Submitted:
09 April 2026
Posted:
10 April 2026
You are already at the latest version
Abstract
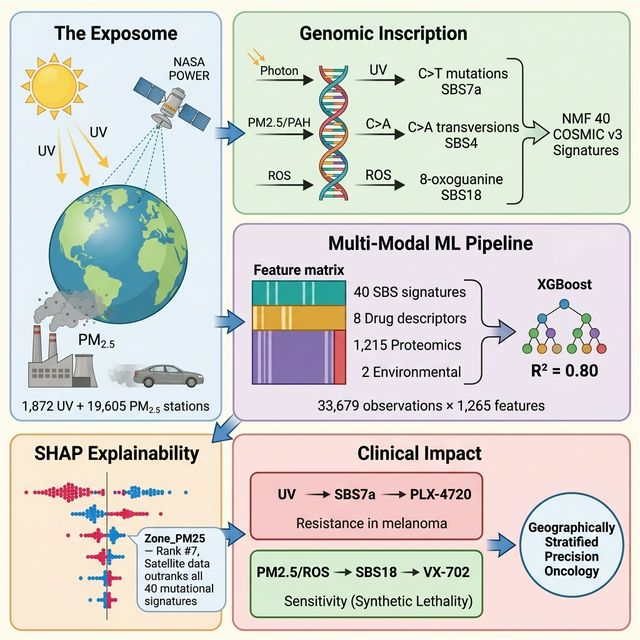
Keywords:
1. Introduction
2. Materials and Methods
2.1. Drug Sensitivity Data
2.2. Mutational Signature Features
2.3. Drug Scaffold Chemistry Features
2.4. Proteomic and Epigenetic Features
2.5. Environmental Satellite Features: Global Kriging Raster
2.5.1. Data Sources
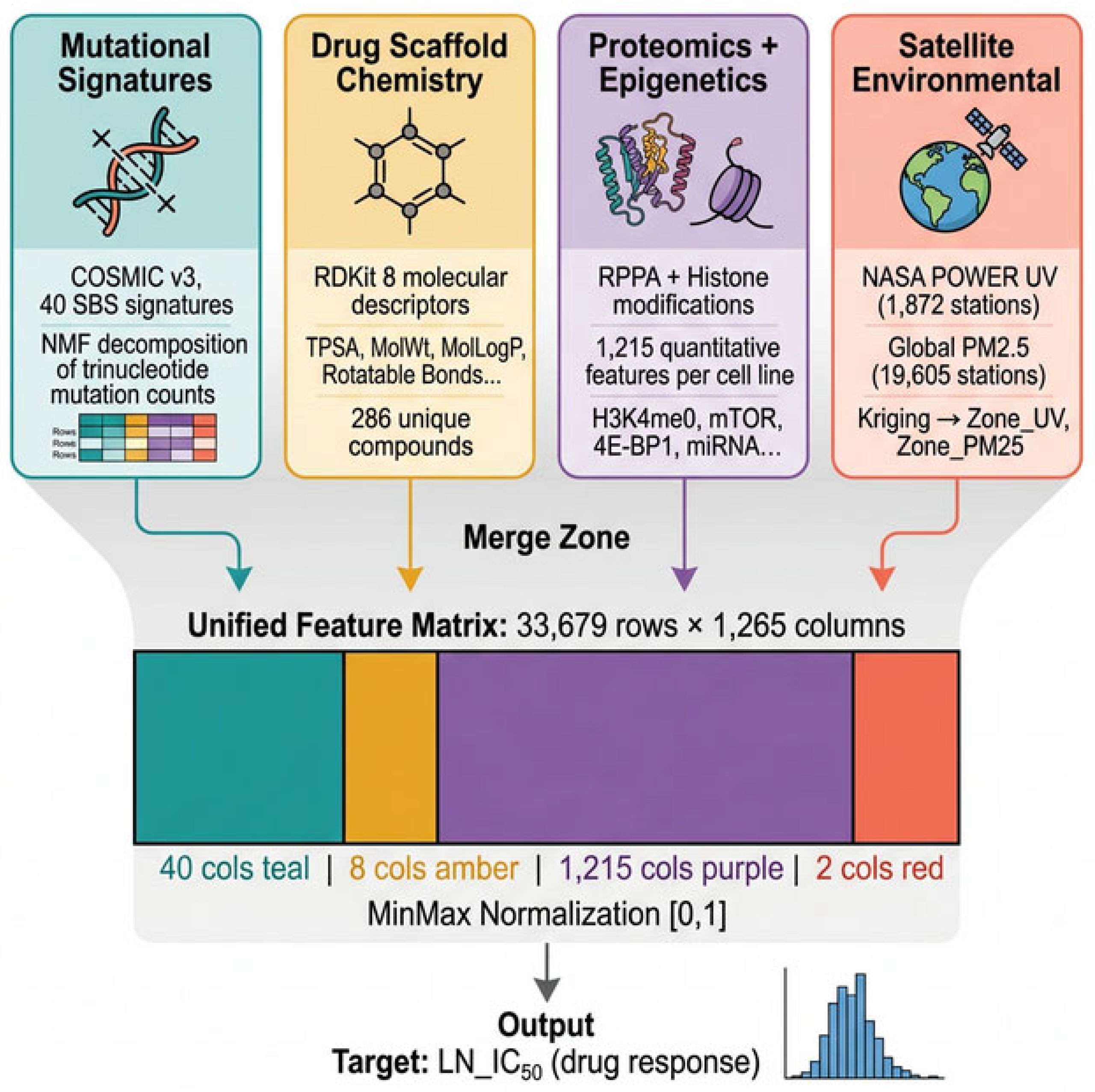
2.5.2. Spatial Interpolation (Kriging)
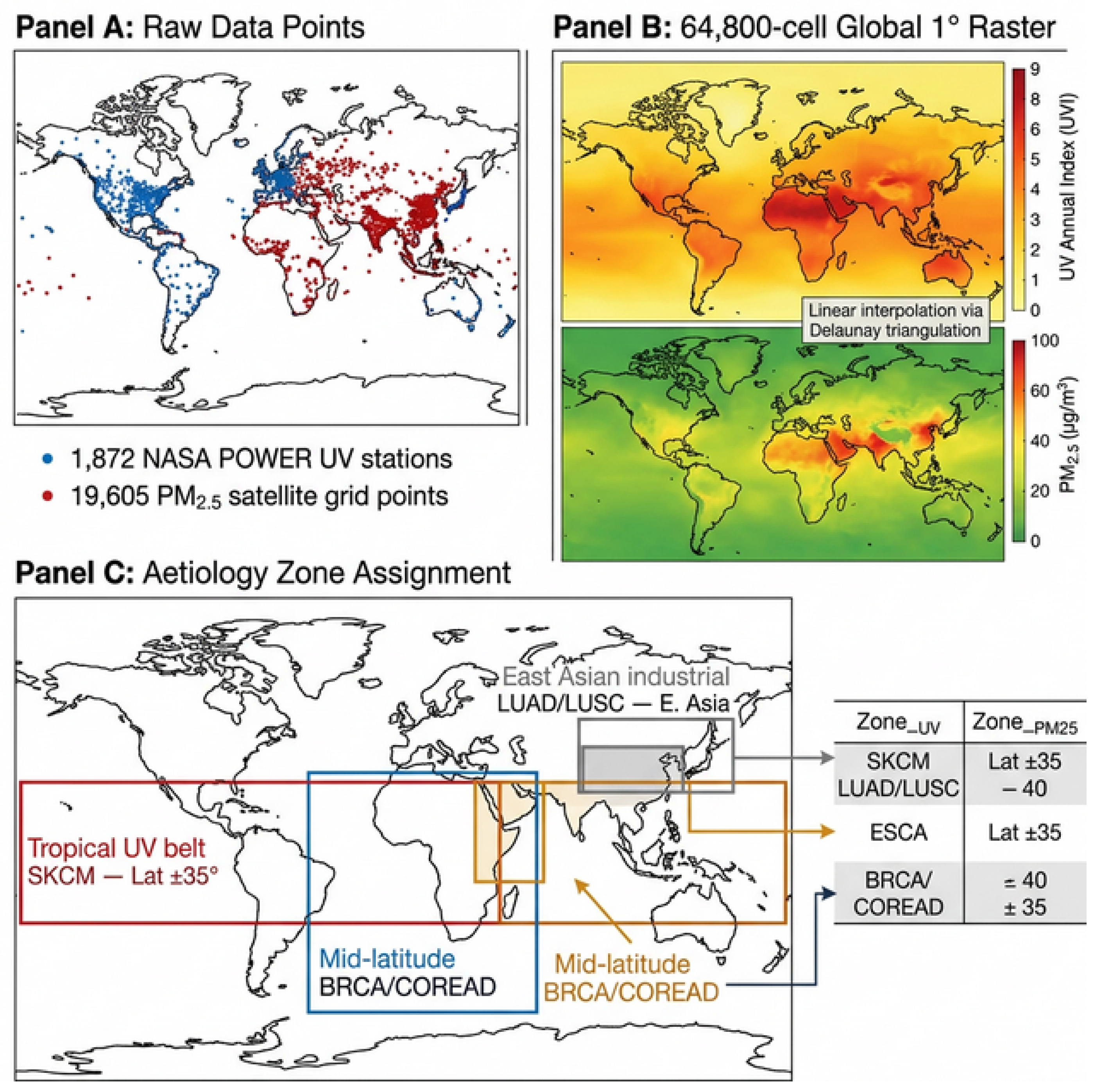
2.5.3. Cancer-Type–Environment Assignment
| TCGA | Cancer Type | Bounding Box | Rationale |
| SKCM | Cutaneous melanoma | Lat −35 to 35, all Lon | Tropical UV belt |
| LUAD | Lung adenocarcinoma | Lat 20–55 N, Lon 70–130 E | E. Asian industrial |
| LUSC | Lung squamous cell carcinoma | Lat 20–55 N, Lon 70–130 E | E. Asian industrial |
| ESCA | Oesophageal | Lat 5–40 N, Lon 25–80 E | E. Africa/C. Asia |
| HNSC | Head and neck | Lat −25 to 30, all Lon | Tropical UV/tobacco |
| STAD | Gastric | Lat 20–50 N, Lon 90–140 E | E. Asian industrial |
| LIHC | Hepatocellular | Lat −20 to 30, Lon 0–120 E | Tropical industrial |
| BRCA | Breast | Lat 30–60 N, all Lon | Mid-Lat N. Hemisphere |
| COREAD | Colorectal | Lat 30–55 N, all Lon | Mid-Lat N. Hemisphere |
| BLCA | Bladder | Lat 30–55 N, all Lon | Mid-Lat N. Hemisphere |
| UCEC | Uterine endometrial | Lat 30–60 N, Lon −100 to 50 | N. Hemisphere mixed |
2.6. Feature Matrix and Preprocessing
2.7. Model Training
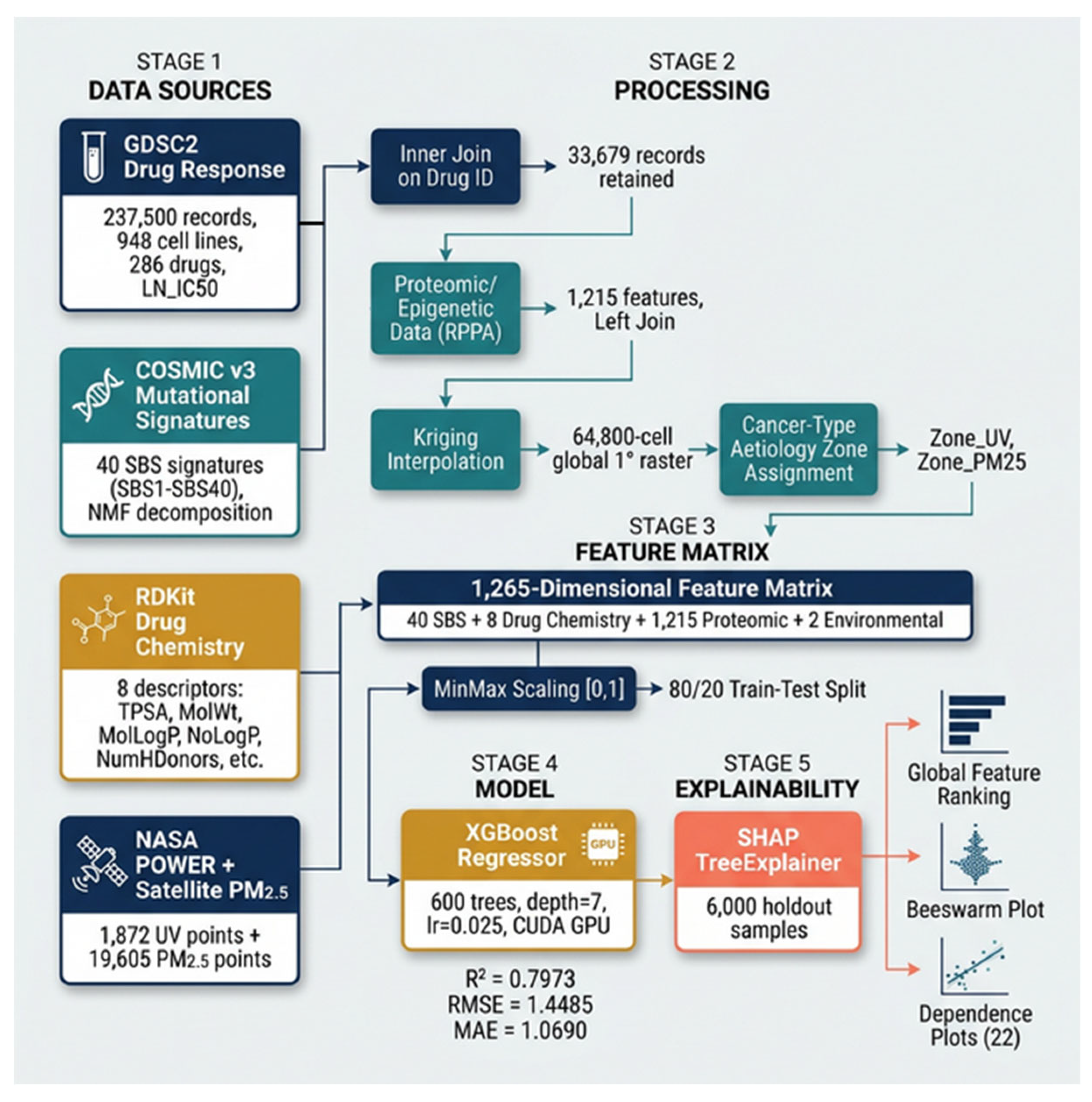
2.8. SHAP Explainability Analysis
2.9. Software
3. Results
3.1. Dataset Characteristics
3.2. Global Environmental Kriging Raster
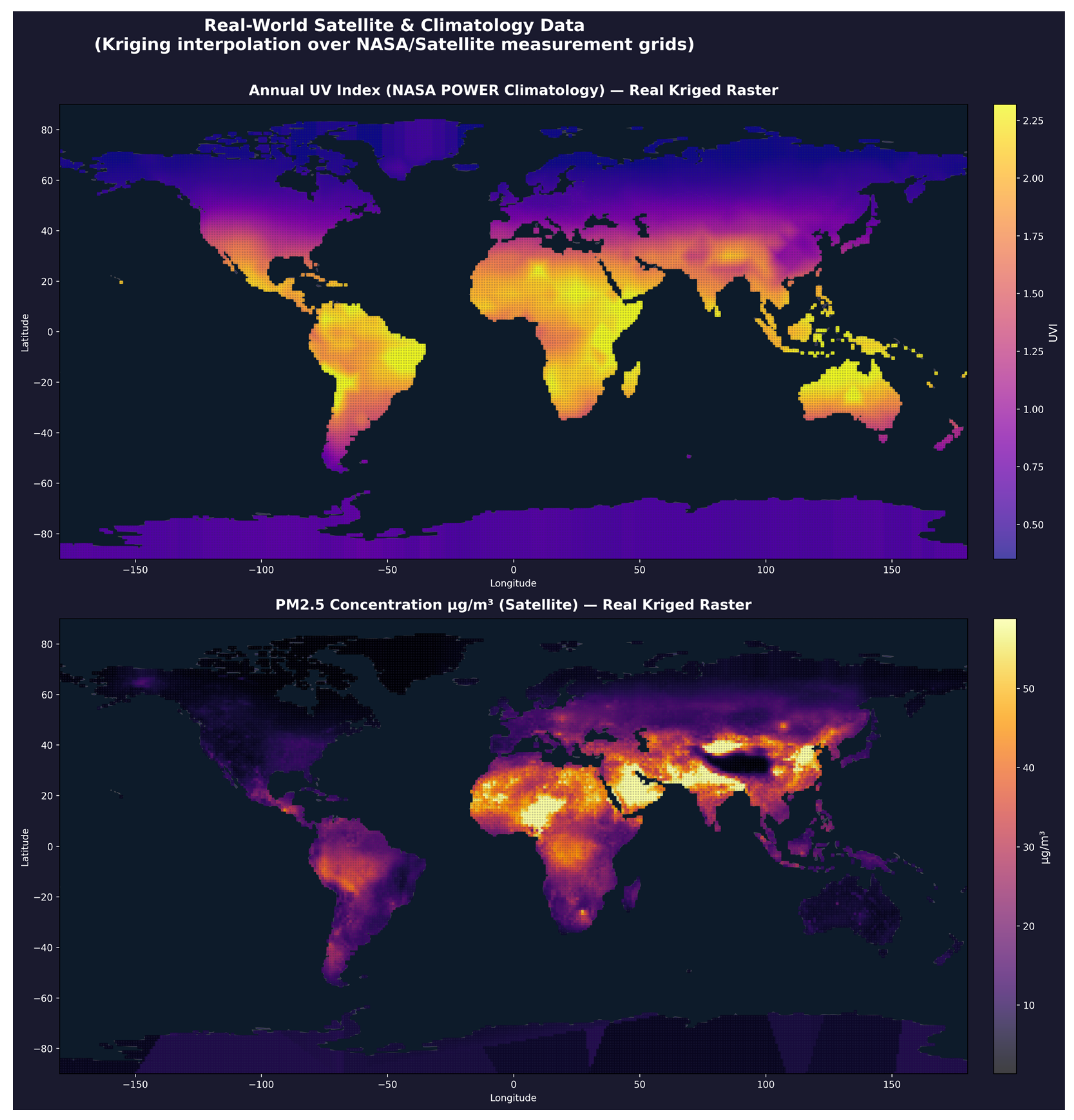
3.3. Cancer-Type Environmental Zone Values
3.4. Model Performance
3.5. SHAP Feature Importance Rankings
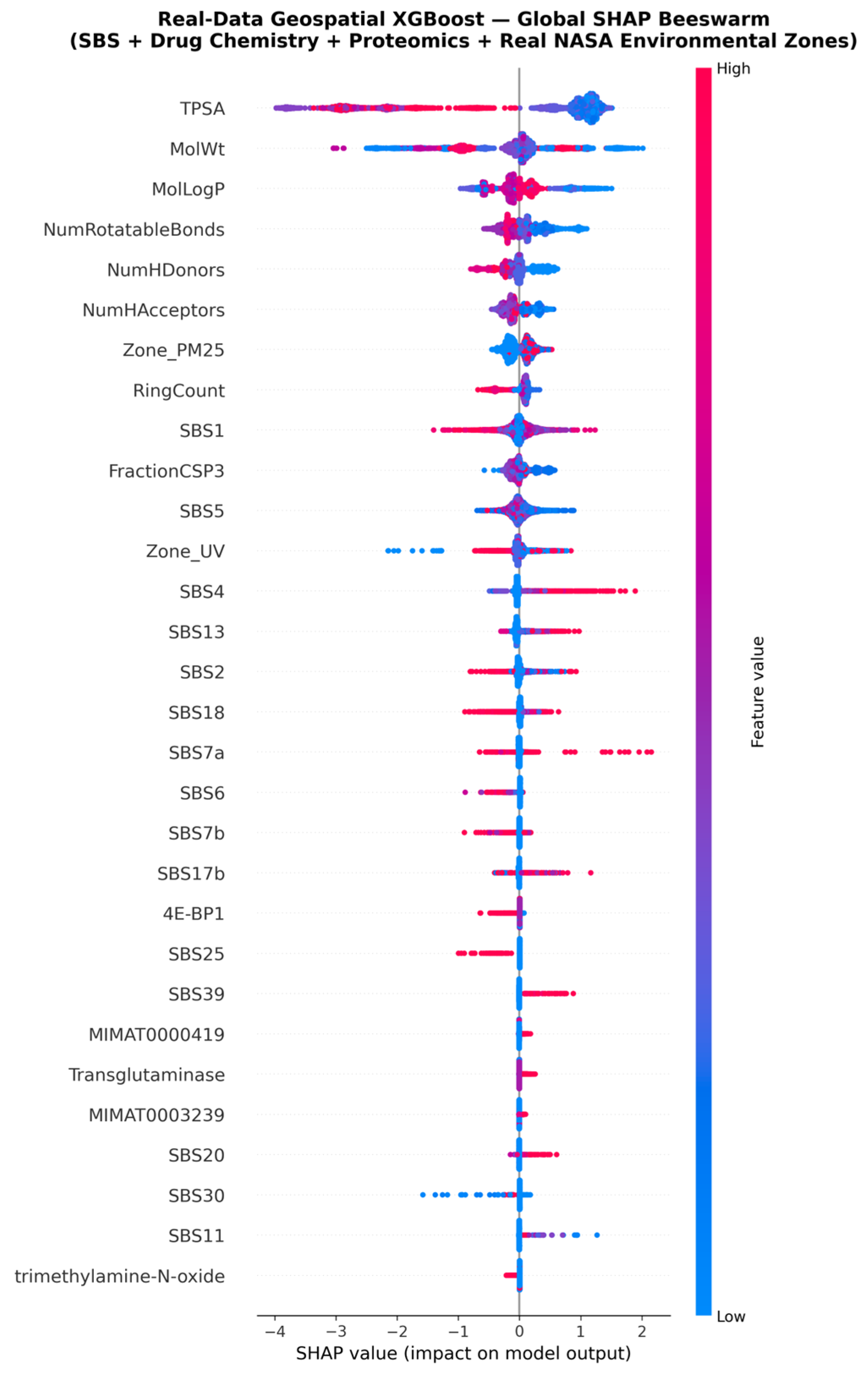
3.6. Satellite Environmental Features in the Global Ranking
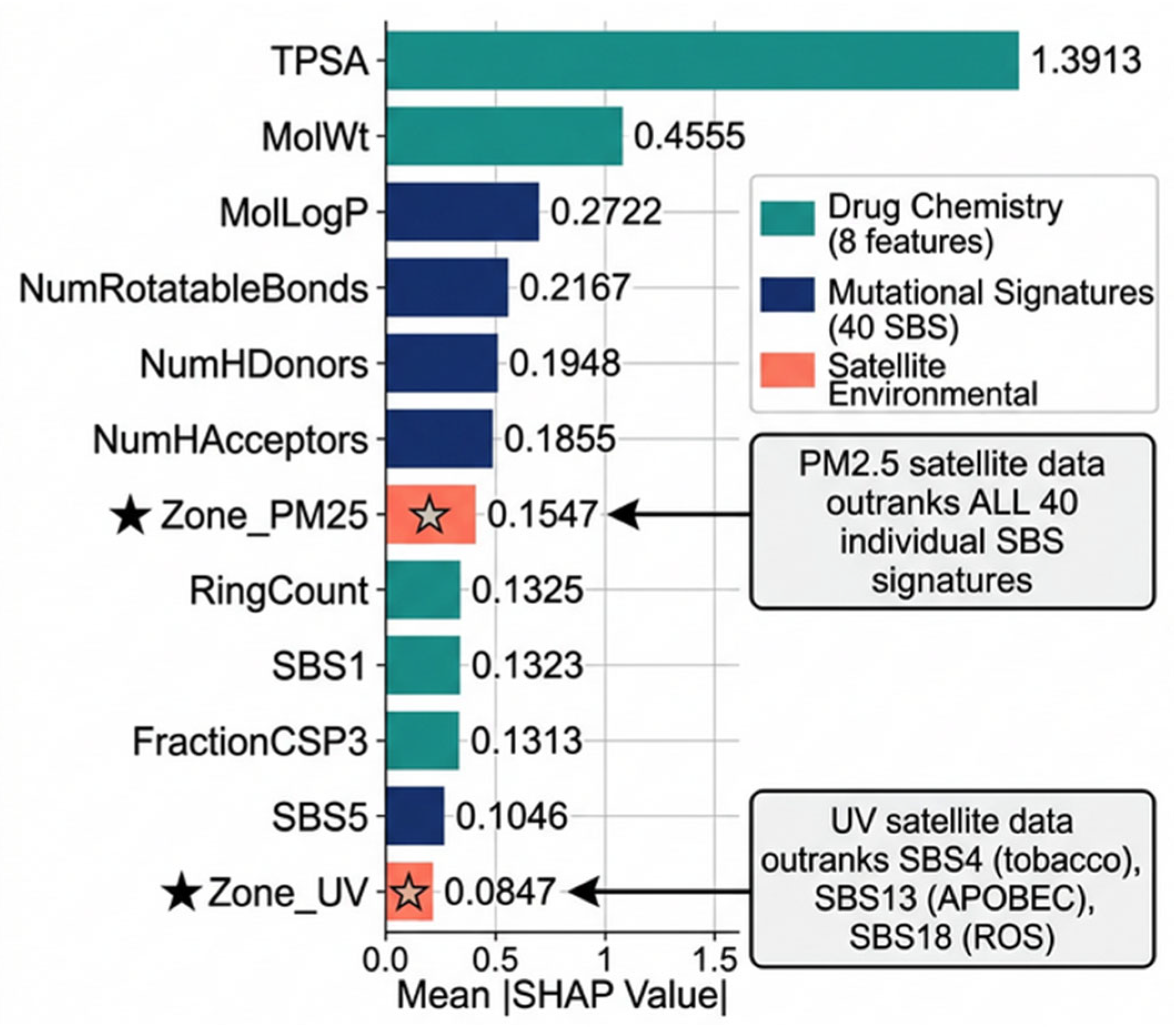
3.7. SHAP Dependence: Zone_PM25
3.8. SHAP Dependence: Zone_UV
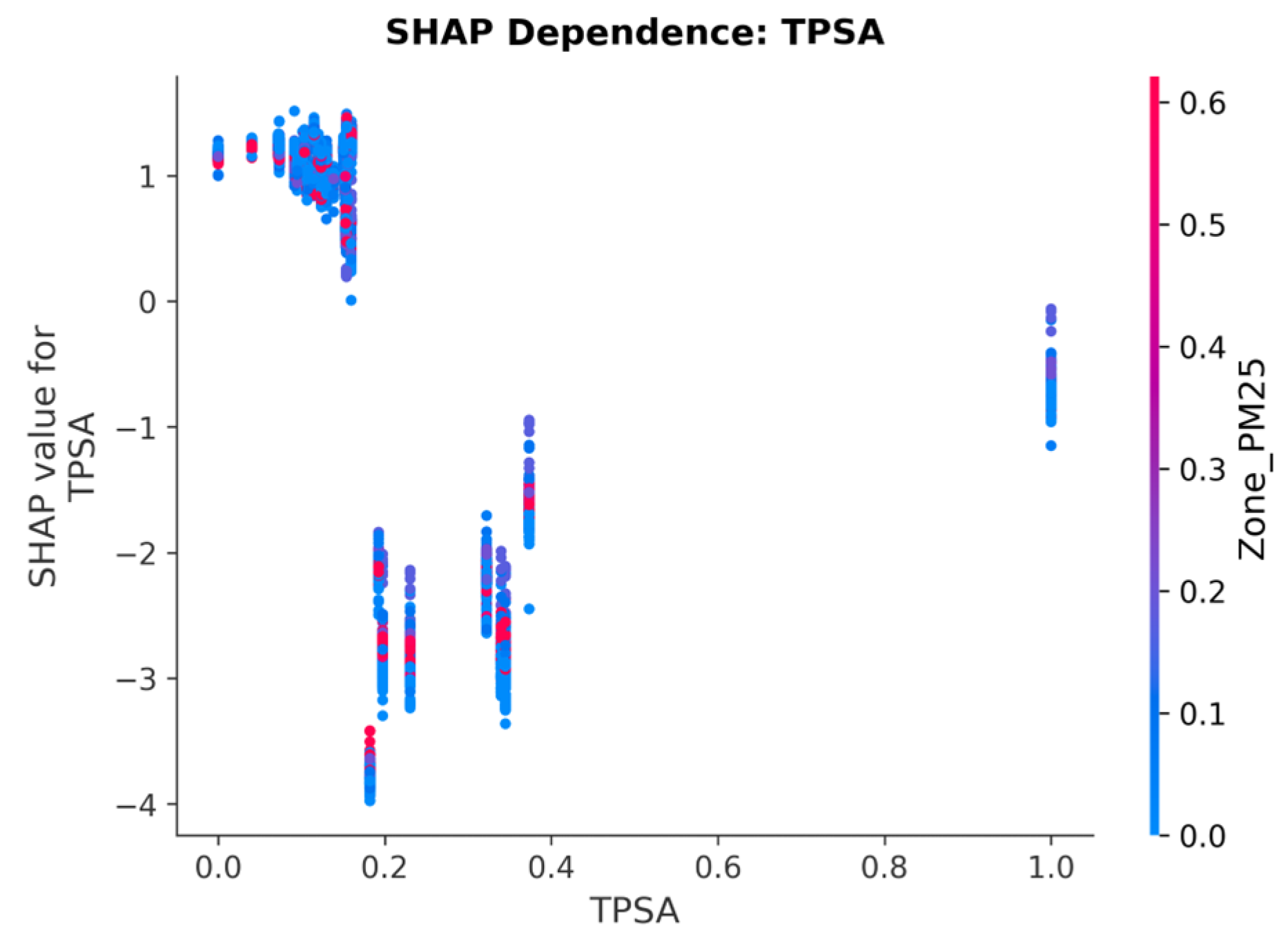
3.9. SHAP Dependence: Drug Chemistry (TPSA)
3.10. Mutational Signature Rankings
| Rank | Feature | SHAP value | Environmental Factor |
| 13 | SBS4 | 0.0783 | tobacco/air pollution |
| 16 | SBS18 | 0.0391 | oxidative stress/ROS |
| 17 | SBS7a | 0.0156 | UV radiation |
| 19 | SBS7b | 0.0120 | UV radiation |
3.11. SHAP Dependence: Environmental Mutational Signatures
3.11.1. SBS4 (Tobacco/Air Pollution)
3.11.2. SBS18 (Oxidative Stress/ROS)
3.11.3. SBS7a (UV Radiation)
3.11.4. SBS1 (Aging/Clock-Like)
4. Discussion
4.1. Summary of Main Findings
4.2. Drug Physicochemistry as the Primary Predictor
4.3. Environmental PM2.5 Exceeds Individual Mutational Signatures
4.4. UV Radiation and SHAP Interaction Effects
4.5. Aging Signatures as Confounders or Baseline Noise
4.6. Limitations
4.7. Future Directions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AOD | Aerosol Optical Depth |
| APOBEC | Apolipoprotein B mRNA Editing Enzyme Catalytic subunit |
| BER | Base Excision Repair |
| BRCA | Breast Cancer susceptibility gene |
| CCLE | Cancer Cell Line Encyclopedia |
