Submitted:
14 April 2026
Posted:
16 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. System architecture
3.1. Modules
3.2. Physiological Modules
- CAP (Compound Action Potential) Simulation Module: simulates the stimulation of the afferent peripheral nerve trunk, with sensory fibre responses recorded at various distances from the stimulation point (e.g., wrist, elbow, axilla, Erb’s point) (Figure 9)
- CMAP (Compound Motor Action Potential) Simulation Module: simulates the stimulation of the efferent peripheral nerve trunks, recording muscle responses across different segments and distances.
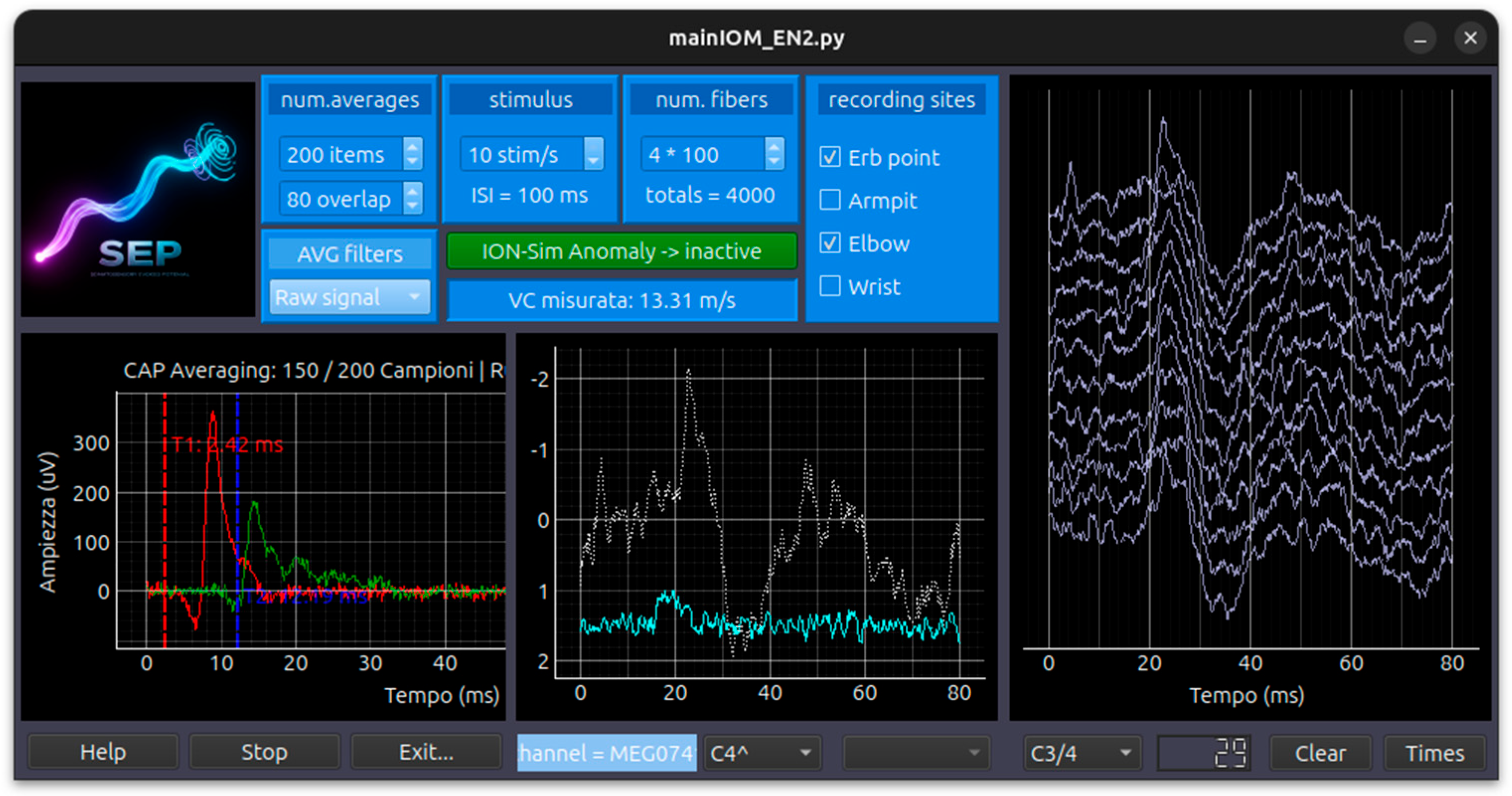
- SEP (Somatosensory Evoked Potential) Simulation Module: involves peripheral nerve stimulation with recording montages at Erb’s point, the CV6 level, and scalp electrodes over the contralateral S1 and M1 sites. Key controllable parameters include the Stimulation Side (Right, Left), Block Average (sweep count), Overlay Percentage, Noise Amplitude (μV), and Gain Control Figure 7
- VEP (Visual Evoked Potential) Simulation Module: simulates retinal flash stimulation with scalp recording over ipsilateral and contralateral occipital sites. Key controllable parameters include the Stimulation Mode (Right, Left, Bilateral), the Noise Level and the number of sweeps per block (update) Figure 8
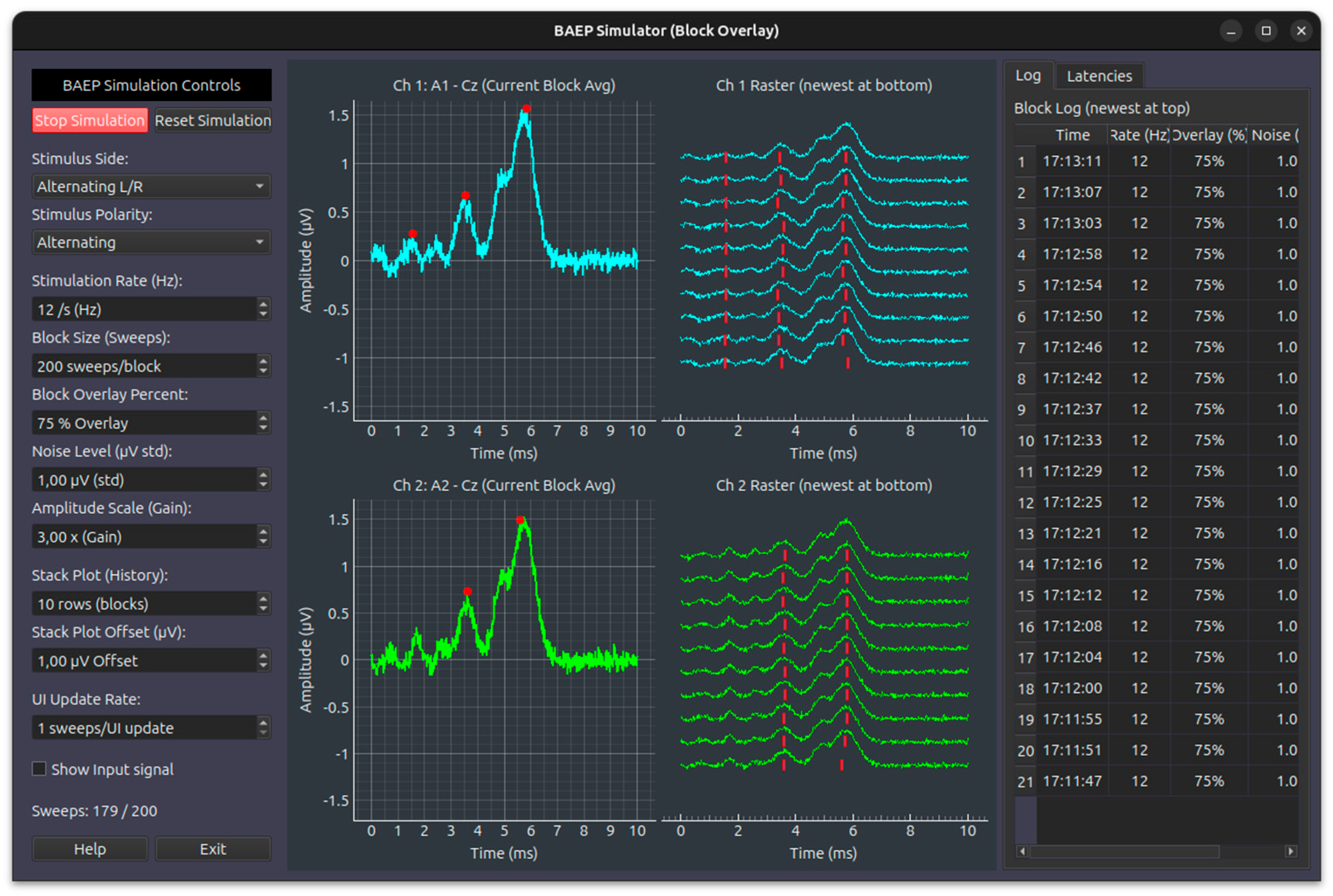
- BAEP (Brainstem Auditory Evoked Potential) Simulation Module: utilizes click-based acoustic stimulation with contralateral masking noise, recorded via scalp electrodes using a bilateral mastoid montage. The module features automatic identification of latency and amplitude peaks for principal wave components (Waves I, III, V). Configurable parameters include: Stimulation Side (Right, Left, Alternating), Stimulus Polarity (Rarefaction, Condensation, Alternating), Stimulus Frequency (Hz), Block Average (sweep count), Overlay Percentage, Noise Amplitude (μV), and Gain Control. Figure 6
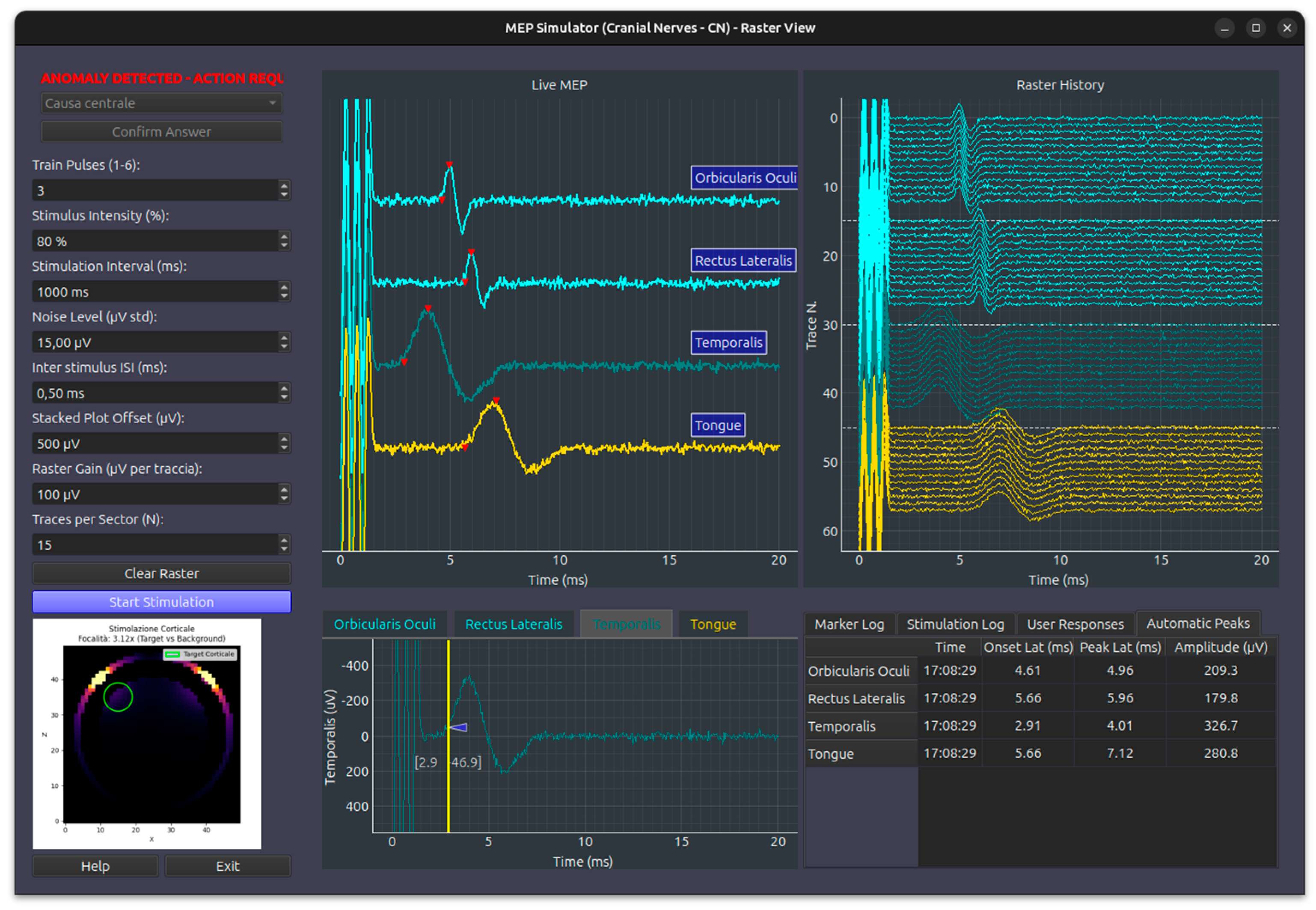
