Submitted:
03 April 2026
Posted:
07 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
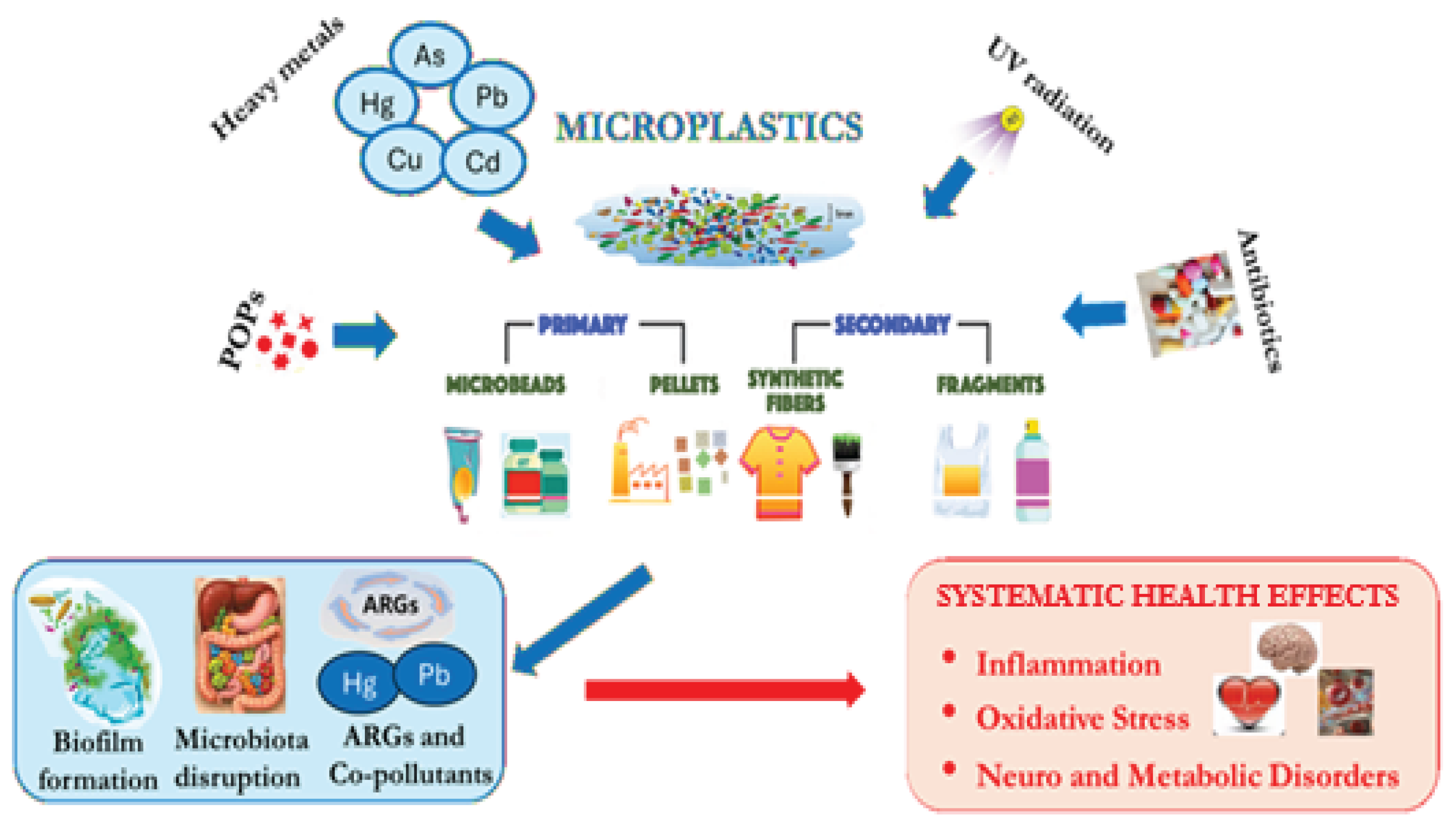
2. Sources and Characteristics of Microplastics
3. Mechanisms of Microplastic–Microbiota Interactions
4. Multi-Omics Approaches in Microplastic Research
5. Health Implications of Microplastic–Microbiota Interactions
6. Knowledge Gaps and Future Directions
Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Thin, Z.S.; Chew, J.; Ong, T.Y.Y.; Ali, R.A.R.; Gew, L.T. Impact of microplastics on the human gut microbiome: a systematic review of microbial composition, diversity, and metabolic disruptions. BMC Gastroenterol. 2025, 25, 583. [Google Scholar] [CrossRef]
- Tang, K.H.D.; Li, R. Aged microplastics and antibiotic resistance genes: a review of aging effects on their interactions. Antibiotics 2024, 13, 941. [Google Scholar] [CrossRef]
- Zhang, Z.; Xu, M.; Wang, L.; Gu, W.; Li, X.; Han, Z.; Fu, X.; Wang, X.; Li, X.; Su, Z. Continuous oral exposure to micro- and nanoplastics induced gut microbiota dysbiosis, intestinal barrier and immune dysfunction in adult mice. Environ. Int. 2023, 182, 108353. [Google Scholar] [CrossRef]
- Pelegrini, K.; Pereira, T.C.B.; Maraschin, T.G.; Teodoro, L.S.; Basso, N.R.S.; De Galland, G.L.B.; Ligabue, R.A.; Bogo, M.R. Micro- and nanoplastic toxicity: A review on size, type, source, and test-organism implications. Sci. Total Environ. 2023, 878, 162954. [Google Scholar] [CrossRef]
- Lin, W.; Cao, S.; Wu, Q.; Xu, F.; Li, R.; Cui, L. Size effects of microplastics on antibiotic resistome and core microbiome in an urban river. Sci. Total Environ. 2024, 919, 170716. [Google Scholar] [CrossRef]
- Vanetti, C.; Brogiatto, M.; Pezzana, S.; Clerici, M.; Fenizia, C. Effects on microplastics on the immmune system: How much should we worry? Immunol. Lett. 2025, 272, 106976. [Google Scholar] [CrossRef]
- Guo, Y.; Zhang, Z.; Wang, J.; Ye, X.; Zhao, J.; Liu, P.; Jia, H. Microplastic-mediated interactions with antibiotics and antibiotic resistance genes in sludge: combined effects and environmental implications. Environ. Geochem. Health 2025, 47, 506. [Google Scholar] [CrossRef]
- Zhang, Z.; Chen, W.; Chan, H.; Peng, J.; Zhu, P.; Li, H.; Jiang, X.; Zhang, Z.; Wang, Y.; Tan, Z.; Peng, Y.; Zhang, Sh; Lin, K.; Yung, K.K.-L. Polystyrene microplastics induce size-dependent multi-organ damage in mice: Insights into gut microbiota and fecal metabolites. J. Hazard. Mater. 2024, 461, 132503. [Google Scholar] [CrossRef]
- Chen, X.; Xu, L.; Chen, Q.; Su, S.; Zhuang, J.; Qiao, D. Polystyrene micro- and nanoparticles exposure induced anxiety-like behaviors, gut microbiota dysbiosis and metabolism disorder in adult mice Ecotoxicol. Environ. Saf. 2023, 259, 15000. [Google Scholar] [CrossRef]
- Li, Y.; Xu, M.; Zhang, Z.; Halimu, G.; Li, Y.; Li, Y.; Gu, W.; Zhang, B.; Wang, X. In vitro study on the toxicity of nanoplastics with different charges to murine splenic lymphocytes. J. Hazard. Mater. 2022, 424, 127508. [Google Scholar] [CrossRef]