| CTRP | Cancer Therapeutics Response Portal |
| COSMIC | Catalogue of Somatic Mutations in Cancer |
| DBS | Doublet Base Substitution |
| EGFR | Epidermal Growth Factor Receptor |
| GDSC | Genomics of Drug Sensitivity in Cancer |
| GIS | Geographic Information System |
| GPU | Graphics Processing Unit |
| HRDetect | Homologous Recombination Deficiency Detect |
| IARC | International Agency for Research on Cancer |
| IC50 | Half-Maximal Inhibitory Concentration |
| iPSC | Induced Pluripotent Stem Cell |
| KRAS | Kirsten Rat Sarcoma viral proto-oncogene |
| LN_IC50 | Natural logarithm of IC50 |
| LUAD | Lung Adenocarcinoma (TCGA code) |
| MAPK | Mitogen-Activated Protein Kinase |
| MAE | Mean Absolute Error |
| NMF | Non-negative Matrix Factorisation |
| NGS | Next-Generation Sequencing |
| PAH | Polycyclic Aromatic Hydrocarbon |
| PAR | Population Attributable Risk |
| PCAWG | Pan-Cancer Analysis of Whole Genomes |
| PM₂.₅ | Particulate Matter ≤2.5 µm aerodynamic diameter |
| RMSE | Root Mean Square Error |
| ROS | Reactive Oxygen Species |
| RPPA | Reverse-Phase Protein Array |
| SBS | Single Base Substitution |
| SHAP | Shapley Additive exPlanations |
| SKCM | Cutaneous Melanoma (TCGA code) |
| TCGA | The Cancer Genome Atlas |
| TPSA | Topological Polar Surface Area |
| TP53 | Tumour Protein 53 |
| UVI | UV Index |
| WGS | Whole-Genome Sequencing |
| XAI | Explainable Artificial Intelligence |
| XGBoost | Extreme Gradient Boosting |
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R. L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians 2024, 74(3), 229–263. [Google Scholar]
- Wild, C. P. Complementing the genome with an “exposome”: the outstanding challenge of environmental exposure measurement in molecular epidemiology. Cancer Epidemiology Biomarkers & Prevention 2005, 14(8), 1847–1850. [Google Scholar]
- Tomasetti, C.; Li, L.; Vogelstein, B. Stem cell divisions, somatic mutations, cancer etiology, and cancer prevention. Science 2017, 355(6331), 1330–1334. [Google Scholar] [CrossRef] [PubMed]
- Clapp, R. W.; Jacobs, M. M.; Loechler, E. L. Environmental and occupational causes of cancer new evidence, 2005–2007. Reviews on environmental health 2008, 23(1), 1. [Google Scholar] [PubMed]
- Wilson, L. F.; Antonsson, A.; Green, A. C.; Jordan, S. J.; Kendall, B. J.; Nagle, C. M.; Whiteman, D. C. How many cancer cases and deaths are potentially preventable? Estimates for Australia in 2013. International journal of cancer 2018, 142(4), 691–701. [Google Scholar] [CrossRef]
- Doll, R.; Peto, R. The causes of cancer: quantitative estimates of avoidable risks of cancer in the United States today. JNCI: Journal of the National Cancer Institute 1981, 66(6), 1192–1308. [Google Scholar] [CrossRef]
- United States. Congress. Office of Technology Assessment. Risks to students in school . US Government Printing Office, 1995. [Google Scholar]
- Boffetta, P.; Nyberg, F. Contribution of environmental factors to cancer risk. British medical bulletin 2003, 68(1), 71–94. [Google Scholar]
- Rushton, L.; Hutchings, S. J.; Fortunato, L.; Young, C.; Evans, G. S.; Brown, T.; Van Tongeren, M. Occupational cancer burden in Great Britain. British journal of cancer 2012, 107 (Suppl 1), S3. [Google Scholar] [CrossRef]