- MEPcb (Motor Evoked Potential - Cranio-Bulbar) Simulation Module: simulates transcranial scalp stimulation using a pulse train, recording from muscles innervated by cranial nerves (facial, trigeminal, glossopharyngeal, hypoglossal, spinal accessory). It supports up to 8 channels for CMAP recording and features a raster plot to track response variations over time, offering automatic marker localization for peak latency and amplitude. User-adjustable parameters include stimulation pulse count, amplitude (%), inter-stimulus interval, and noise levels. Both stack plot and raster display scales are fully customizable Figure 4
- MEPas (Motor Evoked Potential - Upper Limbs) Simulation Module: models muscle activation in the upper limbs following transcranial electrical pulse train stimulation, specifically targeting the 1st DI (first dorsal interosseous), ECD (extensor communis digitorum), and BB (biceps brachii) muscles. This module inherits the control interface and parameter definitions of the MEPcb module, presenting anomalies governed by predefined configuration schemas Figure 4
- MEPai (Motor Evoked Potential - Lower Limbs) Simulation Module: models muscle activation in the lower limbs following transcranial electrical pulse train stimulation, targeting the FHB (flexor hallucis brevis), TA (tibialis anterior), and VL (vastus lateralis) muscles. Similar to the upper limb module, it utilizes the standard MEP control interface and presents anomalies based on predefined configuration schemas Figure 4
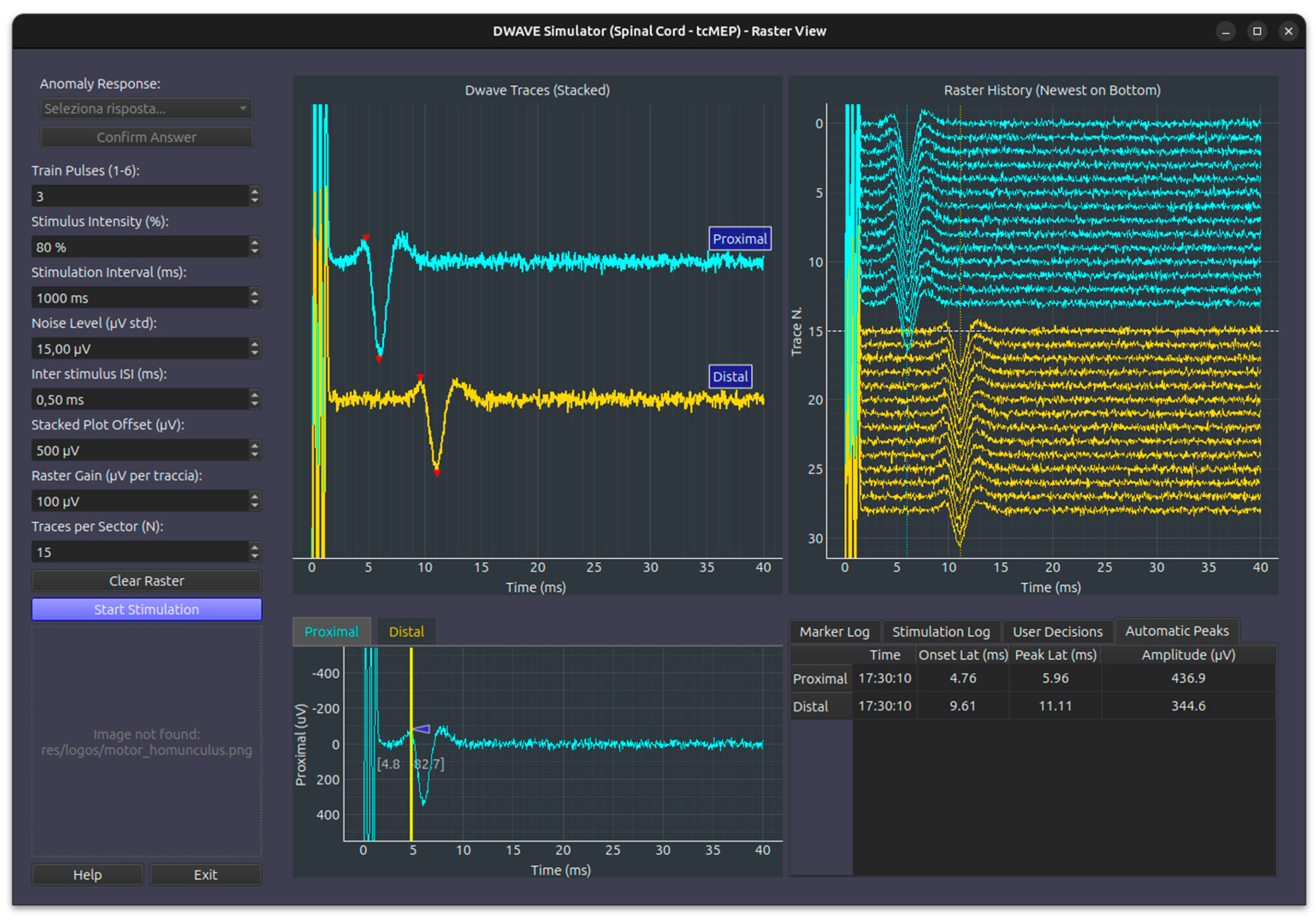
- D-Wave (Direct Wave) Simulation Module: simulates corticospinal tract activation recorded directly from the spinal cord. While stimulation remains transcranial, recording is performed at two distinct spinal sites: proximal and distal to the surgical intervention level. The module incorporates standard stimulation controls and features a specialized “anomaly detection” interface, which manages introduced faults and queries the user for diagnostic responses Figure 5
- DCS (Direct Cortical Stimulation) Simulation Module: models diverse responses secondary to direct cortical stimulation, including motor responses, disruption of language function (speech arrest/aphasia), and alterations in cognitive processing (in progress).
