Submitted:
01 April 2026
Posted:
02 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
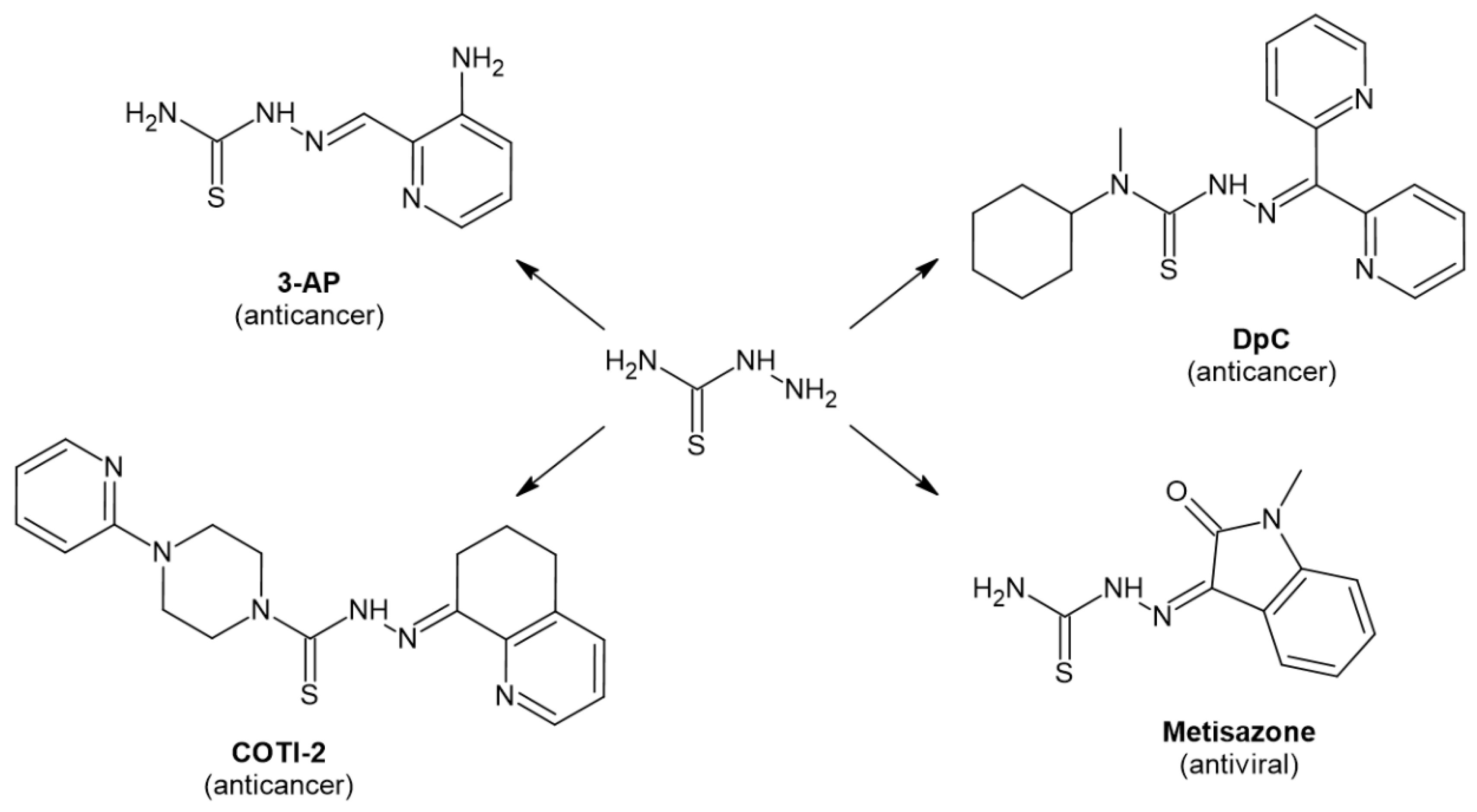
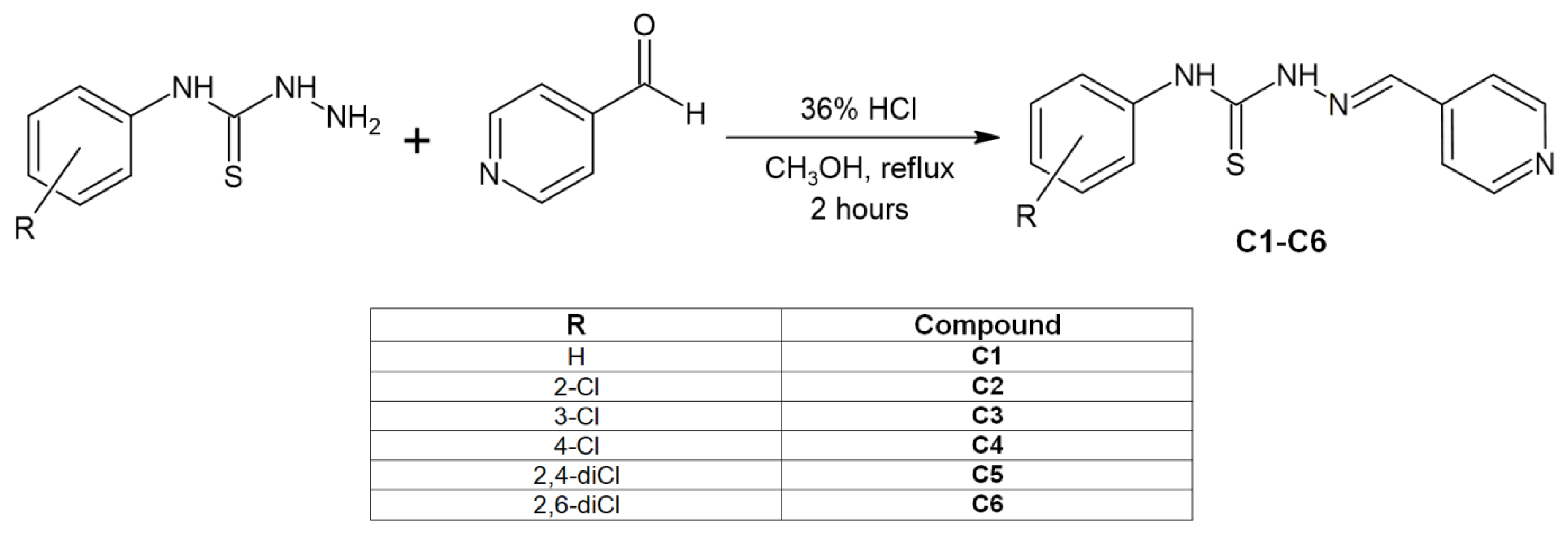
2. Results and Discussion
2.1. Synthesis and Structural Characterization
Single-Crystal X-Ray Diffraction
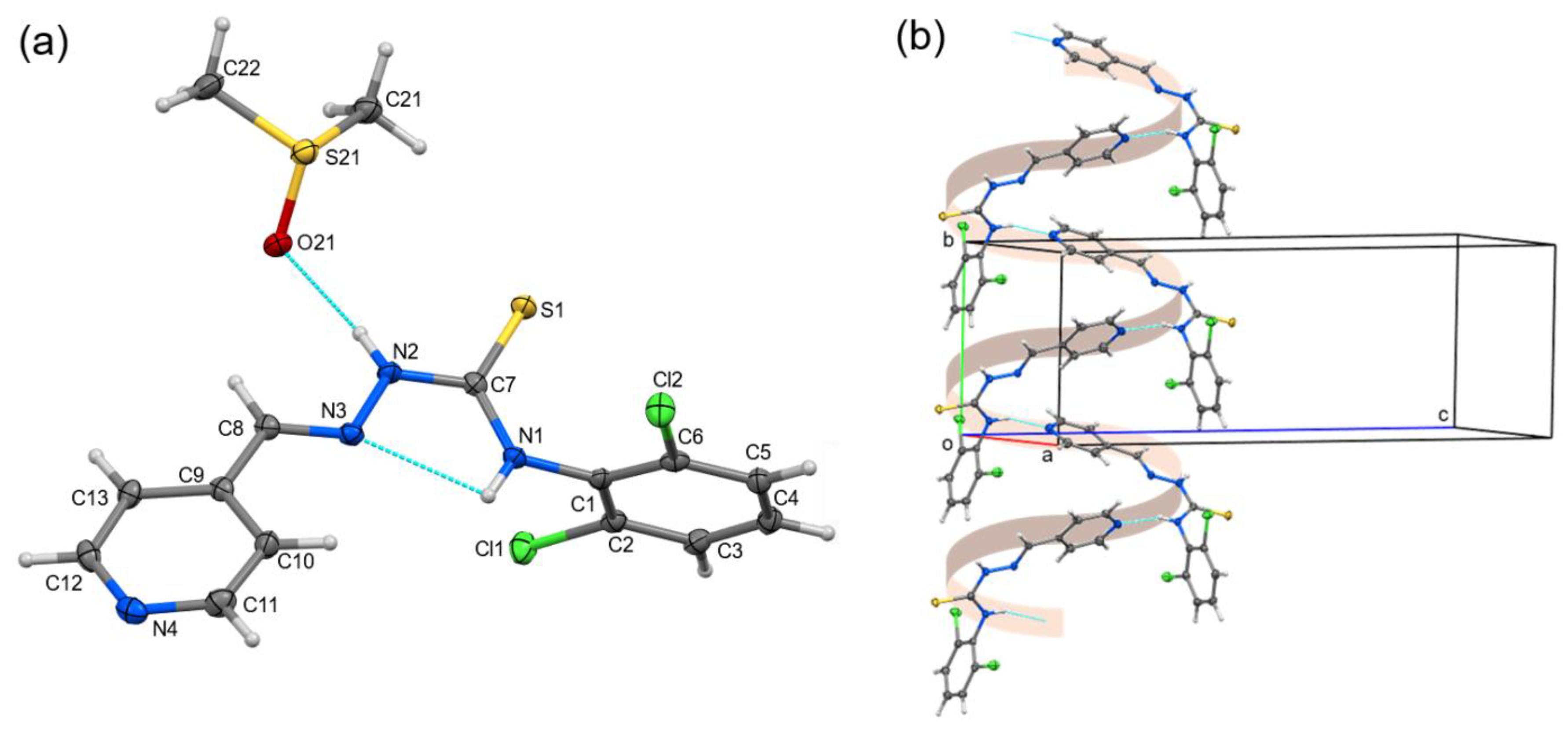
| i—j | dij (Å) | i—j—k | αijk (°) | i—j—k—l | τijkl (°) |
| C1—N1 | 1.416(3) | C1—N1—C7 | 120.5(2) | C1—N1—C7—N2 | 173.7(2) |
| N1—C7 | 1.338(3) | N1—C7—S1 | 124.02(19) | N1—C7—N2—N3 | 2.8(3) |
| C7—S1 | 1.682(3) | N1—C7—N2 | 116.6(2) | C7—N2—N3—C8 | -169.7(2) |
| C7—N2 | 1.359(3) | S1—C7—N2 | 119.40(19) | N2—N3—C8—C9 | -178.6(2) |
| N2—N3 | 1.371(3) | C7—N2—N3 | 119.2(2) | ||
| N3—C8 | 1.282(3) | N2—N3—C8 | 115.9(2) | ||
| C8—C9 | 1.470(3) | N3—C8—C9 | 119.1(2) |
| D-H•••A | d(D—H) (Å) | d(H•••A) (Å) | d(D•••A) (Å) | <(DHA) (°) | Gda(n) |
| N1—H1•••N3 | 0.82(3) | 2.31(3) | 2.627(3) | 104(3) | S(5) |
| N1—H1•••N4[i] | 0.82(3) | 2.08(3) | 2.848(3) | 157(3) | C(10) |
| N2—H2•••O21 | 0.85(3) | 1.94(3) | 2.776(3) | 169(3) | D(2) |
2.2. Antioxidant Activity
| Compound | IC50 ± SD (mg/mL) | TC* ± SD (µM/g) | |
| ABTS | DPPH | ORAC-FL | |
| C1 | 0.020 ± 0.001 | 0.043 ± 0.002 | 0.035 ± 0.001 |
| C2 | 0.035 ± 0.006 | 0.030 ± 0.001 | 0.022 ± 0.001 |
| C3 | 0.021 ± 0.003 | 0.036 ± 0.008 | 0.028 ± 0.001 |
| C4 | 0.046 ± 0.002 | 0.077 ± 0.007 | 0.022 ± 0.001 |
| C5 | 0.170 ± 0.060 | 0.043 ± 0.000 | 0.031 ± 0.001 |
| C6 | 0.016 ± 0.001 | - | 0.016 ± 0.001 |
| Trolox (STD) | 0.015 ± 0.002 | 0.013 ± 0.002 | 1.000 ± 0.000 |
2.3. Anticancer Activity
| Compound | IC50 ± SD [µM] | |||||
| A375 | G-361 | LNCaP | Caco-2 | U-87 MG | BJ | |
| C1 | >100 | >100 | 60.25 ± 0.52 | >100 | >100 | >100 |