- Purdue, M. P.; Hutchings, S. J.; Rushton, L.; Silverman, D. T. The proportion of cancer attributable to occupational exposures. Annals of epidemiology 2015, 25(3), 188–192. [Google Scholar] [CrossRef]
- U.S. Environmental Protection Agency. Review and Evaluation of the Evidence for Cancer Associated with Air Pollution; EPA: Washington, DC, USA, 2002. [Google Scholar]
- Parkin, D. M.; Boyd, L. (2011, August). THE FRACTION OF CANCER ATTRIBUTABLE TO LIFESTYLE AND ENVIRONMENTAL FACTORS IN THE UK IN 2010. In JOURNAL OF EPIDEMIOLOGY AND COMMUNITY HEALTH (Vol. 65, pp. A143-A143). BRITISH MED ASSOC HOUSE, TAVISTOCK SQUARE, LONDON WC1H 9JR, ENGLAND: BMJ PUBLISHING GROUP. [CrossRef]
- Spirtas, R.; Heineman, E. F.; Bernstein, L.; Beebe, G. W.; Keehn, R. J.; Stark, A.; Benichou, J. Malignant mesothelioma: attributable risk of asbestos exposure. Occupational and environmental medicine 1994, 51(12), 804–811. [Google Scholar] [CrossRef]
- LaDou, J. The asbestos cancer epidemic. Environmental health perspectives 2004, 112(3), 285. [Google Scholar] [CrossRef] [PubMed]
- Lacourt, A.; Gramond, C.; Rolland, P.; Ducamp, S.; Audignon, S.; Astoul, P.; Brochard, P. Occupational and non-occupational attributable risk of asbestos exposure for malignant pleural mesothelioma. Thorax 2014, 69(6), 532–539. [Google Scholar] [CrossRef] [PubMed]
- Peto, J.; Decarli, A.; La Vecchia, C.; Levi, F.; Negri, E. The European mesothelioma epidemic. British journal of cancer 1999, 79(3), 666–672. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Cai, Y.; Ou, T.; Zhou, H.; Li, H.; Wang, Z.; Cai, K. Global burden of mesothelioma attributable to occupational asbestos exposure in 204 countries and territories: 1990–2019. Journal of cancer research and clinical oncology 2024, 150(5), 282. [Google Scholar] [CrossRef]
- International Agency for Research on Cancer. IARC: Outdoor Air Pollution a Leading Environmental Cause of Cancer Deaths; Press Release No. 221; IARC/WHO: Lyon, France, 2013. [Google Scholar]
- Loomis, D.; Grosse, Y.; Lauby-Secretan, B.; El Ghissassi, F.; Bouvard, V.; Benbrahim-Tallaa, L.; Straif, K. The carcinogenicity of outdoor air pollution. The lancet oncology 2013, 14(13), 1262–1263. [Google Scholar] [CrossRef]
- Turner, M. C.; Krewski, D.; Pope, C. A., III; Chen, Y.; Gapstur, S. M.; Thun, M. J. Long-term ambient fine particulate matter air pollution and lung cancer in a large cohort of never-smokers. American journal of respiratory and critical care medicine 2011, 184(12), 1374–1381. [Google Scholar] [CrossRef]
- Raaschou-Nielsen, O.; Andersen, Z. J.; Beelen, R.; Samoli, E.; Stafoggia, M.; Weinmayr, G.; Hoek, G. Air pollution and lung cancer incidence in 17 European cohorts: prospective analyses from the European Study of Cohorts for Air Pollution Effects (ESCAPE). The lancet oncology 2013, 14(9), 813–822. [Google Scholar] [CrossRef]
- Pope, C. A., Iii; Burnett, R. T.; Thun, M. J.; Calle, E. E.; Krewski, D.; Ito, K.; Thurston, G. D. Lung cancer, cardiopulmonary mortality, and long-term exposure to fine particulate air pollution. Jama 2002, 287(9), 1132–1141. [Google Scholar] [CrossRef]
