Submitted:
15 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Sources
2.2. Technology Selection through Decision Matrices
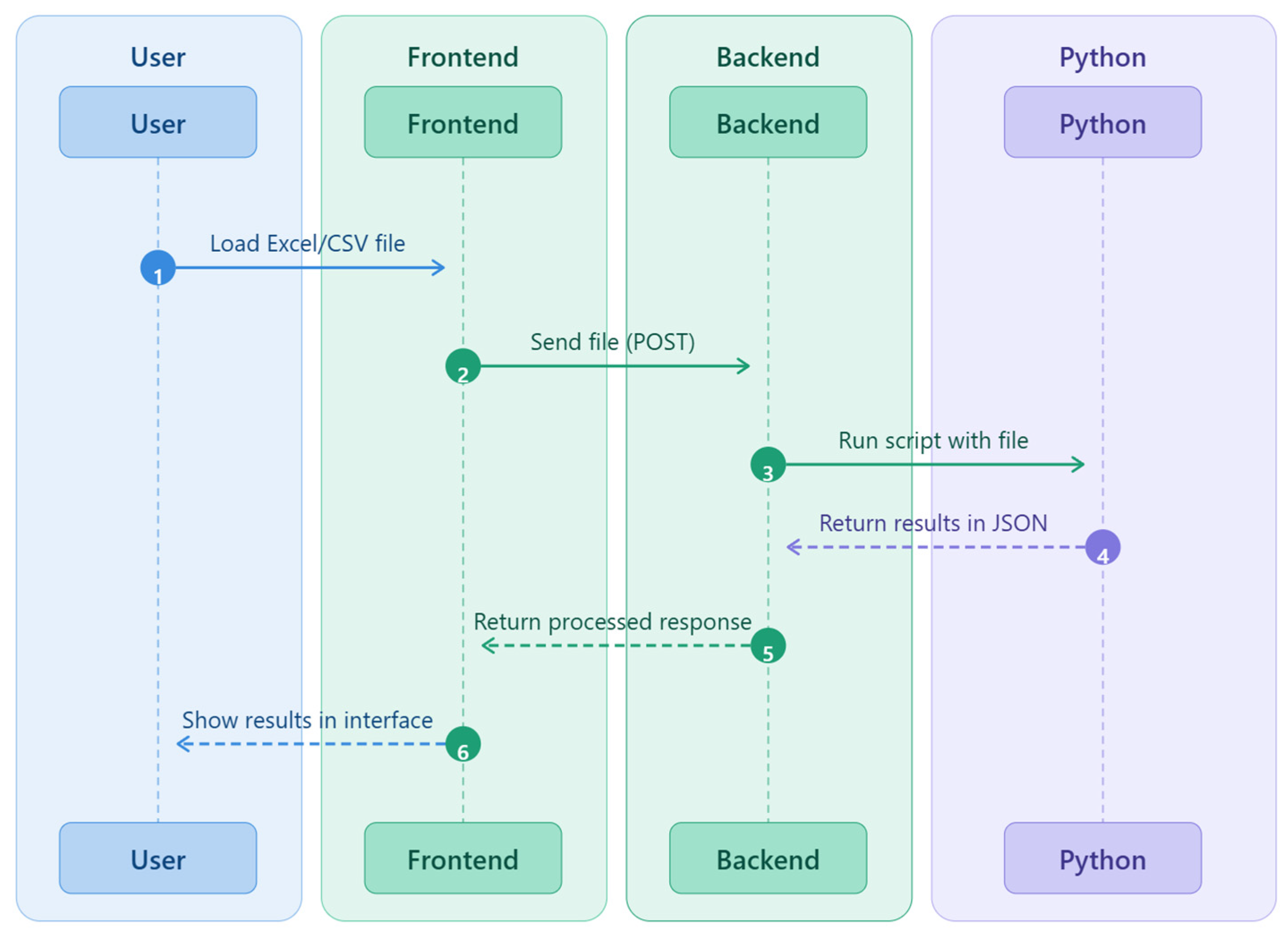
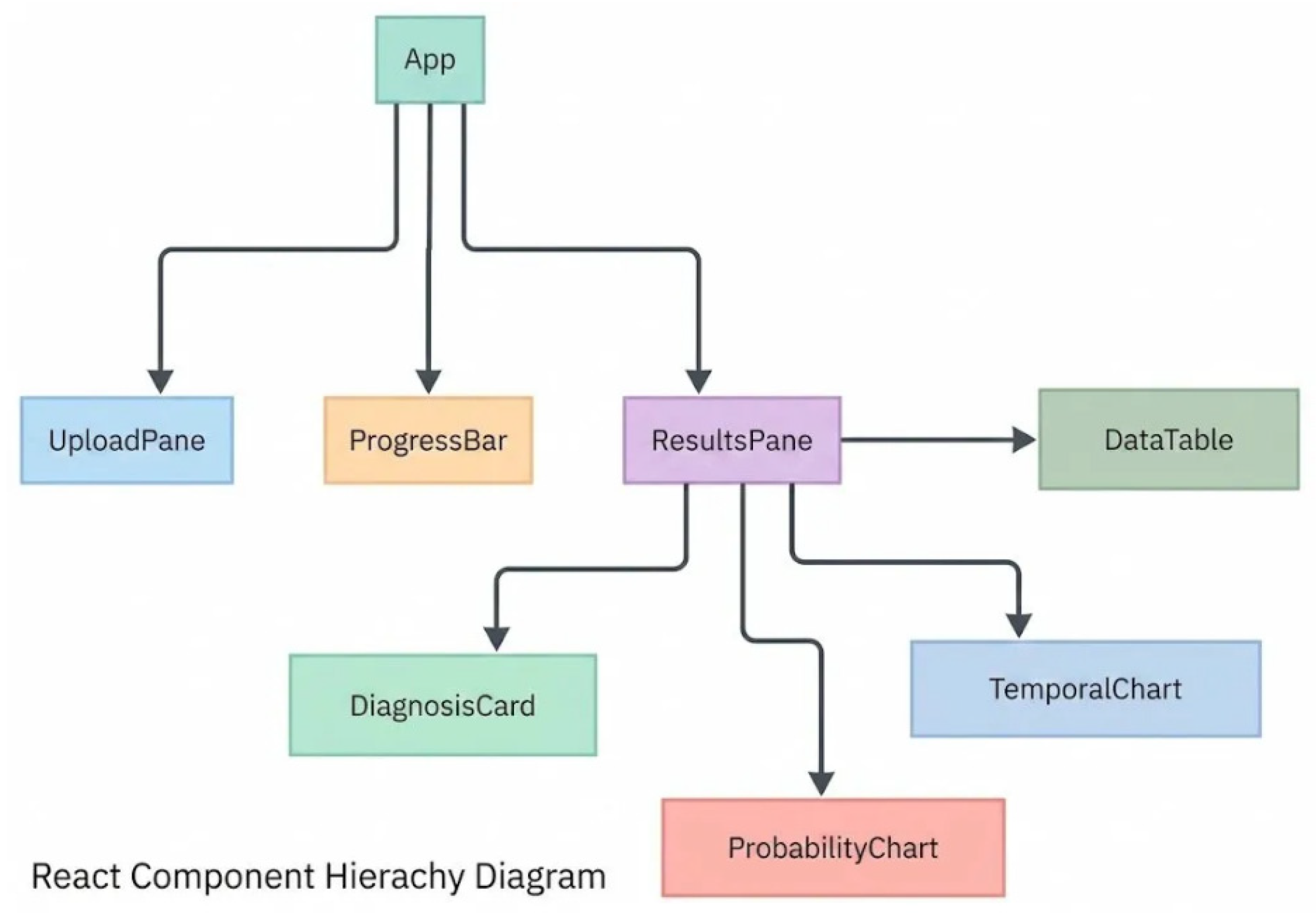
2.3. Software Architecture
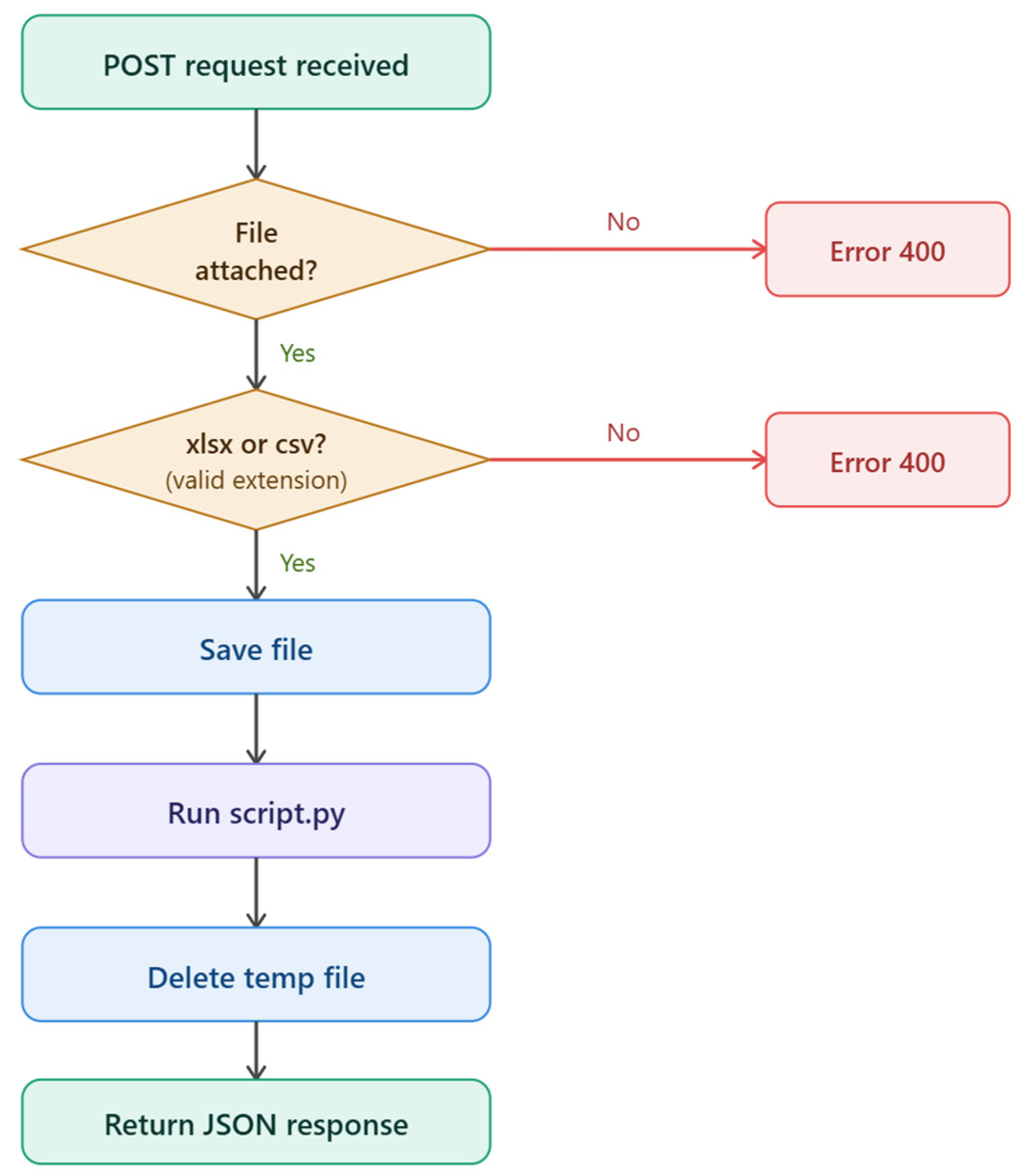
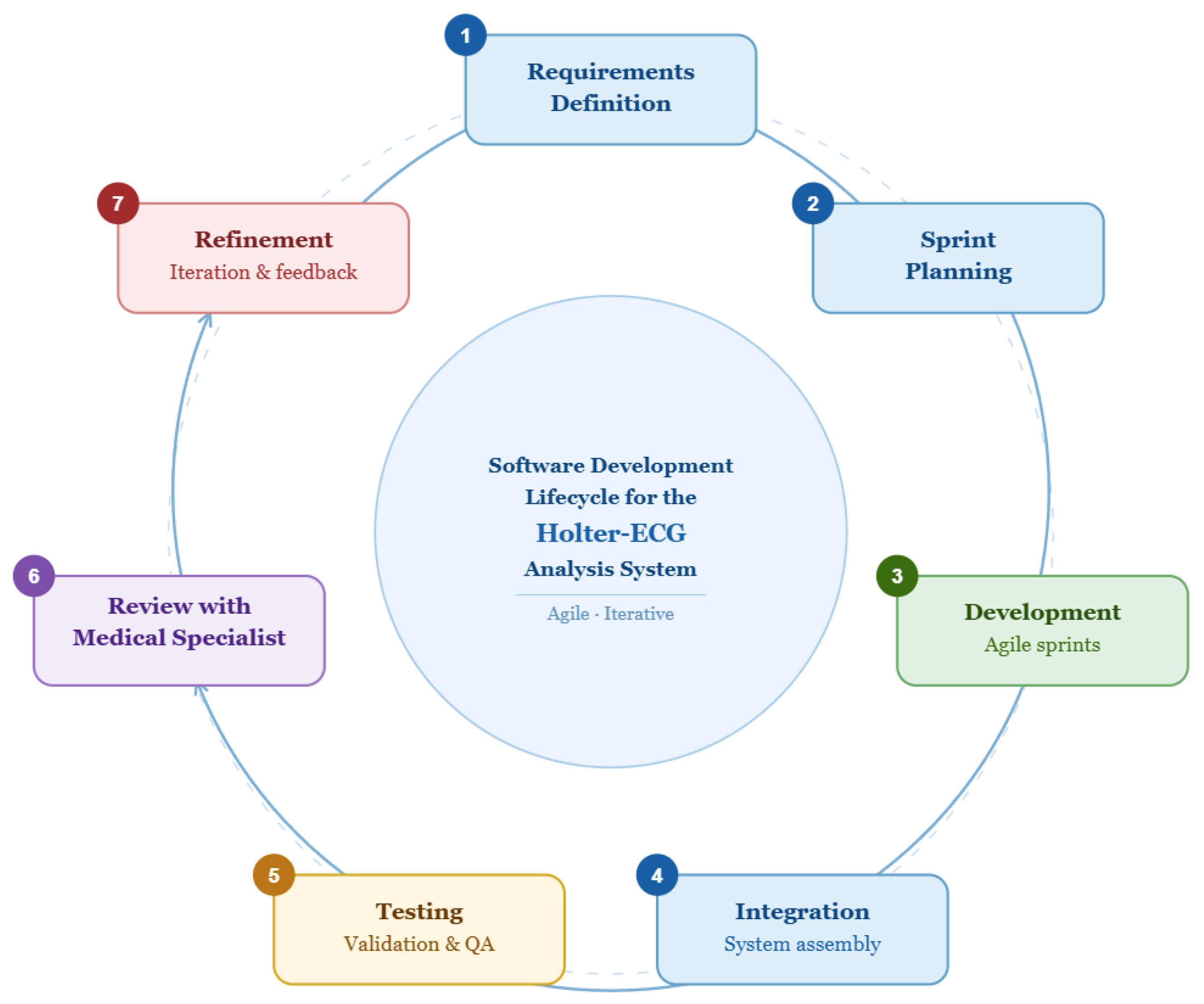
