Submitted:
29 March 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methodology
2.1. Experimental Design
- Base model: 30,000 registers were generated using different stoichiometric and kinetic parameters based on the real bio-digester by means of the ODEs (1), (2), (3), (4) and features in terms of inflow and the organic substrate concentration to produce biogas calculated through the equation (5). An Extreme Gradient (XGBoost) model was trained and tested using a sequential 80/20 split, respectively.
- Lag-based improved model: 10,000 registers were generated incorporating a dynamic seasonal temperature factor to perturb the microbial kinetic rates and simulate real-world environmental exposure. To capture this physical complexity, integrating thermodynamic interaction variables and historical lags of the biogas production. The inclusion of this auto-regressive feature is valid given that this information is available in real world applications, it is calculated through equation (5). An XGBoost model was trained and tested using a sequential 80/20 split to evaluate the system’s inertial memory and prevent data leakage.
2.2. Mathematical Modeling and Mass Balance
2.3. Microbial Kinetics and Thermodynamic Perturbation
2.4. Synthetic Data Generation
- Input flow rate (): Modeled as and physically constrained to the range m3/d.
- Organic substrate loading (): Modeled as and physically constrained to the range mg/L.
- Base model: A dataset with 30,000 records with the exported feature space consisted of Simulation Time (Time), Input Flow Rate (Q_in_m3_d), Organic Substrate Loading(S1_in_mg_L), and the target variable Methane Biogas Production (CH4_Prod_L_d).
- Lag-based improved model: A dataset with 10,000 records with the exported feature space included the thermodynamic perturbation, consisting of Simulation Time (Time), Input Flow Rate (Q_in_m3_d), Organic Substrate Loading (S1_in_mg_L), the Seasonal Temperature Factor (Temp_Factor), and the target variable methane biogas Production (CH4_Prod_L_d).
2.5. Feature Engineering and Machine Learning Architecture
2.6. Model Training and Evaluation Metrics
