Submitted:
29 March 2026
Posted:
31 March 2026
You are already at the latest version
Abstract
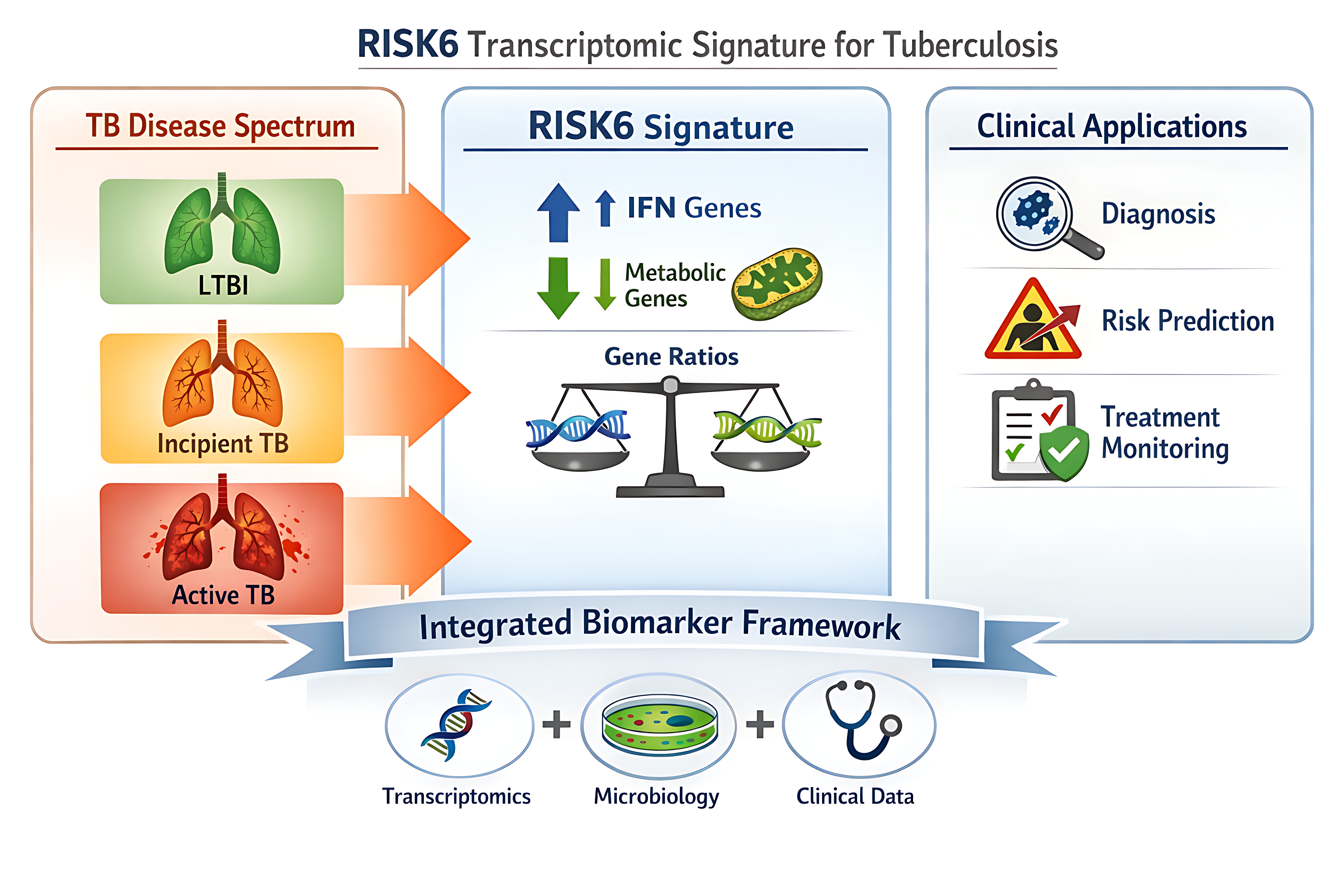
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Literature Search Strategy
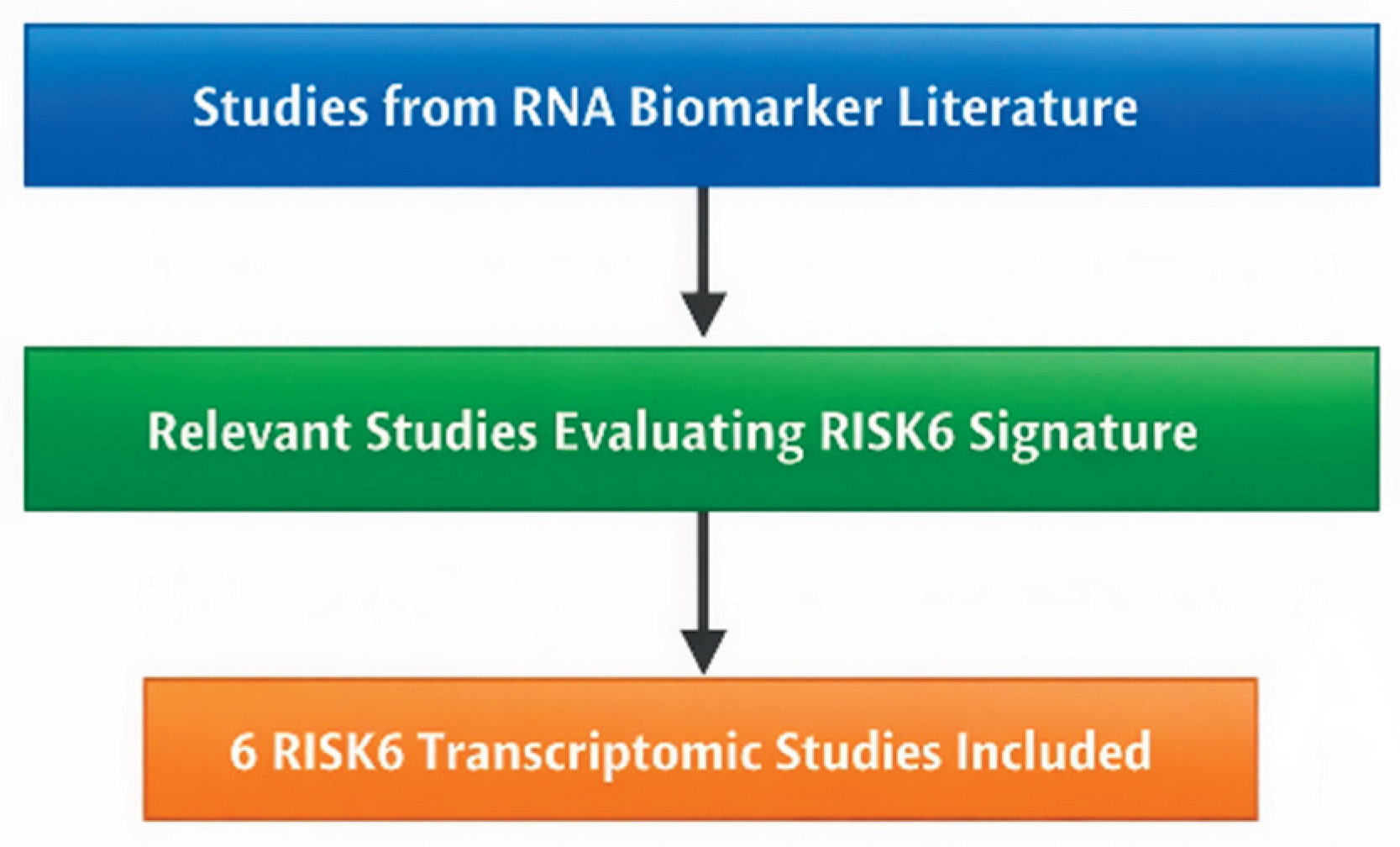
2.3. Study Selection
- Aims to assess the RISK6 transcriptomic signature in TB.
-
Reports of clinical applications such as:
- diagnosis
- prediction of disease progression
- treatment monitoring
- Conducted in human populations.
- did not evaluate RISK6 in particular
- were non-original reports lacking relevant data (e.g., editorials without any analysis)
- were not available in English.
