4. Discussion
The achieved 100% classification accuracy on the test set and 97.6% with five-fold cross-validation convincingly demonstrate that the three-dimensional structure of hemagglutinin (HA) contains all the necessary information for discriminating influenza A virus subtypes. In contrast to existing methods (ClassyFlu [
1], INFINITy [
2]) that work exclusively with DNA or amino acid sequences, the proposed EpitopeGNN approach is the first to use a graph representation of the protein spatial fold and graph neural network mechanisms. This allows it to account not only for the local sequence but also for the global topology of residue interactions and their physicochemical properties, which is critical for understanding antigenic differences.
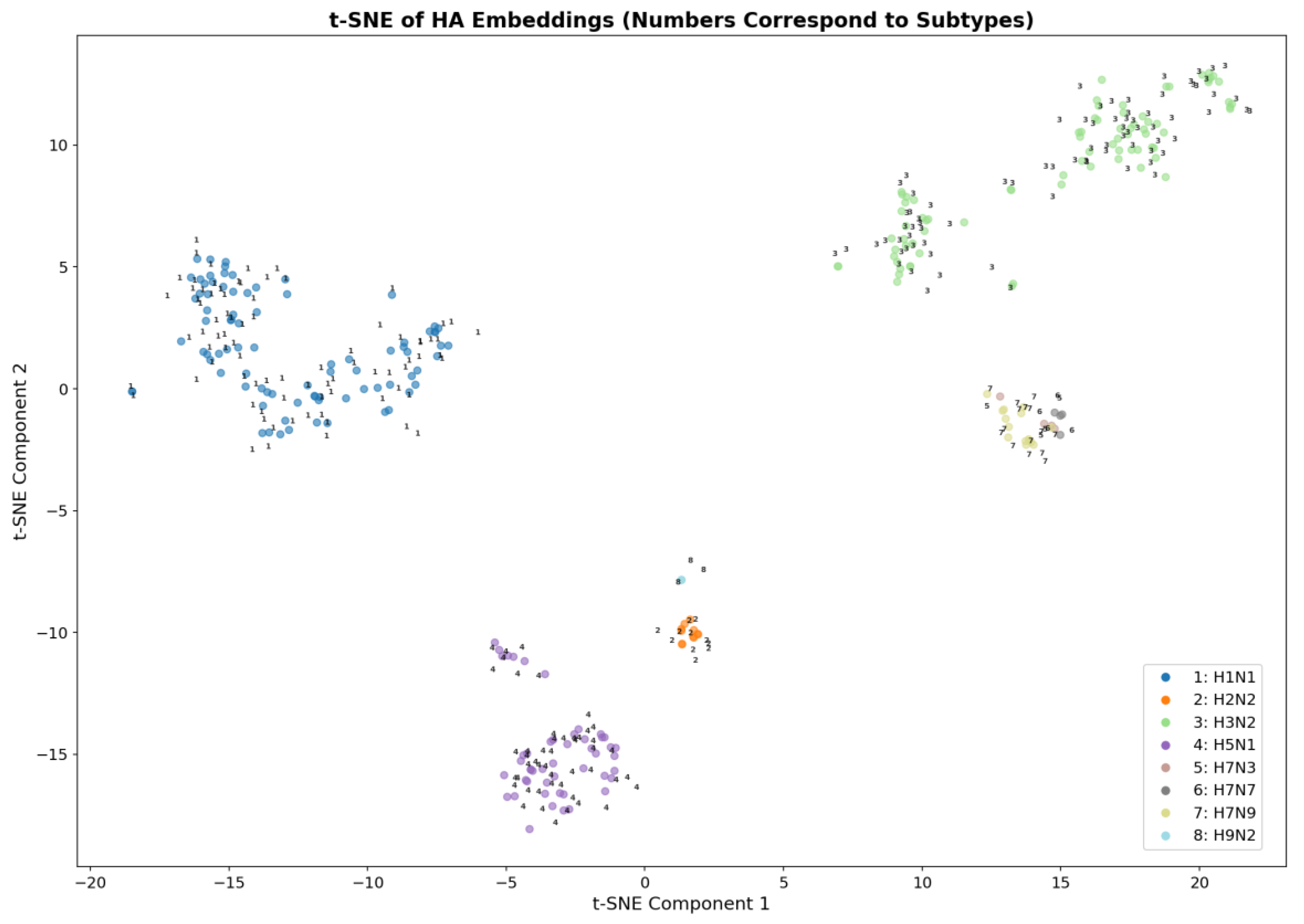
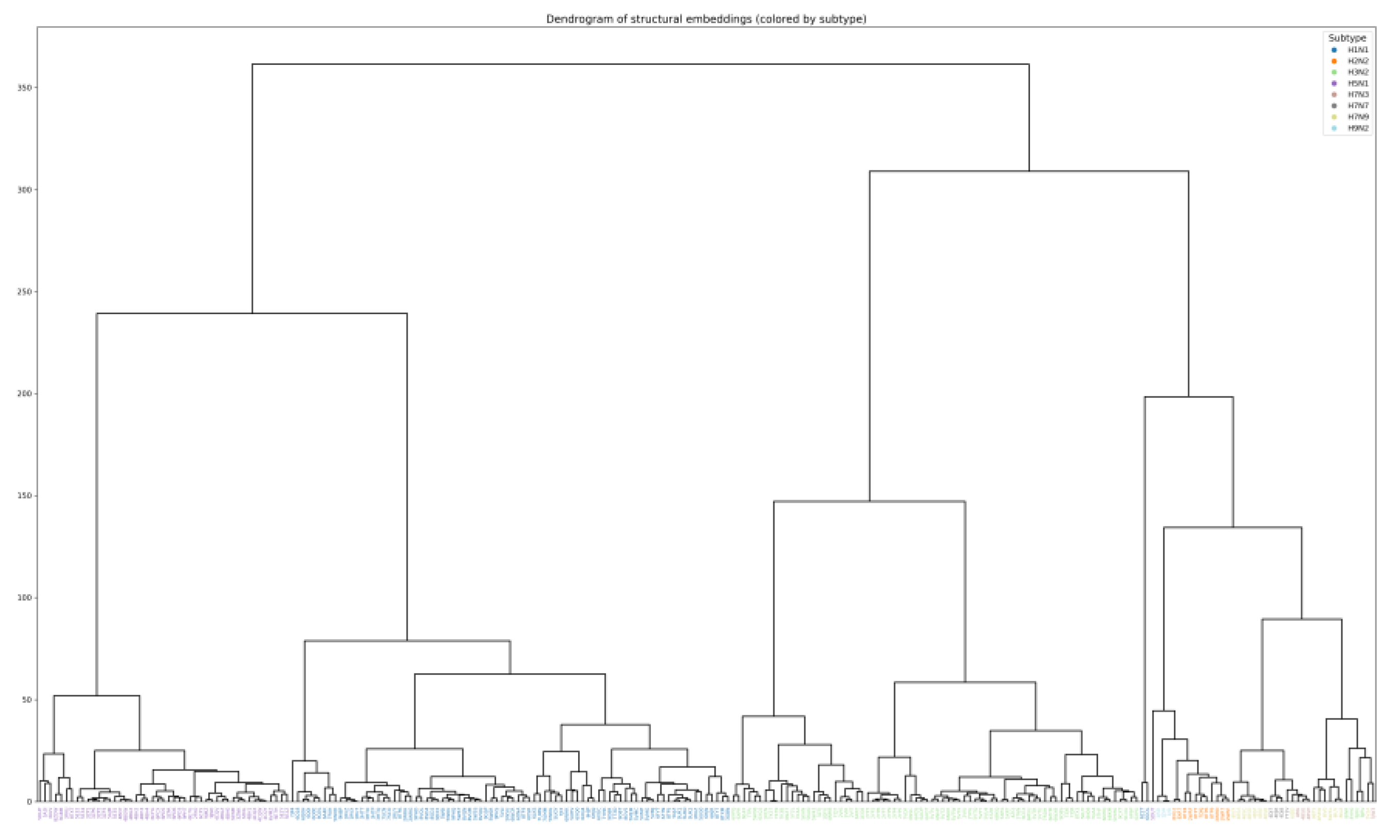
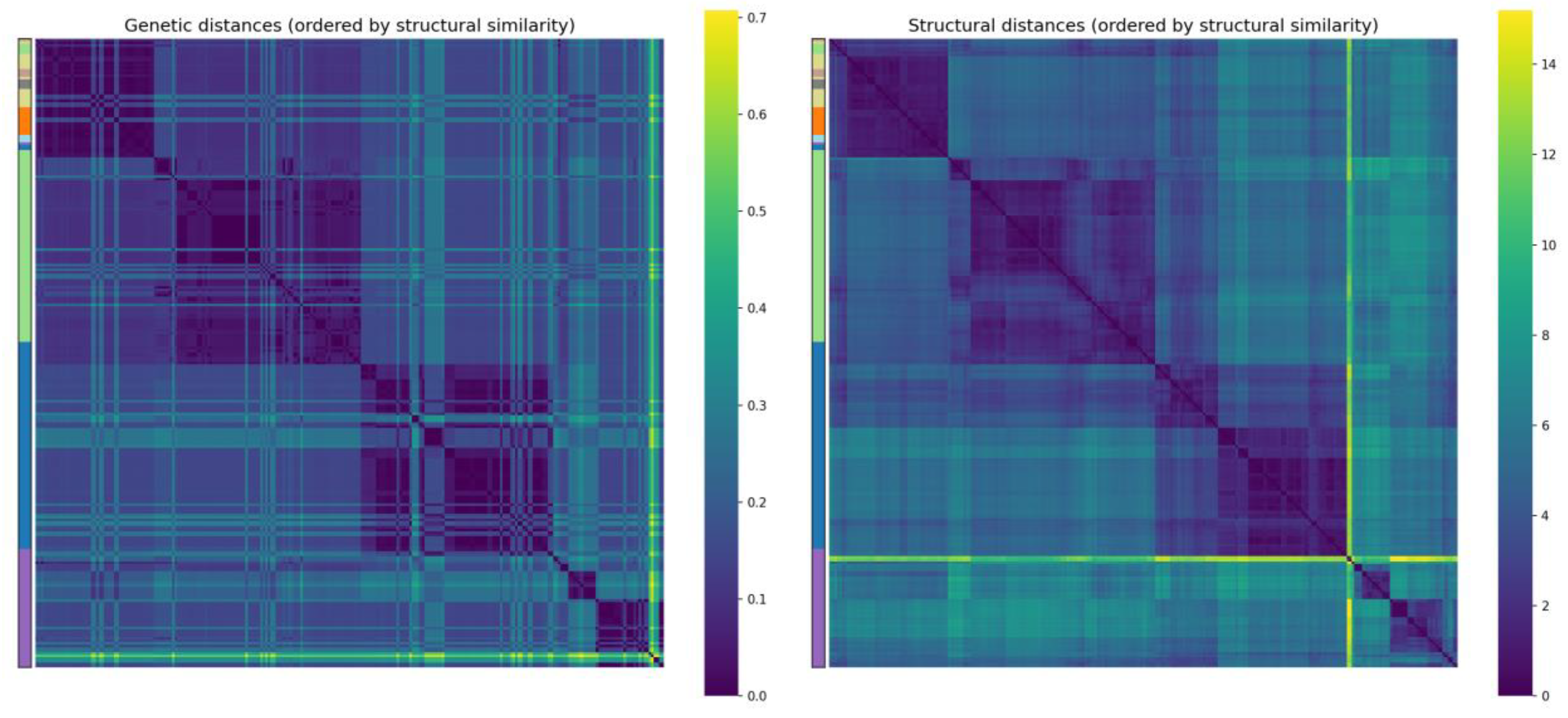
Quantitative analysis of embedding clustering confirmed high separation quality: the overall silhouette coefficient was 0.495, and for most subtypes (H1N1, H2N2, H3N2, H5N1, H9N2) the average silhouette scores exceeded 0.46, reaching 0.96 for H9N2. The proximity of the centroids of H7N3, H7N7 and H7N9 (distances 0.7–1.5) and the partial overlap of these clusters in t-SNE are explained by the small number of samples (4 and 15) and inevitable distortions upon dimensionality reduction. At