Submitted:
17 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Background
2.1. Federated Learning in Healthcare
2.2. Update Confidentiality and Secure Aggregation
2.3. Auditability, Governance Evidence, and Trust in Cross-Organisation Collaboration
2.4. Related Work
3. Proposed System
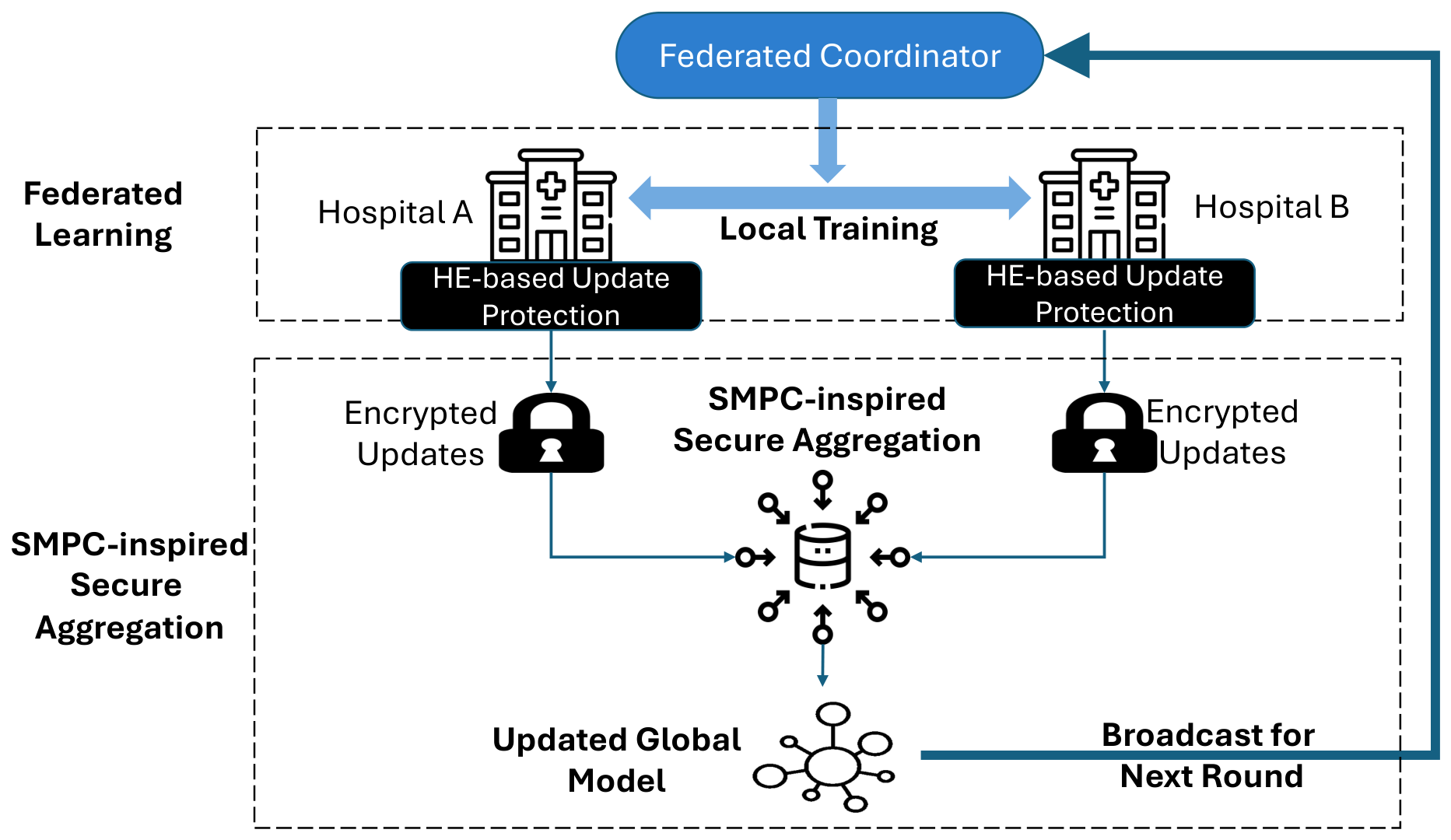
3.1. System Architecture
3.2. Federated Learning Workflow
- Broadcast: The coordinator broadcasts the current global model and round identifier t to all participating peers.
