Submitted:
31 March 2026
Posted:
02 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset and Targeted Markers
2.2. Pipeline Implementation
2.3. Classifiers Evaluation
3. Results
4. Discussion
5. Conclusions
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AUC | Area Under the Curve |
| CBC | Complete Blood Cell Count |
| DOAJ | Directory of Open Access Journals |
| HB | Hemoglobin |
| HT | Hematrocrit |
| LD | Linear dichroism |
| LIS | Laboratory Information System |
| MCH | Mean Corpuscular Hemoglobin |
| MCHC | Mean Corpuscular Hemoglobin Concentration |
| MCV | Mean Corpuscular Volume |
| MDPI | Multidisciplinary Digital Publishing Institute |
| NPV | Negative Predictive Value |
| PPV | Positive Predictive Value |
| RBC | Red Blood Cell |
| RDW | red blood cell distribution width |
| ROC | Receiver Operating Characteristics |
| TLA | Three letter acronym |
References
- Hercberg, S; Preziosi, P; Galan, P. Iron deficiency in Europe. Public Health Nutrition 2001, 4(2b), 537–545. [Google Scholar] [CrossRef] [PubMed]
- Girelli, D; Marchi, G; Camaschella, C. Anemia in the Elderly. Hemasphere Published. 2018, 2(3), e40. [Google Scholar] [CrossRef] [PubMed]
- Wang, D; Sra, M; Glaeser-Khan, S; et al. Cost-Effectiveness of Ferritin Screening Thresholds for Iron Deficiency in Reproductive-Age Women. Am J Hematol. 2025, 100(7), 1132–1140. [Google Scholar] [CrossRef] [PubMed]
- Sholzberg, M; Hillis, C; Crowther, M; Selby, R. Diagnosis and management of iron deficiency in females. CMAJ. Published. 2025, 197(24), E680–E687. [Google Scholar] [CrossRef] [PubMed]
- Lippi, G; Plebani, M. Lights and shadows of artificial intelligence in laboratory medicine. Adv Lab Med. Published. 2025, 6(1), 1–3. [Google Scholar] [CrossRef] [PubMed]