2.4. Data Extraction and Synthesis
- the study design and participants
- clinical utility (diagnostic, prognostic or monitoring)
- key findings
- performance metrics (e.g., AUC, sensitivity, specificity)
- reported limitations
2.5. Analytical Framework
- Diagnostic application
- Prognostic application (progression of disease)
- Treatment monitoring
3. Results
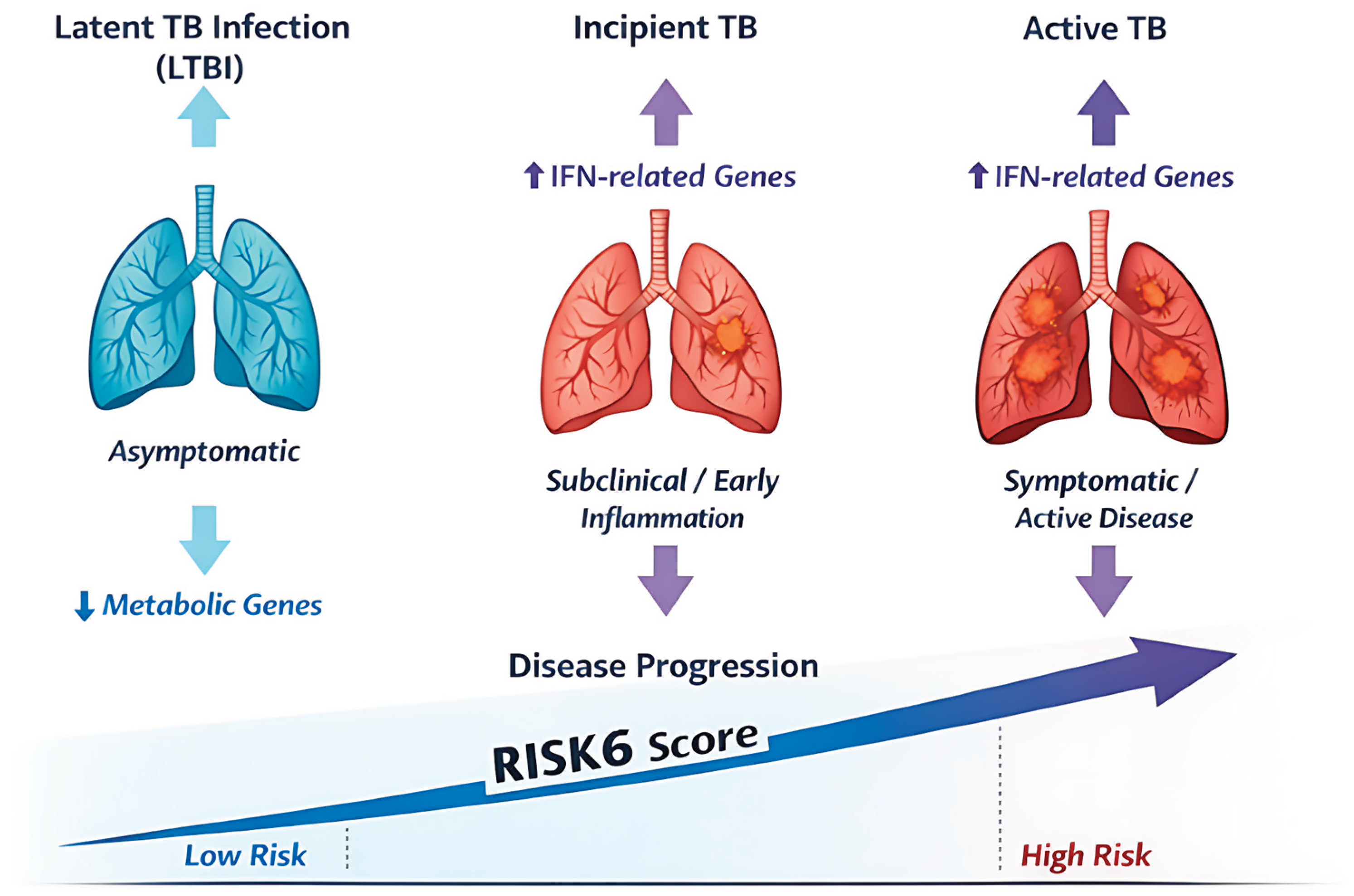
3.1. Biological Basis of RISK6
4. Discussion
4.1. Clinical Applications of RISK6
4.1.1. Diagnostic Application
4.1.2. Prognostic Application (Prediction of Disease Progression)
4.1.3. Treatment Monitoring
4.1.4. Integration into Clinical Practice
4.1.5. Relevance to Drug-Resistant TB
4.1.6. Translational Perspective
4.2. Limitations and Challenges
4.2.1. Variability Across Populations
4.2.2. Limited Long-Term Predictive Performance
4.2.3. Limited Specificity of Interferon-Driven Signatures
4.2.4. Lack of Standardized Thresholds
4.2.5. Limited Evidence in Drug-Resistant TB
4.2.6. Need for Multi-Biomarker Approaches
4.2.7. Implementation Challenges
4.2.8. Translational Perspective on Limitations
4.3. Comparison with Other Transcriptomic Signatures
4.3.1. Comparison with Early Multi-Gene Signatures
4.3.2. Comparison in Prognostic Performance
4.3.3. Comparison in Diagnostic Application
4.3.4. Comparison in Treatment Monitoring
4.3.5. Emerging Role of Composite Signatures
4.3.6. Overall Positioning of RISK6
4.4. Future Perspectives and Relevance to Drug-Resistant TB
4.4.1. Toward Precision TB Medicine
4.4.2. Expanding the Role in MDR-TB
4.4.3. Integration with Microbiological Markers
4.4.4. Toward Multi-Biomarker Models
4.4.5. Translation into Clinical Practice
4.4.6. Future Directions
5. Conclusions
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| TB | Tuberculosis |
| MDR-TB | Multidrug-Resistant Tuberculosis |
| LTBI | Latent Tuberculosis Infection |
| RISK6 | Six-gene transcriptomic signature for tuberculosis risk, diagnosis, and treatment monitoring |
| RNA | Ribonucleic Acid |
| PCR | Polymerase Chain Reaction |
| IFN | Interferon |
| IGRA | Interferon-Gamma Release Assay |
| AUC | Area Under the Curve |
| TTP | Time to Positivity |
| WHO | World Health Organization |
| TPP | Target Product Profile |