- Local training: Each peer performs local optimisation for E epochs (or steps) and obtains updated parameters . The local model update is then computed as
- Protected submission: Each peer protects using the protected update pipeline described in Section 3.3 and Section 3.4, and submits only the protected update to the coordinator.
- Aggregation and model update: The coordinator aggregates the protected updates and applies the resulting global update:where is the server learning rate.
3.3. HE-Based Update Protection for Encrypted Aggregation
3.4. SMPC-Inspired Secure Aggregation via Additive Masking
4. Evaluation
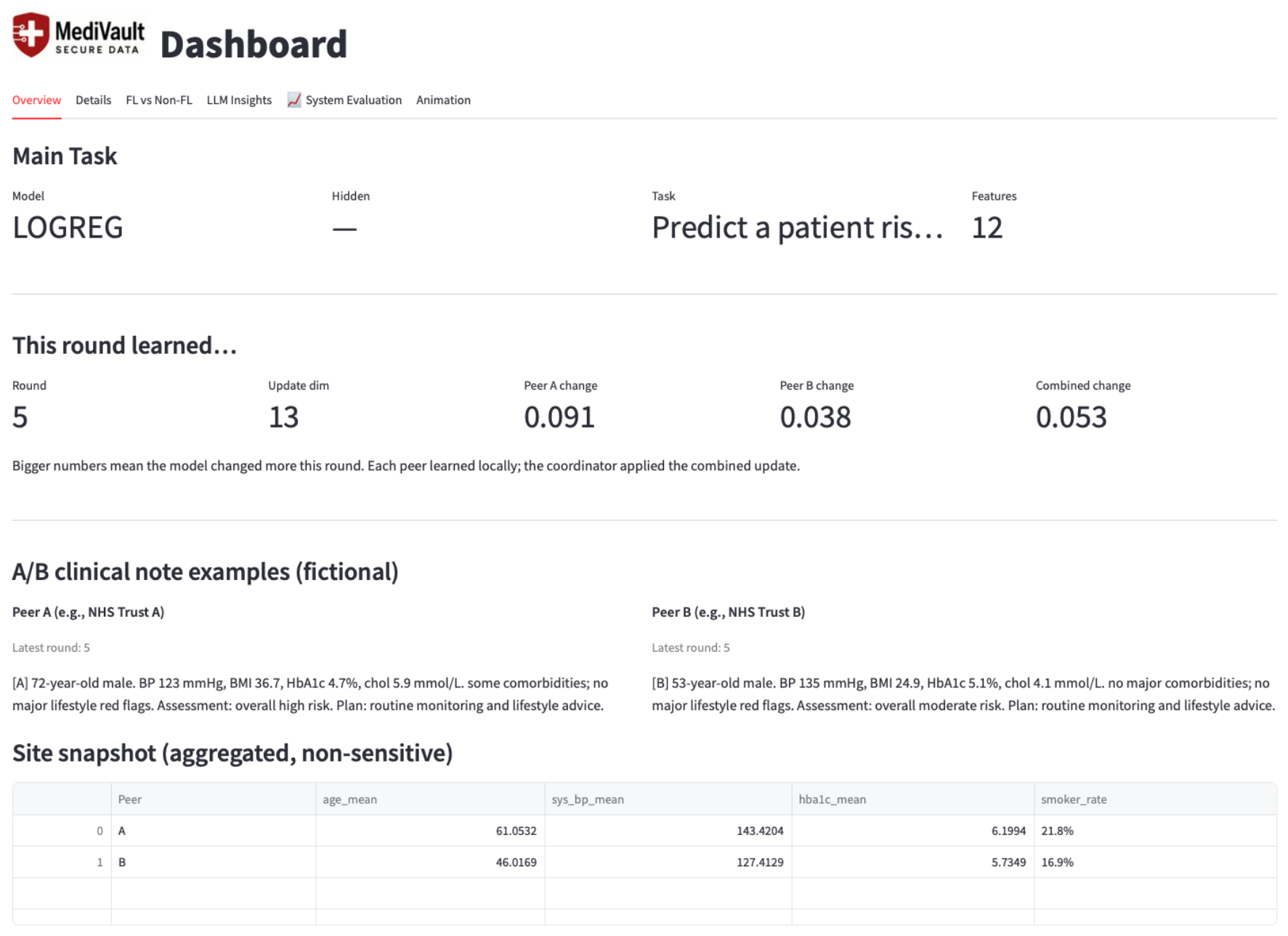
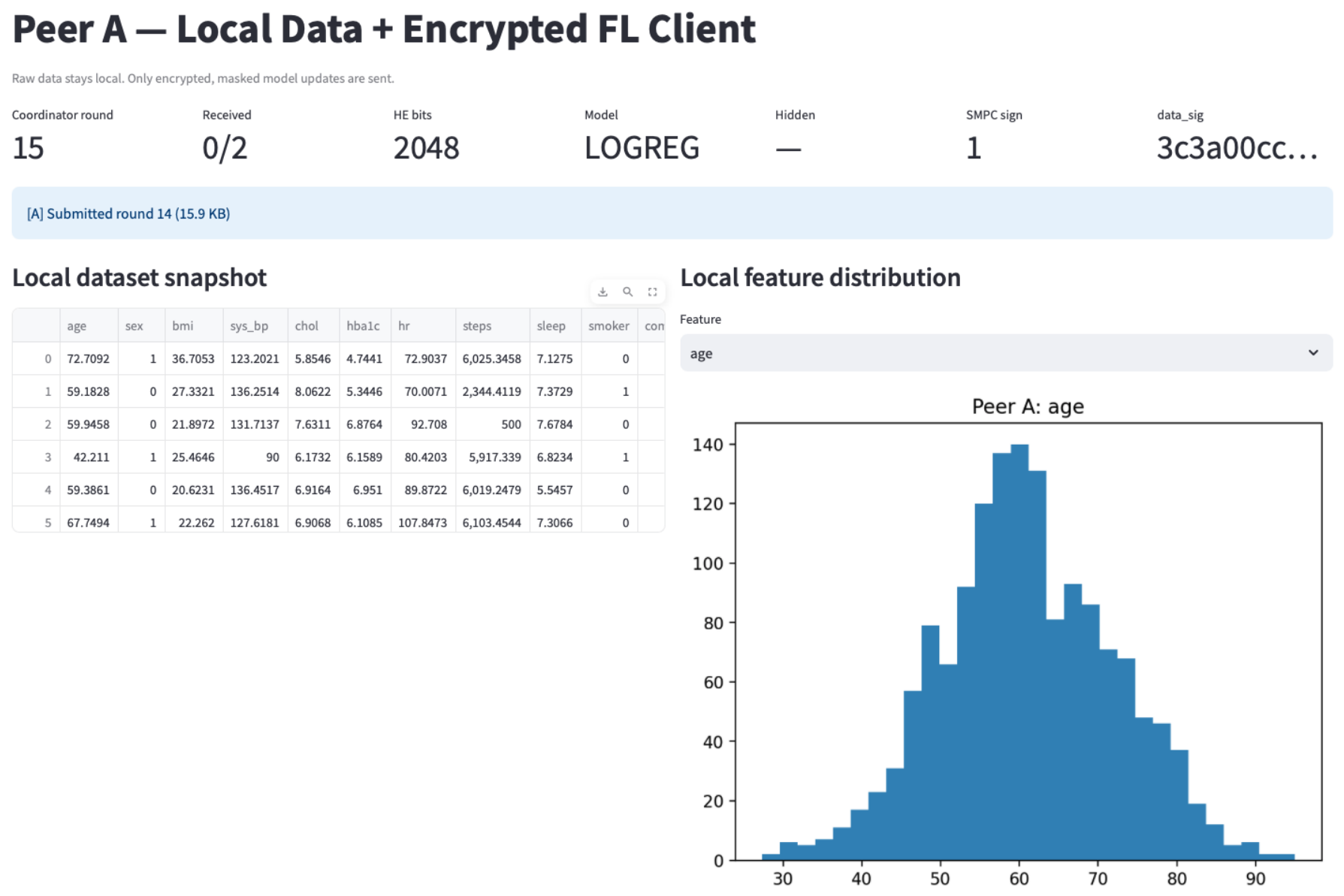
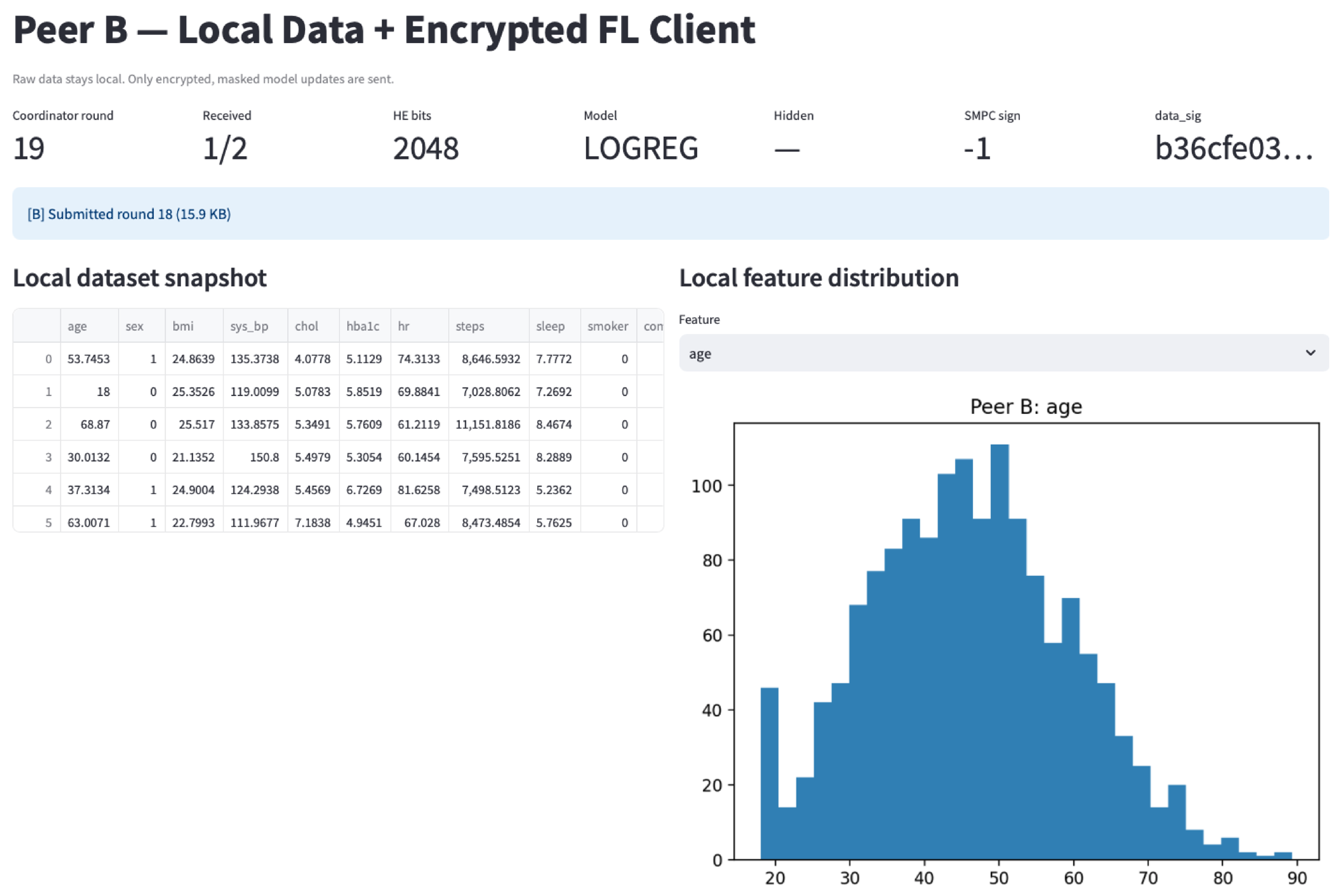
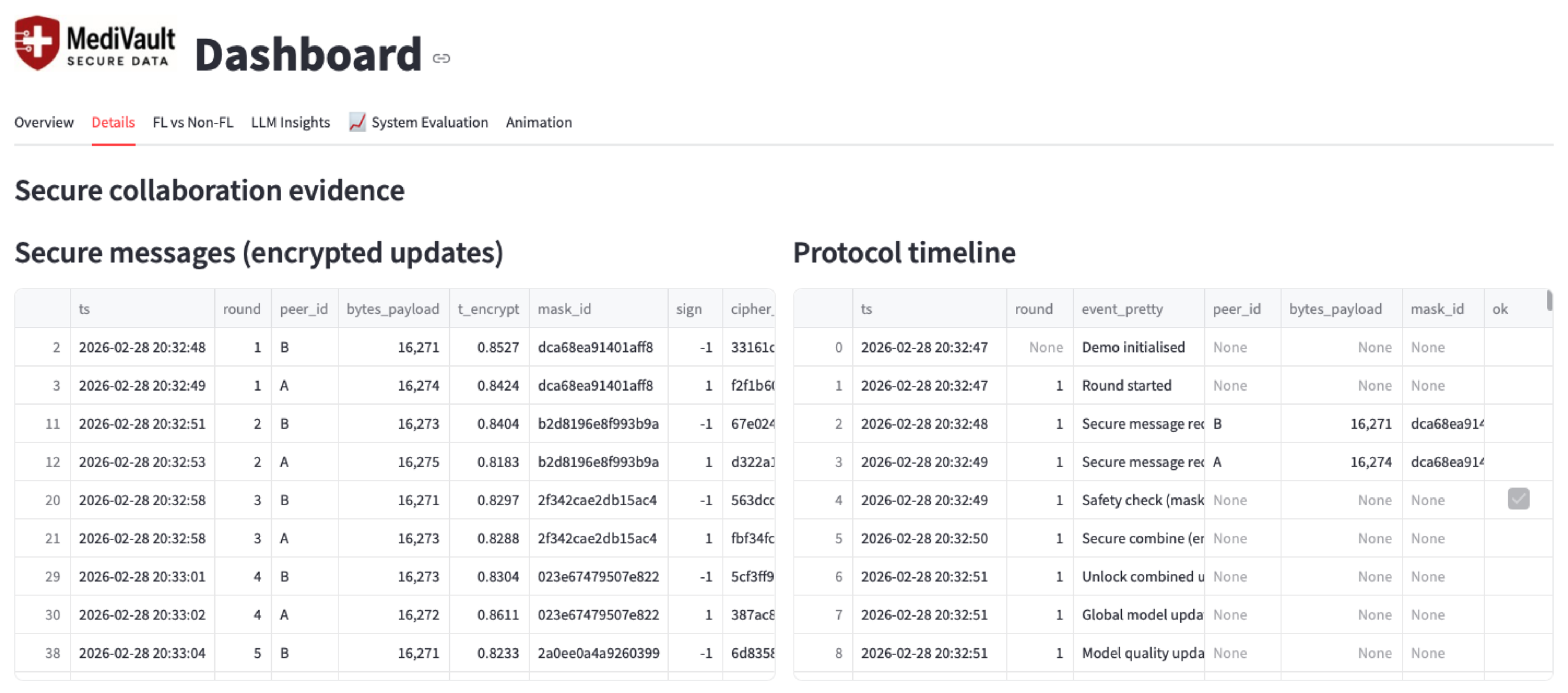
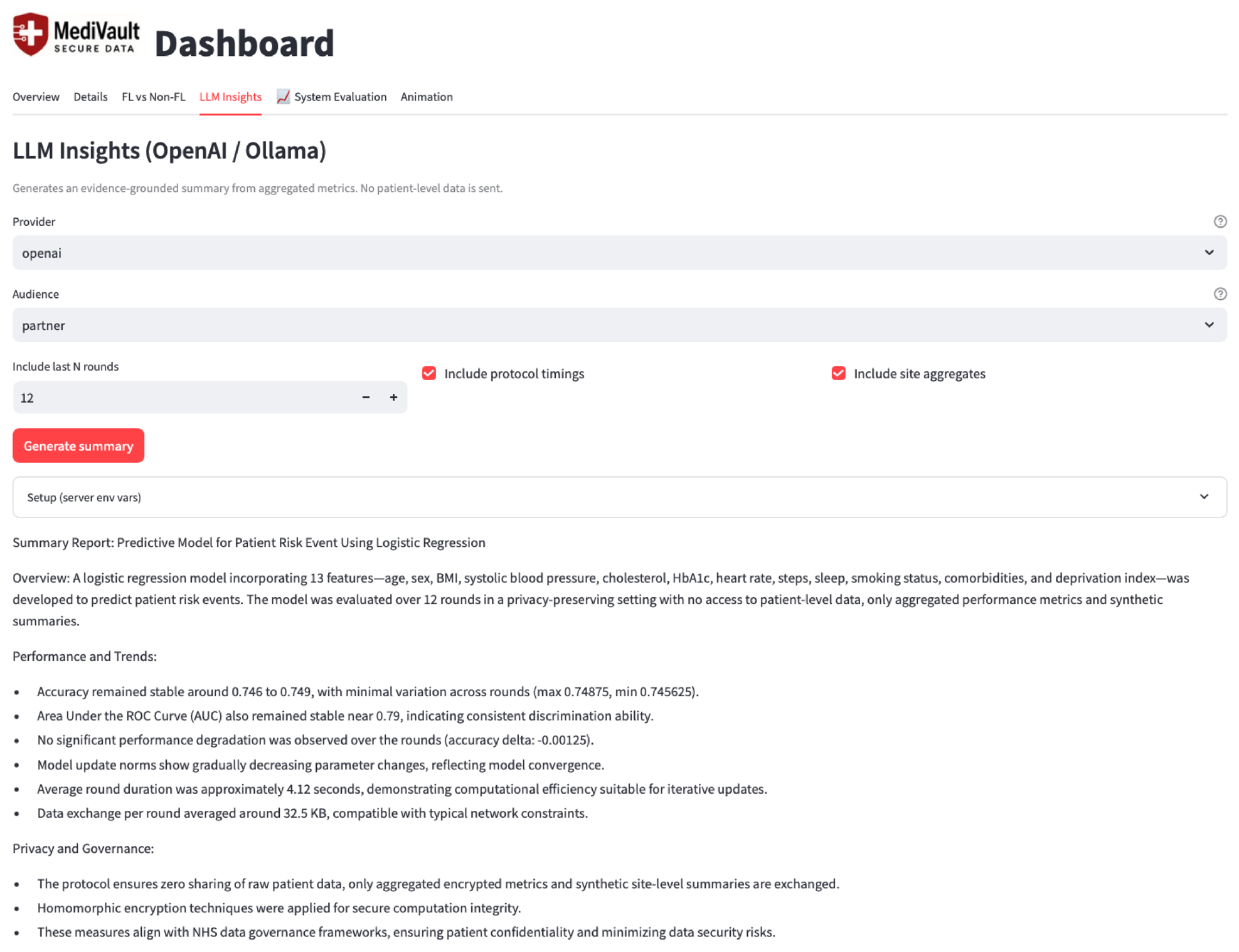
4.1. Implementation and Dashboard Views
4.2. Experimental Setup
4.2.1. Models, Baselines, and FL Setting