- CCEP (Cortico-Cortical Evoked Potentials) Simulation Module: simulates cortical or subcortical stimulation with the recording of evoked responses at specific cortical distances, aimed at mapping the structural connectivity of tracts and cortical areas (in progress).
- EEG (Electroencephalography) Simulation Module: displays resting electrical activity and physiological variations associated with eyes open/closed states and motor activity (e.g., mu-rhythm desynchronization) and incorporates physiological cyclical variations derived from real-world clinical recordings. Figure 10
- ECoG (Electrocorticography) Simulation Module: simulates spontaneous corticographic activity, incorporating physiological cyclical variations derived from real-world clinical recordings Figure 11
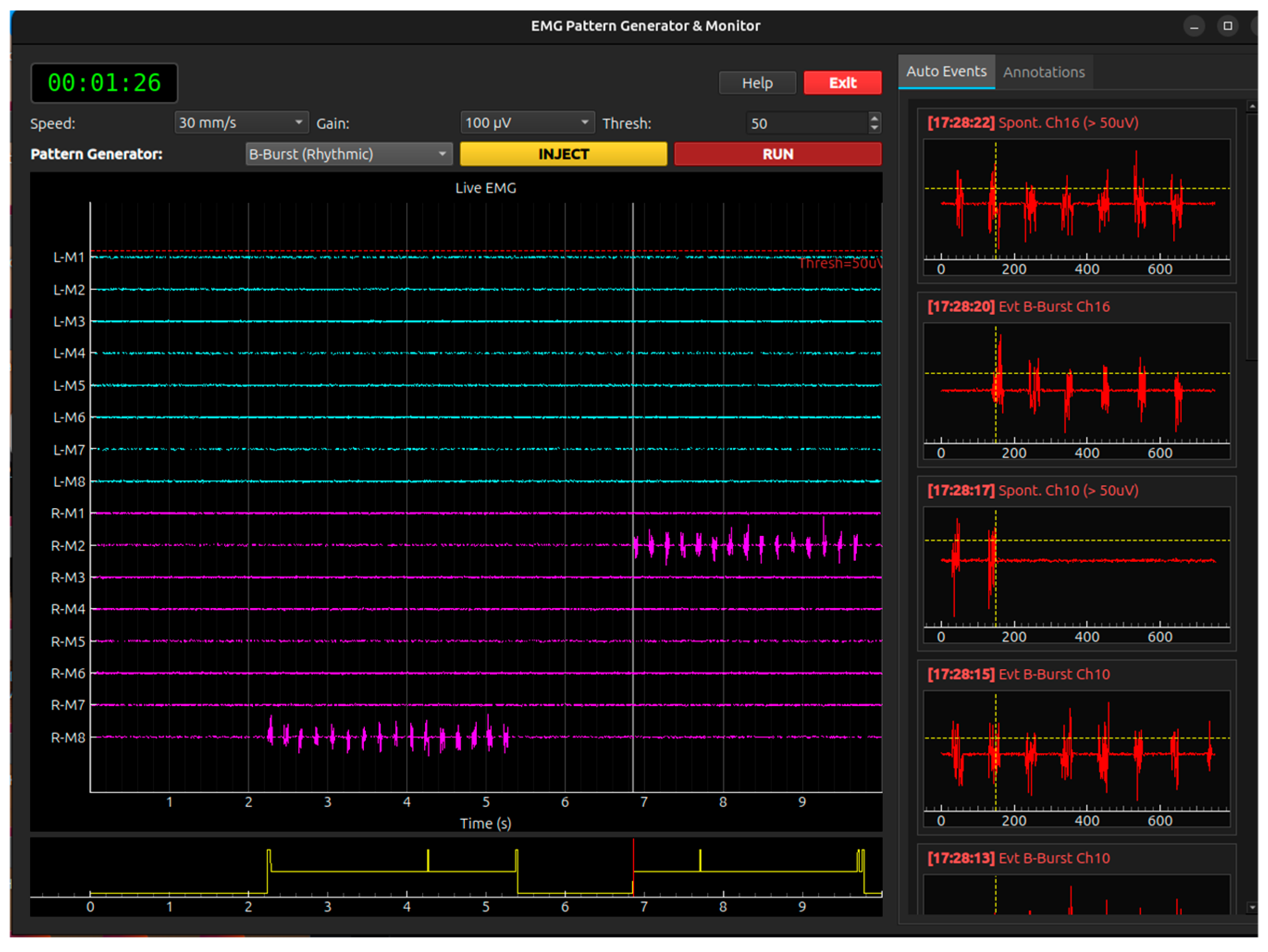
- EMG (Electromyography) Simulation Module: models spontaneous electromyographic activity, including fluctuations and voluntary contraction phenomena, typical of awake patient procedures (e.g., awake craniotomy) and different patterns of spontaneous discharges Figure 12
- ANESTHESIA Simulation Module: displays patient metadata (age, sex, weight, height, BMI, LBM), monitors vital signs (SpO₂, HR, R-R interval, SDNN, RMSSD, NIBP, MAP, temperature, RPM), and manages drug infusion via target-controlled infusion (TCI) for Sevoflurane, Propofol, Remifentanil, and Ketamine Figure 13.