- Rabbani, N; Kim, GYE; Suarez, CJ; Chen, JH. Applications of machine learning in routine laboratory medicine: Current state and future directions. Clin Biochem. 2022, 103, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Ning, W; Wang, Z; Gu, Y; et al. Machine learning models based on routine blood and biochemical test data for diagnosis of neurological diseases. Sci Rep. Published. 2025, 15(1), 27857. [Google Scholar] [CrossRef] [PubMed]
- Negrini, D; Zecchin, P; Ruzzenente, A; et al. Machine Learning Model Comparison in the Screening of Cholangiocarcinoma Using Plasma Bile Acids Profiles. Diagnostics (Basel) Published. 2020, 10(8), 551. [Google Scholar] [CrossRef] [PubMed]
- Farrell, CJ; Makuni, C; Keenan, A; Maeder, E; Davies, G; Giannoutsos, J. A Machine Learning Model for the Routine Detection of "Wrong Blood in Complete Blood Count Tube" Errors. Clin Chem. 2023, 69(9), 1031–1037. [Google Scholar] [CrossRef] [PubMed]
- Spies, NC; Hubler, Z; Azimi, V; et al. Automating the Detection of IV Fluid Contamination Using Unsupervised Machine Learning. Clin Chem. 2024, 70(2), 444–452. [Google Scholar] [CrossRef] [PubMed]
- Auerbach, M; Adamson, JW. How we diagnose and treat iron deficiency anemia. Am J Hematol 2016, 91(1), 31–8. [Google Scholar] [CrossRef] [PubMed]
- Cook, JD. Clinical evaluation of iron deficiency. Semin Hematol 1982, 19(1), 6–18. [Google Scholar] [PubMed]
- Kuhn, M. Building Predictive Models in R Using the caret Package. J Stat Soft 2008, 28(5), 1–26. [Google Scholar] [CrossRef]
- Robin, X; Turck, N; Hainard, A; Tiberti, N; Lisacek, F; Sanchez, JC; Müller, M. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 2011, 12, 77. [Google Scholar] [CrossRef] [PubMed]
- DeLong, ER; DeLong, DM; Clarke-Pearson, DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 1988, 44(3), 837–45. [Google Scholar] [CrossRef] [PubMed]