- Halimu, G.; Zhang, Q.; Liu, L.; Zhang, Z.; Wang, X.; Gu, W.; Zhang, B.; Dai, Y.; Zhang, H.; Zhang, C.; Xu, M. Toxic effects of nanoplastics with different sizes and surface charges on epithelial-to-mesenchymal transition in A549 cells and the potential toxicological mechanism. J. Hazard. Mater. 2022, 430, 128485. [Google Scholar]
- Xu, D.; Ma, Y.; Han, X.; Chen, Y. Systematic toxicity evaluation of polystyrene nanoplastics on mice and molecular mechanism investigation about their internalization into Caco-2 Cells. J. Hazard. Mater. 2021, 417, 126092. [Google Scholar] [CrossRef]
- Deng, B.D.; Sinha, S.R.; Lear, G.; Tropini, C. Microplastics and nanoplastics in the human gut: from signals to standards. Nat. Rev. Gastroenterol. Hepatol. 2026; published 04 March. [Google Scholar]
- Xia, Y.; Lan, Y.; Xu, Y.; Liu, F.; Chen, X.; Luo, J.; Xu, H.; Liu, Y. Effects of microplastics and tetracycline induced intestinal damage, intestinal microbiota dysbiosis, and antibiotic resistome: metagenomic analysis in young mice. Environ. Int. 2025, 199, 109512. [Google Scholar] [CrossRef]
- Wang, K.; Wang, F.; Yu, Y.; Yang, S.; Han, Y.; Yao, H. Microplastics and soil microbiomes. BMC Biol. 2025, 23, 273. [Google Scholar] [CrossRef]
- Bora, S.S.; Gogoi, R.; Sharma, M.R.; Anshu; Borah, M.P.; Deka, P.; Bora, J.; Naorem, R.S.; Das, J.; Teli, A.B. Microplastics and human health: unveiling the gut microbiome disruption and chronic disease risks. Front. Cell. Infect. Microbiol. 2024, 14, 1492759. [Google Scholar] [CrossRef]
- Sun, J.; Peng, S.; Yang, Q.; Yang, J.; Dai, Y.; Xing, L. Microplastics/nanoplastic and neurological health: An overview of neurological defects and mechanisms. Toxicology 2025, 511, 154030. [Google Scholar] [CrossRef]
- Jia, J.; Liu, Q.; Zhao, E.; Li, X.; Xiong, X.; Wu, C. Biofilm formation on microplastics and interactions with antibiotics, antibiotic resistance genes and pathogens in aquatic environment. EEH 2024, 3, 516–528. [Google Scholar] [CrossRef]
- Zhang, B.; Li, Y.; Zhao, Z.; Lyu, H.; Wang, l.; Welden, N.; Tang, J. Microplastics mediated antibiotic resistance gene enrichment and transfer in environment: Different types, microplastic antiobiotic resistance gene ecological island and nano-size effect. Ecotoxicol. Environ. Saf. 2026, 309, 119596. [Google Scholar] [CrossRef]
- Hongjin, C.; ur Rahman, S.; Rehman, A.; Khan, A.A.; Khalid, M. Microplastics and antibiotic resistance genes as rising threats: their interaction represents an urgent environmental concern. Curr. Res. Microb. Sci. 2025, 9, 100447. [Google Scholar] [CrossRef]
- Gao, B.; Chen, L.; Wu, L.; Zhang, S.; Zhao, S.; Mo, Z.; Chen, Z.; Tu, P. Association between microplastics and the functionalities of the human gut microbiome. Ecotoxicol. Environ. Saf. 2025, 290, 117497. [Google Scholar] [CrossRef]
- Bao, Y.; Liu, G.; Yao, H. (2025). Microplastic aging mediates bacterial and antibiotic resistance gene composition in plastisphere and the associated soil solution. Environ.Pollut 2025, 385, 127134. [Google Scholar] [CrossRef]
- Silverstein, J.; Yuang, D. Metagenomic analysis reveals the effects of microplastics on antibiotic resistance genes in sludge anaerobic digestion. Toxics 2024, 12, 920. [Google Scholar] [CrossRef]