References
- World Health Organization, Global Tuberculosis Report 2025, no. September. Geneva, 2025. [Online]. Available: https://iris.who.int/server/api/core/bitstreams/e97dd6f4-b567-4396-8680-717bac6869a9/content.
- Pai, M.; Behr, M.A.; Dowdy, D.; Dheda, K.; Divangahi, M.; Boehme, C.C., “Tuberculosis,” Nat Rev Dis Prim., vol. 2, p. 16076, 2016. [CrossRef]
- Lawn, S.D.; Zumla, A.I., “Tuberculosis,” Lancet, vol. 378, no. 9785, pp. 57–72, 2011. [CrossRef]
- Berry, M.P.R.; Graham, C.M.; McNab, F.W.; Xu, Z.; Bloch, S.A.A.; Oni, T., “An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis,” Nature, vol. 466, no. 7309, pp. 973–977, 2010. [CrossRef]
- O’Garra, A.; Redford, P.S.; McNab, F.W.; Bloom, C.I.; Wilkinson, R.J.; Berry, M.P.R., “The Immune Response in Tuberculosis,” Annu Rev Immunol, vol. 31, pp. 475–527, 2013. [CrossRef]
- Zak, D.E.; Penn-Nicholson, A.; Scriba, T.J.; Thompson, E.; Suliman, S.; Amon, L.M., “A prospective blood RNA signature for tuberculosis disease risk,” Lancet, vol. 387, no. 10035, pp. 2312–22, 2016. [CrossRef]
- Suliman et al, S., “Four-gene pan-African blood signature predicts progression to tuberculosis,” Am. J. Respir. Crit. Care Med., vol. 197, no. 9, pp. 1198–1208, 2018. [CrossRef]
- Gupta et al, R.K., “Concise whole blood transcriptional signatures for incipient tuberculosis: A systematic review and patient-level pooled meta-analysis,” Lancet Respir. Med., vol. 8, no. 4, pp. 395–406, 2020. [CrossRef]
- Penn-Nicholson et al, A., “RISK6, a 6-gene transcriptomic signature of TB disease risk, diagnosis and treatment response,” Sci. Rep., vol. 10, no. 1, pp. 1–21, 2020. [CrossRef]
- Bayaa et al, R., “Multi-country evaluation of RISK6, a 6-gene blood transcriptomic signature, for tuberculosis diagnosis and treatment monitoring,” Sci. Rep., vol. 11, no. 1, pp. 1–12, 2021. [CrossRef]
- Mendelsohn et al, S.C., “Transcriptomic Signatures of Progression to Tuberculosis Disease among Close Contacts in Brazil,” J. Infect. Dis., vol. 230, no. 6, pp. e1355–e1365, 2024. [CrossRef]
- Muwanga et al, V.M., “Blood transcriptomic signatures for symptomatic tuberculosis in an African multicohort study,” Eur. Respir. J., vol. 64, no. 2, 2024. [CrossRef]