| C2 | 76.80 ± 7.62 | 81.66 ± 2.36 | 53.85 ± 0.83 | >100 | 97.28 ± 2.79 | >100 |
| C3 | >100 | >100 | >100 | >100 | >100 | >100 |
| C4 | >100 | >100 | >100 | >100 | >100 | >100 |
| C5 | >100 | >100 | >100 | >100 | >100 | >100 |
| C6 | >100 | >100 | >100 | >100 | >100 | >100 |
2.4. Antibacterial Activity
| MIC [mg/L] | |||||||
| C1 | C2 | C3 | C4 | C5 | C6 | Reference (Van/Cip)* |
|
| Gram-negative bacteria | |||||||
| Escherichia coli ATCC 10530 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 1 |
| Salmonella Typhimurium ATCC 14028 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 1 |
| Gram-positive bacteria | |||||||
| Bacillus cereus ŁOCK 0807 | 1000 | 50 | 1000 | 50 | >1000 | >1000 | 1 |
| Bacillus subtilis ATCC 6633 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 1 |
| Enterococcus faecalis ATCC 29212 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 2 |
| Staphylococcus aureus ATCC 25923 | >1000 | >1000 | 1000 | >1000 | >1000 | >1000 | 1 |
| Staphylococcus aureus ATCC 6538 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 1 |
| Staphylococcus epidermidis ATCC 12228 | >1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 1 |
| Listeria monocytogenes ATCC 19115 | 1000 | >1000 | >1000 | >1000 | >1000 | >1000 | 2 |
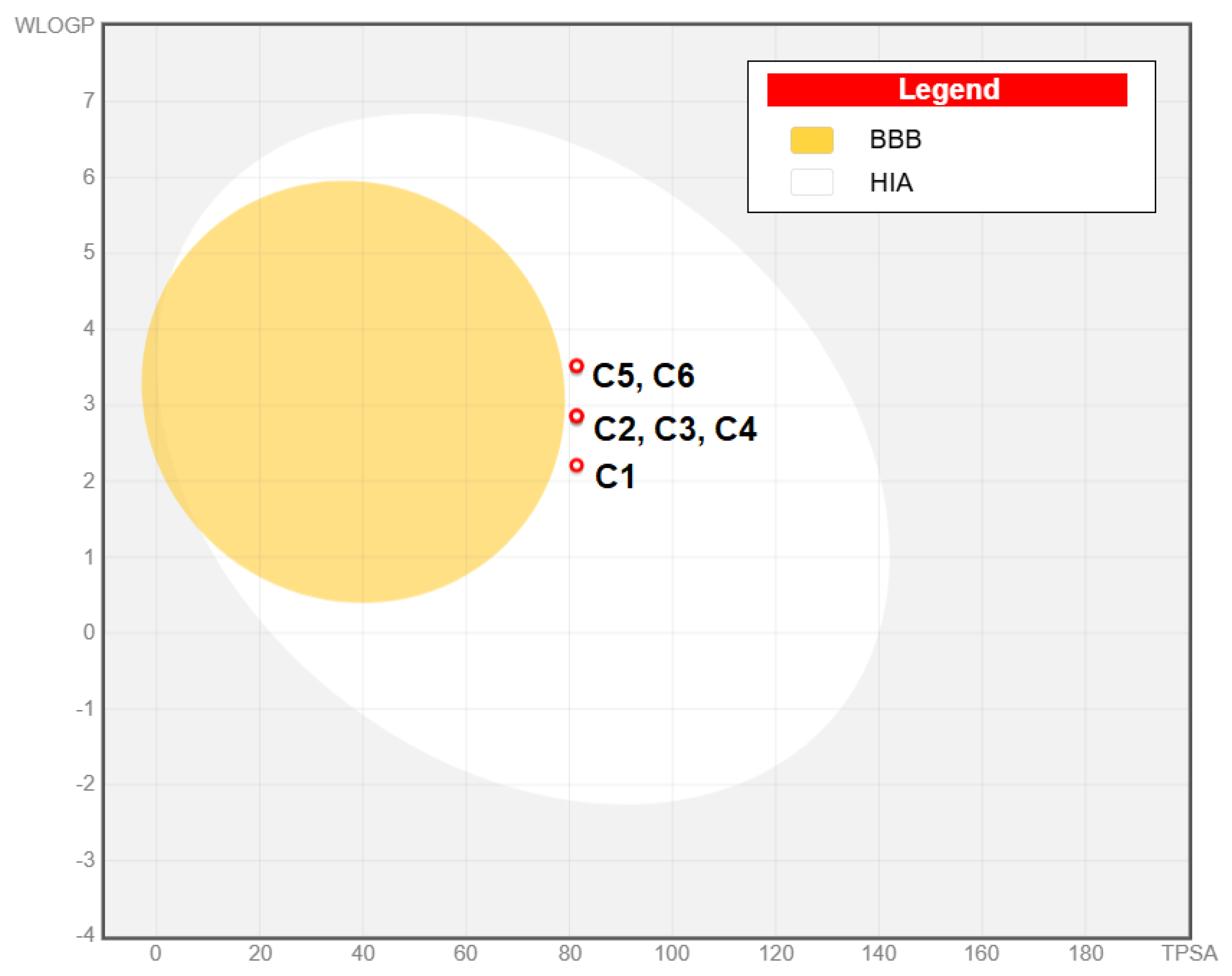
2.4. ADME Analysis
| Molecular weight [g/mol] | TPSA [Å2] | Log S | Log P | CYP1A2 inhibitor | CYP2C19 inhibitor | CYP2C9 inhibitor | CYP2D6 inhibitor | CYP3A4 inhibitor | |
| C1 | 256.33 | 81.40 | -4.83 | 2.58 | Yes | Yes | Yes | No | No |
| C2 | 290.77 | 81.40 | -4.36 | 2.73 | Yes | Yes | Yes | No | No |
| C3 | 290.77 | 81.40 | -5.45 | 3.12 | Yes | Yes | Yes | No | Yes |