3.3. Optimisation of Evoked Response Updates
3.4. Simulation of Spontaneous “Free-Running” Activity
3.5. Anomaly Simulation Module
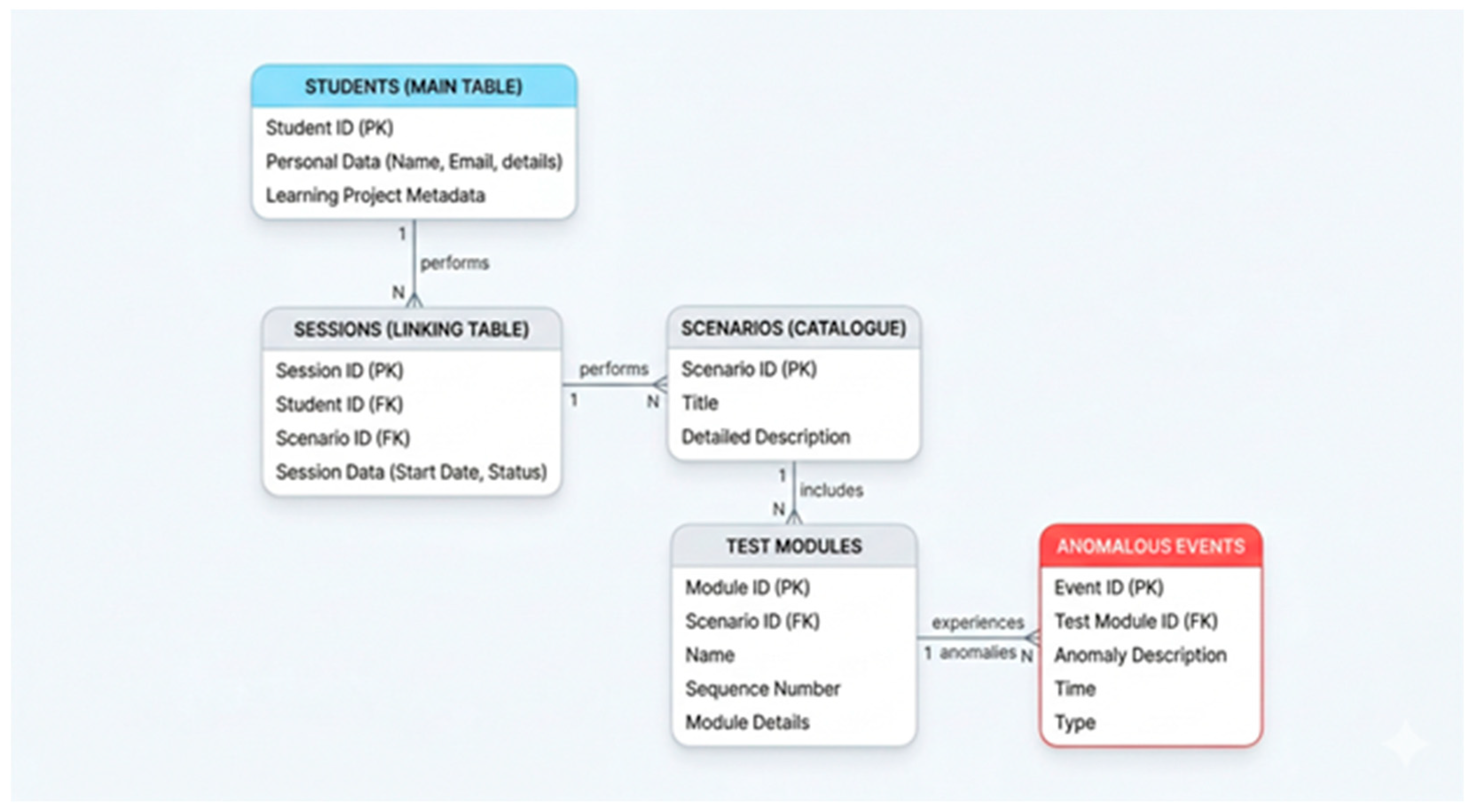
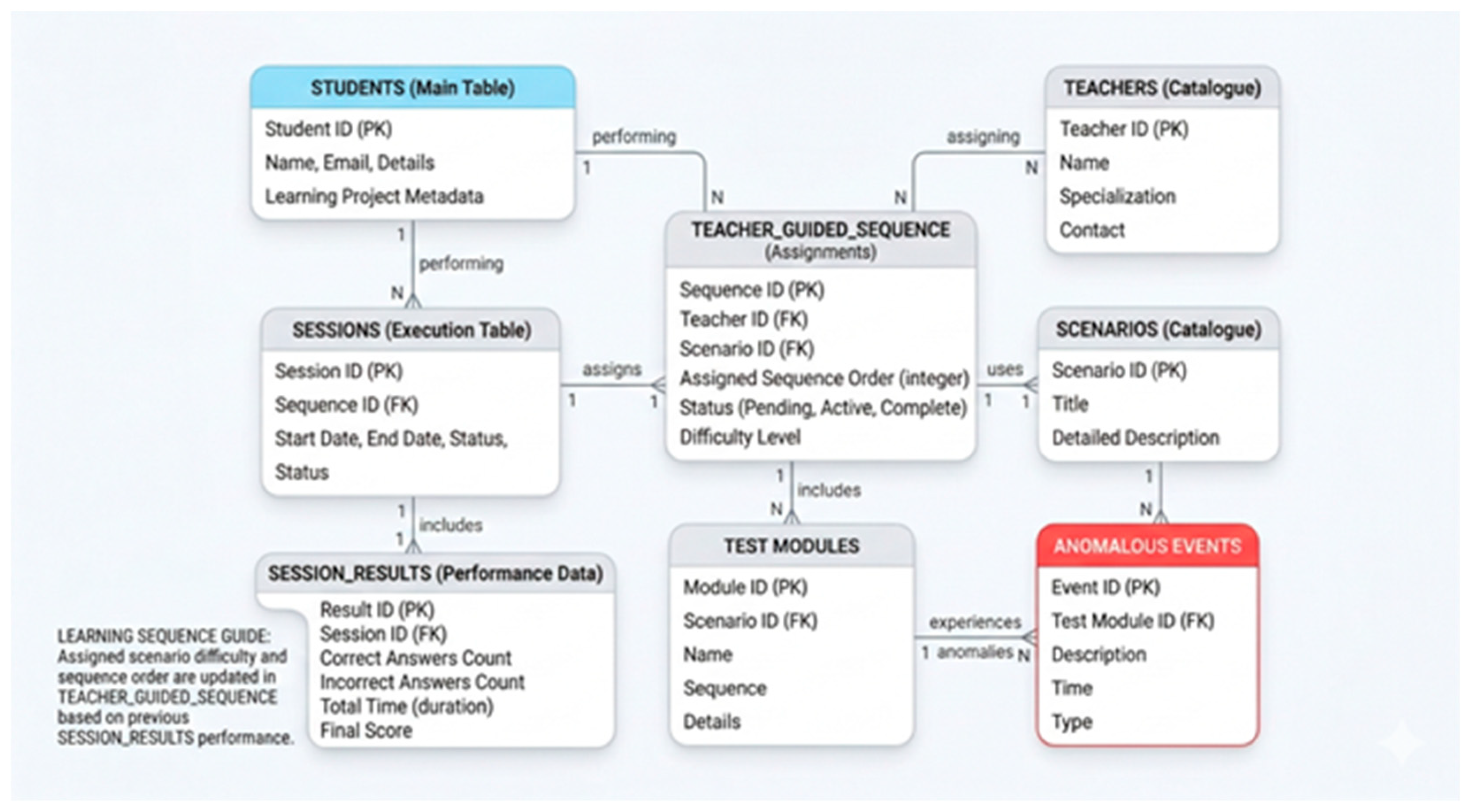
3.6. Learning Module
4. Software Implementation and Availability
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| EEG | Electro Encephalography |
| ECoG | Electro Corticography |
| MEP | Motor Evoked Potential |
| SEP | Somatosensory Evoked Potential |
| VEP | Visual Evoked Potential |
| Dwave | D wave |
| CAP | Compound Action Potential |
| CMAP | Compound Motor Action Potential |
| SFAP | Single Fiber Action Potential |
| MUAP | Motor Unit Action Potential |
| BAEP | Brain Stem Acoustic Evoked Potential |
Appendix A

Appendix A.1 Module MEP
- Dual Gaussian Modeling: A single MEP response is created by summing two Gaussian peaks—a primary negative wave and a secondary “polyphasic” shift. This mimics the complex morphology of real muscle potentials better than a single peak.
- Muscle-Specific Parameters: The simulator uses a global dictionary (ALL_MUSCLE_PARAMS) to assign unique latencies, amplitudes, and widths to each muscle. For example, the Deltoideus has a short latency (8 ms), while the FPA (foot) has a long latency (37 ms), reflecting the distance from the motor cortex.
- Train-of-Pulses Facilitation: The simulator applies a facilitation_map where the final amplitude scale is determined by the number of pulses in the stimulator “train” (1 to 6 pulses).
- Stimulation Artifacts: It generates a series of high-frequency stimulation artifacts at the beginning of the trace, corresponding to the “Train Pulses” and “ISI” (Inter-Stimulus Interval) settings, providing a realistic visual reference for the moment of stimulation.
- Stacked View: Displays the current MEP for each muscle vertically offset, allowing for easy identification of morphology and peak markers (onset and peak latency).
- Raster History: Implements a “waterfall” or scrolling history where the last 15 traces for each muscle are displayed. This is critical in IOM to detect subtle trends or sudden signal loss over time
- Physical Signal Modification: When an anomaly is active, the injector modifies the physical data in real-time (e.g., causing a 50% drop in amplitude or a latency shift).
- Decision-Making Assessment: Once the user detects a change, the UI enables an “Anomaly Response” panel. The user must select the correct clinical action from a list of choices.
Appendix A.2 Module BAEP
- •
- Gaussian Model: The worker uses the formula Amp⋅e−2σ2(t−Lat)2 to generate smooth, physiologically realistic peaks.
- •
- Anatomical Differentiation: The simulator distinguishes between two recording channels:
- ○
- Channel 1 (Ipsilateral, A1-Cz): Contains the full complex (Waves I, II, III, IV, and V).
- ○
- Channel 2 (Contralateral, A2-Cz): Only simulates Waves III, IV, and V, with Wave III reduced in amplitude to 30%, reflecting the far-field nature of the contralateral recording.
- Rarefaction/Condensation: These introduce a small latency shift (0.08 ms) and control the polarity of the Cochlear Microphonic (CM).
- Alternating: The worker alternates polarity every sweep; when these are averaged, the phase-inverted CM is cancelled out, which is a standard clinical technique to isolate the neural response.
- CM Generation: The CM is modeled as a sine wave that decays exponentially over time.
- Noise Injection: Every individual sweep is combined with random Gaussian noise.
- Block Averaging: Signals are processed in “blocks” (e.g., 200 sweeps).