- LOGREG: logistic regression (probabilistic linear classifier).
- LINSVM: linear SVM (margin-based linear classifier).
- Centralised baseline (Non-FL): the model trained on the union of all training data.
- Federated learning (FL): peers train locally and submit model updates to a coordinator. The coordinator applies a FedAvg-style aggregation over received updates and evaluates the global model each round.
4.2.2. Peer Partitions (IID vs Non-IID)
- IID: each peer receives a roughly representative sample of the overall data distribution.
- Non-IID: peer data distributions are intentionally skewed so that different peers no longer follow the same underlying distribution, reflecting realistic site heterogeneity.
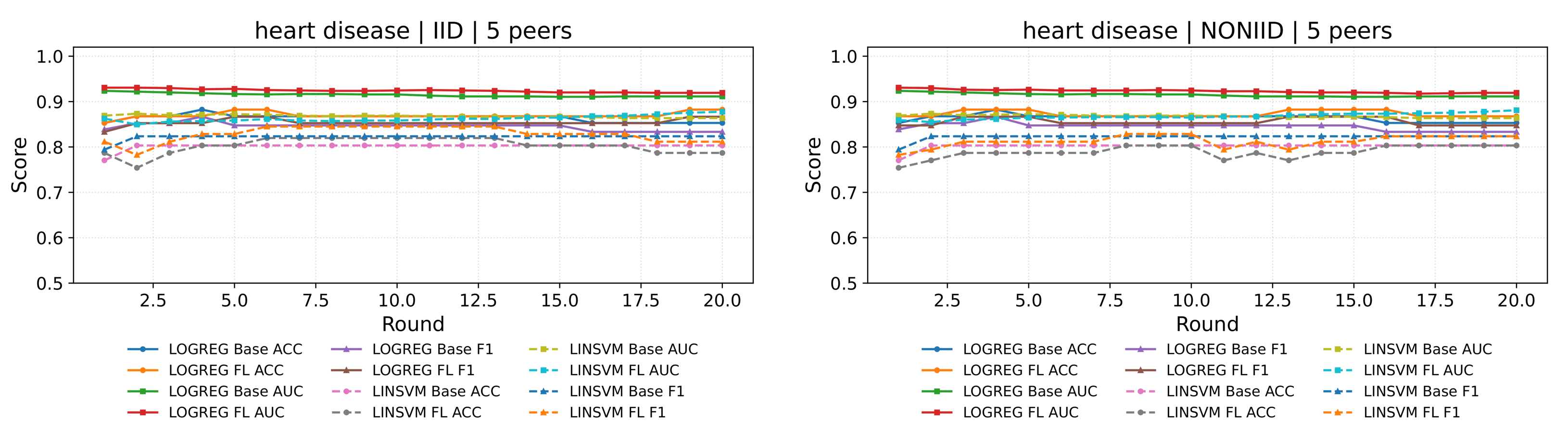
4.2.3. Metrics and Reporting Protocol
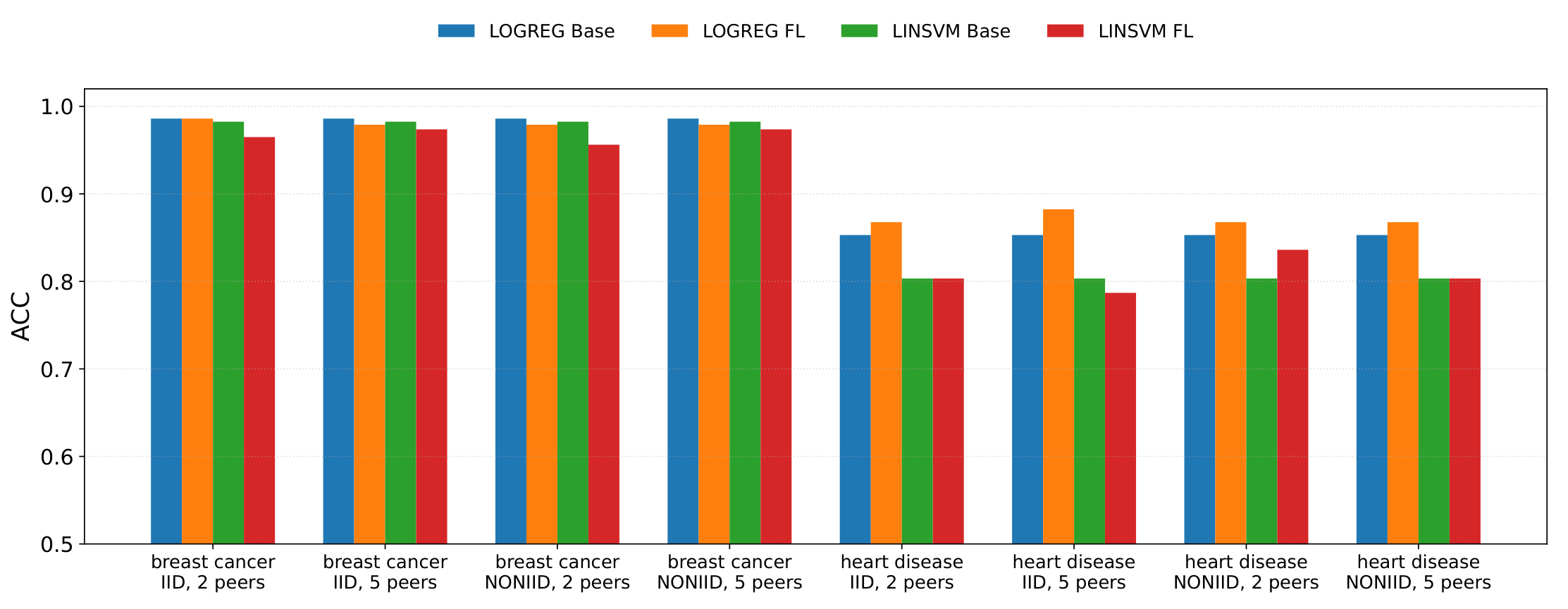
- Accuracy (ACC): overall classification correctness.
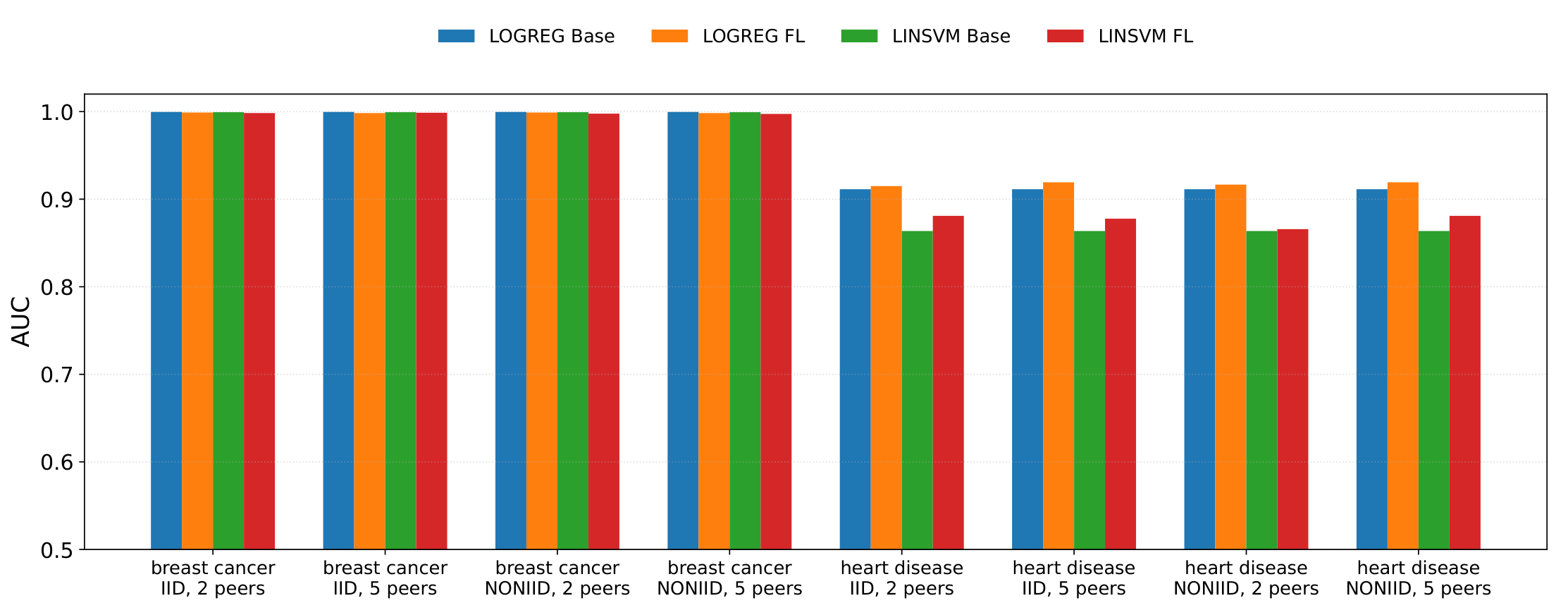
- Area Under the ROC Curve (AUC): threshold-independent ranking quality.