- Neumann et al, M., “The molecular bacterial load assay predicts treatment responses in patients with pre-XDR/XDR-tuberculosis more accurately than GeneXpert Ultra MTB/Rif,” J. Infect., vol. 90, no. 2, p. 106399, 2025. [CrossRef]
- Lange, C.; Dheda, K.; Chesov, D.; Mandalakas, A.M.; Udwadia, Z.; Jr, C.R.H., “Management of drug-resistant tuberculosis,” vol. 394, no. 10202, pp. 953–966, 2019. [CrossRef]
- Cliff et al, J.M., “Distinct phases of blood gene expression pattern through tuberculosis treatment reflect modulation of the humoral immune response,” J. Infect. Dis., vol. 207, no. 1, pp. 18–29, 2013. [CrossRef]
- Esmail et al, H., “Complement pathway gene activation and rising circulating immune complexes characterize early disease in HIV-associated tuberculosis,” Proc. Natl. Acad. Sci. U. S. A., vol. 115, no. 5, pp. E964–E973, 2018. [CrossRef]
- Warsinske, H.; Vashisht, R.; Khatri, P., “Host-response-based gene signatures for tuberculosis diagnosis: A systematic comparison of 16 signatures,” PLoS Med., vol. 16, no. 4, pp. 1–19, 2019. [CrossRef]
- Turner et al, C.T., “Blood transcriptional biomarkers for active pulmonary tuberculosis in a high-burden setting: A prospective, observational, diagnostic accuracy study,” Lancet Respir. Med., vol. 8, no. 4, pp. 407–419, 2020. [CrossRef]
- Burel et al, J.G., “Distinct blood transcriptomic signature of treatment in latent tuberculosis infected individuals at risk of developing active disease,” Tuberculosis, vol. 131, pp. 1–24, 2021. [CrossRef]
- Lundell, S.; Kaipilyawar, V.; Johnson, W.E.; Dietze, R.; Jerrold, J., “Combining blood transcriptomic signatures improves the prediction of progression to tuberculosis among household contacts in Brazil .,” 2025.
- Pitaloka, D.A.E.; Syamsunarno, M.R.A.A.A.; Abdulah, R.; Chaidir, L., “Omics Biomarkers for Monitoring Tuberculosis Treatment: A Mini-Review of Recent Insights and Future Approaches,” Infect. Drug Resist., vol. 15, no. May, pp. 2703–2711, 2022. [CrossRef]
- World Health Organization, High-priority target product profiles for new tuberculosis diagnostics: Report of a consensus meeting, no. April. Geneva, 2014.
- World Health Organization, WHO Consolidated Guidelines on Tuberculosis Module 4: Treatment and care. Geneva, 2025.