- Base model configuration: The model used 1000 boosting rounds (estimators), a learning rate of , and a maximum tree depth of 5. Subsampling and column sampling by tree were both set to . Early stopping was triggered if the validation loss did not improve for 50 consecutive rounds.
- Lag-based improved model configuration: The model used 5000 estimators with a learning rate of and a maximum tree depth of 12. Subsampling and column samplings were set to , with an early stopping patience of 100 rounds.
3. Results
3.1. Synthetic Data Generation
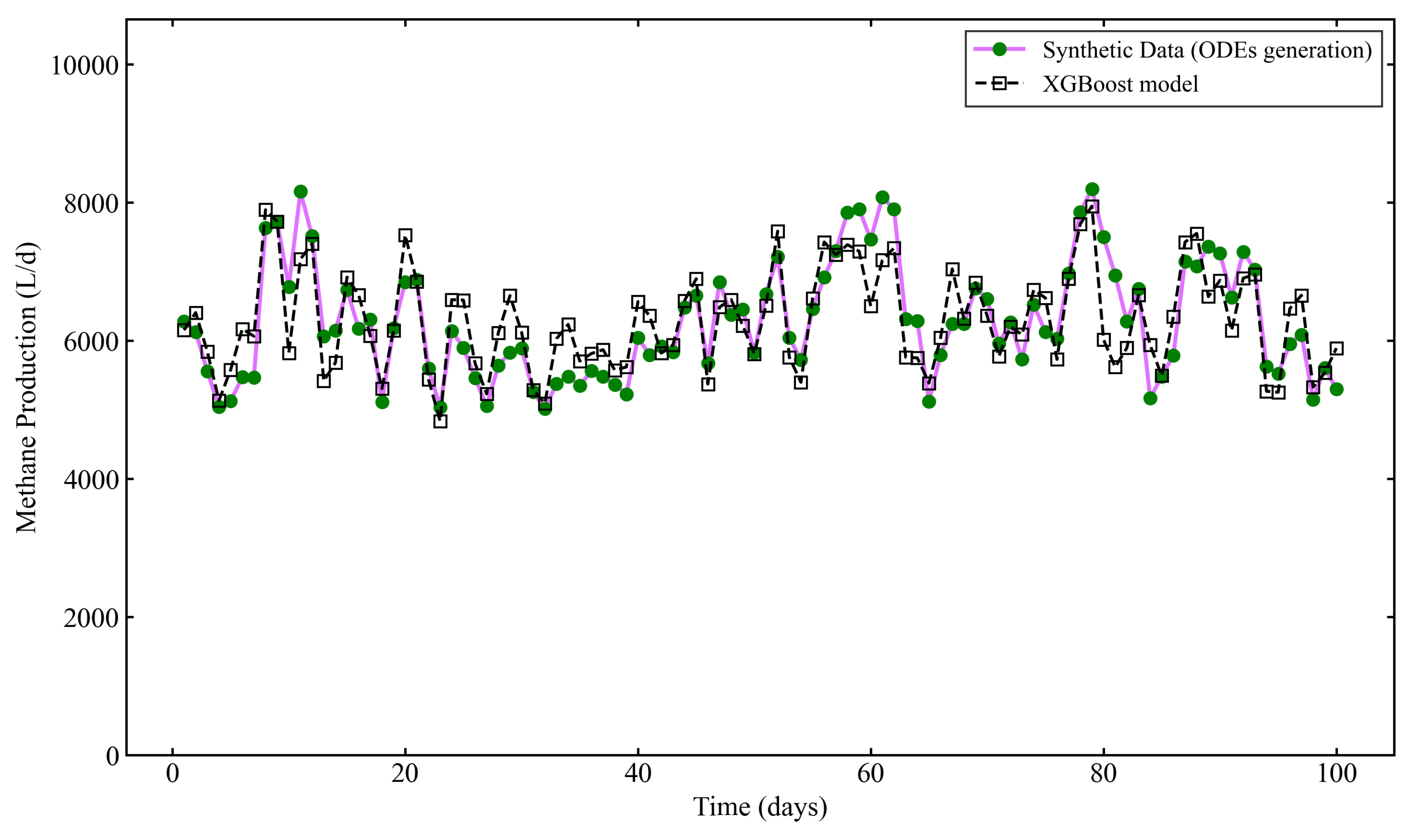
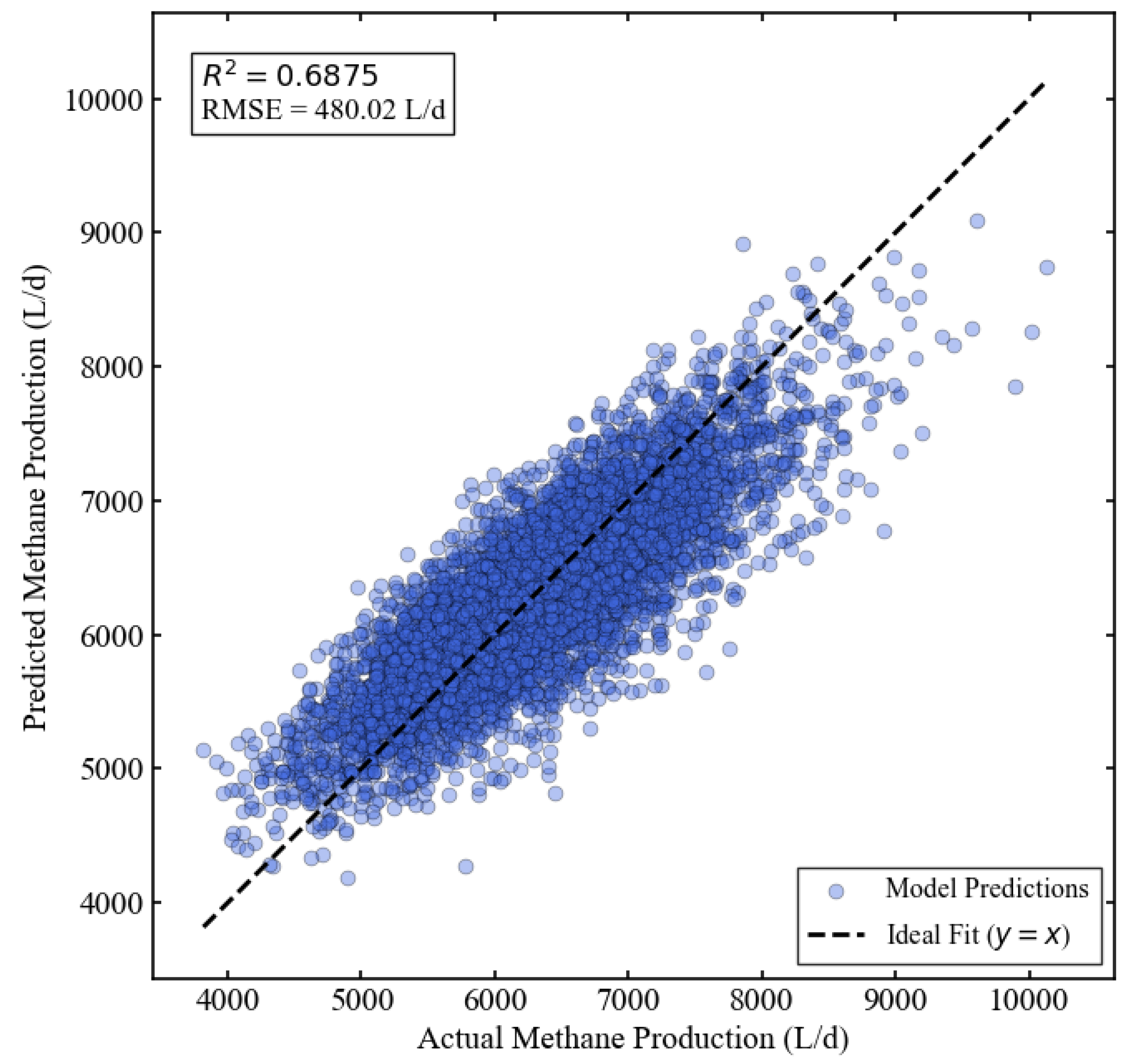
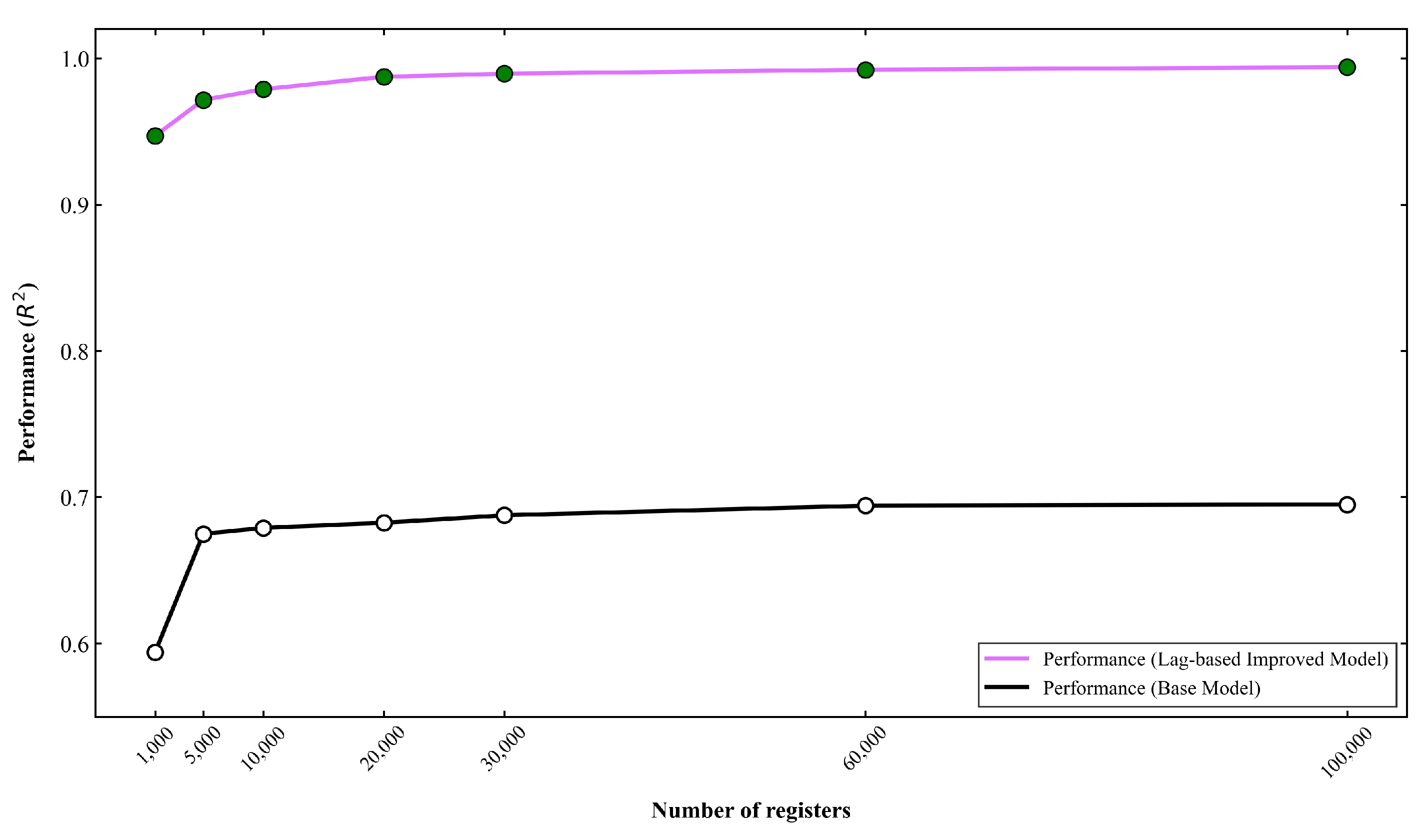
3.2. Performance of the Base Model
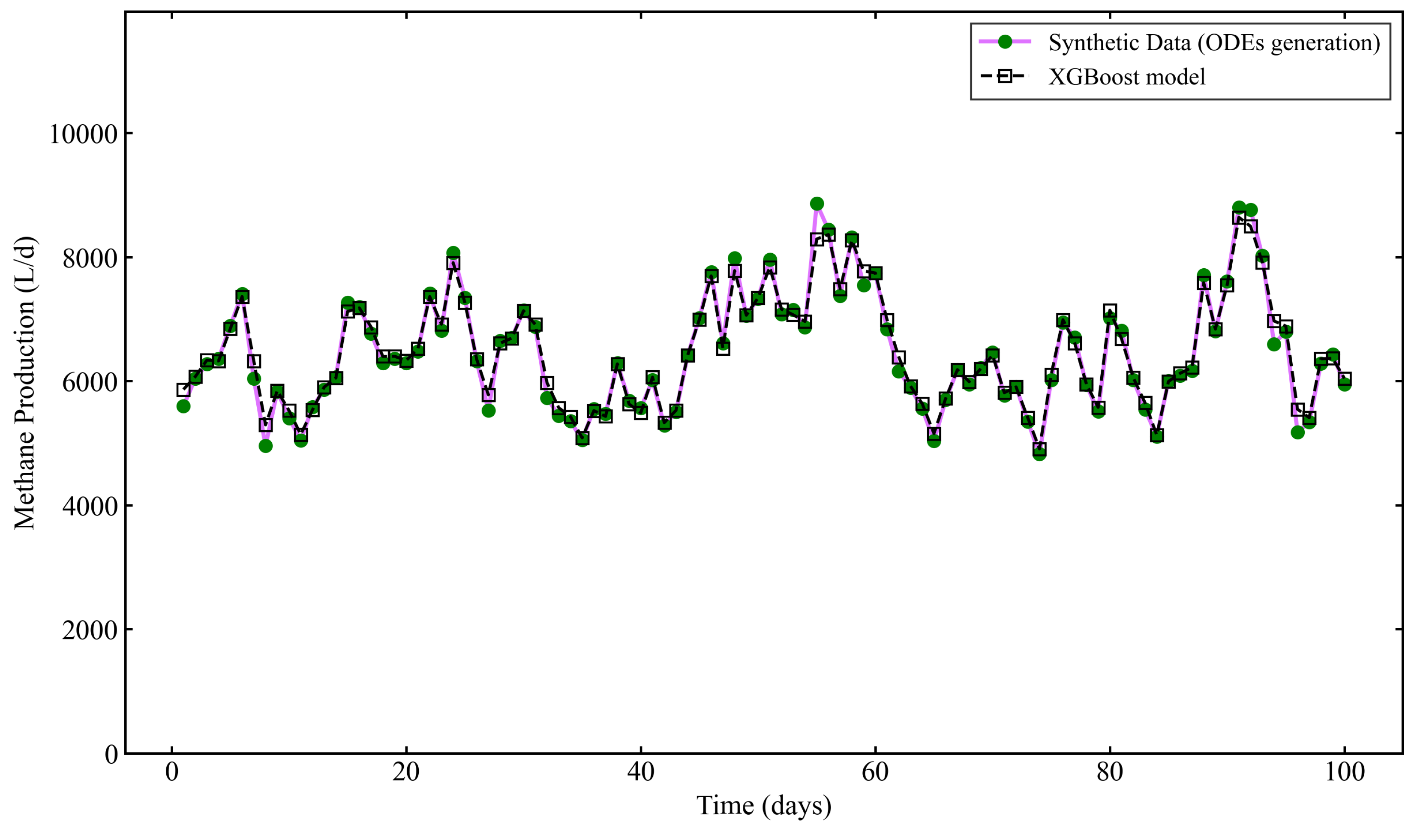