- Hamra, G. B.; Guha, N.; Cohen, A.; Laden, F.; Raaschou-Nielsen, O.; Samet, J. M.; Loomis, D. Outdoor particulate matter exposure and lung cancer: a systematic review and meta-analysis. Environmental health perspectives 2014, 122(9), 906. [Google Scholar] [CrossRef]
- International Agency for Research on Cancer. Outdoor Air Pollution. In IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; IARC: Lyon, France, 2016; Volume 109. [Google Scholar]
- Cohen, A. J.; Pope, C. A., 3rd. Lung cancer and air pollution. Environmental health perspectives 1995, 103 (Suppl 8), 219. [Google Scholar]
- Lim, S. S.; Vos, T.; Flaxman, A. D.; Danaei, G.; Shibuya, K.; Adair-Rohani, H.; Pelizzari, P. M. A comparative risk assessment of burden of disease and injury attributable to 67 risk factors and risk factor clusters in 21 regions, 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. The lancet 2012, 380(9859), 2224–2260. [Google Scholar]
- Carrot-Zhang, J.; Soca-Chafre, G.; Patterson, N.; Thorner, A. R.; Nag, A.; Watson, J.; Meyerson, M. Genetic ancestry contributes to somatic mutations in lung cancers from admixed Latin American populations. Cancer Discovery 2021, 11(3), 591–598. [Google Scholar] [CrossRef] [PubMed]
- Rueda-Zarazua, B.; Gutiérrez, H.; García-Ortiz, H.; Orozco, L.; Ramírez-Martínez, G.; Jiménez-Alvarez, L.; Melendez-Zajgla, J. A pilot study: contrasting genomic profiles of lung adenocarcinoma between patients of european and latin american ancestry. International Journal of Molecular Sciences 2025, 26(10), 4865. [Google Scholar] [CrossRef] [PubMed]
- da Cunha Santos, G.; Shepherd, F. A.; Tsao, M. S. EGFR mutations and lung cancer. Annual Review of Pathology: Mechanisms of Disease 2011, 6(1), 49–69. [Google Scholar] [CrossRef]
- Díaz-Gay, M.; Zhang, T.; Hoang, P. H.; Khandekar, A.; Zhao, W.; Steele, C. D.; Landi, M. T. The mutagenic forces shaping the genomic landscape of lung cancer in never smokers. medRxiv 2024. [Google Scholar] [CrossRef]
- Arrieta, O.; Cardona, A. F.; Martín, C.; Más-López, L.; Corrales-Rodríguez, L.; Bramuglia, G.; Cuello, M. Updated frequency of EGFR and KRAS mutations in nonsmall-cell lung cancer in Latin America: the Latin-American Consortium for the Investigation of Lung Cancer (CLICaP). Journal of Thoracic Oncology 2015, 10(5), 838–843. [Google Scholar] [CrossRef]
- Schabath, M. B.; Cote, M. L. Cancer progress and priorities: lung cancer. Cancer epidemiology, biomarkers & prevention 2019, 28(10), 1563–1579. [Google Scholar]
- Kehm, R. D.; Lloyd, S. E.; Burke, K. R.; Terry, M. B. Advancing environmental epidemiologic methods to confront the cancer burden. American Journal of Epidemiology 2025, 194(1), 195–207. [Google Scholar] [CrossRef]
- Liu, J.; Gan, T.; Hu, W.; Li, Y. Current status and perspectives in environmental oncology. Chronic Diseases and Translational Medicine 2024, 10(04), 293–301. [Google Scholar] [CrossRef]
- Chevalier, A.; Guo, T.; Gurevich, N. Q.; Xu, J.; Yajima, M.; Campbell, J. D. Characterization of mutational signatures in tumors from a large Chinese population. Cancer Research Communications 2025, 5(8), 1466–1476. [Google Scholar] [CrossRef]