- Overlay Method: Instead of resetting to zero after every block, the system can retain a percentage of the previous block’s data (e.g., 75% overlay). This creates a smooth “moving average” effect in the history (Stack Plot), allowing trainees to see signals evolve as noise is reduced.
Appendix A.3 Module SEP
- Single Fiber Modeling: Each individual nerve fiber’s action potential (SFAP) is modeled as a biphasic pulse.
- Fiber Dispersion: The simulator generates a population of fibers (default 100) with varying Conduction Velocities (CV) ranging from 20 to 65 m/s.
- Temporal Summation: The CAP at a specific distance d is the sum of SFAPs, where each fiber’s delay is calculated as Delay=VelocityDistance. As the distance increases, the individual fiber potentials “spread out” (dispersion), accurately mimicking nerve physiology.
- Static Field Potential: It calculates the potential using a Source-Sink dipole model with a fixed conductivity (σe).
- Dynamic Current Pulse: The temporal evolution is governed by a current pulse function that models the rise and decay of the signal as the volley passes the cervical electrode.
- MNE Integration: The simulator loads a pre-recorded dataset from the “mne” library.
- Dynamic Interpolation: It interpolates and resamples the real MEG data to match the simulator’s specific time vector. Users can switch between different MEG channels (e.g., C3, C4) to see how the cortical SEP changes based on electrode location.
- Averaging Engine: To mimic clinical practice, the simulator uses a FIFO (First-In-First-Out) buffer to perform signal averaging. This reduces injected Gaussian noise to reveal the evoked response.
- Waterfall Plotting: The system generates “Waterfall” views, which are historical stacks of averaged blocks, allowing users to monitor signal stability over time.
- Anomaly Injection and AI: The system includes an AnomalyInjector that can physically alter the signals (e.g., adding latency or decreasing amplitude) based on a JSON configuration. This is used for Learning Assessment, where the user must identify the problem and select the correct clinical action, which is then analyzed by an AI Agent for feedback.
Appendix A.4 Module VEP
- N75: A negative peak at ~75 ms.
- P100: The most clinically significant positive peak at ~100 ms.
- N145: A negative peak at ~145 ms.
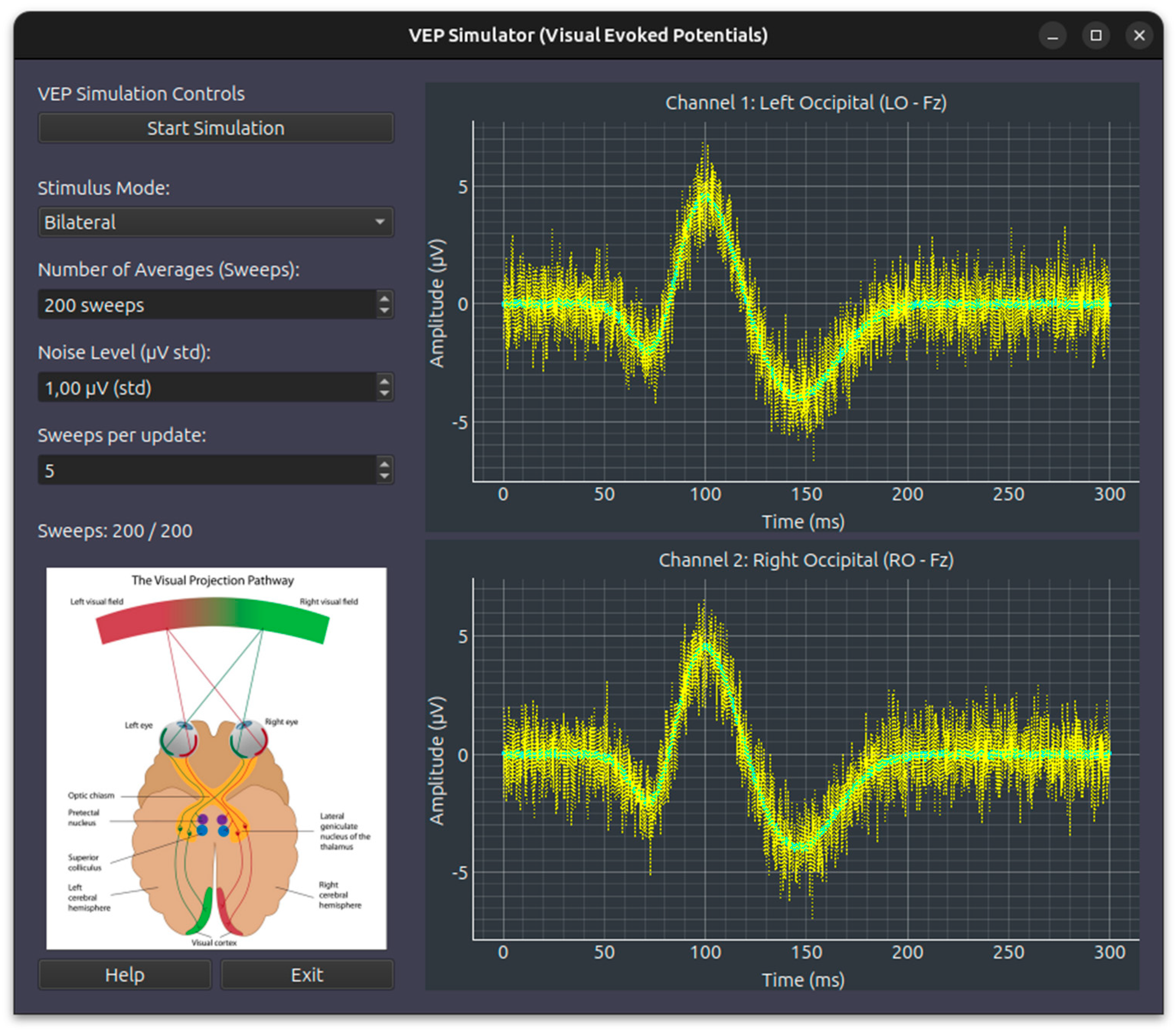
- Bilateral: Full signal strength (100%) is applied to both the Left (Ch1) and Right (Ch2) Occipital channels.
- Monolateral (Left/Right): To simulate the partial crossing of fibers at the optic chiasm, the simulator applies full amplitude (1.0) to the contralateral hemisphere and a reduced scale factor (0.6) to the ipsilateral hemisphere.
- Gaussian Noise: Every “sweep” (individual stimulus response) is injected with random Gaussian noise based on a user-adjustable standard deviation.
- Moving Average: The simulator calculates a continuous mean across sweeps. As the sweep count increases toward the target (e.g., 200 sweeps), the random noise cancels out, and the stable VEP waveform emerges from the baseline.
- Dynamic Simulation Speed: Users can adjust “Sweeps per update” to accelerate the averaging process for training purposes.
Appendix A.5 Module CAP
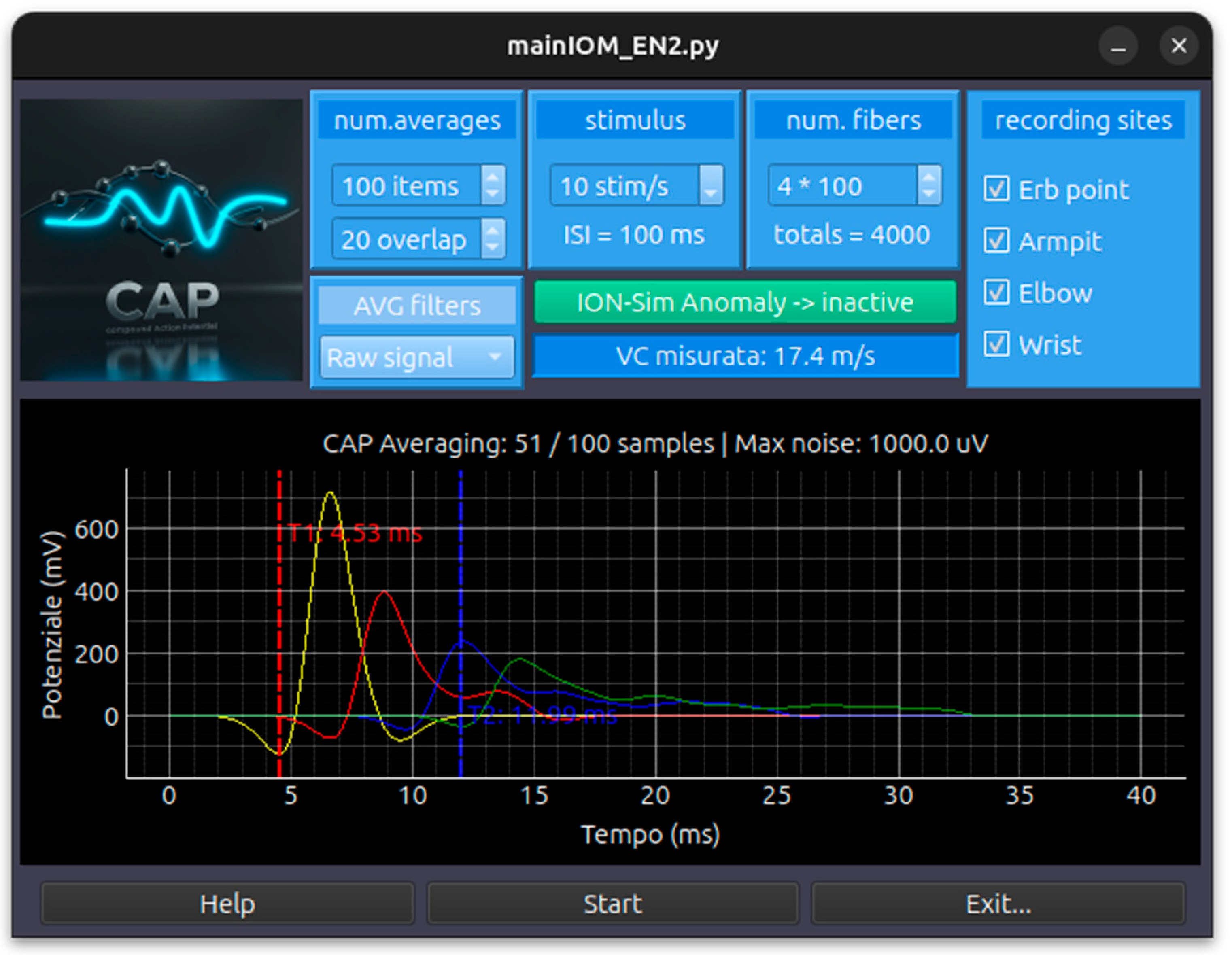
- Bipolar Gaussian Model: Combines two Gaussian functions (one positive and one negative) to create the characteristic biphasic shape of an extracellular recording.