the same time, the high classification accuracy and the absence of errors on the test set indicate that in the original 128-dimensional embedding space these subtypes remain completely separable. Furthermore, a significant correlation was quantitatively demonstrated between structural embeddings and phylogenetic distances, indirectly confirming the presence of an evolutionary signal in the spatial protein fold.
Comparison with existing services highlights the fundamental novelty of this work: none of them use three-dimensional structures. Thus, EpitopeGNN does not replace but complements traditional genetic classifiers by providing a fundamentally different level of information – structural. This opens new possibilities for monitoring virus evolution, enabling tracking not only of genetic changes but also of rearrangements in protein spatial folding that may affect antigenic properties and pandemic potential.
Comparison with sequence-based methods. Traditional approaches for influenza subtype classification, such as ClassyFlu [
1], INFINITy [
2], operate exclusively on nucleotide or amino acid sequences. Although they are fast and widely applicable, they cannot account for the spatial folding of the protein, which is directly linked to antigenic properties. A recent method, CLBTope [
6], also uses only sequence information to predict B-cell epitopes, but its accuracy for hemagglutinin is naturally limited because conformational epitopes are defined by spatial proximity rather than linear sequence. Our structural embedding-based approach overcomes this limitation by explicitly modeling residue interactions and surface accessibility, enabling not only accurate subtype classification but also the identification of potential antigenic determinants. This structural perspective is critical for tracking antigenic drift and for guiding vaccine strain selection.
Limitations and prospects
The model requires an experimentally determined or reliably predicted three-dimensional structure, which is not always available for new strains. However, the rapid development of protein structure prediction methods such as AlphaFold will soon remove this limitation.
Rare subtypes (H7N3, H7N7, H9N2) are represented by a small number of samples, which may affect clustering stability. Enlarging the dataset with new experimental structures or high-quality predicted models will improve classification reliability for these subtypes.
Future work will include enriching the node feature representation with evolutionary information (PSSM), as well as exploring the use of the obtained embeddings for finding structural analogs and predicting other functional properties of HA, such as receptor specificity (human α2,6 vs. avian α2,3).