- Alexandrov, L. B.; Nik-Zainal, S.; Wedge, D. C.; Aparicio, S. A.; Behjati, S.; Biankin, A. V.; Stratton, M. R. Signatures of mutational processes in human cancer. nature 2013, 500(7463), 415–421. [Google Scholar] [CrossRef] [PubMed]
- Helleday, T.; Eshtad, S.; Nik-Zainal, S. Mechanisms underlying mutational signatures in human cancers. Nature reviews genetics 2014, 15(9), 585–598. [Google Scholar] [PubMed]
- Nik-Zainal, S.; Alexandrov, L. B.; Wedge, D. C.; Van Loo, P.; Greenman, C. D.; Raine, K.; Stratton, M. R. Mutational processes molding the genomes of 21 breast cancers. Cell 2012, 149(5), 979–993. [Google Scholar] [CrossRef] [PubMed]
- Greenman, C.; Stephens, P.; Smith, R.; Dalgliesh, G. L.; Hunter, C.; Bignell, G.; Stratton, M. R. Patterns of somatic mutation in human cancer genomes. Nature 2007, 446(7132), 153–158. [Google Scholar] [CrossRef]
- Meyerson, M.; Gabriel, S.; Getz, G. Advances in understanding cancer genomes through second-generation sequencing. Nature Reviews Genetics 2010, 11(10), 685–696. [Google Scholar] [CrossRef]
- Alexandrov, L. B.; Nik-Zainal, S.; Wedge, D. C.; Campbell, P. J.; Stratton, M. R. Deciphering signatures of mutational processes operative in human cancer. Cell reports 2013, 3(1), 246–259. [Google Scholar] [CrossRef]
- Fischer, A.; Illingworth, C. J.; Campbell, P. J.; Mustonen, V. EMu: probabilistic inference of mutational processes and their localization in the cancer genome. Genome biology 2013, 14(4), R39. [Google Scholar] [CrossRef]
- Forbes, S. A.; Beare, D.; Boutselakis, H.; Bamford, S.; Bindal, N.; Tate, J.; Campbell, P. J. COSMIC: somatic cancer genetics at high-resolution. Nucleic acids research 2017, 45(D1), D777–D783. [Google Scholar]
- Petljak, M.; Alexandrov, L. B.; Brammeld, J. S.; Price, S.; Wedge, D. C.; Grossmann, S.; Stratton, M. R. Characterizing mutational signatures in human cancer cell lines reveals episodic APOBEC mutagenesis. Cell 2019, 176(6), 1282–1294. [Google Scholar] [CrossRef]
- Alexandrov, L. B.; Jones, P. H.; Wedge, D. C.; Sale, J. E.; Campbell, P. J.; Nik-Zainal, S.; Stratton, M. R. Clock-like mutational processes in human somatic cells. Nature genetics 2015, 47(12), 1402–1407. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, G. P.; Denissenko, M. F.; Olivier, M.; Tretyakova, N.; Hecht, S. S.; Hainaut, P. Tobacco smoke carcinogens, DNA damage and p53 mutations in smoking-associated cancers. Oncogene 2002, 21(48), 7435–7451. [Google Scholar] [CrossRef] [PubMed]
- Brash, D. E. UV signature mutations. Photochemistry and photobiology 2015, 91(1), 15–26. [Google Scholar] [CrossRef] [PubMed]
- Roberts, S. A.; Lawrence, M. S.; Klimczak, L. J.; Grimm, S. A.; Fargo, D.; Stojanov, P.; Gordenin, D. A. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nature genetics 2013, 45(9), 970–976. [Google Scholar] [CrossRef]
- The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-Cancer Analysis of Whole Genomes. Nature 2020, 578, 82–93. [CrossRef]
- Alexandrov, L. B.; Kim, J.; Haradhvala, N. J.; Huang, M. N.; Tian Ng, A. W.; Wu, Y.; Stratton, M. R. The repertoire of mutational signatures in human cancer. Nature 2020, 578(7793), 94–101. [Google Scholar] [CrossRef]