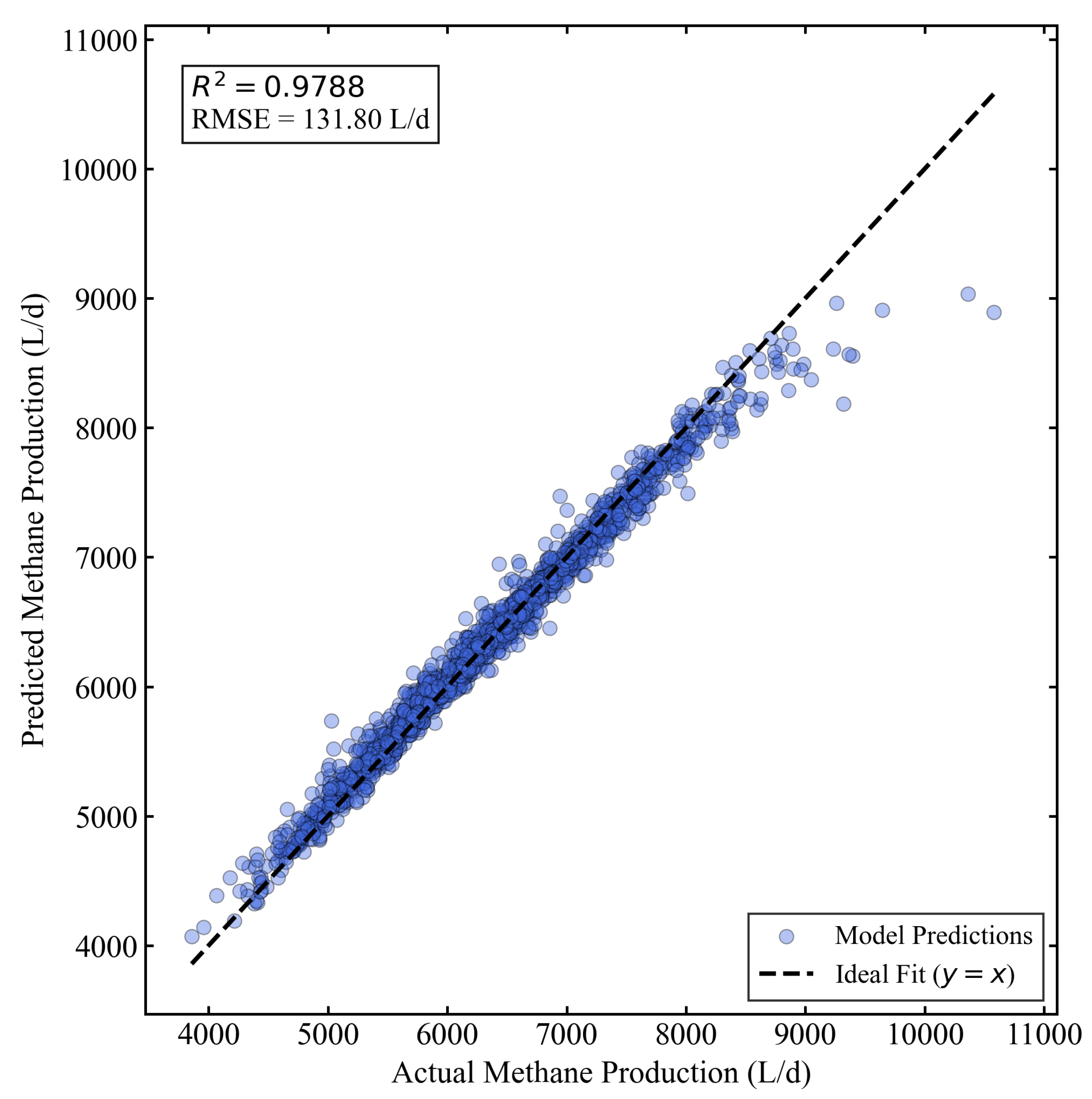
3.3. Performance of the Lag-based Improved Model
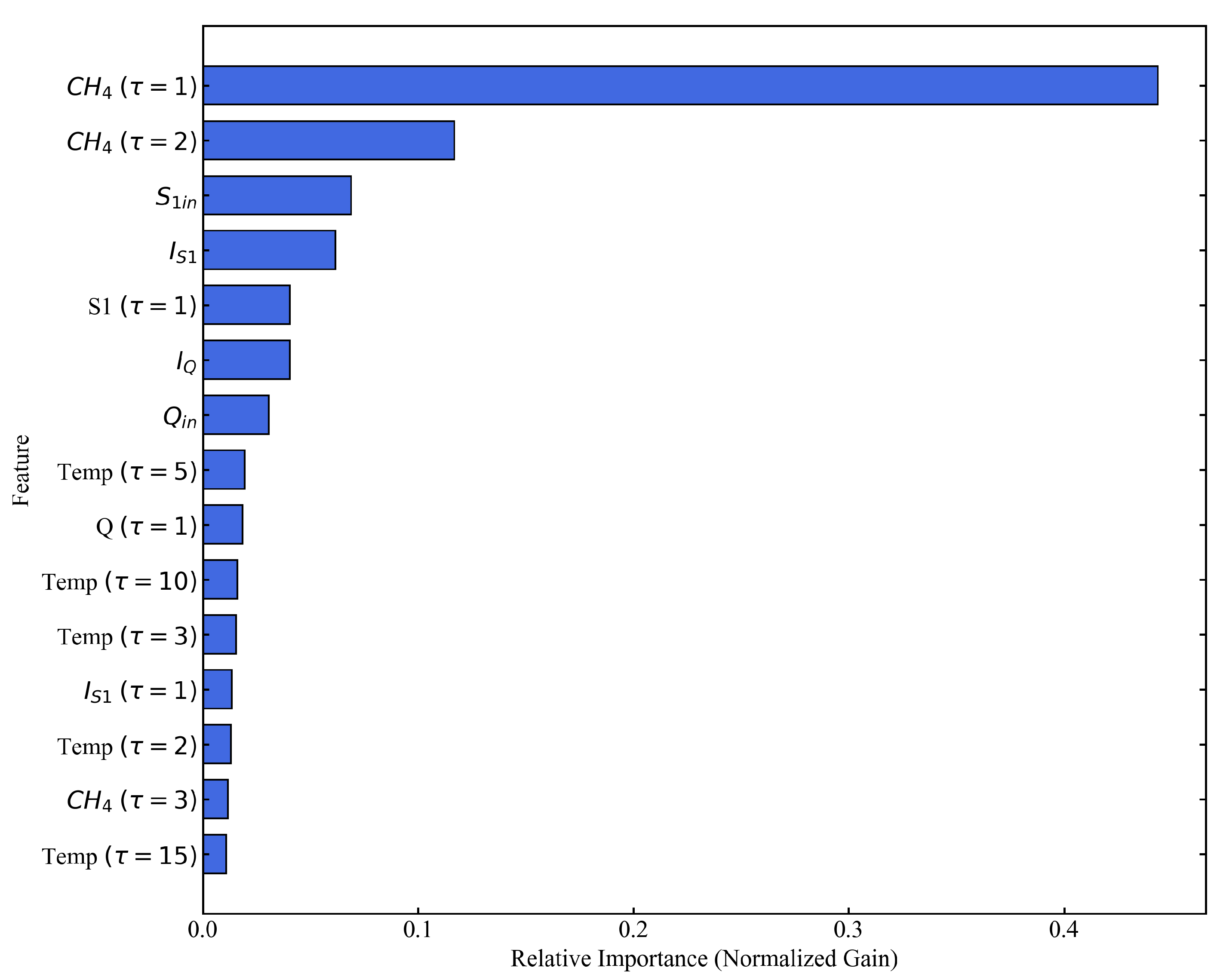
3.3.1. Feature Importance and Microbial Memory
3.3.2. Five-Fold Cross-Validation
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tabatabaei, M.; Ghanavati, H. Biogas. Springer International Publishing 2018.
- Ashokkumar, V.; Kumar, G.; Lakshmanan, H.; Chandramughi, V.; Flora, G.; Kothari, R.; Piechota, G. A critical review of biogas production and upgrading from organic wastes: Recent advances, challenges and opportunities. Biomass and Bioenergy 2025, 194, 107566.
- Nayeri, D.; Mohammadi, P.; Bashardoust, P.; Eshtiaghi, N. A comprehensive review on the recent development of