- Ricker Wavelet Model: An alternative mathematical model (often called the “Mexican Hat”) used to simulate the SFAP pulse.
- Conduction Velocity (VC) Distribution: Each fiber is assigned a velocity within a specific range (typically 20.0 to 65.0 m/s).
- Spatial Propagation: The potential for a specific distance (d) is calculated by applying a time delay to each fiber (Delay=VCd).
- Compound Summation: The CAP displayed on the screen is the linear summation of all individual SFAPs. This naturally simulates temporal dispersion: as the recording site moves further from the stimulus, the signal becomes wider and lower in amplitude because faster and slower fibers arrive at increasingly different times.
- Gaussian Noise Injection: Random noise is added to the raw signal to simulate background interference.
- FIFO Averaging: The simulator uses a First-In-First-Out (FIFO) buffer system (default 100 averages) to perform signal averaging, which improves the signal-to-noise ratio over time.
- Digital Filtering: Users can apply various filters in real-time, including Moving Average (Smooth), Spline Interpolation, and Butterworth Bandpass filters (e.g., 5-3000 Hz or 10-1500 Hz).
- Velocity Measurement: The GUI includes interactive vertical cursors. By placing cursors on the peaks of signals from different distances (e.g., Wrist at 7cm and Elbow at 20cm), the software automatically calculates the measured Conduction Velocity in m/s.
- Anomaly Injection: The code integrates an AnomalyInjector that can physically alter the signal in real-time based on a JSON configuration. This is used for training scenarios where a student must identify clinical changes (such as ischemia or compression) and choose the correct corrective action from a dropdown menu.
Appendix A.6 Module CMAP
- Triphasic Morphology: The single_muap_waveform function models a triphasic (Positive-Negative-Positive) wave. This is achieved by combining three Gaussian-like phases (P1, N1, and P2) with specific amplitudes and time offsets.
- Stochastic Variation: For every motor unit in the simulation, the code randomly varies the amplitude (simulating different motor unit sizes) and the duration/sigma (simulating the physical characteristics of the muscle fibers).
- Conduction Velocity (VC) Distribution: Each motor unit is assigned a specific velocity based on a normal distribution (e.g., mean of 60 m/s with a standard deviation).
- Propagation Delay: The software calculates a unique delay for each unit based on the anatomical distance (Delay=VelocityDistance).
- Temporal Dispersion: Because different fibers conduct at different speeds, the individual MUAPs arrive at the recording electrode at slightly different times. This causes the resulting CMAP to “spread out” and lower in amplitude as the distance increases, accurately reflecting nerve physiology.
- CMAP Methodology: Primarily focuses on peripheral conduction velocity dispersion and a fixed synaptic/junctional delay (e.g., 2.3 ms) to model the time it takes for the signal to cross the neuromuscular junction.
- MEP Methodology: Introduces the concept of Central Jitter (tccm_spread_ms). Unlike the synchronized stimulation of a peripheral nerve, a cortical stimulation (tcMEP) results in desynchronized firing of the corticospinal tract. The code models this by adding a “jitter” or temporal spread to each MUAP, which creates the complex, multi-peaked morphology typical of clinical MEPs.
Appendix A.7 Module EEG
- MNE CNT Loading: The simulator loads Neuroscan .cnt files using the mne.io.read_raw_cnt module.
- Real-time Resampling: If the original file’s sampling frequency differs from the target (default 512 Hz), the code performs on-the-fly resampling.
- Microvolt Scaling: Data is scaled by a factor of −1×106 to ensure signals are displayed in standard microvolts (μV).
- Average Referencing: The spatial mean of all 16 EEG channels is subtracted from each channel to minimize common-mode noise.
- Frequency Filtering: The worker applies a bandpass filter (typically 1.6–35 Hz for EEG and 1.0–25 Hz for ECG) to remove slow drifts and high-frequency muscle interference.
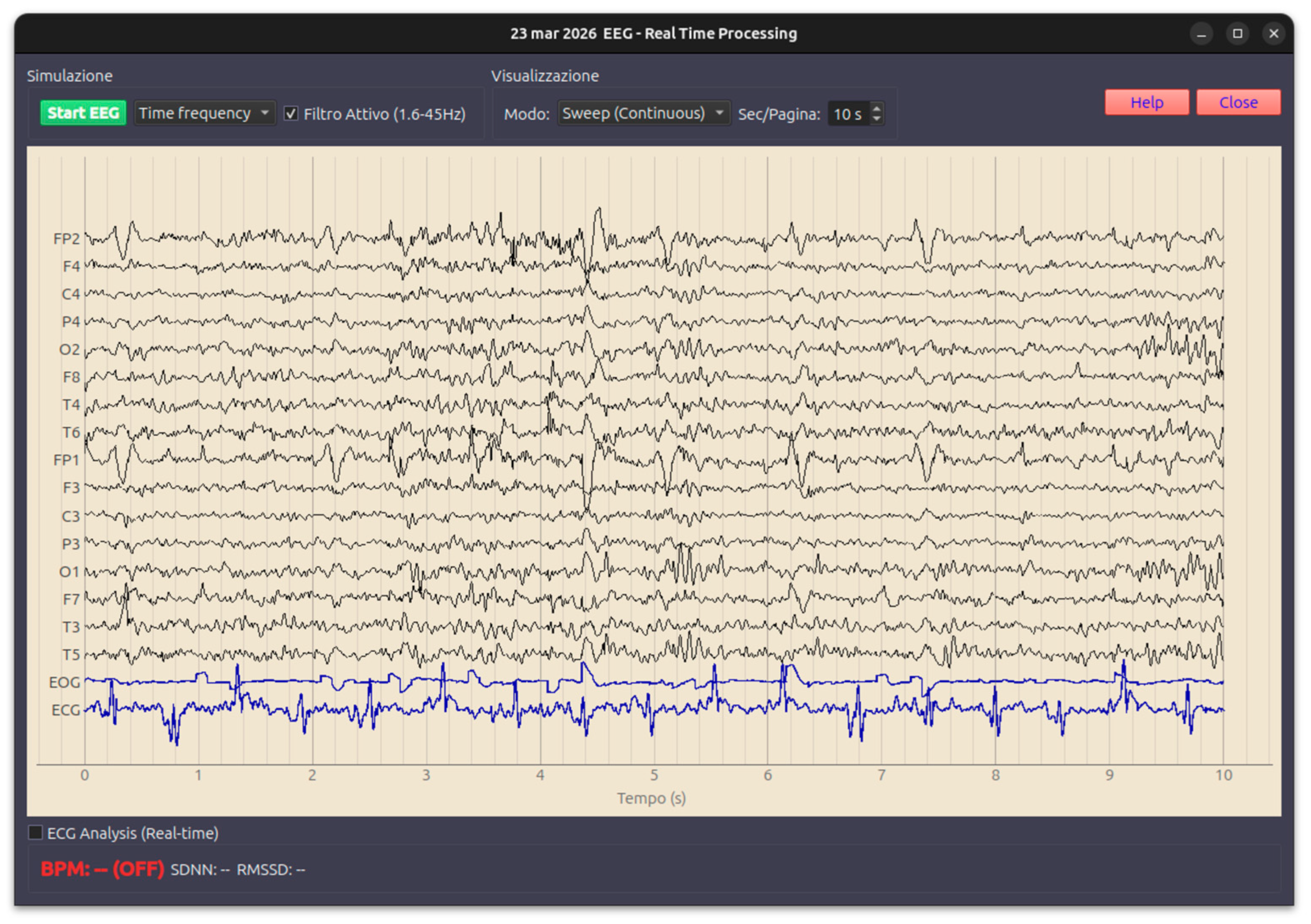
- EOG Blinks: The simulator injects vertical eye movement artifacts into the frontal channels (FP1, FP2) by scaling the loaded EOG template.