| C4 | 290.77 | 81.40 | -5.45 | 3.13 | Yes | Yes | Yes | No | Yes |
| C5 | 325.22 | 81.40 | -4.97 | 3.27 | Yes | Yes | Yes | No | Yes |
| C6 | 325.22 | 81.40 | -4.97 | 3.25 | Yes | Yes | Yes | No | Yes |
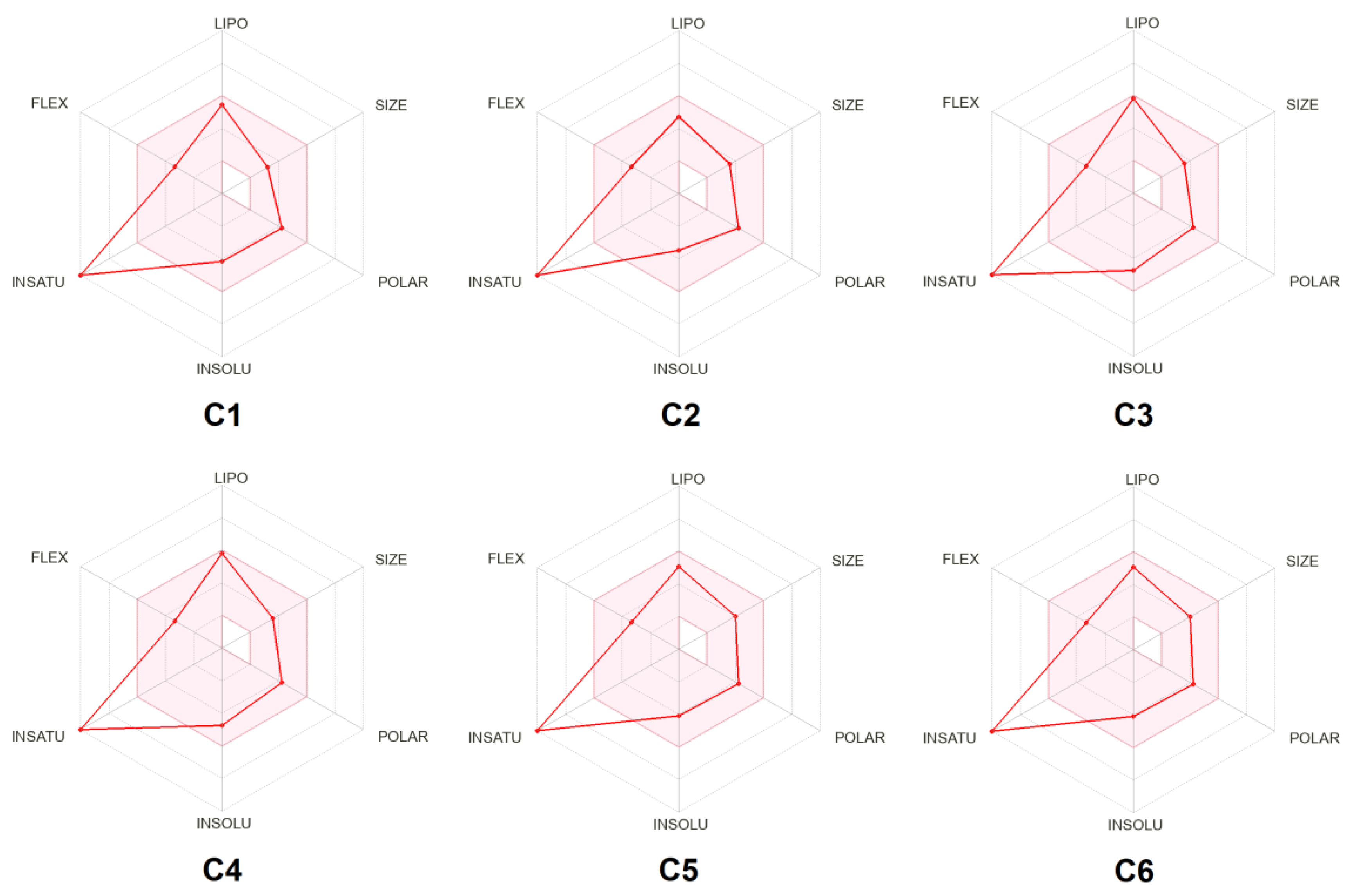
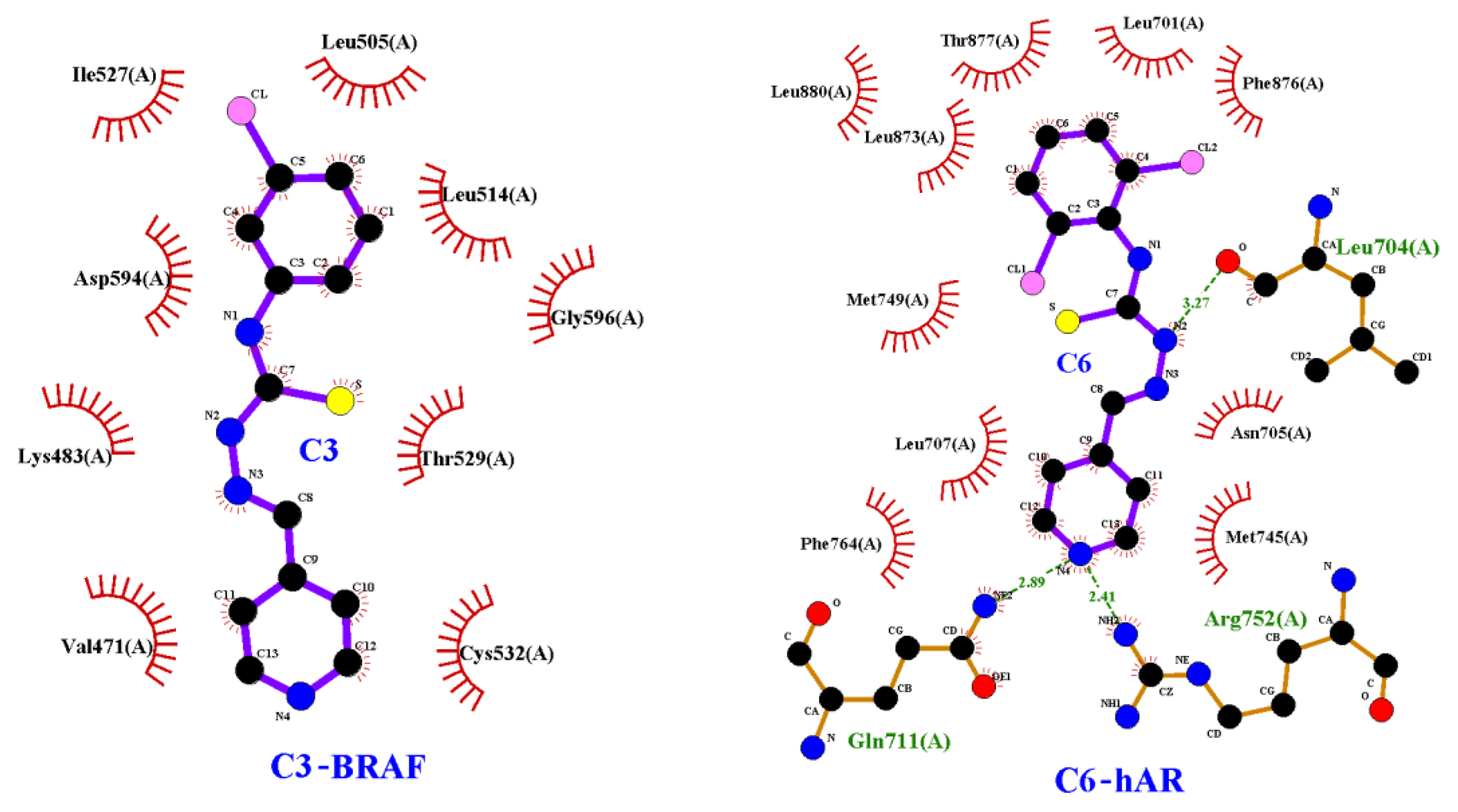
2.5. Molecular Docking
| Compound | Docking score | |||
| BRAF (3OG7) | hAR (2AM9) | EGFR (1M17) | PI3K (4JPS) | |
| C1 | 56.44 | 44.30 | 52.25 | 55.85 |
| C2 | 59.12 | 44.54 | 52.48 | 55.96 |
| C3 | 60.28 | 38.37 | 52.89 | 57.01 |
| C4 | 55.76 | 31.17 | 54.32 | 58.23 |
| C5 | 56.92 | 30.40 | 55.12 | 61.10 |
| C6 | 55.66 | 46.33 | 53.24 | 57.28 |
| Inhibitor* | 107.72 (0.4129) | 76.81 (0.4126) | 76.80 (1.9274) | 91.19 (1.1852) |
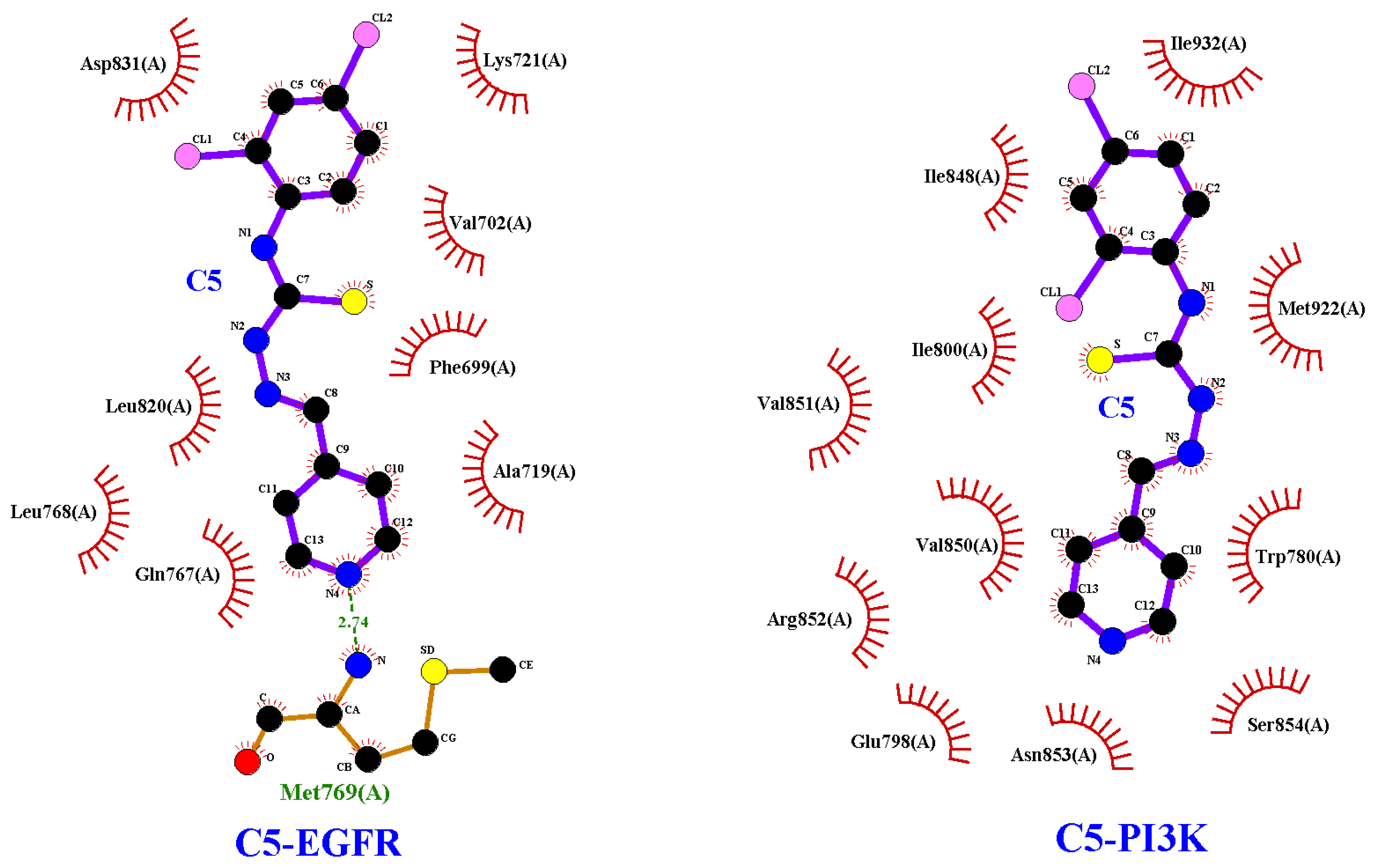
3. Materials and Methods
3.1. Chemicals and Synthesis
3.2. NMR Spectroscopy
3.3. Mass Spectrometry
3.4. Infrared Spectroscopy
3.5. Single-Crystal X-Ray Diffraction
| Empirical formula | C15H16N4OS2Cl2 |
| Formula weight | 403.34 |
| Temperature (K) | 100.00(10) |
| Crystal system | orthorhombic |
| Space group | P212121 |
| a (Å) | 8.25920(10) |
| b (Å) | 8.64490(10) |
| c (Å) | 25.9037(2) |
| α (°) | 90 |
| β (°) | 90 |
| γ (°) | 90 |
| Volume (Å3) | 1849.52(3) |
| Z | 4 |
| ρcalc (g/cm3) | 1.448 |
| μ (mm-1) | 5.358 |
| F(000) | 832.0 |
| Crystal size (mm3) | 0.14 × 0.08 × 0.06 |
| Radiation | Cu Kα (λ = 1.54184) |
| 2θ range for data collect. (°) | 6.824 to 159.422 |
| Index ranges | -7 ≤ h ≤ 10, -10 ≤ k ≤ 10, -33 ≤ l ≤ 29 |
| Reflections collected / independent | 18368 / 3838 |
| Rint | 0.0288 |
| Data/restraints/parameters | 3838/0/225 |
| Goodness-of-fit on F2 | 1.108 |
| Final R indexes [I>2s(I)] | R1 = 0.0219, wR2 = 0.0561 |
| R indexes (all data) | R1 = 0.0233, wR2 = 0.0601 |