anaerobic sludge digestions: Performance, mechanism, operational factors, and future challenges. Results in Engineering 2024, 22, 102292. [CrossRef]
- Wang, Z.; Liu, Y.; Zhang, A.; Liu, Z.; Gai, H. A review of process development, mechanistic insights, and enhancement technologies for anaerobic digestion in industrial wastewater treatment. Journal of Environmental Chemical Engineering 2025, 13, 118217. [CrossRef]
- Simeonov, I.; Chorukova, E.; Kabaivanova, L. Two-stage anaerobic digestion for green energy production: A review. Processes 2025, 13, 294.
- Gavala, H.N.; Angelidaki, I.; Ahring, B.K. Kinetics and modeling of anaerobic digestion process. Biomethanation I 2003, pp. 57–93.
- Farid, M.U.; Olbert, I.A.; Bück, A.; Ghafoor, A.; Wu, G. CFD modelling and simulation of anaerobic digestion reactors for energy generation from organic wastes: A comprehensive review. Heliyon 2025, 11.
- Lucas, D.; Oliveira, P.; Bessa, A.; Marcondes, F.S.; Rodrigues, M. Towards Efficient Biogas Production: Deep Learning-Based Methane Forecasting in Anaerobic Digesters of Wastewater Treatment Plants. In Proceedings of the International Conference on Practical Applications of Agents and Multi-Agent Systems. Springer, 2025, pp. 154–165.
- Shamshad, J.; Rehman, R.U. Innovative approaches to sustainable wastewater treatment: a comprehensive exploration of conventional and emerging technologies. Environmental Science: Advances 2025, 4, 189–222.