- Editorial. Microplastic research must consider microbes. Nature Microbiology 2025, 10, 603. [Google Scholar] [CrossRef]
- Huang, X.; Lu, B.; Liu, H.; Wu, X.; Liu, Y. Ecotoxicological impacts of microplastics to gut microbiota: response mechanisms, challenges, and environmental sustainability — A review. Ecotoxicol. Environ. Saf. 2025, 302, 118748. [Google Scholar] [CrossRef]
- Khan, T.; Khanem, A.; Batool, I.; Ullah, I.; Younas, F. Microplastics: Disseminators of antibiotic resistance genes and pathogenic bacteria. Aquat. Toxicol. 2025, 289, 107591. [Google Scholar] [CrossRef]
- Zhao, C.; Li, Ch.; Li, F. Research progress on the origin, fate, impacts and harm of microplastics and antibiotic resistance genes in wastewater treatment plants. Sci. Rep. 2024, 14, 9719. [Google Scholar] [CrossRef]
- Barbosa, F.; Adeyemi, J.A.; Bocato, M.Z.; Comas, A.; Campiglia, A. A critical viewpoint on current issues, limitations, and future research needs on micro- and nanoplastic studies: from the detection to the toxicological assessment. Environ. Res. 2020, 182, 109089. [Google Scholar] [CrossRef]
- Shruti, V.C.; P’erez-Guevara, F.; Elizalde-Martínez, I.; Kutralam-Muniasamy, G. First study of its kind on the microplastic contamination of soft drinks, cold tea and energy drinks - future research and environmental considerations. Sci. Total Environ. 2020, 726, 138580. [Google Scholar] [CrossRef]
- Sharma, N.; Kumar, V.; Vimal, S.; Umesh, M.; Chakraborty, P.; Basheer, T.; Sarojini, S.; Sharma, P.; Pasrija, R.; Barcelo, D. Microplastic residues in clinical samples: A retrospection on sources, entry routes, detection methods and human toxicity. TrAC 2024, 173, 117618. [Google Scholar] [CrossRef]
- Tiwari, M.; Rathod, T.D.; Ajmal, P.Y.; Bhangare, R.C.; Sahu, S.K. Distribution and characterization of microplastics in beach sand from three different Indian coastal environments. Mar. Pollut. Bull. 2019, 140, 262–273. [Google Scholar] [CrossRef]
- Sutton, S.C.; Hills, R.D., Jr. Role of nanoplastics in decreasing the intestinal microbiome ratio: a review of the scope of polystyrene. Toxics 2025, 13, 1036. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ren, J.; Zheng, B.; Sun, J.; Zhang, J.; Niu, Y.; Shao, B.; Jin, Y. Microplastic toxicity: mechanisms, assessment methods, and future research directions. Front. Toxicol. 2026, 8, 766103. [Google Scholar] [CrossRef]
- Gigault, J.; el Hadri, H.; Nguyen, B.; Grassl, B.; Rowenczyk, L.; Tufenkji, N.; Feng, S.; Wiesner, M. Nanoplastics are neither microplastics nor engineered nanoparticles. Nat. Nanotechnol. 2021, 16, 501–507. [Google Scholar] [CrossRef]
- Mitrano, D.M.; Wick, P.; Nowack, B. Placing nanoplastics in the context of global plastic pollution. Nat. Nanotechnol. 2021, 16, 491–500. [Google Scholar] [CrossRef]
- Shukla, S.; Pei, Y.; Li, W.-G.; Pei, D.-S. Toxicological Research on Nano and Microplastics in Environmental Pollution: Current Advances and Future Directions. Aquat. Toxicol. 2024, 270, 106894. [Google Scholar] [CrossRef]