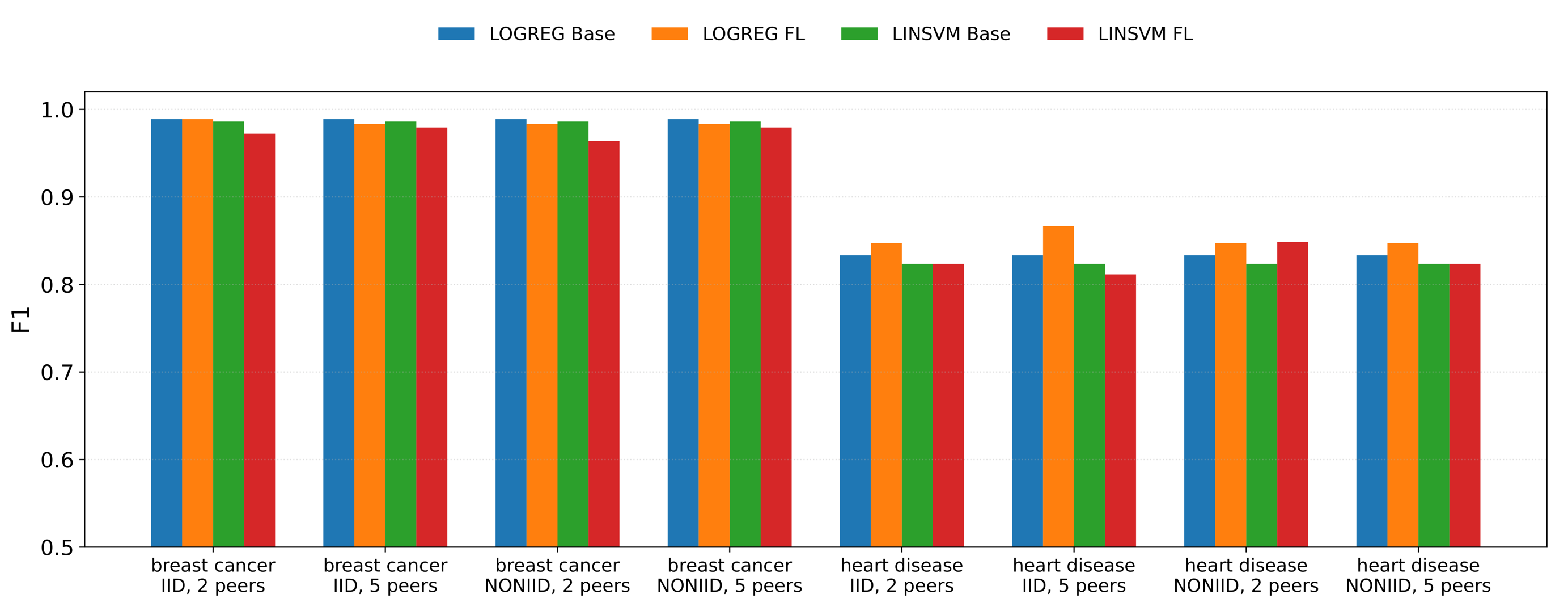
- F1-score (F1): balances precision and recall, which is useful under potential class imbalance.
4.2.4. Secure Update Confidentiality and Secure Aggregation
4.3. Auditability and Governance Evidence
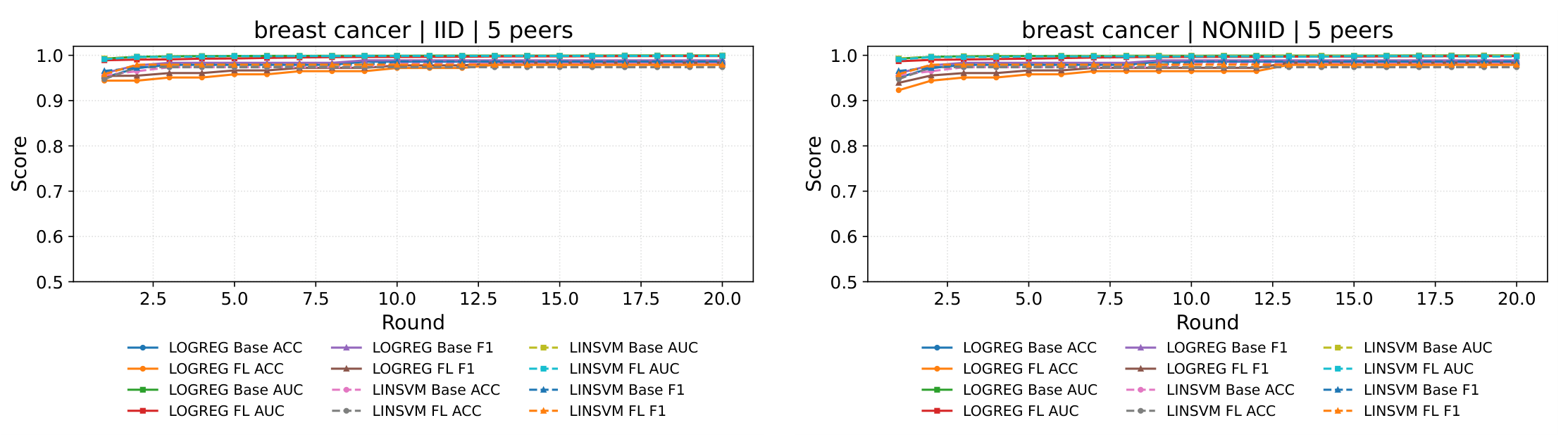
4.4. Results: Performance Comparison (Centralised vs Federated)
4.5. Discussion
5. Conclusion
Funding
References
- Zuo, Z.; Li, J.; Xu, H.; Al Moubayed, N. Curvature-based feature selection with application in classifying electronic health records. Technological Forecasting and Social Change 2021, 173, 121127. [CrossRef]
- Brasil, S.; Pascoal, C.; Francisco, R.; dos Reis Ferreira, V.; A. Videira, P.; Valadão, G. Artificial intelligence (AI) in rare diseases: is the future brighter? Genes 2019, 10, 978.
- Lee, J.; Liu, C.; Kim, J.; Chen, Z.; Sun, Y.; Rogers, J.R.; Chung, W.K.; Weng, C. Deep learning for rare disease: A scoping review. Journal of biomedical informatics 2022, 135, 104227. [CrossRef]
- Visibelli, A.; Roncaglia, B.; Spiga, O.; Santucci, A. The impact of artificial intelligence in the odyssey of rare diseases. Biomedicines 2023, 11, 887. [CrossRef]
- Decherchi, S.; Pedrini, E.; Mordenti, M.; Cavalli, A.; Sangiorgi, L. Opportunities and challenges for machine learning in rare diseases. Frontiers in medicine 2021, 8, 747612.
- Schaefer, J.; Lehne, M.; Schepers, J.; Prasser, F.; Thun, S. The use of machine learning in rare diseases: a scoping review. Orphanet journal of rare diseases 2020, 15, 145.
- McMahan, B.; Moore, E.; Ramage, D.; Hampson, S.; y Arcas, B.A. Communication-efficient learning of deep networks from decentralized data. In Proceedings of the Artificial intelligence and statistics. Pmlr, 2017, pp. 1273–1282.
- Li, T.; Sahu, A.K.; Zaheer, M.; Sanjabi, M.; Talwalkar, A.; Smith, V. Federated optimization in heterogeneous networks. Proceedings of Machine learning and systems 2020, 2, 429–450.
- Kairouz, P.; McMahan, H.B. Advances and open problems in federated learning. Foundations and trends in machine learning 2021, 14, 1–210.
- Sheller, M.J.; Reina, G.A.; Edwards, B.; Martin, J.; Bakas, S. Multi-institutional deep learning modeling without sharing patient data: A feasibility study on brain tumor segmentation. In Proceedings of the International MICCAI Brainlesion Workshop. Springer, 2018, pp. 92–104.
- Rieke, N.; Hancox, J.; Li, W.; Milletarì, F.; Roth, H.R.; Albarqouni, S.; Bakas, S.; Galtier, M.N.; Landman, B.A.; Maier-Hein, K.; et al. The future of digital health with federated learning. NPJ Digital Medicine 2020, 3. [CrossRef]
- Xu, J.; Glicksberg, B.S.; Su, C.; Walker, P.; Bian, J.; Wang, F. Federated learning for healthcare informatics. Journal of healthcare informatics research 2021, 5, 1–19. [CrossRef]
- Austin, J.A.; Lobo, E.H.; Samadbeik, M.; Engstrom, T.; Philip, R.; Pole, J.D.; Sullivan, C.M. Decades in the making: the evolution of digital health research infrastructure through synthetic data, common data models, and federated learning. Journal of Medical Internet Research 2024, 26, e58637.