- Ling, J.Y.X.; Chan, Y.J.; Chen, J.W.; Chong, D.J.S.; Tan, A.L.L.; Arumugasamy, S.K.; Lau, P.L. Machine learning methods for the modelling and optimisation of biogas production from anaerobic digestion: a review. Environmental Science and Pollution Research 2024, 31, 19085–19104.
- Kumar, S.; Kumar, S.; Kumar, D.R.; Sharma, D.; Wipulanusat, W. Machine learning-based optimization of biogas and methane yields in UASB reactors for treating domestic wastewater. Biodegradation 2025, 36, 55.
- Schultz, J.; Scherzinger, M.; Elbanhawy, A.Y.; Kaltschmitt, M. Long-term continuous anaerobic co-digestion of residual biomass—model validation and model-based Investigation of different carbon-to-nitrogen ratios. BioEnergy Research 2025, 18, 58.
- Marycz, M.; Turowska, I.; Glazik, S.; Jasiński, P. Artificial Intelligence in Anaerobic Digestion: A Review of Sensors, Modeling Approaches, and Optimization Strategies. Sensors 2025, 25, 6961.
- Kim, M.; Ghobadi, F.; Tayerani Charmchi, A.S.; Lee, M.; Lee, J. Digital Twins for Clean Energy Systems: A State-of-the-Art Review of Applications, Integrated Technologies, and Key Challenges. Sustainability 2025, 18, 43.
- Cardona Acuña, L.D. Development of an approximate mathematical model for rural biodigesters (Desarrollo de un modelo matemático aproximado para biodigestores rurales). Master thesis in spanish, Universidad de Ibagué, Ibagué, Colombia. Available at: https://hdl.handle.net/20.500.12313/3983, 2021.
- Cheng, M.; Zhao, X.; Dhimish, M.; Qiu, W.; Niu, S. A Review of Data-Driven Surrogate Models for Design Optimization of Electric Motors. IEEE Transactions on Transportation Electrification 2024, 10, 8413–8431. [CrossRef]
- Bergmeir, C.; Hyndman, R.J.; Koo, B. A note on the validity of cross-validation for evaluating autoregressive time series prediction. Computational Statistics & Data Analysis 2018, 120, 70–83. [CrossRef]
| Parameter | Description | Value | Reference |
|---|---|---|---|
| Organic matter consumption yield | 1.30 g/g | [15] | |
| VFA generation yield | 1.06 mmol/g | [15] | |
| VFA consumption yield | 6.30 mmol/g | [15] | |
| Maximum specific growth rate (acidogenic) | 1.70 d−1 | [15] | |
| Maximum specific growth rate (methanogenic) | 0.84 d−1 | [15] | |
| Saturation constant for VFA | 12.0 mmol/L | [15] | |
| Total bio-digester volume | 61.0 m3 | [15] | |
| Number of CSTRs in series | 3 | [15] | |
| Methane yield coefficient | 0.18 mmol/g | Calibrated 1 | |
| Saturation constant for organic matter | 6000 mg/L | Calibrated 2 | |
| Haldane inhibition constant | 150 mmol/L | Calibrated 3 |
| Analysis | Model | RMSE (L/d) | MAE (L/d) | |
|---|---|---|---|---|
| Base Model | XGBoost (16 features) | 0.6875 | 480.02 | 381.19 |
| Lag-based Model | XGBoost (40 features) | 0.9788 | 131.80 | 85.48 |
| Fold | RMSE (L/d) | |
|---|---|---|
| 1 | 0.9550 | 190.90 |
| 2 | 0.9774 | 130.79 |
| 3 | 0.9797 | 122.00 |
| 4 | 0.9796 | 121.08 |
| 5 | 0.9784 | 135.36 |
| Average | 0.9740 ± 0.0095 | 140.03 ± 26.00 |
| Temperature Noise Level | RMSE (L/d) | |
|---|---|---|
| 0% (Deterministic control) | 0.9788 | 131.80 |
| 10% Stochastic noise | 0.9682 | 169.50 |
| 20% Stochastic noise | 0.9599 | 180.90 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