2.4. Software Development
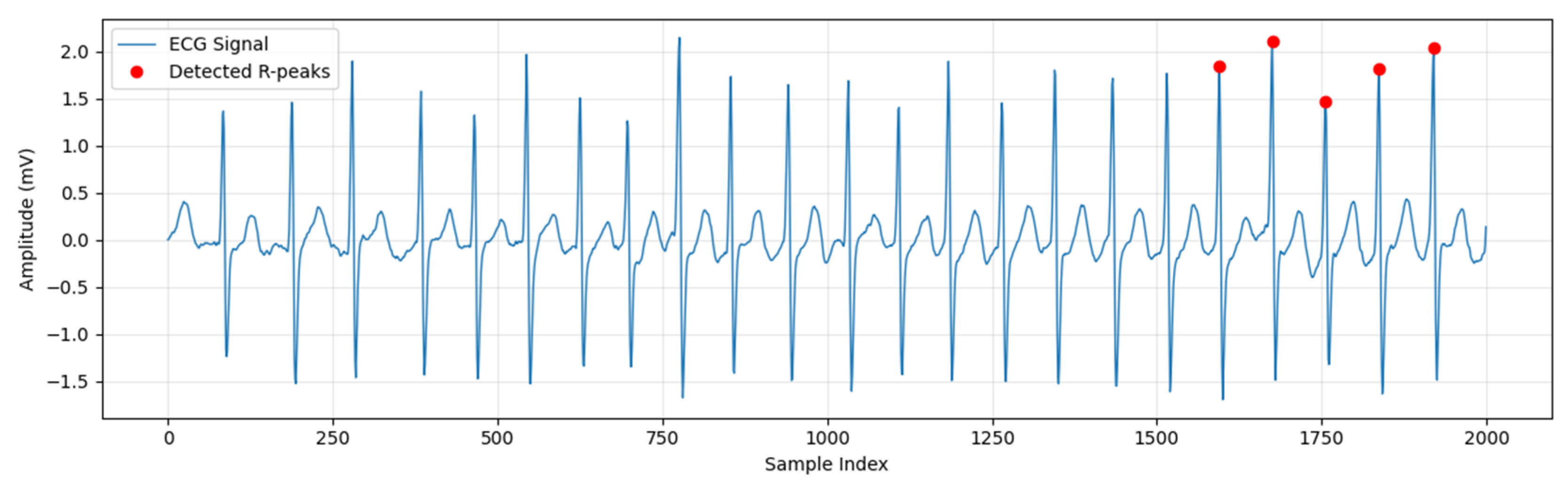
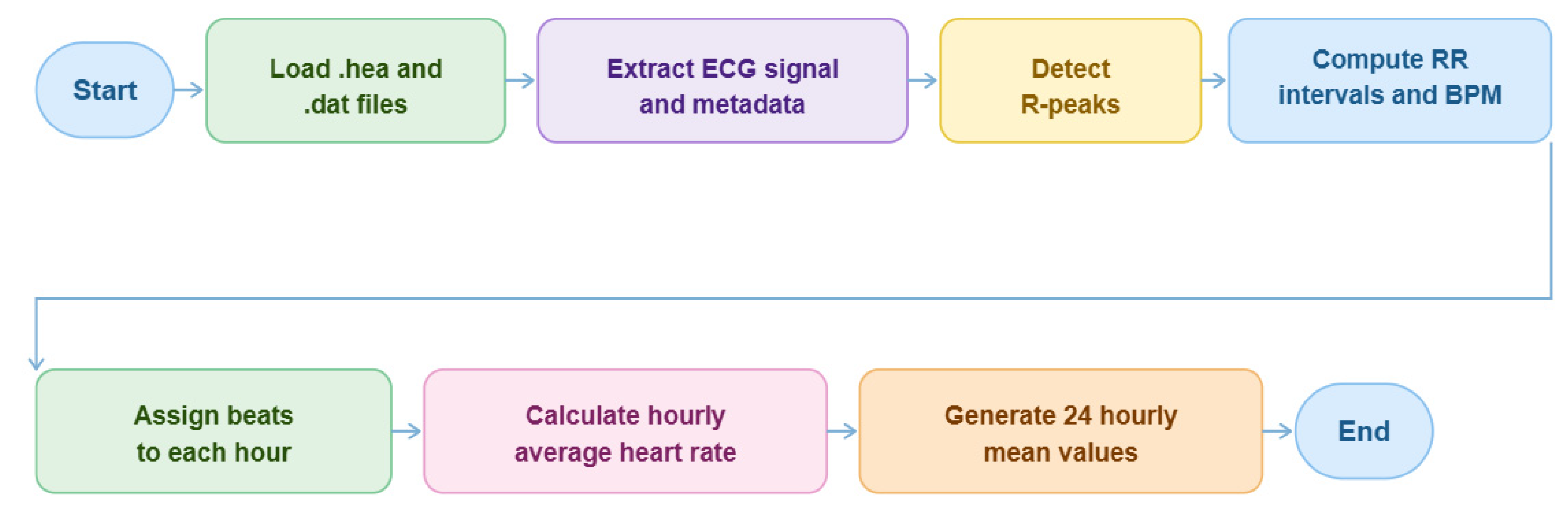
2.5. Raw ECG Signal Preprocessing
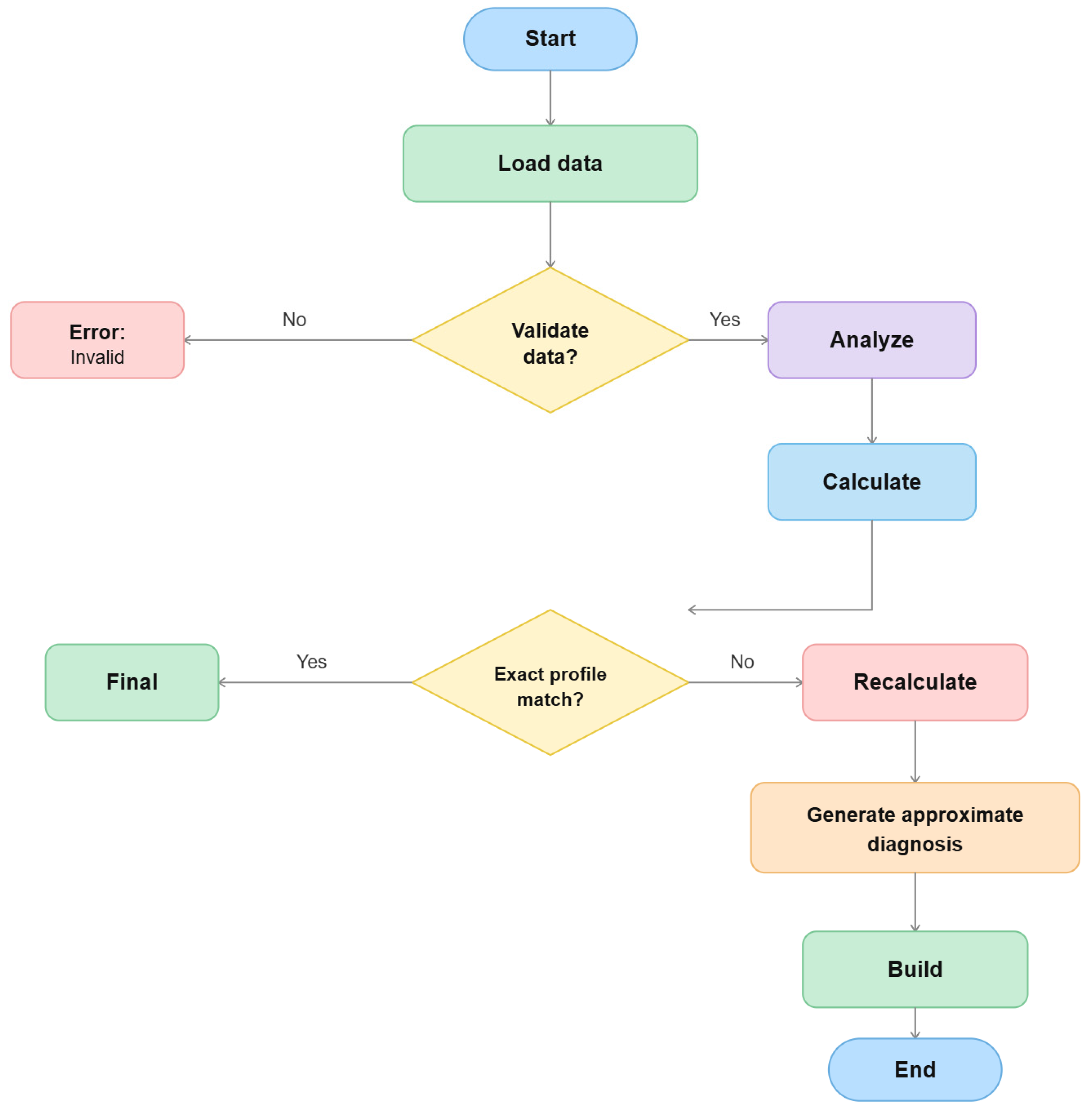
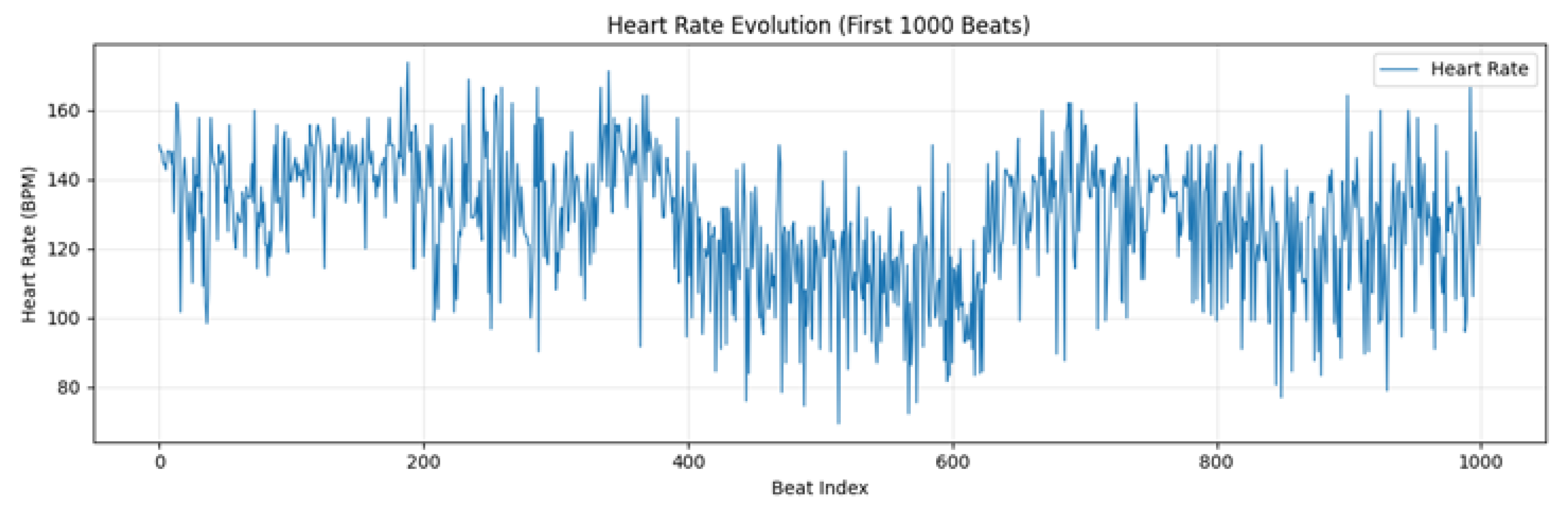
2.6. Probabilistic Analysis Framework
3. Results
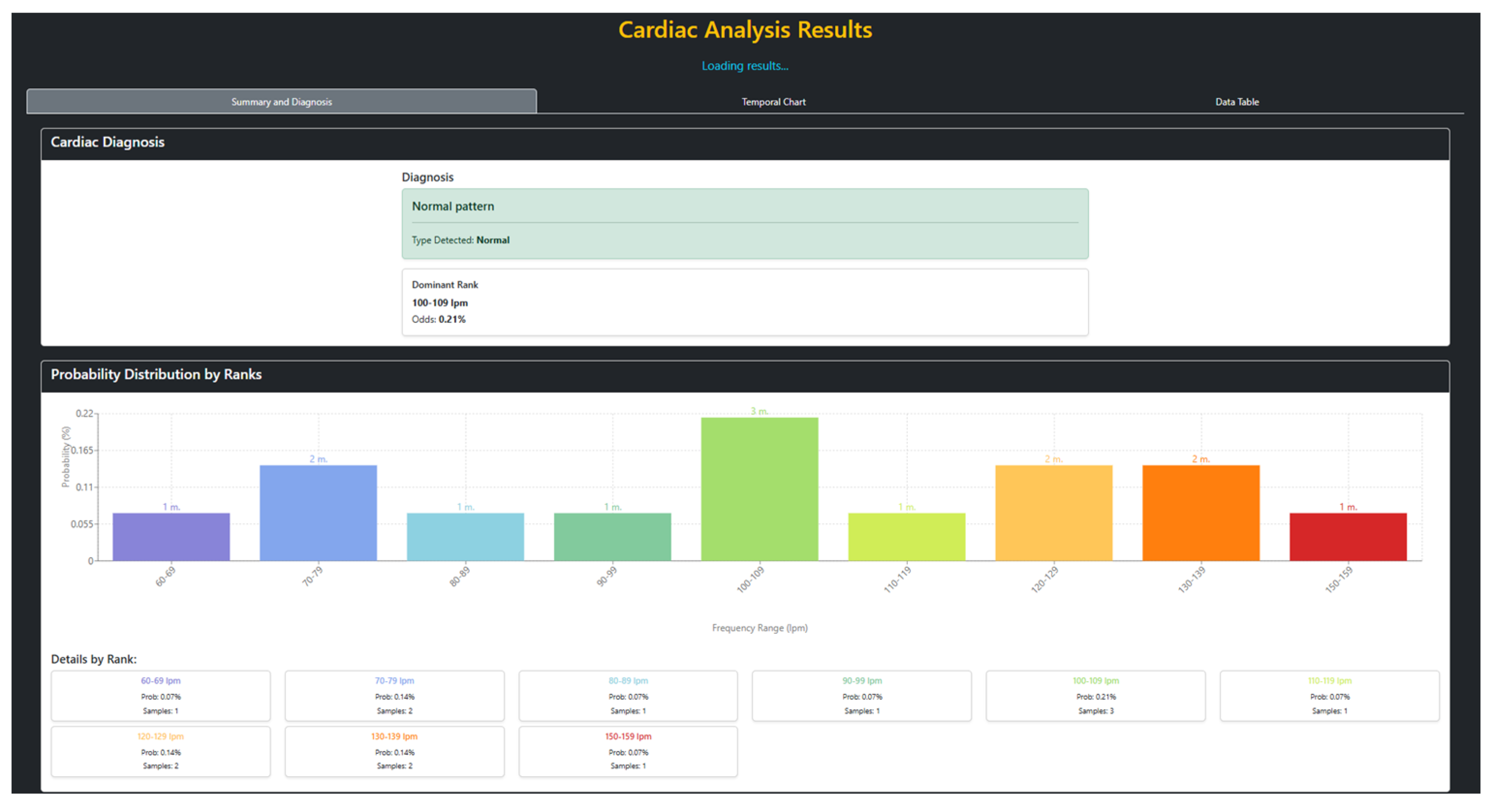