- Shafik, W. Digital healthcare systems in a federated learning perspective. In Federated learning for digital healthcare systems; Elsevier, 2024; pp. 1–35.
- Bashir, A.K.; Victor, N.; Bhattacharya, S.; Huynh-The, T.; Chengoden, R.; Yenduri, G.; Maddikunta, P.K.R.; Pham, Q.V.; Gadekallu, T.R.; Liyanage, M. Federated learning for the healthcare metaverse: Concepts, applications, challenges, and future directions. IEEE Internet of Things Journal 2023, 10, 21873–21891. [CrossRef]
- Milasheuski, U.; Barbieri, L.; Tedeschini, B.C.; Nicoli, M.; Savazzi, S. On the impact of data heterogeneity in federated learning environments with application to healthcare networks. In Proceedings of the 2024 IEEE conference on artificial intelligence (CAI). IEEE, 2024, pp. 1017–1023.
- Bonawitz, K.; Ivanov, V.; Kreuter, B.; Marcedone, A.; McMahan, H.B.; Patel, S.; Ramage, D.; Segal, A.; Seth, K. Practical secure aggregation for privacy-preserving machine learning. In Proceedings of the proceedings of the 2017 ACM SIGSAC Conference on Computer and Communications Security, 2017, pp. 1175–1191.
- Beutel, D.J.; Topal, T.; Mathur, A.; Qiu, X.; Fernandez-Marques, J.; Gao, Y.; Sani, L.; Li, K.H.; Parcollet, T.; de GusmÃĢo, P.P.B.; et al. Flower: A friendly federated learning research framework. arXiv preprint arXiv:2007.14390 2020.
- He, C.; Li, S.; So, J.; Zeng, X.; Zhang, M.; Wang, H.; Wang, X.; Vepakomma, P.; Singh, A.; Qiu, H.; et al. Fedml: A research library and benchmark for federated machine learning. arXiv preprint arXiv:2007.13518 2020.
- Zhu, L.; Liu, Z.; Han, S. Deep Leakage from Gradients. Advances in Neural Information Processing Systems 2019, 32, 1–11. Placeholder entry. Please update with full details if needed.
- Melis, L.; Song, C.; De Cristofaro, E.; Shmatikov, V. Exploiting unintended feature leakage in collaborative learning. In Proceedings of the 2019 IEEE symposium on security and privacy (SP). IEEE, 2019, pp. 691–706.
- Bonawitz, K.; Ivanov, V.; Kreuter, B.; Marcedone, A.; McMahan, H.B.; Patel, S.; Ramage, D.; Segal, A.; Seth, K. Practical secure aggregation for privacy-preserving machine learning. In Proceedings of the proceedings of the 2017 ACM SIGSAC Conference on Computer and Communications Security, 2017, pp. 1175–1191.
- Gentry, C. Fully homomorphic encryption using ideal lattices. In Proceedings of the Proceedings of the forty-first annual ACM symposium on Theory of computing, 2009, pp. 169–178.
- Antunes, R.S.; André da Costa, C.; Küderle, A.; Yari, I.A.; Eskofier, B. Federated learning for healthcare: Systematic review and architecture proposal. ACM Transactions on Intelligent Systems and Technology (TIST) 2022, 13, 1–23.
- Chaddad, A.; Wu, Y.; Desrosiers, C. Federated learning for healthcare applications. IEEE internet of things journal 2023, 11, 7339–7358.
- Nguyen, D.C.; Pham, Q.V.; Pathirana, P.N.; Ding, M.; Seneviratne, A.; Lin, Z.; Dobre, O.; Hwang, W.J. Federated learning for smart healthcare: A survey. ACM Computing Surveys (Csur) 2022, 55, 1–37.
- Dhade, P.; Shirke, P. Federated learning for healthcare: a comprehensive review. Engineering Proceedings 2024, 59, 230.
- Zwitter, M.; Soklic, M. Breast Cancer. UCI Machine Learning Repository, 1988. [CrossRef]
- Janosi, Andras, S.W.P.M.; Detrano, R. Heart Disease. UCI Machine Learning Repository, 1989. [CrossRef]

| Final | Best | Mean±Std | |||||||||
| Dataset | Model | Part. | Peers | Base | FL | Base | FL | Base | FL | ||
| BC | LINSVM | IID | 2 | 0.982 | 0.965 | -0.018 | 0.982 | 0.974 | -0.009 | 0.977±0.007 | 0.969±0.008 |
| BC | LINSVM | IID | 5 | 0.982 | 0.974 | -0.009 | 0.982 | 0.974 | -0.009 | 0.977±0.007 | 0.972±0.006 |
| BC | LINSVM | Non | 2 | 0.982 | 0.956 | -0.026 | 0.982 | 0.965 | -0.018 | 0.977±0.007 | 0.951±0.013 |
| BC | LINSVM | Non | 5 | 0.982 | 0.974 | -0.009 | 0.982 | 0.974 | -0.009 | 0.977±0.007 | 0.969±0.009 |
| BC | LOGREG | IID | 2 | 0.986 | 0.986 | +0.000 | 0.986 | 0.986 | +0.000 | 0.981±0.008 | 0.976±0.011 |
| BC | LOGREG | IID | 5 | 0.986 | 0.979 | -0.007 | 0.986 | 0.979 | -0.007 | 0.981±0.008 | 0.967±0.012 |
| BC | LOGREG | Non | 2 | 0.986 | 0.979 | -0.007 | 0.986 | 0.979 | -0.007 | 0.981±0.008 | 0.974±0.010 |
| BC | LOGREG | Non | 5 | 0.986 | 0.979 | -0.007 | 0.986 | 0.979 | -0.007 | 0.981±0.008 | 0.965±0.015 |
| HD | LINSVM | IID | 2 | 0.803 | 0.803 | +0.000 | 0.836 | 0.820 | -0.016 | 0.812±0.016 | 0.805±0.006 |