| Largest diff. peak and hole (e•Å-3) | 0.26/-0.18 |
| Flack parameter | -0.015(9) |
3.6. Antioxidant Activity
3.7. Anticancer Activity
Cell Culture
MTT Cytotoxicity Assay
3.8. Antibacterial Activity
3.9. ADME Analysis
3.10. Molecular Docking
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hassan, A.A.; Shawky, A.M.; Shehatta, H.S. Chemistry and Heterocyclization of Thiosemicarbazones. Journal of Heterocyclic Chemistry 2012, 49, 21–37. [Google Scholar] [CrossRef]
- Prajapati, N.P.; Patel, H.D. Novel Thiosemicarbazone Derivatives and Their Metal Complexes: Recent Development. Synthetic Communications 2019, 49, 2767–2804. [Google Scholar] [CrossRef]
- Rogalewicz, B.; Świątkowski, M.; Iwan, M.; Michalczuk, M.; Kubik, J.; Korga-Plewko, A.; Pitucha, M.; Gajda, A.; Ścieszka, S.; Kordialik-Bogacka, E.; et al. Structural Studies and Biological Activity of the New Ni(II), Cu(II), Zn(II), Pd(II), and Ag(I) Thiosemicarbazone-Based Complexes. Journal of Inorganic Biochemistry 2025, 271, 112962. [Google Scholar] [CrossRef]
- Kizilcikli, İ.; Kurt, Y.D.; Akkurt, B.; Genel, A.Y.; Birteksöz, S.; Ötük, G.; Ülküseven, B. Antimicrobial Activity of a Series of Thiosemicarbazones and Their ZnII and PdII Complexes. Folia Microbiol 2007, 52, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Mbah, J.A.; Ayimele, G.A.; Eyonganyoh, E.N.; Nfor, E.N. Synthesis, Molecular Structure and Antibacterial Activity of Benzylmethyl-4-Methyl-3-Thiosemicarbazone. International Journal of Organic Chemistry 2017, 7, 83–90. [Google Scholar] [CrossRef]