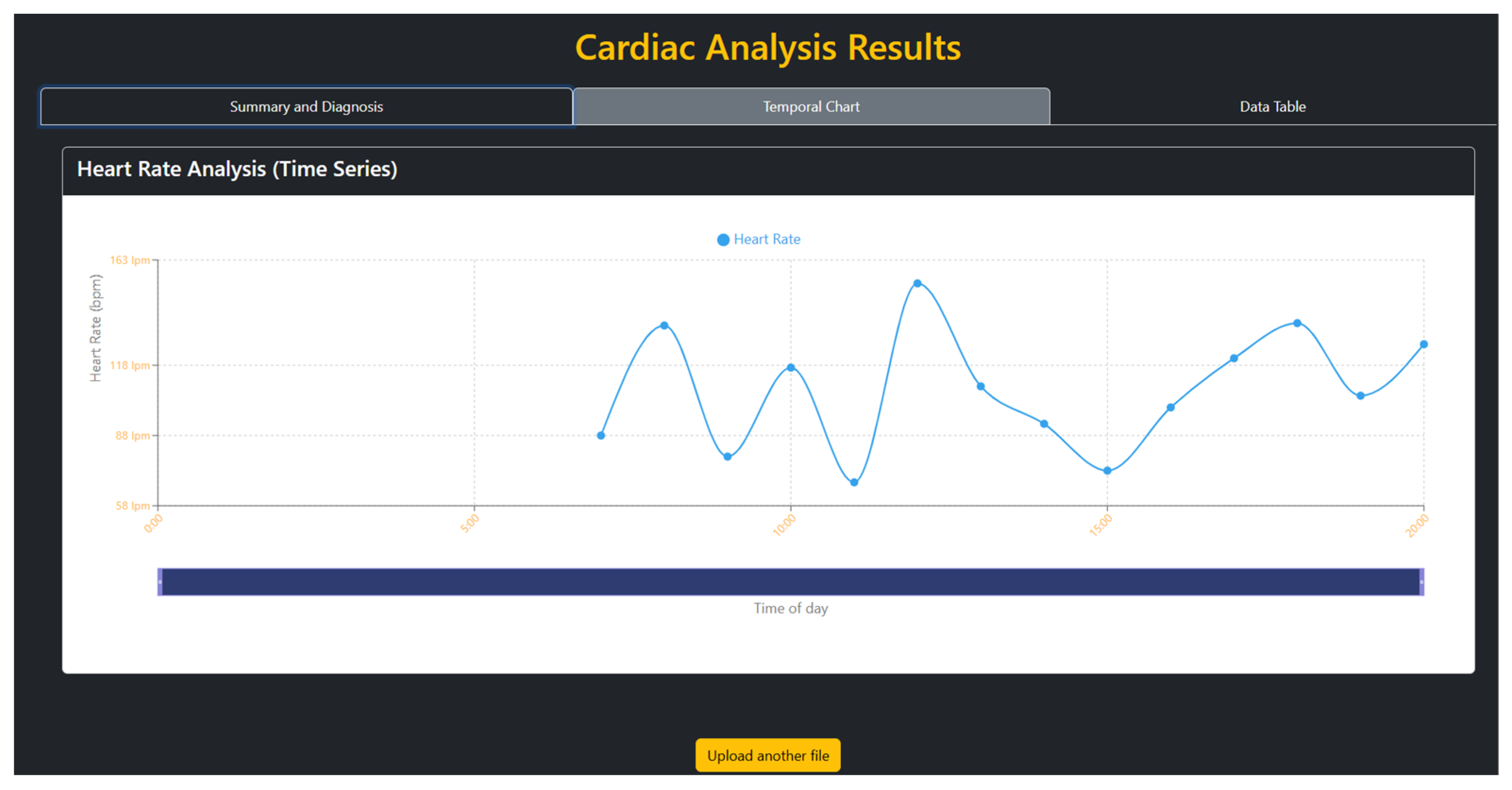
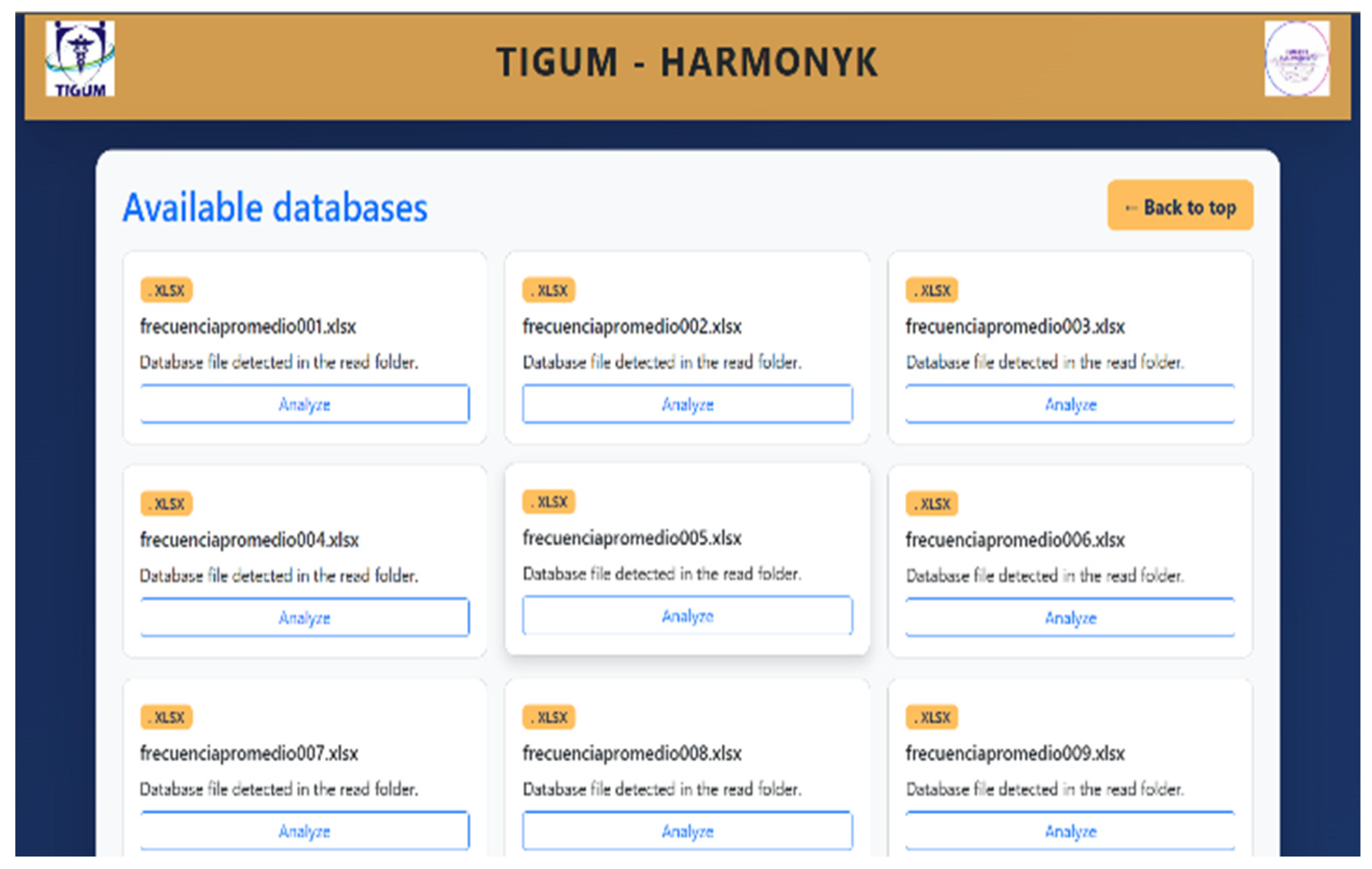
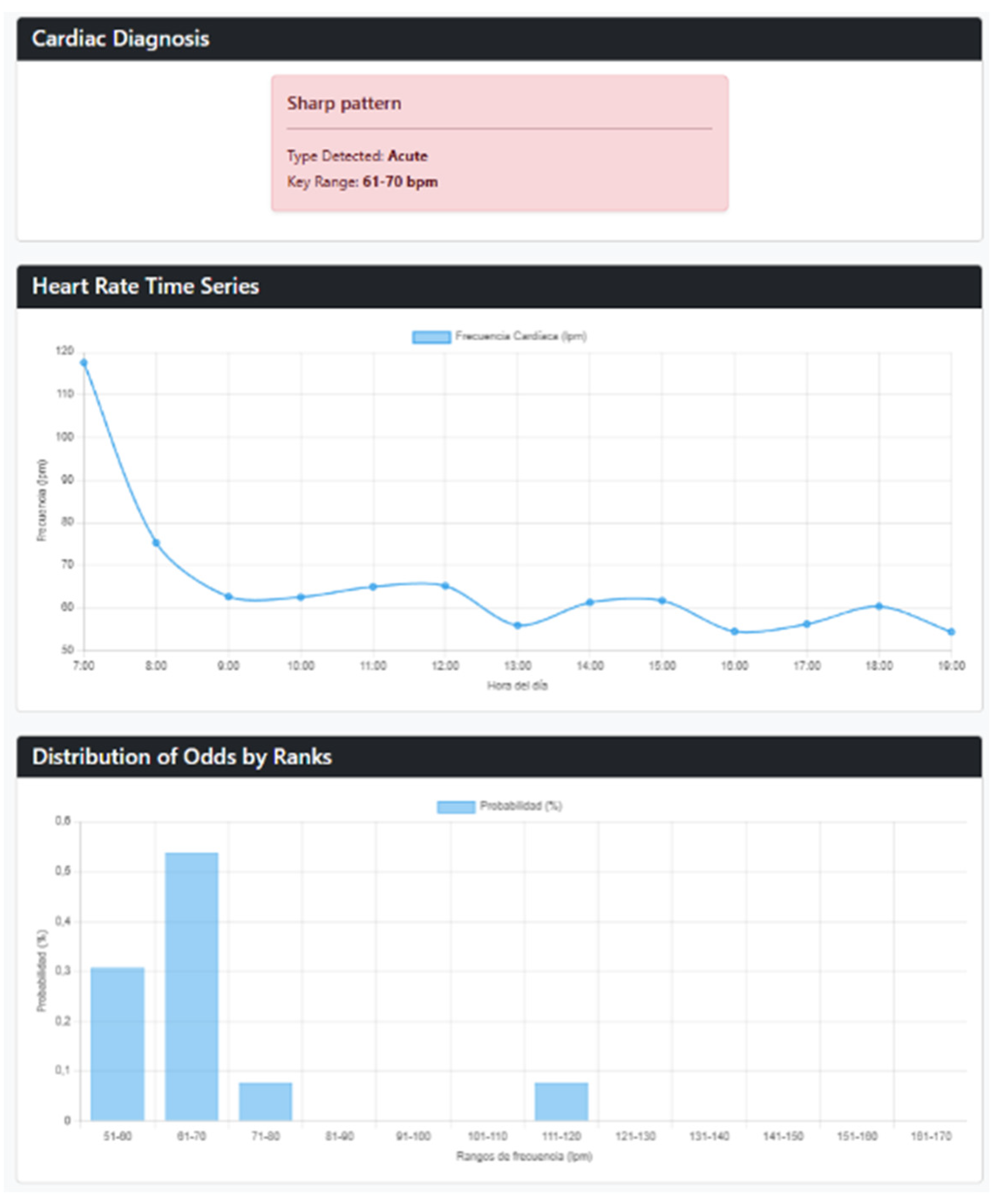
3.1. Web Interface and Execution Results
3.2. Database-Based Validation Results
4. Discussion
5. Conclusions
6. Patents
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- World Health