| HD | LINSVM | IID | 5 | 0.803 | 0.787 | -0.016 | 0.836 | 0.820 | -0.016 | 0.812±0.016 | 0.797±0.010 |
| HD | LINSVM | Non | 2 | 0.803 | 0.836 | +0.033 | 0.836 | 0.852 | +0.016 | 0.812±0.016 | 0.833±0.011 |
| HD | LINSVM | Non | 5 | 0.803 | 0.803 | +0.000 | 0.836 | 0.820 | -0.016 | 0.812±0.016 | 0.802±0.008 |
| HD | LOGREG | IID | 2 | 0.853 | 0.868 | +0.015 | 0.882 | 0.882 | +0.000 | 0.864±0.008 | 0.867±0.006 |
| HD | LOGREG | IID | 5 | 0.853 | 0.868 | +0.015 | 0.882 | 0.882 | +0.000 | 0.864±0.008 | 0.869±0.006 |
| HD | LOGREG | Non | 2 | 0.853 | 0.838 | -0.015 | 0.882 | 0.882 | +0.000 | 0.864±0.008 | 0.855±0.018 |
| HD | LOGREG | Non | 5 | 0.853 | 0.838 | -0.015 | 0.882 | 0.882 | +0.000 | 0.864±0.008 | 0.864±0.012 |
| Final | Best | Mean±Std | |||||||||
| Dataset | Model | Part. | Peers | Base | FL | Base | FL | Base | FL | ||
| BC | LINSVM | IID | 2 | 0.999 | 0.998 | -0.001 | 0.999 | 0.998 | -0.001 | 0.999±0.001 | 0.998±0.001 |
| BC | LINSVM | IID | 5 | 0.999 | 0.999 | -0.001 | 0.999 | 0.999 | -0.001 | 0.999±0.001 | 0.999±0.001 |
| BC | LINSVM | Non | 2 | 0.999 | 0.998 | -0.002 | 0.999 | 0.998 | -0.001 | 0.999±0.001 | 0.998±0.001 |
| BC | LINSVM | Non | 5 | 0.999 | 0.997 | -0.002 | 0.999 | 0.999 | -0.001 | 0.999±0.001 | 0.998±0.002 |
| BC | LOGREG | IID | 2 | 1.000 | 0.999 | -0.001 | 1.000 | 0.999 | -0.001 | 1.000±0.000 | 0.998±0.002 |
| BC | LOGREG | IID | 5 | 1.000 | 0.998 | -0.001 | 1.000 | 0.998 | -0.001 | 1.000±0.000 | 0.996±0.003 |
| BC | LOGREG | Non | 2 | 1.000 | 0.999 | -0.001 | 1.000 | 0.999 | -0.001 | 1.000±0.000 | 0.997±0.002 |
| BC | LOGREG | Non | 5 | 1.000 | 0.998 | -0.001 | 1.000 | 0.998 | -0.001 | 1.000±0.000 | 0.995±0.003 |
| HD | LINSVM | IID | 2 | 0.864 | 0.881 | +0.017 | 0.915 | 0.916 | +0.001 | 0.894±0.018 | 0.903±0.013 |
| HD | LINSVM | IID | 5 | 0.864 | 0.878 | +0.014 | 0.915 | 0.916 | +0.001 | 0.894±0.018 | 0.897±0.012 |
| HD | LINSVM | Non | 2 | 0.864 | 0.866 | +0.002 | 0.915 | 0.900 | -0.015 | 0.894±0.018 | 0.878±0.021 |
| HD | LINSVM | Non | 5 | 0.864 | 0.881 | +0.017 | 0.915 | 0.916 | +0.001 | 0.894±0.018 | 0.900±0.014 |
| HD | LOGREG | IID | 2 | 0.911 | 0.915 | +0.004 | 0.924 | 0.926 | +0.003 | 0.912±0.008 | 0.918±0.004 |
| HD | LOGREG | IID | 5 | 0.911 | 0.920 | +0.009 | 0.924 | 0.931 | +0.007 | 0.912±0.008 | 0.920±0.004 |
| HD | LOGREG | Non | 2 | 0.911 | 0.917 | +0.006 | 0.924 | 0.931 | +0.007 | 0.912±0.008 | 0.917±0.007 |
| HD | LOGREG | Non | 5 | 0.911 | 0.920 | +0.009 | 0.924 | 0.931 | +0.007 | 0.912±0.008 | 0.920±0.004 |
| Final | Best | Mean±Std | |||||||||
| Dataset | Model | Part. | Peers | Base | FL | Base | FL | Base | FL | ||
| BC | LINSVM | IID | 2 | 0.986 | 0.972 | -0.014 | 0.986 | 0.978 | -0.008 | 0.981±0.007 | 0.976±0.007 |
| BC | LINSVM | IID | 5 | 0.986 | 0.979 | -0.007 | 0.986 | 0.979 | -0.007 | 0.981±0.007 | 0.978±0.005 |
| BC | LINSVM | Non | 2 | 0.986 | 0.964 | -0.022 | 0.986 | 0.970 | -0.016 | 0.981±0.007 | 0.959±0.012 |
| BC | LINSVM | Non | 5 | 0.986 | 0.979 | -0.007 | 0.986 | 0.979 | -0.007 | 0.981±0.007 | 0.976±0.009 |
| BC | LOGREG | IID | 2 | 0.989 | 0.989 | +0.000 | 0.989 | 0.989 | +0.000 | 0.982±0.009 | 0.981±0.009 |
| BC | LOGREG | IID | 5 | 0.989 | 0.983 | -0.006 | 0.989 | 0.983 | -0.006 | 0.982±0.009 | 0.972±0.010 |
| BC | LOGREG | Non | 2 | 0.989 | 0.983 | -0.006 | 0.989 | 0.983 | -0.006 | 0.982±0.009 | 0.980±0.008 |
| BC | LOGREG | Non | 5 | 0.989 | 0.983 | -0.006 | 0.989 | 0.983 | -0.006 | 0.982±0.009 | 0.971±0.012 |
| HD | LINSVM | IID | 2 | 0.824 | 0.824 | +0.000 | 0.849 | 0.836 | -0.013 | 0.825±0.010 | 0.824±0.006 |
| HD | LINSVM | IID | 5 | 0.824 | 0.812 | -0.012 | 0.849 | 0.836 | -0.013 | 0.825±0.010 | 0.814±0.010 |
| HD | LINSVM | Non | 2 | 0.824 | 0.848 | +0.025 | 0.849 | 0.862 | +0.013 | 0.825±0.010 | 0.845±0.011 |
| HD | LINSVM | Non | 5 | 0.824 | 0.824 | +0.000 | 0.849 | 0.836 | -0.013 | 0.825±0.010 | 0.823±0.008 |
| HD | LOGREG | IID | 2 | 0.839 | 0.847 | +0.009 | 0.867 | 0.867 | +0.000 | 0.844±0.009 | 0.848±0.007 |
| HD | LOGREG | IID | 5 | 0.839 | 0.852 | +0.014 | 0.867 | 0.867 | +0.000 | 0.844±0.009 | 0.851±0.007 |
| HD | LOGREG | Non | 2 | 0.839 | 0.820 | -0.019 | 0.867 | 0.867 | +0.000 | 0.844±0.009 | 0.835±0.019 |
| HD | LOGREG | Non | 5 | 0.839 | 0.847 | +0.009 | 0.867 | 0.867 | +0.000 | 0.844±0.009 | 0.847±0.011 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