- Finch, R.A.; Liu, M.; Grill, S.P.; Rose, W.C.; Loomis, R.; Vasquez, K.M.; Cheng, Y.; Sartorelli, A.C. Triapine (3-Aminopyridine-2-Carboxaldehyde- Thiosemicarbazone): A Potent Inhibitor of Ribonucleotide Reductase Activity with Broad Spectrum Antitumor Activity. Biochem Pharmacol 2000, 59, 983–991. [Google Scholar] [CrossRef]
- Finch, R.A.; Liu, M.-C.; Cory, A.H.; Cory, J.G.; Sartorelli, A.C. Triapine (3-Aminopyridine-2-Carboxaldehyde Thiosemicarbazone; 3-AP): An Inhibitor of Ribonucleotide Reductase with Antineoplastic Activity. Advances in Enzyme Regulation 1999, 39, 3–12. [Google Scholar] [CrossRef]
- Niso, M.; Kopecka, J.; Abatematteo, F.S.; Berardi, F.; Riganti, C.; Abate, C. Multifunctional Thiosemicarbazones Targeting Sigma Receptors: In Vitro and in Vivo Antitumor Activities in Pancreatic Cancer Models. Cell Oncol. 2021, 44, 1307–1323. [Google Scholar] [CrossRef]
- El Majzoub, R.; Fayyad-kazan, M.; Nasr El Dine, A.; Makki, R.; Hamade, E.; Grée, R.; Hachem, A.; Talhouk, R.; Fayyad-Kazan, H.; Badran, B. A Thiosemicarbazone Derivative Induces Triple Negative Breast Cancer Cell Apoptosis: Possible Role of miRNA-125a-5p and miRNA-181a-5p. Genes Genom 2019, 41, 1431–1443. [Google Scholar] [CrossRef]
- Bai, C.; Wu, S.; Ren, S.; Zhu, M.; Luo, G.; Xiang, H. Synthesis and Evaluation of Novel Thiosemicarbazone and Semicarbazone Analogs with Both Anti-Proliferative and Anti-Metastatic Activities against Triple Negative Breast Cancer. Bioorganic & Medicinal Chemistry 2021, 37, 116107. [Google Scholar] [CrossRef]
- Sever, B.; Çiftçi, G.A.; Özdemir, A.; Altıntop, M.D. Design, Synthesis and Biological Evaluation of New Bis(Thiosemicarbazone) Derivatives as Potential Targeted Anticancer Agents for Non-Small Cell Lung Cancer. J. Res. Pharm. 2025, 24, 670–680. [Google Scholar] [CrossRef]