- Rashidi, HH; Tran, N; Albahra, S; Dang, LT. Machine learning in health care and laboratory medicine: General overview of supervised learning and Auto-ML. Int J Lab Hematol. 2021, 43 Suppl 1, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Tepakhan, W; Srisintorn, W; Penglong, T; Saelue, P. Machine learning approach for differentiating iron deficiency anemia and thalassemia using random forest and gradient boosting algorithms. Sci Rep. Published. 2025, 15(1), 16917. [Google Scholar] [CrossRef] [PubMed]
- Barton, JC; Barton, JC; Acton, RT. Factors related to mean corpuscular volume in HFE p.C282Y homozygotes. EJHaem 2024, 6(1), e1063. [Google Scholar] [CrossRef] [PubMed]
- Garduno-Rapp, NE; Ng, YS; Weon, JL; Saleh, SN; Lehmann, CU; Tian, C; Quinn, A. Early identification of patients at risk for iron-deficiency anemia using deep learning techniques. Am J Clin Pathol 2024, 162(3), 243–251. [Google Scholar] [CrossRef] [PubMed]
- Efros, O; Soffer, S; Mudrik, A; et al. Predictive machine-learning model for screening iron deficiency without anaemia: a retrospective cohort study. BMJ Open. Published. 2025, 15(8), e097016. [Google Scholar] [CrossRef]
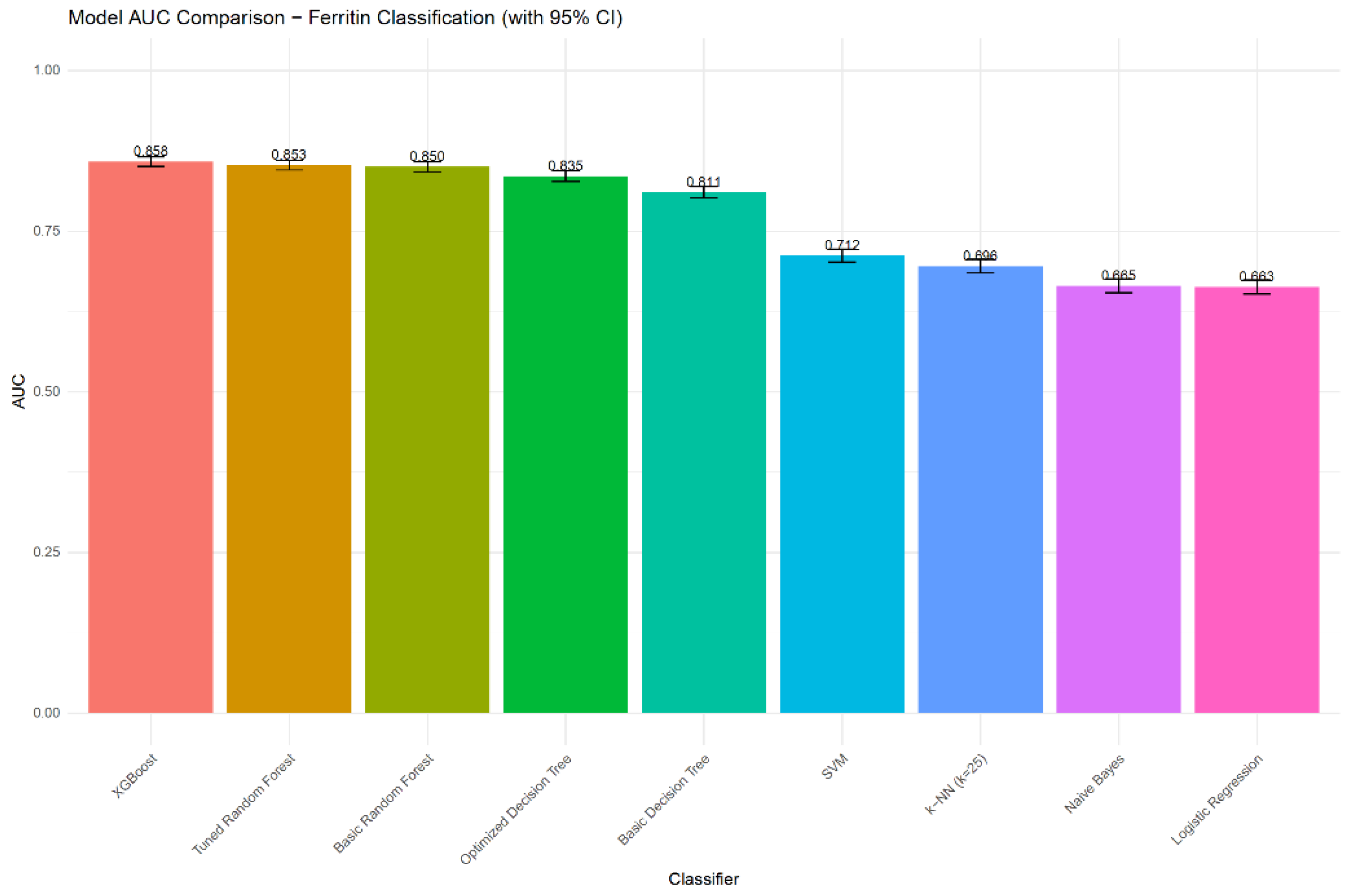
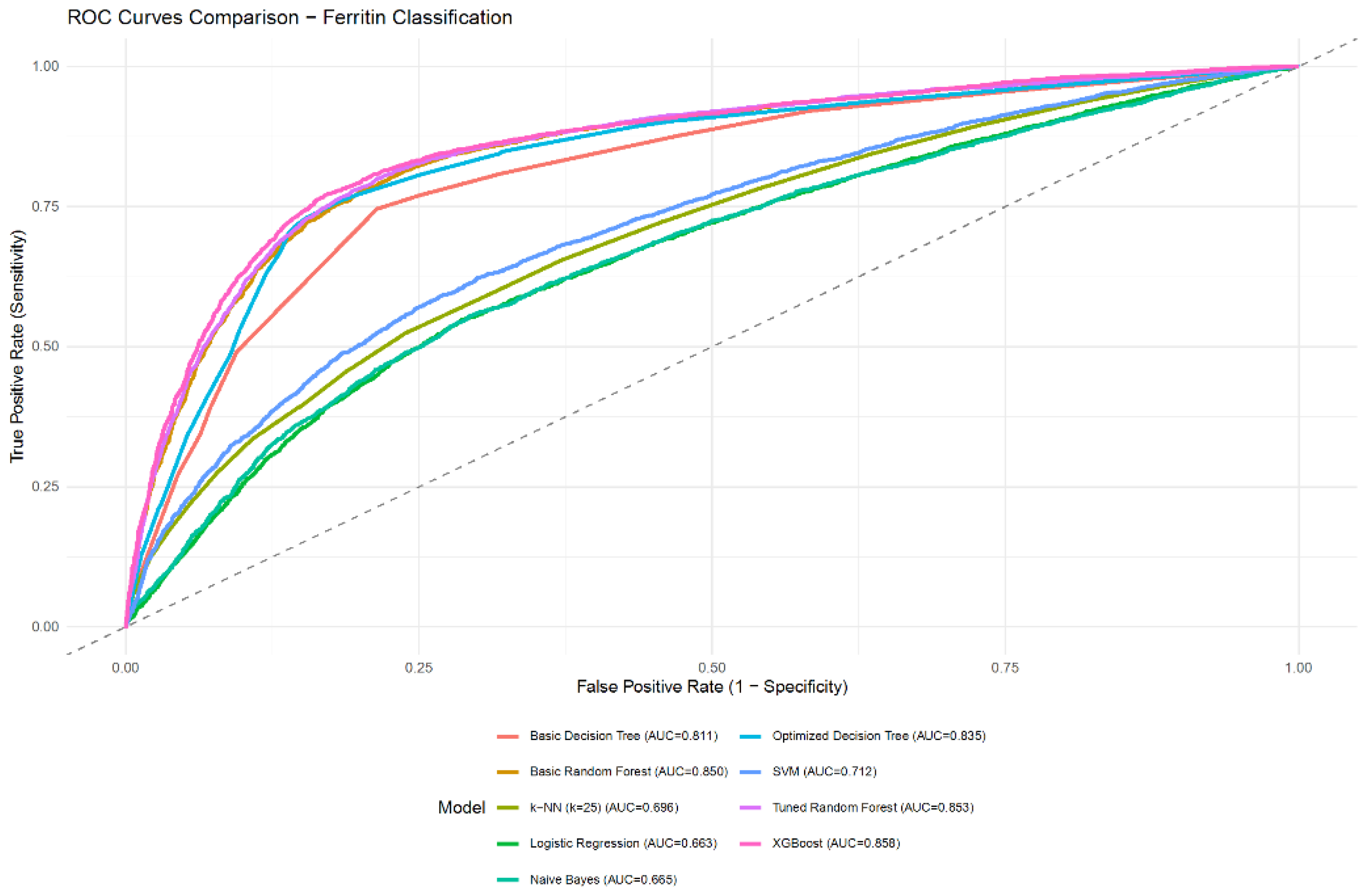
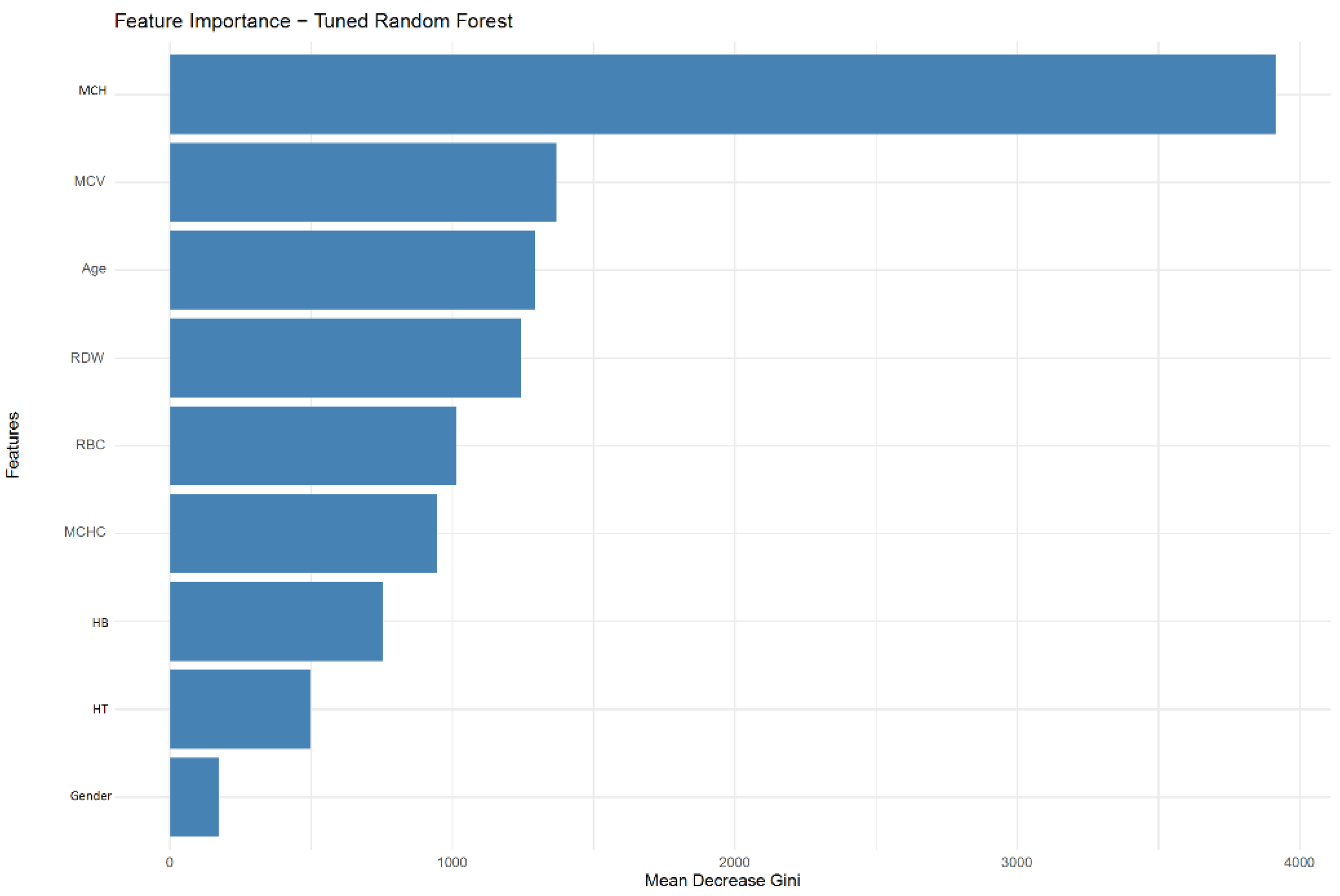
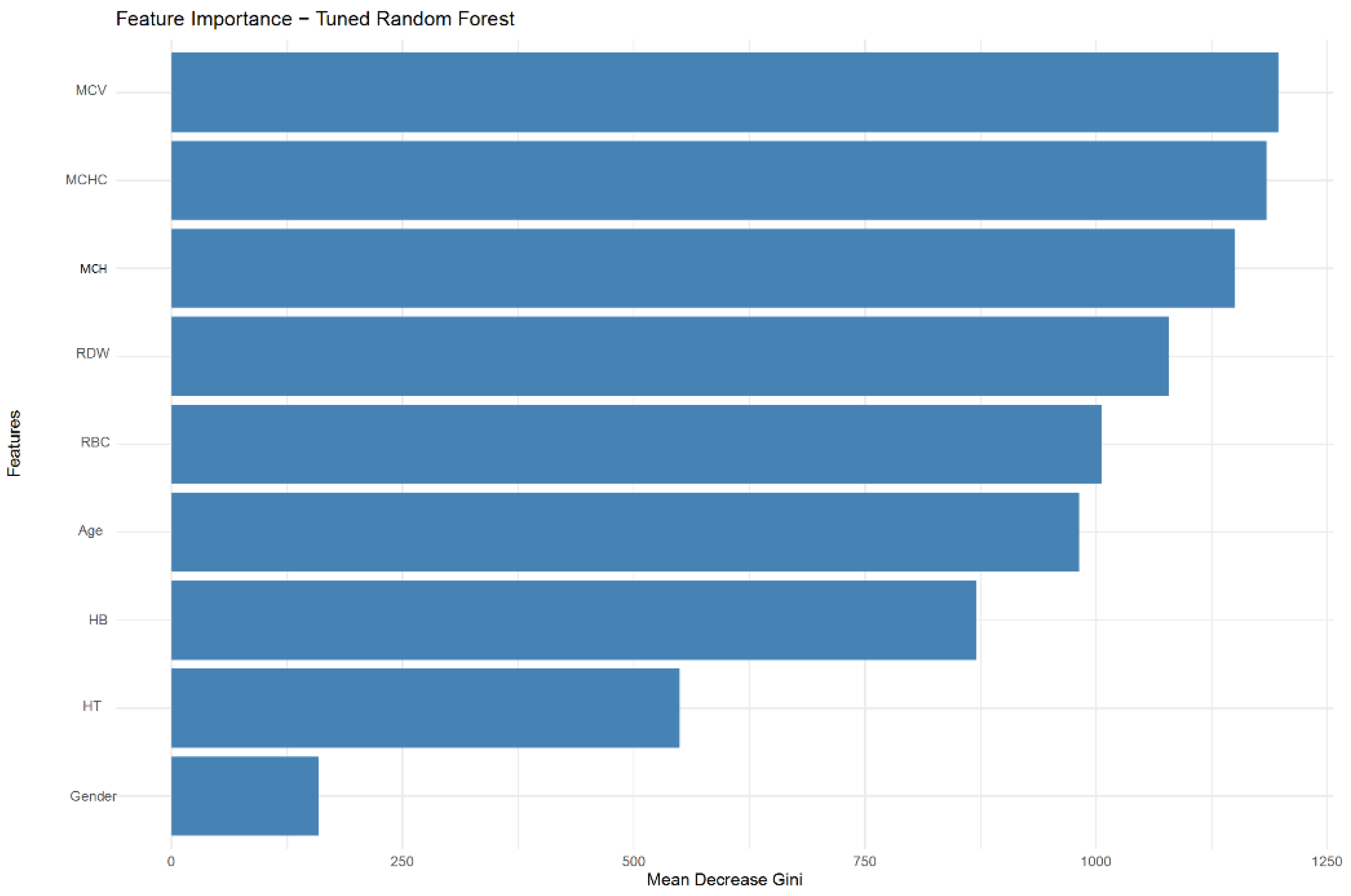
| Model | Accuracy | Balanced accuracy | Sensitivity | Specificity | PPV | NPV | F1-score | AUC | AUC 95% CI |
|---|---|---|---|---|---|---|---|---|---|
| Logistic Regression | 0.6383 | 0.6196 | 0.4578 | 0.7815 | 0.6242 | 0.6451 | 0.5282 | 0.6634 | 0.6525 - 0.6742 |
| Basic Decision Tree | 0.7681 | 0.7658 | 0.7458 | 0.7859 | 0.7342 | 0.7959 | 0.7399 | 0.8107 | 0.8021 - 0.8194 |
| Optimized Decision Tree | 0.7947 | 0.7880 | 0.7300 | 0.8460 | 0.7898 | 0.7980 | 0.7587 | 0.8354 | 0.8271 - 0.8436 |
| Basic Random Forest | 0.7902 | 0.7851 | 0.7402 | 0.8299 | 0.7753 | 0.8011 | 0.7573 | 0.8500 | 0.8422 - 0.8578 |