- Tate, J. G.; Bamford, S.; Jubb, H. C.; Sondka, Z.; Beare, D. M.; Bindal, N.; Forbes, S. A. COSMIC: the catalogue of somatic mutations in cancer. Nucleic acids research 2019, 47(D1), D941–D947. [Google Scholar] [CrossRef]
- Kucab, J. E.; Zou, X.; Morganella, S.; Joel, M.; Nanda, A. S.; Nagy, E.; Nik-Zainal, S. A compendium of mutational signatures of environmental agents. Cell 2019, 177(4), 821–836. [Google Scholar] [CrossRef]
- Nik-Zainal, S.; Kucab, J. E.; Morganella, S.; Glodzik, D.; Alexandrov, L. B.; Arlt, V. M.; Phillips, D. H. The genome as a record of environmental exposure. Mutagenesis 2015, 30(6), 763–770. [Google Scholar] [CrossRef]
- Boysen, G.; Alexandrov, L. B.; Rahbari, R.; Nookaew, I.; Ussery, D.; Chao, M. R.; Cooke, M. S. Investigating the origins of the mutational signatures in cancer. Nucleic acids research 2025, 53(1), gkae1303. [Google Scholar] [CrossRef]
- Maura, F.; Degasperi, A.; Nadeu, F.; Leongamornlert, D.; Davies, H.; Moore, L.; Bolli, N. A practical guide for mutational signature analysis in hematological malignancies. Nature communications 2019, 10(1), 2969. [Google Scholar] [CrossRef] [PubMed]
- Islam, S. A.; Díaz-Gay, M.; Wu, Y.; Barnes, M.; Vangara, R.; Bergstrom, E. N.; Alexandrov, L. B. Uncovering novel mutational signatures by de novo extraction with SigProfilerExtractor. Cell genomics 2022, 2(11). [Google Scholar] [CrossRef] [PubMed]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V. E.; Zhou, S.; Diaz, L. A., Jr.; Kinzler, K. W. Cancer genome landscapes. science 2013, 339(6127), 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Stratton, MR; Campbell, PJ; Futreal, PA. The cancer genome. Nature 2009, 458(7239), 719–724. [Google Scholar] [CrossRef]
- McFarland, CD; Korolev, KS; Kryukov, GV; Sunyaev, SR; Mirny, LA. Impact of deleterious passenger mutations on cancer progression. Proc Natl Acad Sci U S A 2013, 110(8), 2910–2915. [Google Scholar] [CrossRef]
- McFarland, C. D.; Mirny, L. A.; Korolev, K. S. Tug-of-war between driver and passenger mutations in cancer and other adaptive processes. Proceedings of the National Academy of Sciences 2014, 111(42), 15138–15143. [Google Scholar]
- Bozic, I.; Antal, T.; Ohtsuki, H.; Carter, H.; Kim, D.; Chen, S.; Nowak, M. A. Accumulation of driver and passenger mutations during tumor progression. Proceedings of the National Academy of Sciences 2010, 107(43), 18545–18550. [Google Scholar] [CrossRef]
- Kumar, S.; Warrell, J.; Li, S.; McGillivray, P. D.; Meyerson, W.; Salichos, L.; Gerstein, M. B. Passenger mutations in more than 2,500 cancer genomes: overall molecular functional impact and consequences. Cell 2020, 180(5), 915–927. [Google Scholar] [CrossRef]
- Rheinbay, E.; Nielsen, M. M.; Abascal, F.; Wala, J. A.; Shapira, O.; Tiao, G.; Getz, G. Analyses of non-coding somatic drivers in 2,658 cancer whole genomes. Nature 2020, 578(7793), 102–111. [Google Scholar] [CrossRef]
- Shuai, S.; Gallinger, S.; Stein, L. D. Combined burden and functional impact tests for cancer driver discovery using DriverPower. Nature communications 2020, 11(1), 734. [Google Scholar] [CrossRef]