- Kunos, C.A.; Radivoyevitch, T.; Waggoner, S.; Debernardo, R.; Zanotti, K.; Resnick, K.; Fusco, N.; Adams, R.; Redline, R.; Faulhaber, P.; et al. Radiochemotherapy plus 3-Aminopyridine-2-Carboxaldehyde Thiosemicarbazone (3-AP, NSC #663249) in Advanced-Stage Cervical and Vaginal Cancers. Gynecologic Oncology 2013, 130, 75–80. [Google Scholar] [CrossRef]
- Kunos, C.A.; Waggoner, S.; von Gruenigen, V.; Eldermire, E.; Pink, J.; Dowlati, A.; Kinsella, T.J. Phase I Trial of Pelvic Radiation, Weekly Cisplatin, and 3-Aminopyridine-2-Carboxaldehyde Thiosemicarbazone (3-AP, NSC #663249) for Locally Advanced Cervical Cancer. Clin Cancer Res 2010, 16, 1298–1306. [Google Scholar] [CrossRef]
- He, Z.-X.; Huo, J.-L.; Gong, Y.-P.; An, Q.; Zhang, X.; Qiao, H.; Yang, F.-F.; Zhang, X.-H.; Jiao, L.-M.; Liu, H.-M.; et al. Design, Synthesis and Biological Evaluation of Novel Thiosemicarbazone-Indole Derivatives Targeting Prostate Cancer Cells. European Journal of Medicinal Chemistry 2021, 210, 112970. [Google Scholar] [CrossRef]
- Altintop, M.D.; Sever, B.; Özdemir, A.; Kuş, G.; Oztopcu-Vatan, P.; Kabadere, S.; Kaplancikli, Z.A. Synthesis and Evaluation of Naphthalene-Based Thiosemicarbazone Derivatives as New Anticancer Agents against LNCaP Prostate Cancer Cells. Journal of Enzyme Inhibition and Medicinal Chemistry 2016, 31, 410–416. [Google Scholar] [CrossRef]
- Shahi, N.; Yadav, P.N.; Chaudhary, U.; Saad, M.; Mahiya, K.; Khan, A.; Shafi, S.; Pokharel, Y.R. 5-Methoxyisatin N(4)-Pyrrolidinyl Thiosemicarbazone (MeOIstPyrd) Restores Mutant P53 and Inhibits the Growth of Skin Cancer Cells, In Vitro. ACS Omega 2023, 8, 31998–32016. [Google Scholar] [CrossRef]
- Cancer Tomorrow. Available online: https://gco.iarc.who.int/today/ (accessed on 18 February 2026).
