Submitted:
26 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Challenges in Replicating the Clinician Workflow
2.1. Limitations in History Collection and Triage
2.2. Limitations in Differential Diagnosis
2.3. Limitations in Documentation
2.4. A Clinical Example


3. Why the Data Cannot Support Reliable Diagnosis
3.1. The Setup
3.2. Three Sources of Permanent Bias
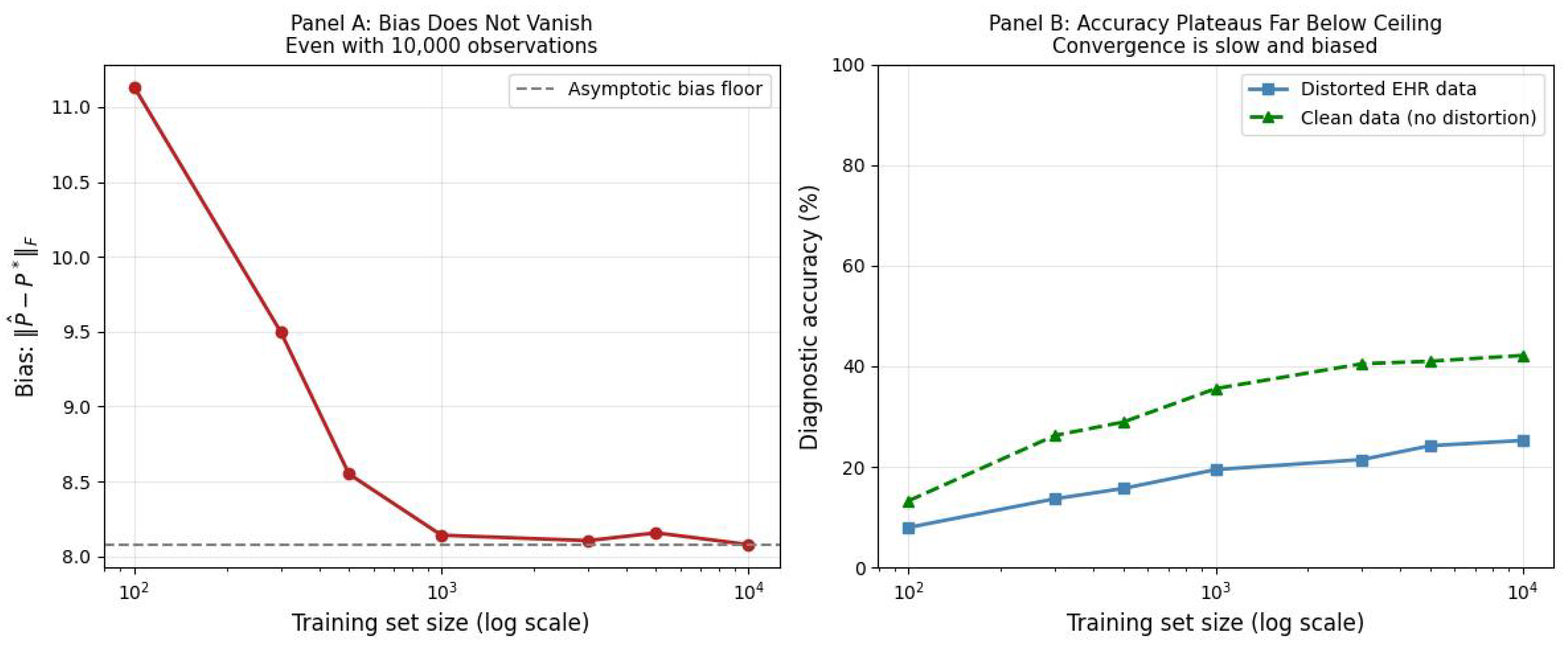
3.3. The Lower Bound Interpretation
4. Empirical Evidence: GPT-4o Diagnostic Instability
4.1. Experiment: Physician-Validated Clinical Cases from the Bangladesh Pilot
Dataset and Sample:
Distortion Operators:
Results:
5. Future Directions for AI in Healthcare
What LLMs Can Do Well
What Requires a Different Approach
References
- Barnett, Karen, Stewart W Mercer, Michael Norbury, Graham Watt, Sally Wyke, and Bruce Guthrie. 2012. Epidemiology of multimorbidity and implications for health care, research, and medical education: a cross-sectional study. The Lancet 380, 9836: 37–43. [Google Scholar] [CrossRef]
- Bélisle-Pipon, J. C., and et al. 2024. Why we need to be careful with LLMs in medicine. Frontiers in Medicine 11, 1495582. [Google Scholar] [CrossRef] [PubMed]
- Bell, Sigall K, Tom Delbanco, Joann G Elmore, Patricia S Fitzgerald, Alan Fossa, Kendall Harcourt, Suzanne G Leveille, Thomas H Payne, Rebecca A Stametz, Jan Walker, and et al. 2020. Frequency and types of patient-reported errors in electronic health record ambulatory care notes. JAMA network open 3, 6: e205867. [Google Scholar] [CrossRef] [PubMed]
- Bundy, A., N. Chater, and S. Muggleton. 2023. Introduction to cognitive artificial intelligence. Philosophical Transactions of the Royal Society A 381, 20220051. [Google Scholar] [CrossRef] [PubMed]
- Elstein, A. S., L. S. Shulman, and S. A. Sprafka. 1978. Medical Problem Solving: An Analysis of Clinical Reasoning. Cambridge, MA: Harvard University Press. [Google Scholar]
- Espe, S. 2018. MalaCards: the human disease database. Journal of the Medical Library Association: JMLA 106, 1: 140. [Google Scholar] [CrossRef]