- Chen, G.; Li, X.; Wang, Z.; Li, M.; Wang, W.; Lu, R.; Wang, S.; Li, Q.; Hu, Z.; Wu, Y.; Li, Z.; Wang, P.; Cao, Y. Human exposure to micro(nano)plastics: Health risks and analysis methods. TrAC 2024, 178, 117835. [Google Scholar] [CrossRef]
- Jiang, R.; Lu, G.; Yan, Z.; Liu, J.; Wu, D.; Wang, Y. Microplastic degradation by hydroxy-rich bismuth oxychloride. J. Hazard Mater. 2021, 405, 124247. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Yan, D.; Fu, J.; Chen, Y.; Ou, H. Ultraviolet-C and vacuum ultraviolet inducing surface degradation of microplastics. Water Res. 2020, 186, 116360. [Google Scholar] [CrossRef] [PubMed]
- Miao, F.; Liu, Y.; Gao, M.; Yu, X.; Xiao, P.; Wang, M.; Wang, S.; Wang, X. Degradation of polyvinyl chloride microplastics via an electro-Fenton-like system with a TiO2/graphite cathode. J. Hazard Mater. 2020, 399, 123023. [Google Scholar] [CrossRef]
- Pham, T.H.; Do, H.T.; Phan Thi, L.A.; Singh, P.; Raizada, P.; Chi-Sheng Wu, J.; Nguyen, V. H. Global challenges in microplastics: from fundamental understanding to advanced degradations toward sustainable strategies. Chemosphere 2021, 267, 129275. [Google Scholar] [CrossRef]
- Yuan, J.; Ma, J.; Sun, Y.; Zhou, T.; Zhao, Y.; Yu, F. Microbial degradation and other environmental aspects of microplastics/plastics. Sci. Total Environ. 2020, 715, 136968. [Google Scholar] [CrossRef]
- Nabi, I.; Bacha, A.U.R.; Li, K.; Cheng, H.; Wang, T.; Liu, Y.; Ajmal, S.; Yang, Y.; Feng, Y.; Zhang, L. Complete photocatalytic mineralization of microplastic on TiO2 nanoparticle film. iScience 2020, 23, 101326. [Google Scholar] [CrossRef]
- Uheida, A.; Mejía, H.G.; Abdel-Rehim, M.; Hamd, W.; Dutta, J. 2021. Visible light photocatalytic degradation of polypropylene microplastics in a continuous water flow system. J. Hazard Mater. 2021, 406, 124299. [Google Scholar] [CrossRef]
- Kasar, P.; Sharma, D.K.; Ahmaruzzaman, M. Thermal and catalytic decomposition of waste plastics and its co-processing with petroleum residue through pyrolysis process. J. Clean. Prod. 2020, 265, 121639. [Google Scholar] [CrossRef]
- Domínguez-Jaimes, L.P.; Cedillo-González, E.I.; Luévano-Hipólito; Acuña-Bedoya, J. D.; Hernández-López, J.M. Degradation of primary nanoplastics by photocatalysis using different anodized TiO2 structures. J. Hazard Mater. 2021, 413, 125452. [Google Scholar] [CrossRef] [PubMed]
- Chandra, P.; Singh, D.P.; Enespa. Microplastic degradation by bacteria in aquatic ecosystem. Microorganisms for Sustainable Environment and Health. Elsevier, 2020, 431–467.
- Sánchez, C. Fungal potential for the degradation of petroleum-based polymers: an overview of macro- and microplastics biodegradation. Biotechnol. Adv. 2020, 40, 107501. [Google Scholar] [CrossRef]
- Shabbir, S.; Faheem, M.; Ali, N.; Kerr, P.G.; Wang, L.F.; Kuppusamy, S.; Li, Y. 2020. Periphytic biofilm: an innovative approach for biodegradation of microplastics. Sci. Total Environ. 2020, 717, 137064. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Ma, J.; Sun, Y.; Zhou, T.; Zhao, Y.; Yu, F. Microbial degradation and other environmental aspects of microplastics/plastics. Sci. Total Environ. 2020, 715, 136968. [Google Scholar] [CrossRef] [PubMed]