- Murray, C.J.L.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; Johnson, S.C.; et al. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. The Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Shakya, B.; Yadav, P.N. Thiosemicarbazones as Potent Anticancer Agents and Their Modes of Action. Mini-Reviews in Medicinal Chemistry 20 638–661. [CrossRef] [PubMed]
- Synnott, N.C.; O’Connell, D.; Crown, J.; Duffy, M.J. COTI-2 Reactivates Mutant P53 and Inhibits Growth of Triple-Negative Breast Cancer Cells. Breast Cancer Res Treat 2020, 179, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Quach, P.; Gutierrez, E.; Basha, M.T.; Kalinowski, D.S.; Sharpe, P.C.; Lovejoy, D.B.; Bernhardt, P.V.; Jansson, P.J.; Richardson, D.R. Methemoglobin Formation by Triapine, Di-2-Pyridylketone-4,4-Dimethyl-3-Thiosemicarbazone (Dp44mT), and Other Anticancer Thiosemicarbazones: Identification of Novel Thiosemicarbazones and Therapeutics That Prevent This Effect. Molecular Pharmacology 2012, 82, 105–114. [Google Scholar] [CrossRef]
- Rogalewicz, B.; Iwan, M.; Świątkowski, M.; Michalczuk, M.; Kubik, J.; Humeniuk, E.; Korga-Plewko, A.; Pitucha, M.; Boruta, T.; Ścieszka, S.; et al. Structure-Activity Relationship of Cu(II) and Pd(II) Thiosemicarbazone Complexes with Anticancer and Antibacterial Properties. ChemRxiv 2025. [Google Scholar] [CrossRef]
- Joshi, S.; Srivastava, R. Effect of “Magic Chlorine” in Drug Discovery: An in Silico Approach. RSC Adv. 2023, 13, 34922–34934. [Google Scholar] [CrossRef]
- Chiodi, D.; Ishihara, Y. “Magic Chloro”: Profound Effects of the Chlorine Atom in Drug Discovery. J. Med. Chem. 2023, 66, 5305–5331. [Google Scholar] [CrossRef]
- Vlad, I.M.; Nuță, D.C.; Ancuceanu, R.V.; Caproiou, M.T.; Dumitrascu, F.; Marinas, I.C.; Chifiriuc, M.C.; Măruţescu, L.G.; Zarafu, I.; Papacocea, I.R.; et al. New O-Aryl-Carbamoyl-Oxymino-Fluorene Derivatives with MI-Crobicidal and Antibiofilm Activity Enhanced by Combination with Iron Oxide Nanoparticles. Molecules 2021, 26, 3002. [Google Scholar] [CrossRef]
- Hassanzadeh, F.; Jafari, E.; Hakimelahi, G.H.; Khajouei, M.R.; Jalali, M.; Khodarahmi, G.A. Antibacterial, Antifungal and Cytotoxic Evaluation of Some New Quinazolinone Derivatives. Res Pharm Sci 2012, 7, 87–94. [Google Scholar]
- Demchenko, S.; Lesyk, R.; Yadlovskyi, O.; Zuegg, J.; Elliott, A.G.; Drapak, I.; Fedchenkova, Y.; Suvorova, Z.; Demchenko, A. Synthesis, Antibacterial and Antifungal Activity of New 3-Aryl-5H-Pyrrolo [1,2-a]Imidazole and 5H-Imidazo [1,2-a]Azepine Quaternary Salts. Molecules 2021, 26, 4253. [Google Scholar] [CrossRef]
- Al-Mutairi, A.A.; Al-Alshaikh, M.A.; Al-Omary, F.A.M.; Hassan, H.M.; El-Mahdy, A.M.; El-Emam, A.A. Synthesis, Antimicrobial, and Anti-Proliferative Activities of Novel 4-(Adamantan-1-Yl)-1-Arylidene-3-Thiosemicarbazides, 4-Arylmethyl N′-(Adamantan-1-Yl)Piperidine-1-Carbothioimidates, and Related Derivatives. Molecules 2019, 24, 4308. [Google Scholar] [CrossRef] [PubMed]
- Kawai, A.; Kobashigawa, Y.; Hirata, K.; Morioka, H.; Imoto, S.; Nishi, K.; Chuang, V.T.G.; Yamasaki, K.; Otagiri, M. Chlorine Atoms of an Aripiprazole Molecule Control the Geometry and Motion of Aripiprazole and Deschloro-Aripiprazole in Subdomain IIIA of Human Serum Albumin. ACS Omega 2022, 7, 29944–29951. [Google Scholar] [CrossRef] [PubMed]
- Naumann, K. Influence of Chlorine Substituents on Biological Activity of Chemicals: A Review. Pest Management Science 2000, 56, 3–21. [Google Scholar] [CrossRef]
- Naumann, K.; Cropscience, B. How Chlorine in Molecules Affects Biological Activity. 2003. [Google Scholar]
- Nagaoka, H.; Miyakoshi, T.; Kasuga, J.; Yamada, Y. Synthesis of a Halogenated Clavulone Analog. Tetrahedron Letters 1985, 26, 5053–5056. [Google Scholar] [CrossRef]
- Czylkowska, A.; Pitucha, M.; Raducka, A.; Fornal, E.; Kordialik-Bogacka, E.; Ścieszka, S.; Smoluch, M.; Burdan, F.; Jędrzejec, M.; Szymański, P. Thiosemicarbazone-Based Compounds: A Promising Scaffold for Developing Antibacterial, Antioxidant, and Anticancer Therapeutics. Molecules 2025, 30, 129. [Google Scholar] [CrossRef]
- Cornea, A.C.; Marc, G.; Ionuț, I.; Moldovan, C.; Stana, A.; Oniga, S.D.; Pîrnău, A.; Vlase, L.; Oniga, I.; Oniga, O. Synthesis, Characterization, and Antioxidant Activity Evaluation of New N-Methyl Substituted Thiazole-Derived Polyphenolic Compounds. Molecules 2025, 30, 1345. [Google Scholar] [CrossRef]
- Neacșu, S.M.; Mititelu, M.; Ozon, E.A.; Musuc, A.M.; Iuga, I.D.M.; Manolescu, B.N.; Petrescu, S.; Pandele Cusu, J.; Rusu, A.; Surdu, V.-A.; et al. Comprehensive Analysis of Novel Synergistic Antioxidant Formulations: Insights into Pharmacotechnical, Physical, Chemical, and Antioxidant Properties. Pharmaceuticals 2024, 17, 690. [Google Scholar] [CrossRef]
- Sharipova, G.M.; Safarova, I.V.; Khairullina, V.R.; Gerchikov, A.Y.; Zimin, Y.S.; Savchenko, R.G.; Limantseva, R.M. Kinetics and Mechanism of Antioxidant Action of Polysubstituted Tetrahydroquinolines in Liquid-Phase Oxidation Reactions of Organic Compounds by Oxygen. International Journal of Chemical Kinetics 2022, 54, 435–443. [Google Scholar] [CrossRef]
- Olicheva, V.; Beloborodov, V.; Sharifi, S.; Dubrovskaya, A.; Zhevlakova, A.; Selivanova, I.; Ilyasov, I. Dihydroquercetin and Related Flavonoids in Antioxidant Formulations with α-Tocopherol. International Journal of Molecular Sciences 2025, 26, 5659. [Google Scholar] [CrossRef]
- Luo, M.; Zhou, L.; Huang, Z.; Li, B.; Nice, E.C.; Xu, J.; Huang, C. Antioxidant Therapy in Cancer: Rationale and Progress. Antioxidants 2022, 11, 1128. [Google Scholar] [CrossRef] [PubMed]
- Antioxidants and Cancer | Health and Medicine | Research Starters | EBSCO Research. Available online: https://www.ebsco.com (accessed on 17 February 2026).