| Tuned Random Forest | 0.7941 | 0.7893 | 0.7476 | 0.8310 | 0.7782 | 0.8059 | 0.7626 | 0.8529 | 0.8451 - 0.8606 |
| XGBoost | 0.8016 | 0.7962 | 0.7495 | 0.8430 | 0.7910 | 0.8093 | 0.7697 | 0.8584 | 0.8507 - 0.8660 |
| SVM | 0.6689 | 0.6514 | 0.5006 | 0.8023 | 0.6675 | 0.6695 | 0.5721 | 0.7119 | 0.7016 - 0.7221 |
| k-NN (tuned as k=25) | 0.6563 | 0.6428 | 0.5254 | 0.7601 | 0.6346 | 0.6689 | 0.5749 | 0.6956 | 0.6852 - 0.7061 |
| Naive Bayes | 0.6405 | 0.6241 | 0.4822 | 0.7660 | 0.6203 | 0.6511 | 0.5426 | 0.6649 | 0.6541 - 0.6758 |
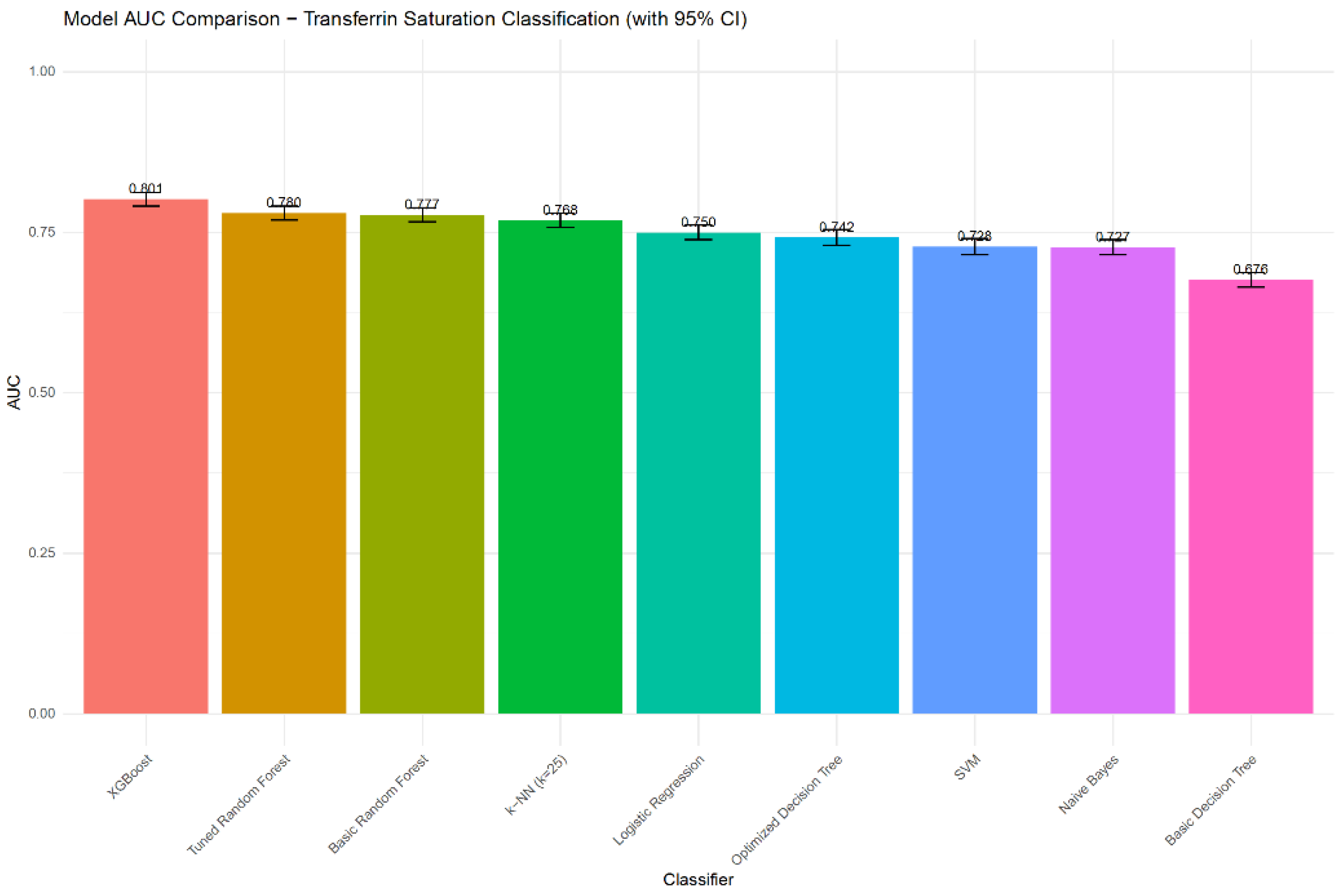
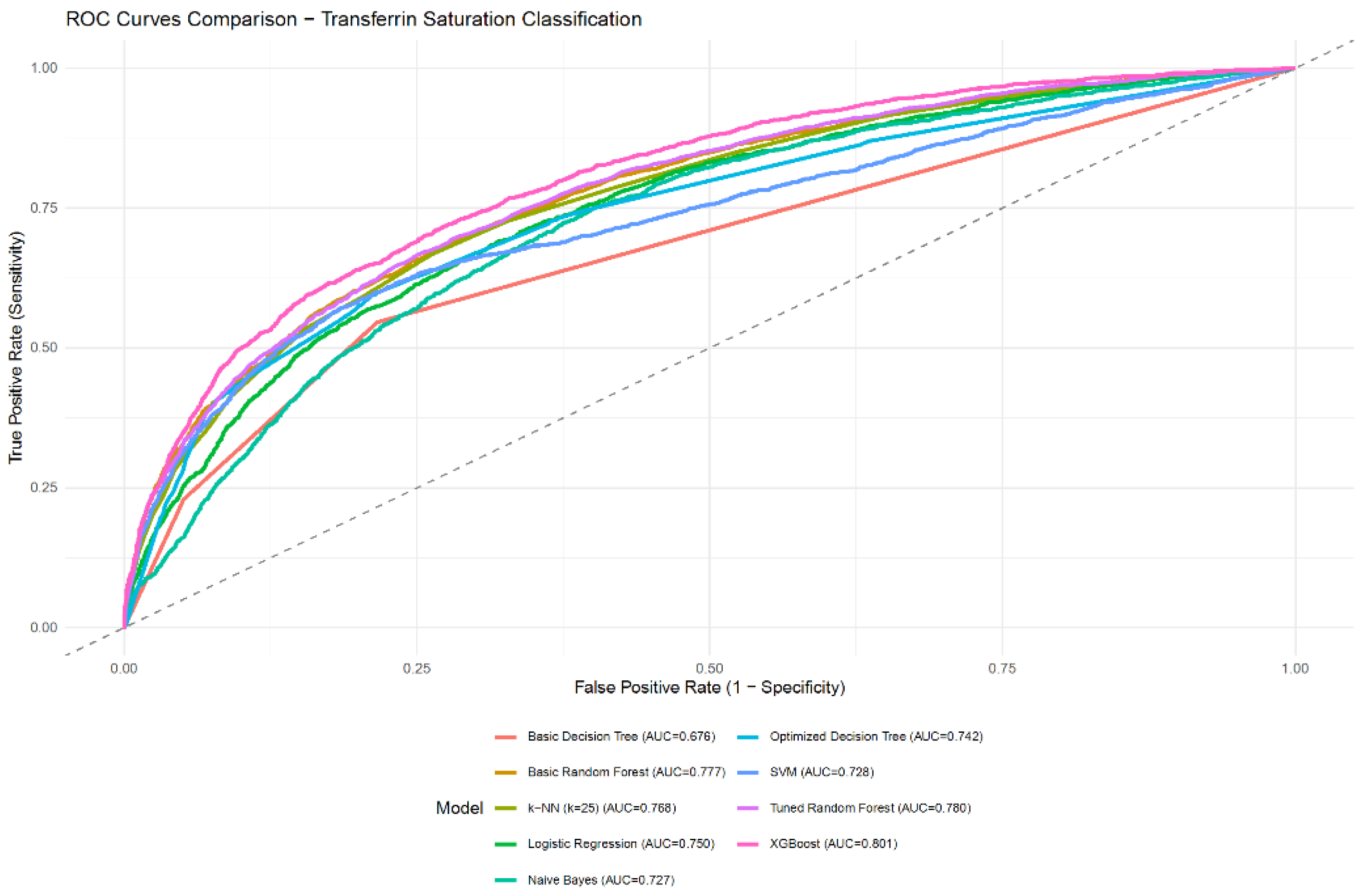
| Model | Accuracy | Balanced accuracy | Sensitivity | Specificity | PPV | NPV | F1-score | AUC | AUC 95% CI |
|---|---|---|---|---|---|---|---|---|---|
| Logistic Regression | 0.7814 | 0.5898 | 0.2212 | 0.9585 | 0.6274 | 0.7956 | 0.3271 | 0.7496 | 0.7383 - 0.7610 |