- Roe et al, J.K., “Blood transcriptomic diagnosis of pulmonary and extrapulmonary tuberculosis,” JCI Insight, vol. 1, no. 16, pp. 1–14, 2016. [CrossRef]
- Sambarey et al, A., “Unbiased Identification of Blood-based Biomarkers for Pulmonary Tuberculosis by Modeling and Mining Molecular Interaction Networks,” EBioMedicine, vol. 15, pp. 112–126, 2017. [CrossRef]
- Maertzdorf et al, J., “Concise gene signature for point-of-care classification of tuberculosis,” EMBO Mol. Med., vol. 8, no. 2, pp. 86–95, 2016. [CrossRef]
- Diacon et al, A.H., “Multidrug-Resistant Tuberculosis and Culture Conversion with Bedaquiline,” N. Engl. J. Med., vol. 371, no. 8, pp. 723–732, 2014. [CrossRef]
| Study | Population / Setting | Design | Application | Key Findings | Performance | Limitations | Clinical Interpretation |
|---|---|---|---|---|---|---|---|
| Penn-Nicholson et al., 2020 [9] | Multicohort | Prospective / validation | Diagnosis, prognosis, monitoring | Correlates with disease activity; declines during treatment | AUC >0.85; dynamic change | Limited MDR-TB data; no standardized cut-off | Multi-purpose biomarker; promising for treatment monitoring |
| Bayaa et al., 2021 [10] | Multicountry | Validation | Diagnosis | Differentiates active TB vs LTBI/healthy | AUC ≈ 0.94; Se ~90.9%; Sp ~87.8% | Variability; overlap with inflammation | Potential triage tool (non-sputum-based) |
| Mendelsohn et al., 2024 [11] | Household contacts | Prospective cohort | Prognosis | Strong short-term prediction (≤6–9 months) | Meets WHO TPP short-term | Weak ≥12 months | Best for short-term risk stratification (incipient TB) |
| Muwanga et al., 2024 [12] | Multicountry | Cross-cohort | Diagnosis, prognosis | Performance varies by geography | AUC ~0.75–0.85 | Heterogeneity | Requires population-specific calibration |
| Burel et al., 2021 [19] | TB vs LTBI | Observational | Diagnosis | Distinguishes TB states (host response) | AUC ~0.85–0.90 | Not RISK6-specific | Supports biological validity of host signatures |
| Lundell et al., 2025 [20] | Integrated cohorts | Modeling | Prognosis (composite) | Combined signatures ↑ accuracy | Se ~90%; Sp ~88% | Complexity | RISK6 may be stronger in composite panels |
| Domain | Key Evidence | Strengths | Limitations | Clinical Role |
|---|---|---|---|---|
| Diagnosis | Bayaa et al. [10]; Muwanga et al. [12]; Burel et al. [19] | High diagnostic accuracy (AUC up to 0.94); non-sputum-based | Reduced specificity in inflammatory conditions; variability across populations | Potential triage tool, especially in paucibacillary TB |
| Prognosis (Disease Progression) | Penn-Nicholson et al. [9]; Mendelsohn et al. [11] | Strong short-term prediction (≤6–9 months); aligns with incipient TB concept | Declining performance over longer intervals | Useful for identifying high-risk individuals for targeted prevention |
| Treatment Monitoring | Penn-Nicholson et al. [9]; Muwanga et al. [12] | Dynamic decline during therapy; reflects disease activity | Lack of standardized thresholds; limited MDR-TB data | Potential early marker of treatment response and failure |
| Drug-Resistant TB (Emerging) | Limited evidence [14] | Theoretical relevance for response monitoring | Insufficient validation in MDR-TB populations | Promising but requires further validation |
| Integrated / Composite Biomarker Models | Lundell et al. [20] | Improved predictive accuracy when combined with other signatures | Increased complexity; reduced scalability | RISK6 may be most effective as part of multi-biomarker strategies |
| Signature | No. of Genes | Primary Application | Performance Characteristics | Strengths | Limitations | Clinical Positioning |
|---|---|---|---|---|---|---|
| RISK6 | 6 | Diagnosis, prognosis, monitoring | AUC up to 0.94; strong short-term prediction; dynamic treatment response | Parsimonious; ratio-based; multi-functional; PCR-compatible | Reduced long-term prediction; variability across populations; lack of standard cut-offs | Most clinically versatile; suitable for integrated diagnostic and monitoring strategies |
| ACS16 / RISK16 | 16 | Prognosis | Strong prediction of progression to active TB | Well-characterized IFN-driven signature | Large gene set; limited scalability | Useful for research and risk prediction; less practical for routine use |
| ACS11 / RISK11 | 11 | Prognosis | Moderate predictive performance | Reduced gene set vs ACS16 | Limited long-term performance; reduced specificity | Potential for targeted risk stratification with further validation |
| PREDICT29 | 29 | Prognosis | High performance in selected cohorts | Broad biological coverage | Complex; limited generalizability | Promising but not yet clinically scalable |
| Composite / Integrated Models | Variable | Prognosis, risk stratification | Improved sensitivity and specificity (~90%) | Highest predictive accuracy; adaptable | High complexity; cost; standardization challenges | Likely future direction; RISK6 may serve as a component of these models |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