Organization. Cardiovascular diseases (CVDs). Available online: https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases-(cvds) (accessed on 11 April 2026).
- Horsky, J.; Schiff, G.D.; Johnston, D.; Mercincavage, L.; Bell, D.; Middleton, B. Interface design principles for usable decision support: A targeted review of best practices for clinical prescribing interventions. J. Biomed. Inform. 2012, 45, 1202–1216.
- Bayor, A.A.; Li, J.; Yang, I.A.; Varnfield, M. Designing Clinical Decision Support Systems (CDSS)—A User-Centered Lens of the Design Characteristics, Challenges, and Implications: Systematic Review. J. Med. Internet Res. 2025, 27, e63733.
- Brito, J.J.; Li, J.; Moore, J.H.; Greene, C.S.; Nogoy, N.A.; Garmire, L.X.; Mangul, S. Recommendations to enhance rigor and reproducibility in biomedical research. GigaScience 2020, 9, giaa056.
- Sattar, Y.; Chhabra, L. Holter Monitor. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2025.
- Giada, F.; Bartoletti, A. Value of ambulatory electrocardiographic monitoring in syncope. Cardiol. Clin. 2015, 33, 361–366.
- Diemberger, I.; Martignani, C.; Biffi, M.; Boriani, G. Holter ECG for pacemaker/defibrillator carriers: What is its role in the era of remote monitoring? Heart 2015, 101, 1272–1278.