- EMG Noise: High-frequency Gaussian noise is added to all channels to simulate patient tension or muscle movement.
- Block-based PSD: Every 2 seconds, the worker triggers a Power Spectral Density (PSD) calculation using the Welch method to display the frequency distribution (Delta, Theta, Alpha, Beta bands).
- Spectral Whitening: A specific feature allows for 1/f “whitening” to flatten the power spectrum, making higher frequency oscillations like Alpha and Beta more visible against the dominant lower frequencies.
- ECG R-Peak Detection: If enabled, the system runs the analyze_r_peaks utility on the ECG channel, calculating Heart Rate (BPM) and Heart Rate Variability (SDNN, RMSSD) in real-time.
Appendix A.8 Module ECoG
- Template-Based Streaming: The simulator loads real ECoG recordings from .mat files (using scipy.io) and converts them into MNE RawArray objects.
- Worker-Timer Architecture: A dedicated ECoGWorker runs on a separate thread to prevent UI freezing. It streams data chunks at specific intervals (default 100ms) to mimic live acquisition from an amplifier.
- Circular Buffer Logic: The _run_simulation_step method manages a circular pointer over the loaded dataset, ensuring continuous data flow by wrapping back to the start once the file ends.
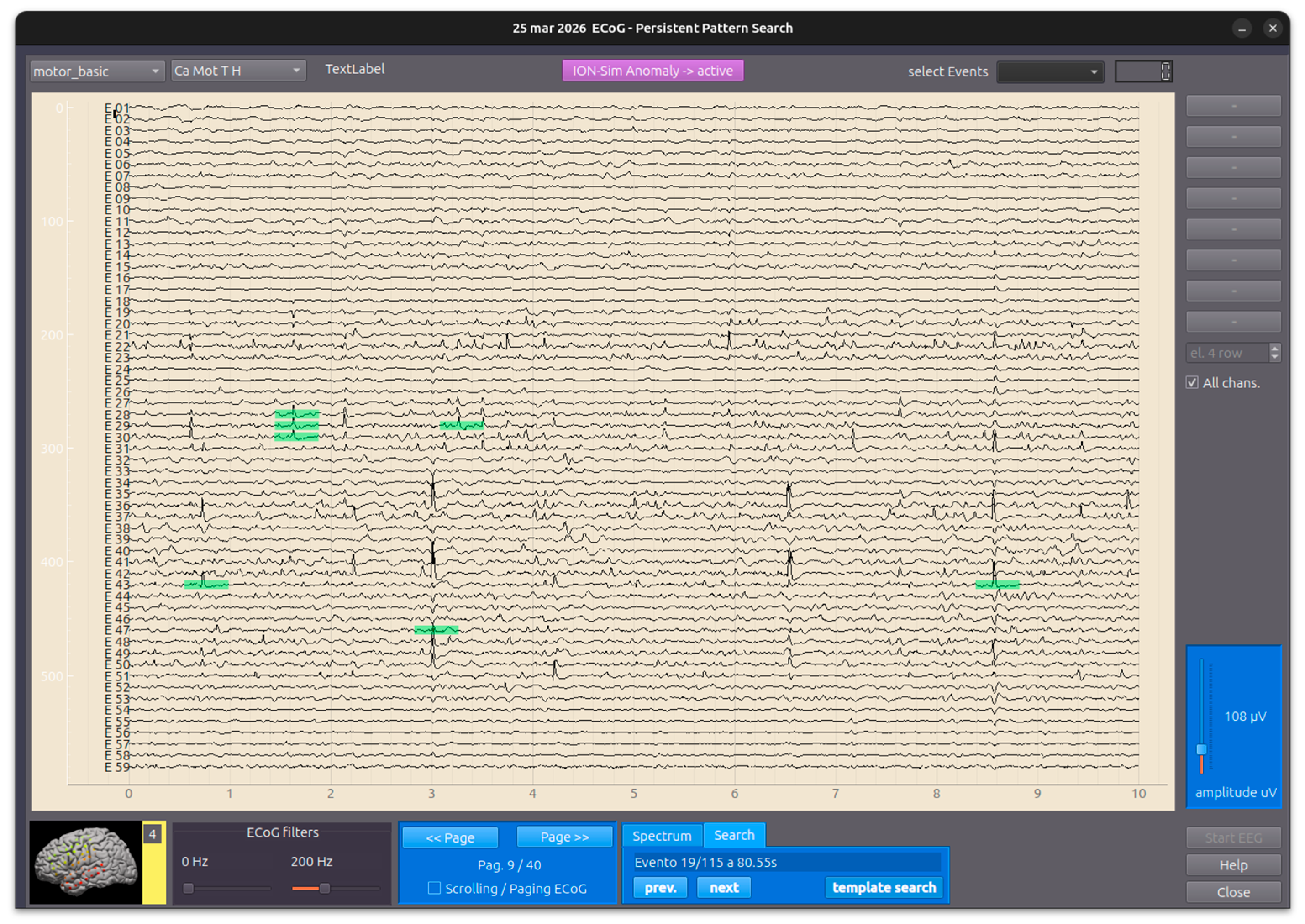
- Common Average Reference (CAR): For each chunk, the spatial mean across all electrodes is subtracted from each channel to remove global noise and artifacts common to all sensors.
- Bandpass Filtering: A dynamic Butterworth filter (applied via scipy) allows users to adjust low-cut (e.g., 2Hz) and high-cut (e.g., 300Hz) frequencies in real-time to focus on specific oscillations like Gamma or Ripple bands.
- ArtifactGenerator Integration: The simulator uses an external ArtifactGenerator class that reads configurations from a JSON file.
- Dynamic Modification: Based on the JSON “scenario,” the worker modifies the raw ECoG signal in real-time. This can include:
- Specific Channel Targeting: Anomalies can be applied to all channels or a subset.
- Time-Locked Events: Using a continuous time vector, the generator can overlay sinusoidal noise, spikes, or baseline drifts.
- ROI Selection: Users select a “Region of Interest” on the plot using a graphical UI element (LinearRegionItem).
- FFT-Based Cross-Correlation: To achieve a “100x speedup,” the code uses Fast Fourier Transforms (via the fast_normalized_cross_correlation utility) to scan the entire recording for matches that exceed a user-defined correlation threshold (e.g., 0.80).
Appendix A.9 Module Electromyography Free Running
Appendix A.9 Module Anesthesia
- Propofol (Schnider Model) [38]: Uses a three-compartment model to calculate how Propofol distributes and is eliminated based on the patient’s age and weight.
- Remifentanil (Minto Model) [39]: A specialized model for high-potency opioids, accounting for rapid onset and offset.
- Ketamine: Modeled as a sympathomimetic agent that counteracts the depressive effects of other drugs on the cardiovascular system.
- Sevoflurane: Simulates inhalation anesthesia measured in MAC (Minimum Alveolar Concentration).