- Zurier, H.S.; Goddard, J.M. Biodegradation of microplastics in food and agriculture. Curr. Opin. Food Sci. 2021, 37, 37–44. [Google Scholar] [CrossRef]
- Wang, K.; Wang, F.; Yu, Y.; Yang, S.; Han, Y.; Yao, H. Microplastics and soil microbiomes. BMC Biol. 2025, 23, 273. [Google Scholar] [CrossRef]
- Menéndez-Pedriza, A.; Jaumot, J. Interaction of environmental pollutants with microplastics: a critical review of sorption factors, bioaccumulation and ecotoxicological effects. Toxics 2020, 8, 40. [Google Scholar] [CrossRef]
- Procopio, A.C.; Soggiu, A.; Urbani, A.; Roncada, P. nteractions between microplastics and microbiota in a One Health perspective. OneHealth 2025, 20, 101002. [Google Scholar]
- Eichinger, J.; Tretola, M.; Seifer, J.; Brugger, D. Interactions between microplastics and the gastrointestinal microbiome. Ital. J. Anim. Sci. 2024, 23, 1044–1056. [Google Scholar] [CrossRef]
- Eichinger, J.; Tretolab, M.; Seifertc, J.; Bruggere, D. Review: interactions between microplastics and the gastrointestinal microbiome. Ital. J. Anim. Sci. 2024, 23, 1044–1056. [Google Scholar] [CrossRef]
- Parsaeimehr, A.; Miller, C.M.; Ozbay, G. Microplastics and their interactions with microbiota. Heliyon 2023, 9, e15104. [Google Scholar] [CrossRef]
- Sun, H.; Chen, N.; Yang, X.; Xia, Y.; Wu, D. Effects induced by polyethylene microplastics oral exposure on colon mucin release, inflammation, gut microflora composition and metabolism in mice. Ecotoxicol. Environ. Saf. 2021, 220, 112340. [Google Scholar] [CrossRef]
- Zhang, X.; Dong, Z.; Zhang, S.; Ma, J.; Liu, S. Microplastic biofilm as hotspots of antibiotic resistance genes and potential pathogens. NPJ Biofilms Microbiomes 2026, 12, ar.n. 24. [Google Scholar] [CrossRef]
- Wang, J.; Yang, Y.; Shi, Y.; Wei, L.; Gao, L.; Liu, M. Oxidized vs. unmodified polyethylene microplastics neurotoxicity via microbiota–gut–brain axis in mice. Environ. Int. 2024, 185, 108523. [Google Scholar] [CrossRef]
- Goswami, P.; Kanda, K.; Taamura-Andoh, Y.; Watanabe, M.; Guruge, K.S. Microplastics: Hidden drivers of antimicrobial resistance in aquatic systems. NanoImpact 2525, 38, 100566. [Google Scholar] [CrossRef]
- Fackelmann, G.; Sommer, S. Microplastics and the gut microbiome: How chronically exposed species may suffer from gut dysbiosis. Mar. Pollut. Bull. 2019, 143, 92–100. [Google Scholar] [CrossRef]
- Su, H.; Xu, W.; Hu, X.; Xu, Y.; Wen, G.; Cao, Y. The impact of microplastics on antibiotic resistance genes, metal resistance genes, and bacterial community in aquaculture environment. J. Hazard. Mater. 2025, 489, 137704. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhang, Z.; Hsueh, Y.; Zhang, C.; Yu, J.; Zhu, J.; Niu, J.; Yin, N.; Zhang, J.; Cui, X.; Liu, X.; Xu, K.; Yuan, Ch. Interactions between environmental pollutants and gut microbiota: a review connecting conventional heavy metals and emerging microplastics. Environ. Res. 2025, 269, 120928. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Zeng, Y.; Cai, Z.; Wu, J.; Chan, L.L.; Zhu, J.; Zhou, J. Polystyrenemicroplastics alter the intestinalmicrobiota function and the hepaticmetabolism status in marine medaka (Oryzias melastigma). Sci. Total Environ. 2021, 759, 143558. [Google Scholar] [CrossRef]
