google Scholar] [CrossRef]
- Lin, M. F.; et al. GLnexus: joint variant calling for large cohort sequencing. In bioRxiv; 11 Jun 2018. [Google Scholar] [CrossRef]
- O’Leary, N. A.; et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016, vol. 44(no. D1), D733–D745. [Google Scholar] [CrossRef]
- Browning, B. L.; Zhou, Y.; Browning, S. R. A One-Penny Imputed Genome from Next-Generation Reference Panels. Am. J. Hum. Genet. 2018, vol. 103(no. 3), 338–348. [Google Scholar] [CrossRef]
- Browning, B. L.; Tian, X.; Zhou, Y.; Browning, S. R. Fast two-stage phasing of large-scale sequence data. Am. J. Hum. Genet. 2021, vol. 108(no. 10), 1880–1890. [Google Scholar] [CrossRef]
- A global reference for human genetic variation | Nature. Available online: https://www.nature.com/articles/nature15393 (accessed on Mar. 25 2026).
- Rentzsch, P.; Witten, D.; Cooper, G. M.; Shendure, J.; Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019, vol. 47(no. D1), D886–D894. [Google Scholar] [CrossRef]
- Andrews, S. FastQC A Quality Control Tool for High Throughput Sequence Data. - References - Scientific Research Publishing. 2010. Available online: https://www.scirp.org/reference/referencespapers?referenceid=4024153 (accessed on Mar. 25 2026).
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, vol. 32(no. 19), 3047–3048. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A. M.; Lohse, M.; Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014, vol. 30(no. 15), 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Ensembl 2022 | Nucleic Acids Research | Oxford Academic. Available online: https://academic.oup.com/nar/article/50/D1/D988/6430486?login=true (accessed on Mar. 25 2026).
- Li, H. , The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, vol. 25(no. 16), 2078–2079. [Google Scholar] [CrossRef]
- “MarkDuplicates (Picard),” GATK. Available online: https://gatk.broadinstitute.org/hc/en-us/articles/360037052812-MarkDuplicates-Picard (accessed on Mar. 25 2026).
- Zhao, H.; Sun, Z.; Wang, J.; Huang, H.; Kocher, J.-P.; Wang, L. CrossMap: a versatile tool for coordinate conversion between genome assemblies. Bioinformatics 2014, vol. 30(no. 7), 1006–1007. [Google Scholar] [CrossRef]
- Poplin, R.; et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 2018, vol. 36(no. 10), 983–987. [Google Scholar] [CrossRef] [PubMed]
- Lin, M. F.; et al. GLnexus: joint variant calling for large cohort sequencing. In bioRxiv; 11 Jun 2018. [Google Scholar] [CrossRef]
- Browning, B. L.; Tian, X.; Zhou, Y.; Browning, S. R. Fast two-stage phasing of large-scale sequence data. Am. J. Hum. Genet. 2021, vol. 108(no. 10), 1880–1890. [Google Scholar] [CrossRef]
- Byrska-Bishop, M.; et al. High coverage whole genome sequencing of the expanded 1000 Genomes Project cohort including 602 trios. In bioRxiv; 07 Feb 2021. [Google Scholar] [CrossRef]
- Leitwein, M.; Duranton, M.; Rougemont, Q.; Gagnaire, P.-A.; Bernatchez, L. Using Haplotype Information for Conservation Genomics. Trends Ecol. Evol. 2020, vol. 35(no. 3), 245–258. [Google Scholar] [CrossRef]
- Wang; Li, M.; Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, vol. 38(no. 16), e164. [Google Scholar] [CrossRef]
- Schubach, M.; Maass, T.; Nazaretyan, L.; Röner, S.; Kircher, M. CADD v1.7: using protein language models, regulatory CNNs and other nucleotide-level scores to improve genome-wide variant predictions. Nucleic Acids Res. 2024, vol. 52(no. D1), D1143–D1154. [Google Scholar] [CrossRef]
- Pedregosa, F.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, vol. 12(no. 85), 2825–2830. [Google Scholar]
- Stassen, S. V.; Siu, D. M. D.; Lee, K. C. M.; Ho, J. W. K.; So, H. K. H.; Tsia, K. K. PARC: ultrafast and accurate clustering of phenotypic data of millions of single cells. Bioinformatics 2020, vol. 36(no. 9), 2778–2786. [Google Scholar] [CrossRef] [PubMed]
- Zyla; Szumala, K.; Polanski, A.; Polanska, J.; Marczyk, M. dpGMM: A new R package for efficient and robust Gaussian mixture modeling of 1D and 2D data. J. Comput. Sci. 2026, vol. 95, 102811. [Google Scholar] [CrossRef]
- Bengtsson, H. A Unifying Framework for Parallel and Distributed Processing in R using Futures. R J. 2021, vol. 13(no. 2), 273–291. [Google Scholar] [CrossRef]
- Büttner; Miao, Z.; Wolf, F. A.; Teichmann, S. A.; Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 2019, vol. 16(no. 1), 43–49. [Google Scholar] [CrossRef]
- Jefferis, G.; Kemp, S.; Arya, S.; Mount, D. RANN: Fast Nearest Neighbour Search (Wraps ANN Library) Using L2 Metric. R. Available online: https://github.com/jefferislab/rann.