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and Computational Approaches to Estimate Solubility and Permeability in Drug Discovery and Development settings1PII of Original Article: S0169-409X(96)00423-1. The Article Was Originally Published in Advanced Drug Delivery Reviews 23 (1997) 3–25.1. Advanced Drug Delivery Reviews 2001, 46, 3–26. [Google Scholar] [CrossRef] [PubMed]
- Ghose, A.K.; Viswanadhan, V.N.; Wendoloski, J.J. A Knowledge-Based Approach in Designing Combinatorial or Medicinal Chemistry Libraries for Drug Discovery. 1. A Qualitative and Quantitative Characterization of Known Drug Databases. J. Comb. Chem. 1999, 1, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Veber, D.F.; Johnson, S.R.; Cheng, H.-Y.; Smith, B.R.; Ward, K.W.; Kopple, K.D. Molecular Properties That Influence the Oral Bioavailability of Drug Candidates. J. Med. Chem. 2002, 45, 2615–2623. [Google Scholar] [CrossRef]
- Egan, W.J.; Merz, Kenneth M.; Baldwin, J.J. Prediction of Drug Absorption Using Multivariate Statistics. J. Med. Chem. 2000, 43, 3867–3877. [Google Scholar] [CrossRef] [PubMed]
- Muegge, I.; Heald, S.L.; Brittelli, D. Simple Selection Criteria for Drug-like Chemical Matter. J. Med. Chem. 2001, 44, 1841–1846. [Google Scholar] [CrossRef]
- Kozyra, P.; Pitucha, M. Revisiting the Role of B-RAF Kinase as a Therapeutic Target in Melanoma. Curr Med Chem 2024, 31, 2003–2020. [Google Scholar] [CrossRef]
- Fujita, K.; Nonomura, N. Role of Androgen Receptor in Prostate Cancer: A Review. World J Mens Health 2019, 37, 288–295. [Google Scholar] [CrossRef]
- Xu, H.; Zong, H.; Ma, C.; Ming, X.; Shang, M.; Li, K.; He, X.; Du, H.; Cao, L. Epidermal Growth Factor Receptor in Glioblastoma. Oncol Lett 2017, 14, 512–516. [Google Scholar] [CrossRef]
- He, Y.; Sun, M.M.; Zhang, G.G.; Yang, J.; Chen, K.S.; Xu, W.W.; Li, B. Targeting PI3K/Akt Signal Transduction for Cancer Therapy. Sig Transduct Target Ther 2021, 6, 425. [Google Scholar] [CrossRef]
- Agilent. CrysAlis PRO; Agilent Technologies Ltd.: Yarnton, Oxfordshire, England, 2014. [Google Scholar]
- Sheldrick, G.M. SHELXT – Integrated Space-Group and Crystal-Structure Determination. Acta Cryst A 2015, 71, 3–8. [Google Scholar] [CrossRef]
- Sheldrick, G.M. Crystal Structure Refinement with SHELXL. Acta Cryst C 2015, 71, 3–8. [Google Scholar] [CrossRef]
- Dolomanov, O.V.; Bourhis, L.J.; Gildea, R.J.; Howard, J. a. K.; Puschmann, H. OLEX2: A Complete Structure Solution, Refinement and Analysis Program. J Appl Cryst 2009, 42, 339–341. [Google Scholar] [CrossRef]
- Macrae, C.F.; Sovago, I.; Cottrell, S.J.; Galek, P.T.A.; McCabe, P.; Pidcock, E.; Platings, M.; Shields, G.P.; Stevens, J.S.; Towler, M.; et al. Mercury 4.0: From Visualization to Analysis, Design and Prediction. J Appl Cryst 2020, 53, 226–235. [Google Scholar] [CrossRef] [PubMed]
- Spek, A.L. Structure Validation in Chemical Crystallography. Acta Cryst D 2009, 65, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Yoon, J. UV Direct Photolysis of 2,2′-Azino-Bis(3-Ethylbenzothiazoline-6-Sulfonate) (ABTS) in Aqueous Solution: Kinetics and Mechanism. Journal of Photochemistry and Photobiology A: Chemistry 2008, 197, 232–238. [Google Scholar] [CrossRef]
- Sharma, O.P.; Bhat, T.K. DPPH Antioxidant Assay Revisited. Food Chemistry 2009, 113, 1202–1205. [Google Scholar] [CrossRef]
- Maciejewska, K.; Czarnecka, K.; Kręcisz, P.; Niedziałek, D.; Wieczorek, G.; Skibiński, R.; Szymański, P. Novel Cyclopentaquinoline and Acridine Analogs as Multifunctional, Potent Drug Candidates in Alzheimer’s Disease. International Journal of Molecular Sciences 2022, 23, 5876. [Google Scholar] [CrossRef]
- Giske, C.G.; Turnidge, J.; Cantón, R.; Kahlmeter, G. Update from the European Committee on Antimicrobial Susceptibility Testing (EUCAST). J Clin Microbiol 60 e00276-21. [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci Rep 2017, 7, 42717. [Google Scholar] [CrossRef]
- Daina, A.; Zoete, V. A BOILED-Egg To Predict Gastrointestinal Absorption and Brain Penetration of Small Molecules. ChemMedChem 2016, 11, 1117–1121. [Google Scholar] [CrossRef]
- Jones, G.; Willett, P.; Glen, R.C.; Leach, A.R.; Taylor, R. Development and Validation of a Genetic Algorithm for Flexible docking11Edited by F. E. Cohen. Journal of Molecular Biology 1997, 267, 727–748. [Google Scholar] [CrossRef]
- Verdonk, M.L.; Cole, J.C.; Hartshorn, M.J.; Murray, C.W.; Taylor, R.D. Improved Protein–Ligand Docking Using GOLD. Proteins: Structure, Function, and Bioinformatics 2003, 52, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Groom, C.R.; Bruno, I.J.; Lightfoot, M.P.; Ward, S.C. The Cambridge Structural Database. Acta Cryst B 2016, 72, 171–179. [Google Scholar] [CrossRef]
- Bollag, G.; Hirth, P.; Tsai, J.; Zhang, J.; Ibrahim, P.N.; Cho, H.; Spevak, W.; Zhang, C.; Zhang, Y.; Habets, G.; et al. Clinical Efficacy of a RAF Inhibitor Needs Broad Target Blockade in BRAF-Mutant Melanoma. Nature 2010, 467, 596–599. [Google Scholar] [CrossRef]
- Pereira de Jésus-Tran, K.; Côté, P.-L.; Cantin, L.; Blanchet, J.; Labrie, F.; Breton, R. Comparison of crystal structures of human androgen receptor ligand-binding domain complexed with various agonists reveals molecular determinants responsible for binding affinity. Protein Science 2006, 15, 987–999. [Google Scholar] [CrossRef]
- Stamos, J.; Sliwkowski, M.X.; Eigenbrot, C. Structure of the Epidermal Growth Factor Receptor Kinase Domain Alone and in Complex with a 4-Anilinoquinazoline Inhibitor*. Journal of Biological Chemistry 2002, 277, 46265–46272. [Google Scholar] [CrossRef]
- Huet, J.; Teinkela Mbosso, E.J.; Soror, S.; Meyer, F.; Looze, Y.; Wintjens, R.; Wohlkönig, A. High-Resolution Structure of a Papaya Plant-Defence Barwin-like Protein Solved by in-House Sulfur-SAD Phasing. Acta Cryst D 2013, 69, 2017–2026. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Cheeseman, J.R.; Scalmani, G.; Caricato, M.; Hratchian, H.P.; Li, X.; Barone, V.; Bloino, J.; Zheng, G. Gaussian 09.
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple Ligand–Protein Interaction Diagrams for Drug Discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef] [PubMed]
- Wallace, A.C.; Laskowski, R.A.; Thornton, J.M. LIGPLOT: A Program to Generate Schematic Diagrams of Protein-Ligand Interactions. Protein Engineering, Design and Selection 1995, 8, 127–134. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