- Yu, J.; Chen, L.; Wu, B. Size-specific effects of microplastics and lead on zebrafish. Chemosphere 2023, 337, 139383. [Google Scholar] [CrossRef]
- Yang, X.Y.; Zhang, Z. W.; Chen, G.D.; Yuan, S. Gut microbiome remodeling induced by microplastic exposure in humans. Gut Microbes 2026, 18, 2617696. [Google Scholar] [CrossRef]
- Wang, J.; Guo, X.; Xue, J. Biofilm-developed microplastics as vectors of pollutants in aquatic environments. Environ Sci Technol. 2021, 55, 12780–12790. [Google Scholar] [CrossRef] [PubMed]
- Tuvo, B.; Scarpaci, M.; Bracaloni, S; Esposito, E.; Costa, A.L.; Ioppolo, M.; Casini, B. Microplastics and antibiotic resistance: the magnitude of the problem and the emerging role of hospital wastewater. Int J Environ Res Public Health 2023, 20, 5868. [Google Scholar] [CrossRef]
- Zhu, J.; Dong, X.; Zhao, N.; Jiang, S.; Jin, H. Microplastics in polystyrene-made food containers from China:abundance, shape, size, and human intake. Environ. Sci. Pollut. Res. 2023, 30, 40084–40093. [Google Scholar] [CrossRef]
- Chen, g Y.; Lu, J.; Fu, S.; Wang, S.; Senehi, N.; Yuan, Q. Enhanced propagation of intracellular and extracellular antibiotic resistance genes in municipal wastewater by microplastics. Environ Pollut. 2022, 292, 1182841052. [Google Scholar]
- Lu, J.; Zhang, Y.; Wu, J.; Luo, Y. Effects of microplastics on distribution of antibiotic resistance genes in recirculating aquaculture system. Ecotoxicol Environ Saf. 2019, 184, 109631. [Google Scholar] [CrossRef]
- Guruge, K.S.; Goswami, P.; Kanda, K.; Abeynayaka, A.; Kumagai, M.; Watanabe, M.; Tamamura-Andoh, Y. Plastiome: plastisphere-enriched mobile resistome in aquatic environments. J. Hazard. Mater. 2025, 471, 134353. [Google Scholar] [CrossRef]
- Balta, I.; Lemon, J.; Gadaj, A.; Cretescu, I.; Stef, D.; Pet, I.; Stef, L.; McCleery, D.; Douglas, A.; Corcionivoschi, N. The interplay between antimicrobial resistance, heavy metals and microplastics: mechanisms of co-selection and horizontal gene transfer. Front. Microbiol. 2025, 16, 1550587. [Google Scholar] [CrossRef]
- Li, Z.; Yuan, D. Metagenomic analysis of Reveals the Effects of Microplatics on Antibiotic Resistance genes in Sludge Anaerobic Digestion. Toxics 2024, 12, 920. [Google Scholar] [CrossRef]
- Gao, B.; Shi, X.; Li, S.; Xu, W.; Gao, N.; Shan, J.; Shen, W. Size-dependent effects of polystyrene microplastics on gut metagenome and antibiotic resistance in C57BL/6 mice. Ecotoxicol. Environ. Saf. 2023, 254, 114737. [Google Scholar] [CrossRef] [PubMed]
- Ye, G.; Li, M.; Huang, H.; Avell’an-Llaguno, R.D.; Chen, J.; Chen, G.; Huang, Q. Polystyrene microplastic exposure induces selective accumulation of antibiotic resistance genes in gut microbiota and its potential health risks. Int. J. Biol. Macromol. 2025, 309, 142983. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, L.; Wang, J.; Shao, M.; Wei, Z.; Wang, L.; Li, B.; Li, Ch.; Luo, X.; Li, F.; Zheng, H. Microplastics enhance the prevalence of antibiotic resistance genes in mariculture sediments by enriching host bacteria and promoting horizontal gene transfer. Eco Environ. Health 2025, 4, 100136. [Google Scholar] [CrossRef]