| Basic Decision Tree | 0.7764 | 0.5893 | 0.2294 | 0.9493 | 0.5884 | 0.7958 | 0.3300 | 0.6759 | 0.6644 - 0.6874 |
| Optimized Decision Tree | 0.7978 | 0.6519 | 0.3710 | 0.9328 | 0.6356 | 0.8243 | 0.4685 | 0.7419 | 0.7299 - 0.7539 |
| Basic Random Forest | 0.8016 | 0.6591 | 0.3847 | 0.9335 | 0.6463 | 0.8276 | 0.4823 | 0.7773 | 0.7663 - 0.7882 |
| Tuned Random Forest | 0.7996 | 0.6469 | 0.3530 | 0.9408 | 0.6532 | 0.8214 | 0.4583 | 0.7801 | 0.7692 - 0.7909 |
| XGBoost | 0.8063 | 0.6650 | 0.3932 | 0.9368 | 0.6631 | 0.8301 | 0.4937 | 0.8013 | 0.7911 - 0.8115 |
| SVM | 0.7947 | 0.6090 | 0.2516 | 0.9663 | 0.7025 | 0.8033 | 0.3705 | 0.7278 | 0.7150 - 0.7406 |
| k-NN (tuned as k=25) | 0.7937 | 0.6307 | 0.3171 | 0.9444 | 0.6432 | 0.8139 | 0.4248 | 0.7684 | 0.7574 - 0.7795 |
| Naive Bayes | 0.7503 | 0.6198 | 0.3688 | 0.8708 | 0.4744 | 0.8136 | 0.4150 | 0.7266 | 0.7151 - 0.7382 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