- Goldberger, A.L.; West, B.J. Fractals in physiology and medicine. Yale J. Biol. Med. 1987, 60, 421–435.
- Goldberger, A.L.; Amaral, L.A.N.; Hausdorff, J.M.; Ivanov, P.C.; Peng, C.-K.; Stanley, H.E. Fractal dynamics in physiology: Alterations with disease and aging. Proc. Natl. Acad. Sci. USA 2002, 99, 2466–2472.
- Huikuri, H.V.; Mäkikallio, T.H.; Peng, C.-K.; Goldberger, A.L.; Hintze, U.; Møller, M. Fractal correlation properties of R-R interval dynamics and mortality in patients with depressed left ventricular function after an acute myocardial infarction. Circulation 2000, 101, 47–53.
- Rodríguez, J.O. Entropía proporcional de los sistemas dinámicos cardiacos: Predicciones físicas y matemáticas de la dinámica cardiaca de aplicación clínica. Rev. Colomb. Cardiol. 2010, 17, 115–129.
- Rodríguez, J.; Correa, C.; Ortiz, L.; Prieto, S.; Bernal, P.; Ayala, J. Evaluación matemática de la dinámica cardiaca con la teoría de la probabilidad. Rev. Mex. Cardiol. 2009, 20, 183–189.
- Fatovich, D.M.; Phillips, M. The probability of probability and research truths. Emerg. Med. Australas. 2017, 29, 242–244.
- Upshur, R.E.G. A short note on probability in clinical medicine. J. Eval. Clin. Pract. 2013, 19, 463–466.
- Velásquez, J.R.; Ramírez López, L.J.; Torres, S.G. New Predictive Diagnostic Method for Cardiac Dynamics Based on Probability Distributions. Diagnostics 2025, 15, 650.
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.-K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, e215–e220.
- Tsutsui, K.; Biton Brimer, S.; Ben-Moshe, N.; Sellal, J.M.; Oster, J.; Mori, H.; Ikeda, Y.; Arai, T.; Nakano, S.; Kato, R.; et al. SHDB-AF: A Japanese Holter ECG database of atrial fibrillation. Sci. Data 2025, 12, 454.
- Jager, F.; Taddei, A.; Moody, G.B.; Emdin, M.; Antolic, G.; Dorn, R.; Smrdel, A.; Marchesi, C.; Mark, R.G. Long-term ST database: A reference for the development and evaluation of automated ischaemia detectors and for the study of the dynamics of myocardial ischaemia. Med. Biol. Eng. Comput. 2003, 41, 172–183.
- Moody, G.B. MIT-BIH Normal Sinus Rhythm Database v1.0.0. PhysioNet 1999. Available online: https://www.physionet.org/physiobank/database/nsrdb/ (accessed on 11 April 2026).
- Rodríguez Velásquez, J.R.; Oliveros Acosta, D.; Rodríguez Peña, D.S.; Sosa Pinzón, J.A.; Prieto Bohórquez, S.E.; Correa Herrera, S.C. Diagnostic methodology of cardiac dynamics based on the Zipf–Mandelbrot law: Evaluation with 50 patients. Rev. Mex. Cardiol. 2018, 29, 83–89.
- Rodríguez, J.; Oliveros, D.; Correa, C.; Prieto, S. Aplicabilidad clínica de software diagnóstico de la dinámica cardíaca basado en la Ley de Zipf-Mandelbrot. Rev. Haban. Cienc. Méd. 2019, 18, 624–633.
- Laplace, P.S. Théorie analytique des probabilités; Courcier: Paris, France, 1812.
- Kolmogorov, A.N. Grundbegriffe der Wahrscheinlichkeitsrechnung; Springer: Berlin, Germany, 1933.
- Bernoulli, J. Ars Conjectandi; Thurnisius: Basel, Switzerland, 1713.
- Mason, H.T.; Martinez-Cedillo, A.P.; Vuong, Q.C.; Garcia-de-Soria, M.C.; Smith, S.; Geangu, E.; Knight, M.I. A Complete Pipeline for Heart Rate Extraction from Infant ECGs. Signals 2024, 5, 118–146.
- Badr, A.; Badawi, A.; Rashwan, A.; Elgazzar, K. XBeats: A Real-Time Electrocardiogram Monitoring and Analysis System. Signals 2022, 3, 189–208.
| Criterion | React | Vue | Angular | Vanilla JS® |
| Graphics rendering | 5 | 4 | 3 | 2 |
| Complex state management | 5 | 4 | 5 | 1 |
| Development time | 4 | 5 | 3 | 2 |
| Bundle size | 3 | 4 | 2 | 5 |
| Ecosystem maturity | 5 | 4 | 4 | 1 |
| TOTAL | 22 | 21 | 17 | 11 |
| Criterion | PHP | Node.js | Flask | .NET |
| Legacy compatibility | 5 | 3 | 2 | 4 |
| Prototyping speed | 3 | 5 | 5 | 2 |
| Memory efficiency | 4 | 3 | 3 | 2 |
| Database connectivity | 5 | 4 | 4 | 5 |
| Cloud deployment | 4 | 5 | 5 | 3 |
| TOTAL | 21 | 20 | 19 | 16 |
| Criterion | Python® | C++ | R® | MATLAB® |
| Processing speed | 4 | 5 | 2 | 4 |
| Backend compatibility | 4 | 3 | 2 | 4 |
| Memory usage | 4 | 5 | 2 | 4 |
| Statistical functions | 5 | 5 | 5 | 1 |
| License cost | 5 | 5 | 5 | 1 |
| TOTAL | 22 | 20 | 16 | 18 |
| Hour | Average Heart Rate (BPM) | Hour | Average Heart Rate (BPM) |
| 0 | 117.57 | 12 | 54.3 |
| 1 | 75.26 | 13 | 57.81 |
| 2 | 62.61 | 14 | 54.83 |
| 3 | 62.49 | 15 | 52.07 |
| 4 | 64.93 | 16 | 53.33 |
| 5 | 65.13 | 17 | 56.07 |
| 6 | 55.85 | 18 | 50.98 |
| 7 | 61.23 | 19 | 51.19 |
| 8 | 61.66 | 20 | 56.48 |
| 9 | 54.41 | 21 | 54.53 |
| 10 | 56.14 | 22 | 61.87 |
| 11 | 60.32 | 23 | 75.84 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