- Yin, L.; Yang, M.; Teng, A.; Ni, C.; Wang, P.; Tang, Sh. Unraveling microplastic effects on gut microbiota across Various Animals Using machine Learning. ACS Nano 2025, 19, 369–380. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.N.; Gao, C.C.; Ren, H.Y.; Wen, B.; Wang, Z.N.; Gao, J.Z.; Chen, Z.-Z. Multi-omics association pattern between gut microbiota and host metabolism of a filter-feeding fish in situ exposed to microplastics. Environ. Int. 2025, 197, 109360. [Google Scholar] [CrossRef]
- Jain, R.; Gaur, A.; Suravajhala, R.; Chauhan, U.; Pant, M.; Tripathi, V.; Pant, G. Microplastic pollution: Understanding microbial degradation and strategies for pollutant reduction. Sci. Total Environ. 2023, 905, 167098. [Google Scholar] [CrossRef]
- Barboza, L.G.A.; Lopes, C.; Oliveira, P.; Bessa, F.; Otero, V.; Henriques, B.; Raimundo, J.; Caetano, M.; Vale, C.; Guilhermino, L. Microplastics in wild fish from north East Atlantic Ocean and its potential for causing neurotoxic effects, lipid oxidative damage, and human health risks associated with ingestion exposure. Sci. Total Environ. 2020, 717, 134625. [Google Scholar] [CrossRef]
- Santiago, M.S.A.; Avellar, M.Ch.W.; Perobelli, J.E. Could the gut microbiota be capable of making individuals more or less susceptible to environmental toxicants? Toxicology 2024, 503, 153751. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Lu, B.; Liu, H.; Wu, X.; Liu, Y. Ecotoxicological impacts of microplastics to gut microbiota: response mechanisms, challenges, and environmental sustainability — A review. Ecotoxicol. Environ. Saf. 2025, 302, 118748. [Google Scholar] [CrossRef]
- Zhang, Z.; Chen, W.; Chan, H.; Peng, J.; Zhu, P.; Li, J.; Jiang, X.; Zhang, Z.; Wang, Y.; Tan, Z.; Peng, Y.; Zhang, S.; Lin, K.; Yung, K.K. -L Polystyrene microplastics induce size-dependent multi-organ damage in mice: Insights into gut microbiota and fecal metabolites. J. Hazard. Mater. 2024, 461, 132503. [Google Scholar] [CrossRef]
- Tamargo, A.; Molinero, N.; Reinosa, J.J.; Alcolea-Rodriguez, V.; Portela, R.; Bañares, M.A.; Fernández, J.F.; Moreno-SArribas, M.V. PET microplastics affect human gut microbiota communities during simulated gastrointestinal digestion, first evidence of plausible polymer biodegradation during human digestion. Sci. Rep. 2022, 12, 528. [Google Scholar] [CrossRef] [PubMed]
- Bao, L.; Cui, X.; Zeng, T.; Liu, G.; Lai, W.; Zhao, H.; Gao, F.; Wu, J.; Leong, K.W.; Chen, Ch. Incorporation of polylactic acid microplastics into the carbon cycle as a carbon source to remodel the endogenous metabolism of the gut. Proc. Natl. Acad. Sci. USA 2025, 122, e2417104122. [Google Scholar] [CrossRef]
- Farooq, U.; Muhammad, Z.; Yang, Q.; Usman, M.; Qu, Z.; Zou, B.; Liu, N. Microplastics and the gut-brain axis: unraveling neurotoxic mechanisms and health implications. Innov. Med. 2026, 4, 100190. [Google Scholar] [CrossRef]