- Feuerriegel, Stefan, Philipp Spitzer, Daniel Hendriks, Jan Rudolph, Sarah Schlaeger, Jens Ricke, Niklas Kühl, and Boj Hoppe. 2025. The effect of medical explanations from large language models on diagnostic accuracy in radiology. [Google Scholar] [CrossRef]
- Gilbert, S., J. N. Kather, and A. Hogan. 2024. Augmented non-hallucinating large language models as medical information curators. npj Digital Medicine 7, 100. [Google Scholar] [CrossRef]
- Goh, Ethan, Robert Gallo, Jason Hom, Eric Strong, Yingjie Weng, Hannah Kerman, Joséphine A Cool, Zahir Kanjee, Andrew S Parsons, Neera Ahuja, and et al. 2024. Large language model influence on diagnostic reasoning: a randomized clinical trial. JAMA network open 7, 10: e2440969. [Google Scholar] [CrossRef]
- Griot, Maxime, Christophe Hemptinne, Jean Vanderdonckt, and et al. 2025. Large language models lack essential metacognition for reliable medical reasoning. Nature Communications 16, 642. [Google Scholar] [CrossRef]
- Hernán, M. A., and J. M. Robins. 2020. Causal Inference: What If. Boca Raton: Chapman & Hall/CRC. [Google Scholar]
- Herper, Matthew. 2017. MD anderson benches IBM watson in setback for artificial intelligence in medicine. Forbes.
- Jiang, Lavender Yao, Xujin Chris Liu, Nima Pour Nejatian, Mustafa Nasir-Moin, Duo Wang, Anas Abidin, Kevin Eaton, Howard Antony Riina, Ilya Laufer, Paawan Punjabi, and et al. 2023. Health system-scale language models are all-purpose prediction engines. Nature 619, 7969: 357–362. [Google Scholar] [CrossRef]
- Kabir, A., R. Kabir, and J. Nahar. 2024. In pursuit of an expert artificial intelligence system: reproducing human physicians’ diagnostic reasoning and triage decision making. Journal of Artificial Intelligence and Soft Computing Techniques, 1–14. [Google Scholar] [CrossRef]
- Kabir, R., Michael Williams, and Nashif Rayhan. 2026. Cognitive ai-assisted primary care health delivery: A pilot study in bangladesh. Working Paper.