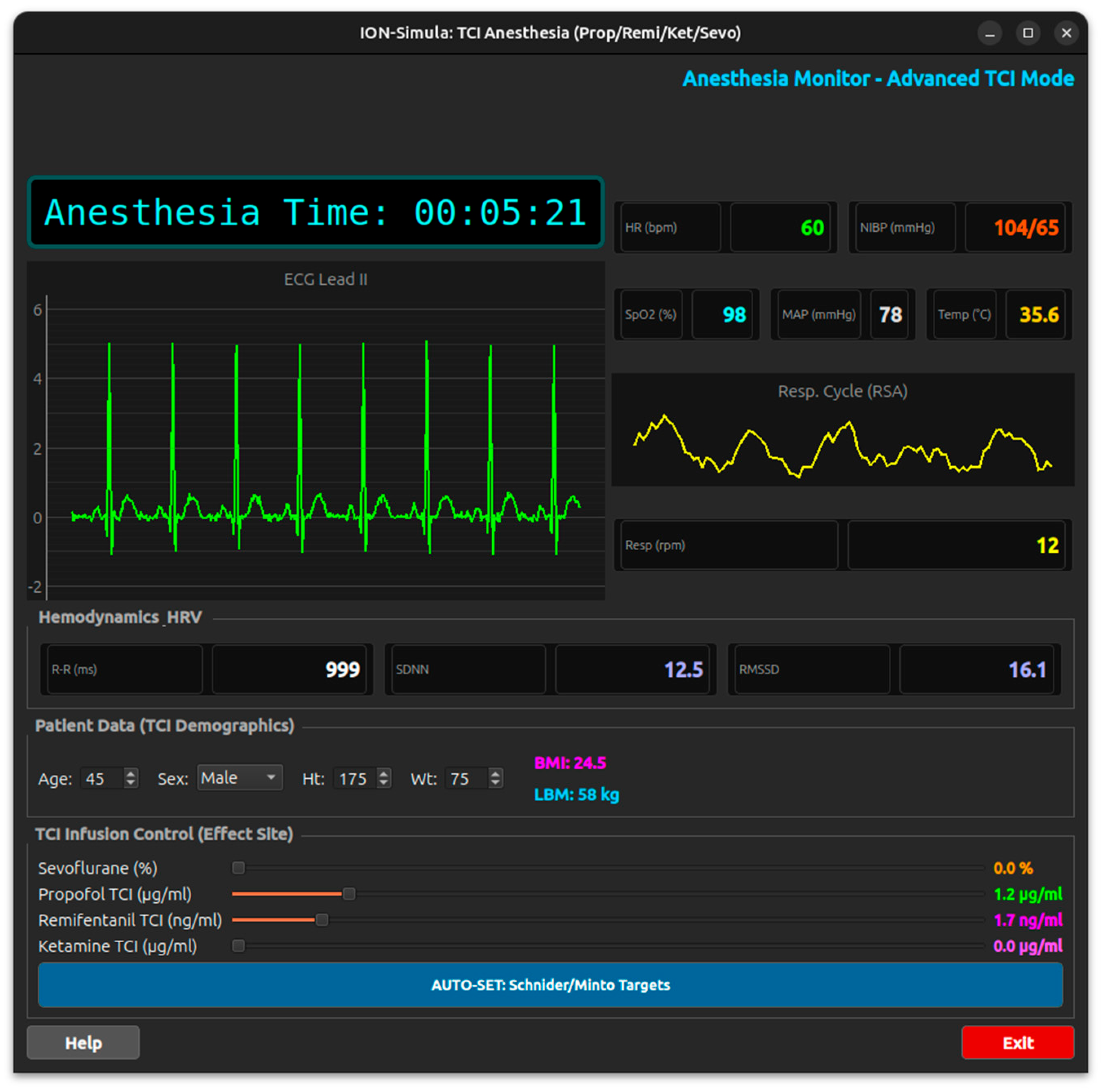
- Synergy vs. Antagonism: It calculates a global depressive effect from Propofol, Remifentanil, and Sevo. This combined effect reduces Heart Rate and MAP.
- Ketamine Offset: Ketamine is programmed with a “sympathomimetic” factor. As the Ketamine concentration increases, it mathematically offsets the bradycardia and hypotension caused by Propofol, raising HR and MAP back toward baseline.
- ECG Waveform: A synthetic ECG signal is generated by concatenating a baseline with a “QRS complex” (a high-frequency triangle wave). The frequency of these complexes is tied directly to the calculated instantaneous Heart Rate.
- Respiratory Sinus Arrhythmia (RSA): The simulator introduces small, natural fluctuations in Heart Rate to mimic the interaction between breathing and the heart.
- HRV Metrics: The code calculates SDNN (Standard Deviation of NN intervals) and RMSSD in real-time. These metrics decrease as the depth of anesthesia increases, providing a neurophysiological indicator of autonomic nervous system depression.
- MAP Calculation: The simulator approximates Mean Arterial Pressure (MAP) using the drug concentrations and a random walk component to simulate blood pressure stability.
- Real-time Controls: Users can adjust drug dosages via sliders, and the simulation immediately updates the decay curves (Ce) and the resulting physiological waveforms.
- Visualization: It uses pyqtgraph to provide a “Vital Signs Monitor” view (ECG) alongside a “Pharmacokinetic” view (Concentration curves), bridging the gap between drug administration and physiological response.
Appendix B
References
- Loftus, C. M.; Biller, J.; Baron, E. M. (Eds.) Intraoperative Neuromonitoring; McGraw-Hill Education Medical: New York, NY, 2014. [Google Scholar]
- Deletis, V.; Shils, J. L.; Sala, F.; Seidel, K. (Eds.) Neurophysiology in Neurosurgery: A Modern Approach, 2ª ed.; Academic Press (Elsevier), 2020. [Google Scholar]
- Verst, S. M.; Barros, M. R.; Maldaun, M. V. C. (Eds.) Intraoperative Monitoring: Neurophysiology and Surgical Approaches; Springer Nature Switzerland AG, 2022. [Google Scholar]
- https://www.st-andrews.ac.uk/~wjh/neurosim/.
- https://apps.microsoft.com/detail/9nc7xnpkmszm?hl=en-US&gl=US.
- https://www.st-andrews.ac.uk/~wjh/neurosim/modules.html.
- Heitler, B. Neurosim: Some Thoughts on Using Computer Simulation in Teaching Electrophysiology. [CrossRef] [PubMed]
- Newman, M. H.; Newman, E. A. MetaNeuron: A Free Neuron Simulation Program for Teaching Cellular Neurophysiology. J Undergrad Neurosci Educ 2013, 12, A11–A17. [Google Scholar]
- https://github.com/lrkrol/SEREEGA.
- https://lrkrol.com/tools.html.
- https://ebrains.eu/data-tools-services/tools?filter=whole-brain-simulation&page=12.
- https://www.thevirtualbrain.org/tvb/zwei/neuroscience-simulation.
- https://wiki.ebrains.eu/bin/view/Collabs/documentation/tutorials/The%20Virtual%20Brain/.
- https://www.ni.com/en/support/downloads/tools-network/download.labview-biomedical-toolkit.html.
- https://www.ni.com/docs/en-US/bundle/labview-biomedical-toolkit-api-ref/page/lvbiomed/bio_sim_eeg.html.
- https://stalbergsoftware.com/emgsimulators.aspx.
- https://stalbergsoftware.com/mncsimulators.aspx.
- https://www.ni.com/docs/en-US/bundle/labview-biomedical-toolkit-api-ref/page/lvbiomed/bio_sim_emg.html.
- https://www.st-andrews.ac.uk/~wjh/neurosim/.
- https://www.adinstruments.com/research/human/neuro/stimulation.
- https://www.adinstruments.com/support/downloads/windows/visual-evoked-potential-vep-%E2%80%93-pre-lab-prep.
- https://www.physoc.org/magazine-articles/new-virtual-adventure-in-physiology-practicals/.
- https://iworx.com/labscribe/.
- https://iworx.com/products/psychological-physiology/pk-tr-psychological-physiology-roam-teaching-kit/.
- https://cadwell.education/biomedical-training/.
- https://www.cadwell.com/clinical-solutions/electrodiagnostics/evoked-potentials/.
- Goetz, S. M., Madhi Alavi, S. M., Deng, Zhi-De, Peterchev, A. V. Statistical Model of Motor Evoked Potentials for Simulation of Transcranial Magnetic and Electric Stimulation . [CrossRef]
- https://github.com/lrkrol/SEREEGA.
- Nunes, P. L.; Nunes, M. D.; Srinivasan, R. Multi-Scale Neural Sources of EEG: Genuine, Equivalent, and Representative. A Tutorial Review Brain Topography 2019, 32, 193–214. [Google Scholar] [CrossRef] [PubMed]
- https://www.inomed.com/.
- https://neurosoft.com/en/catalog/iom.
- https://simtk.org/projects/opensim.
- https://surgeonslab.com/.
- https://www.insimo.com/.
- https://www.ibm.com/think/topics/digital-twin.
- Contento, G., Budai R., Characterisation of the Conduction Properties of Nerves with the Distribution of Fibre Conduction Velocities, in Bioinstrumentation and Biosensors, ed. D.L. Wise, Marcel Dekker, Inc., New York, 1991.
- Contento, G., et al., Conduction Velocity Distributions of Normal and Pathological Fibres in the same Nerve, Proc. VIII Annual Conf. of the IEEE Engineering in Medicine and Biology Society, Fort Worth, 1986.
- Linassi, F.; Zanatta, P.; Spano, L.; Burelli, P.; Farnia, A.; Carron, M. Schnider and Eleveld Models for Propofol Target-Controlled Infusion Anesthesia: A Clinical Comparison Life (Basel) 2023, 13(10), 2065. [CrossRef]
- Eleveld, D. J.; ColinAnthony, P.; Absalom, R.; Struys, M. M.R.F. Target-controlled-infusion models for remifentanil dosing consistent with approved recommendations. British Journal of Anaesthesia 2020, 125, 483–491. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