- Snehamayee, N.; Somya, S.; Kumar, S.Ch.; Niranjan, M.; Ranjan, S.B.; Kumar, M.N. Microplastics and Human Health: A comprehensive Review on Exposure Pathways, toxicity and Emerging Risks. Microplastics 2026, 5, 8. [Google Scholar] [CrossRef]
- Demarquoy, J. Microplastics and probiotics: mechanisms of interaction and their consequences for health. AIMS Microbiol. 2025, 11, 388–409. [Google Scholar] [CrossRef]
| Category | Parameter/ Marker | Direction of Change | Description / Mechanism | Health Implications | References |
|---|---|---|---|---|---|
| Inflammatory markers |
IL-6 | Increased | Upregulation of pro-inflammatory signaling pathways induced by microplastic exposure |
Chronic inflammation, immune dysregulation | [14,60] |
| TNF-α | Increased | Activation of innate immune responses and cytokine production |
Tissue damage, inflame matory disorders | [14,21] | |
| Pro-inflammatory cytokines (general) | Increased | Shift toward pro- inflammatory cytokine profile |
Increased risk of inflame matory diseases | [21,60] | |
| Oxidative stress | Reactive oxygen species (ROS) | Increased | Microplastics induce oxidative stress at cellular level |
Cellular damage, apoptosis | [14,60] |
| Antioxidant defenses | Decreased (relative) | Imbalance or depletion of antioxidant systems |
Accumulation of oxidative damage | [21] | |
| Gut microbiota (beneficial taxa) | Lactobacillus spp. | Decreased | Disruption of gut microbial balance under MP exposure |
Reduced host protection and immune regulation | [14,60] |
| Bifidobacterium spp. | Decreased | Microplastic-induced dysbiosis | Impaired gut homeostasis | [14] | |
| Gut microbiota (opportunistic/pathogenic taxa) | Escherichia coli | Increased | Selective enrichment in altered gut environment |
Increased infection risk and inflammation | [14,60] |
| Clostridium spp. | Increased | Dysbiotic shift favoring opportunistic taxa |
Production of harmful metabolites | [60] | |
| Microbial metabolites | Short-chain fatty acids (SCFAs, e.g., butyrate) | Decreased | Loss of SCFA- producing bacteria |
Impaired gut barrier integrity and immune regulation | [14,60] |
| Other metabolites (e.g., lactate, propionate) | Altered | Microbial metabolic reprogramming |
Disrupted host–microbiome interactions | [21] | |
| Gut barrier function |
Intestinal barrier integrity | Decreased | Physical disruption and inflammation-mediated damage | Increased intestinal permeability (“leaky gut”) | [14,60] |
| Antibiotic resistance |
Antibiotic resistance genes (ARGs) | Increased | Biofilm-mediated horizontal gene transfer on MPs | Spread of antimicrobial resistance | [2,19,60] |
| Biofilm formation |
EPS production, biofilm-associated genes | Increased | MPs serve as substrate for plastisphere formation | Reservoir of pathogens and ARGs | [18,20] |
| Microbial function | Xenobiotic metabolism pathways | Increased | Adaptive response to pollutants adsorbed on MPs | Altered microbial metabolism | [7,21] |
| Systemic effects |
Immune homeostasis | Disrupted | Combined effects of dysbiosis and inflammation | Increased susceptibility to chronic diseases | [21,60] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