- Iorio, F.; Knijnenburg, T. A.; Vis, D. J.; Bignell, G. R.; Menden, M. P.; Schubert, M.; Garnett, M. J. A landscape of pharmacogenomic interactions in cancer. Cell 2016, 166(3), 740–754. [Google Scholar] [CrossRef]
- Ghandi, M.; Huang, F. W.; Jané-Valbuena, J.; Kryukov, G. V.; Lo, C. C.; McDonald, E. R., III; Sellers, W. R. Next-generation characterization of the cancer cell line encyclopedia. Nature 2019, 569(7757), 503–508. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Soares, J.; Greninger, P.; Edelman, E. J.; Lightfoot, H.; Forbes, S.; Garnett, M. J. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic acids research 2012, 41(D1), D955–D961. [Google Scholar] [CrossRef] [PubMed]
- Barretina, J.; Caponigro, G.; Stransky, N.; Venkatesan, K.; Margolin, A. A.; Kim, S.; Garraway, L. A. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012, 483(7391), 603–607. [Google Scholar] [CrossRef] [PubMed]
- Garnett, M. J.; Edelman, E. J.; Heidorn, S. J.; Greenman, C. D.; Dastur, A.; Lau, K. W.; Benes, C. H. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 2012, 483(7391), 570–575. [Google Scholar] [CrossRef]
- Boot, A.; Huang, M. N.; Ng, A. W.; Ho, S. C.; Lim, J. Q.; Kawakami, Y.; Rozen, S. G. In-depth characterization of the cisplatin mutational signature in human cell lines and in esophageal and liver tumors. Genome research 2018, 28(5), 654–665. [Google Scholar] [CrossRef]
- Davies, H.; Glodzik, D.; Morganella, S.; Yates, L. R.; Staaf, J.; Zou, X.; Nik-Zainal, S. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nature medicine 2017, 23(4), 517–525. [Google Scholar] [CrossRef]
- Flaherty, K. T.; Puzanov, I.; Kim, K. B.; Ribas, A.; McArthur, G. A.; Sosman, J. A.; Chapman, P. B. Inhibition of mutated, activated BRAF in metastatic melanoma. New England Journal of Medicine 2010, 363(9), 809–819. [Google Scholar] [CrossRef]
- Davies, H.; Bignell, G. R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Futreal, P. A. Mutations of the BRAF gene in human cancer. Nature 2002, 417(6892), 949–954. [Google Scholar] [CrossRef]
- Kopetz, S.; Desai, J.; Chan, E.; Hecht, J. R.; O’Dwyer, P. J.; Maru, D.; Saltz, L. Phase II pilot study of vemurafenib in patients with metastatic BRAF-mutated colorectal cancer. Journal of clinical oncology 2015, 33(34), 4032–4038. [Google Scholar] [CrossRef]
- Hayward, N. K.; Wilmott, J. S.; Waddell, N.; Johansson, P. A.; Field, M. A.; Nones, K.; Mann, G. J. Whole-genome landscapes of major melanoma subtypes. Nature 2017, 545(7653), 175–180. [Google Scholar] [CrossRef]
- Hayward, NK; Wilmott, JS; Waddell, N; et al. Whole-genome landscapes of major melanoma subtypes. Nature 2017, 545(7653), 175–180. [Google Scholar] [CrossRef] [PubMed]
- Viel, A.; Bruselles, A.; Meccia, E.; Fornasarig, M.; Quaia, M.; Canzonieri, V.; Bignami, M. A specific mutational signature associated with DNA 8-oxoguanine persistence in MUTYH-defective colorectal cancer. EBioMedicine 2017, 20, 39–49. [Google Scholar] [CrossRef]