- Sulonen, A.-M.; et al. Comparison of solution-based exome capture methods for next generation sequencing. Genome Biol. 2011, vol. 12(no. 9), R94. [Google Scholar] [CrossRef]
- Performance comparison of four commercial human whole-exome capture platforms | Scientific Reports. Available online: https://www.nature.com/articles/srep12742 (accessed on Mar. 20 2026).
- López, F. V.; Ashton, J. J.; Cheng, G.; Ennis, S. A systematic analysis of contemporary whole exome sequencing capture kits to optimise high-coverage capture of CCDS regions. NAR Genomics Bioinforma. 2025, vol. 7(no. 3), lqaf115. [Google Scholar] [CrossRef] [PubMed]
- Agilent Technologies | Agilent. Available online: https://www.agilent.com/en/ (accessed on Apr. 10 2026).
- Yu, Y.; Mai, Y.; Zheng, Y.; Shi, L. Assessing and mitigating batch effects in large-scale omics studies. Genome Biol. 2024, vol. 25(no. 1), 254. [Google Scholar] [CrossRef] [PubMed]
- Garrison, E.; Kronenberg, Z. N.; Dawson, E. T.; Pedersen, B. S.; Prins, P. A spectrum of free software tools for processing the VCF variant call format: vcflib, bio-vcf, cyvcf2, hts-nim and slivar. PLOS Comput. Biol. 2022, vol. 18(no. 5), e1009123. [Google Scholar] [CrossRef] [PubMed]
- Tan, A.; Abecasis, G. R.; Kang, H. M. Unified representation of genetic variants. Bioinformatics 2015, vol. 31(no. 13), 2202–2204. [Google Scholar] [CrossRef]
- Yun, T.; Li, H.; Chang, P.-C.; Lin, M. F.; Carroll, A.; McLean, C. Y. Accurate, scalable cohort variant calls using DeepVariant and GLnexus. Bioinformatics 2021, vol. 36(no. 24), 5582–5589. [Google Scholar] [CrossRef]
- Barbitoff, Y. A.; Abasov, R.; Tvorogova, V. E.; Glotov, A. S.; Predeus, A. V. Systematic benchmark of state-of-the-art variant calling pipelines identifies major factors affecting accuracy of coding sequence variant discovery. BMC Genomics 2022, vol. 23(no. 1), 155. [Google Scholar] [CrossRef]
- Pinto, V.; Sousa, L.; Silva, C. Variant calling in genomics: A comparative performance analysis and decision guide. PLOS One 2026, vol. 21(no. 2), e0339891. [Google Scholar] [CrossRef]
- Marth, G. T.; Czabarka, E.; Murvai, J.; Sherry, S. T. The Allele Frequency Spectrum in Genome-Wide Human Variation Data Reveals Signals of Differential Demographic History in Three Large World Populations. Genetics 2004, vol. 166(no. 1), 351–372. [Google Scholar] [CrossRef]
- Taliun, D. , Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 2021, vol. 590(no. 7845), 290–299. [Google Scholar] [CrossRef] [PubMed]
- Savas, P. , The Subclonal Architecture of Metastatic Breast Cancer: Results from a Prospective Community-Based Rapid Autopsy Program ‘CASCADE. PLoS Med. 2016, vol. 13(no. 12), e1002204. [Google Scholar] [CrossRef] [PubMed]
| Dataset | Samples | Exome capture kit | Kit alias | |
| PRJNA412025 | 10 | SureSelect Human All Exon V4 | SureSelectV4 | |
| BEAUTY | 106 | |||
| PRJNA516884 | 9 | SureSelect Human All Exon V5 | SureSelectV5 | |
| EGAD00001002747 | 2051 | |||
| SureSelect XT Clinical Research | SureSelectXTClin | |||
| PRJNA824495 | 38 | |||
| PRJNA1085200 | 19 | SureSelect Human All Exon V6 | SureSelectV6 | |
| PRJNA851929 | 8 | |||
| EGAD50000000770 | 49 | SureSelect XT Human All Exon V7 | SureSelectXTV7 | |
| TCGA | 5662 | SeqCap EZ Exome V2 | SeqCapV2 | |
| SeqCap EZ Exome V3 | SeqCapV3 | |||
| EGAD00001003137 | 4 | |||
| Yale | 61 | IDT xGen Exome Research Panel | IDT |
| Capture kit | 0 | 1 | 2 | 3 | 4 | 5 |
| IDT | 61 | |||||
| SeqCapV2 | 261 | 1 | 195 | |||
| SeqCapV3 | 111 | 2 | ||||
| SureSelectV4 | 1 | 114 | 1 | |||
| SureSelectV5 | 90 | 4 | 3 | 2 | ||
| SureSelectV6 | 26 | 1 | ||||
| SureSelectXTV7 | 49 | |||||
| SureSelectXTClin | 1 | 116 | 37 | 1 | ||
| Total | 374 | 207 | 197 | 118 | 115 | 66 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