- Pilati, C.; Shinde, J.; Alexandrov, L. B.; Assié, G.; André, T.; Hélias-Rodzewicz, Z.; Laurent-Puig, P. Mutational signature analysis identifies MUTYH deficiency in colorectal cancers and adrenocortical carcinomas. The Journal of pathology 2017, 242(1), 10–15. [Google Scholar] [CrossRef] [PubMed]
- Pereira, L.; Igea, A.; Canovas, B.; Dolado, I.; Nebreda, A. R. Inhibition of p38 MAPK sensitizes tumour cells to cisplatin-induced apoptosis mediated by reactive oxygen species and JNK. EMBO molecular medicine 2013, 5(11), 1759–1774. [Google Scholar] [CrossRef] [PubMed]
- Trempolec, N.; Dave-Coll, N.; Nebreda, A. R. SnapShot: p38 MAPK signaling. Cell 2013, 152(3), 656–656. [Google Scholar] [CrossRef]
- Smirnov, P.; Safikhani, Z.; El-Hachem, N.; Wang, D.; She, A.; Olsen, C.; Haibe-Kains, B. PharmacoGx: an R package for analysis of large pharmacogenomic datasets. Bioinformatics 2016, 32(8), 1244–1246. [Google Scholar] [CrossRef]
- Lee, S. J.; Serre, M. L.; Van Donkelaar, A.; Martin, R. V.; Burnett, R. T.; Jerrett, M. Comparison of geostatistical interpolation and remote sensing techniques for estimating long-term exposure to ambient PM2. 5 concentrations across the continental United States. Environmental health perspectives 2012, 120(12), 1727. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. Xgboost: A scalable tree boosting system. In Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining, 2016, August; pp. 785–794. [Google Scholar]
- Lundberg, S. M.; Lee, S. I. A unified approach to interpreting model predictions. Advances in neural information processing systems 2017, 30. [Google Scholar]
- Lundberg, S. M.; Erion, G.; Chen, H.; DeGrave, A.; Prutkin, J. M.; Nair, B.; Lee, S. I. From local explanations to global understanding with explainable AI for trees. Nature machine intelligence 2020, 2(1), 56–67. [Google Scholar] [CrossRef]
- RDKit: Open-Source Cheminformatics. Available online: https://www.rdkit.org.
- NASA Langley Research Center POWER Project funded through the NASA Earth Science Directorate Applied Science Program. Available online: https://power.larc.nasa.gov (accessed on 7 March 2026).
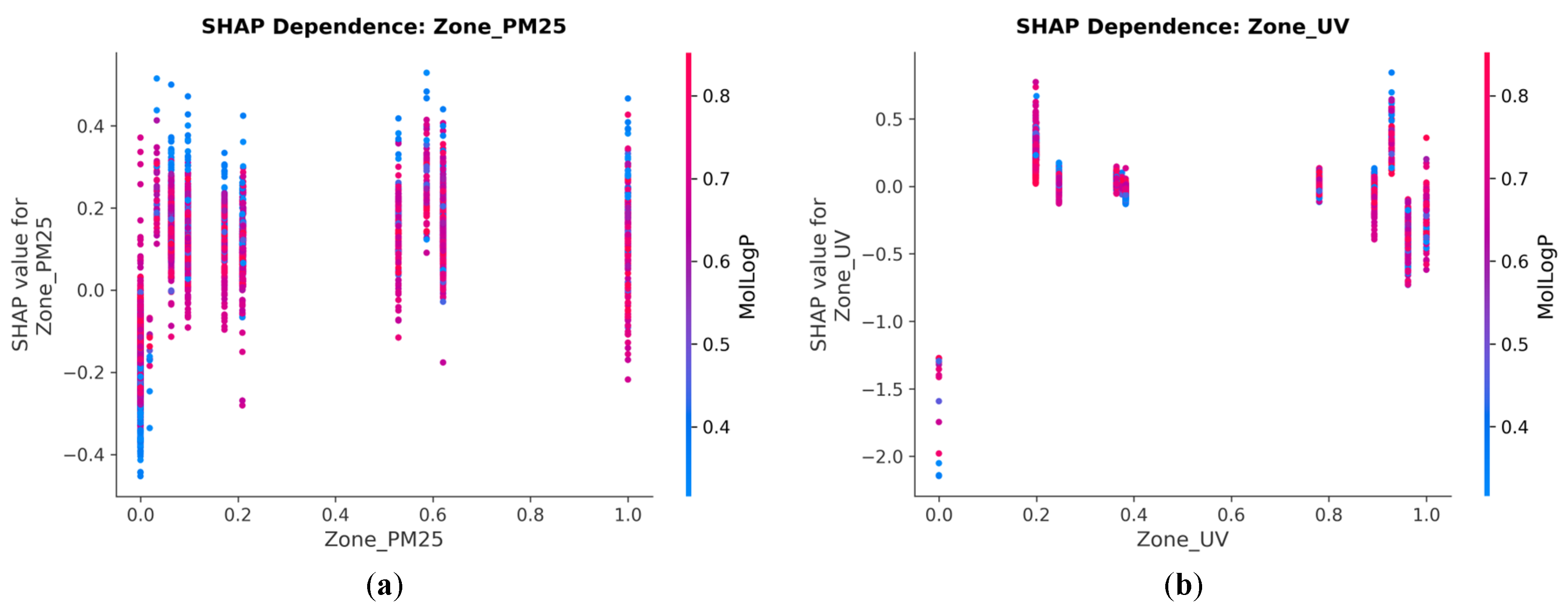
| Rank | Feature | Mean |SHAP| | Category |
| 1 | TPSA | 1.3913 | Drug chemistry |
| 2 | MolWt | 0.4555 | Drug chemistry |
| 3 | MolLogP | 0.2722 | Drug chemistry |
| 4 | NumRotatableBonds | 0.2167 | Drug chemistry |
| 5 | NumHDonors | 0.1948 | Drug chemistry |
| 6 | NumHAcceptors | 0.1855 | Drug chemistry |
| 7 | Zone_PM25 | 0.1547 | NASA Satellite (PM2.5) |
| 8 | RingCount | 0.1325 | Drug chemistry |
| 9 | SBS1 | 0.1323 | SBS signature (aging) |
| 10 | FractionCSP3 | 0.1313 | Drug chemistry |
| 11 | SBS5 | 0.1046 | SBS signature (aging/clock-like) |
| 12 | Zone_UV | 0.0847 | NASA Satellite (UV) |
| 13 | SBS4 | 0.0783 | SBS signature (tobacco/PM2.5) |
| 14 | SBS13 | 0.0725 | SBS signature (APOBEC) |
| 15 | SBS2 | 0.05 | SBS signature (APOBEC) |
| 16 | SBS18 | 0.0391 | SBS signature (oxidative stress) |
| 17 | SBS7a | 0.0156 | SBS signature (UV) |
| 18 | SBS6 | 0.0156 | SBS signature (MMR deficiency) |
| 19 | SBS7b | 0.012 | SBS signature (UV) |
| 20 | SBS17b | 0.0105 | SBS signature (treatment/ROS) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