- Kotseruba, I., and J. K. Tsotsos. 2016. A review of 40 years of cognitive architecture research. Preprint arXiv:1610.08602. [Google Scholar]
- Li, Yihan, Xiyuan Fu, Ghanshyam Verma, Paul Buitelaar, and Mingming Liu. 2025. Mitigating hallucination in large language models (llms): An application-oriented survey on rag, reasoning, and agentic systems. arXiv arXiv:2510.24476. [Google Scholar] [CrossRef]
- Lukac, Paul J, William Turner, Sitaram Vangala, Aaron T Chin, Joshua Khalili, Ya-Chen Tina Shih, Catherine Sarkisian, Eric M Cheng, and John N Mafi. 2025. Ambient ai scribes in clinical practice: a randomized trial. NEJM AI 2, 12: AIoa2501000. [Google Scholar] [CrossRef] [PubMed]
- Obermeyer, Ziad, Brian Powers, Christine Vogeli, and Sendhil Mullainathan. 2019. Dissecting racial bias in an algorithm used to manage the health of populations. Science 366, 6464: 447–453. [Google Scholar] [CrossRef]
- O’Donovan, James, Madeleine Ballard, Rebecca Hope, Jane Wamae, Adriana Viola Miranda, Brian DeRenzi, Richard Kabanda, Nan Chen, Lennie Bazira, Mallika Raghavan, and et al. 2026. Governance, scale, and integration: building community health worker systems ready for artificial intelligence. The Lancet Primary Care. [Google Scholar] [CrossRef]
- Qazi, Ihsan Ayyub, Ayesha Ali, Asad Ullah Khawaja, Muhammad Junaid Akhtar, Ali Zafar Sheikh, and Muhammad Hamad Alizai. 2026. Automation bias in large language model–assisted diagnostic reasoning among physicians trained in ai literacy—a randomized clinical trial. NEJM AI 3, 5: AIoa2501001. [Google Scholar] [CrossRef]
- Ramaswamy, Ashwin, Aditya Tyagi, Hannah Hugo, and et al. 2026. ChatGPT health performance in a structured test of triage recommendations. Nature Medicine. [Google Scholar] [CrossRef]
- Ribeiro, Marco Tulio, Sameer Singh, Carlos Guestrin, Scott M Lundberg, and Su-In Lee. 2016. Why should i trust you?: explaining the predictions of any. classifier. arXiv [cs. LG. [Google Scholar]
- Rosenbacke, R., and et al. 2025. Beyond hallucinations: the illusion of understanding in large language models. Preprint arXiv:2510.14665. [Google Scholar]
- Singhal, Karan, Tao Tu, Juraj Gottweis, Rory Sayres, Ellery Wulczyn, Mohamed Amin, Le Hou, Kevin Clark, Stephen R Pfohl, Heather Cole-Lewis, and et al. 2025. Toward expert-level medical question answering with large language models. Nature medicine 31, 3: 943–950. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y., D. Sheng, Z. Zhou, and Y. Wu. 2024. AI hallucination: towards a comprehensive classification of distorted information in AI-generated content. Humanities and Social Science Communications 11, 1278. [Google Scholar] [CrossRef]
- Thaler, S. 1995. Virtual input phenomena within the death of a simple pattern associator. Neural Networks 8, 1: 55–56. [Google Scholar] [CrossRef]
- Toma, A., S. Senkaiahliyan, P. R. Lawler, B. Rubin, and B. Wang. 2023. Generative AI could revolutionize health care—but not if control is ceded to big tech. Nature 624, 36–38. [Google Scholar] [CrossRef]
- Topol, Eric J. 2019. High-performance medicine: the convergence of human and artificial intelligence. Nature Medicine 25, 1: 44–56. [Google Scholar] [CrossRef]
- U.S. Department of Health and Human Services, Office of Inspector General. 2014, May. Improper payments for evaluation and management services cost Medicare billions in 2010. Technical Report OEI-04-10-00181, U.S. Department of Health and Human Services, Office of Inspector General. Published May 28, 2014.
- Weiskopf, Nicole G, David A Dorr, Christie Jackson, Harold P Lehmann, and Caroline A Thompson. 2023. Healthcare utilization is a collider: an introduction to collider bias in ehr data reuse. Journal of the American Medical Informatics Association 30, 5: 971–977. [Google Scholar] [CrossRef]
- Xu, Z., S. Jain, and M. Kankanhalli. 2024. Hallucination is inevitable: an innate limitation of large language models. Preprint arXiv:2401.11817. [Google Scholar]
- Yang, Xi, Aokun Chen, Nima PourNejatian, Hoo Chang Shin, Kaleb E Smith, Christopher Parisien, Colin Compas, Cheryl Martin, Anthony B Costa, Mona G Flores, and et al. 2022. A large language model for electronic health records. NPJ digital medicine 5, 1: 194. [Google Scholar] [CrossRef]
- Zack, Travis, Eric Lehman, Mirac Suzgun, Jorge A Rodriguez, Leo Anthony Celi, Judy Gichoya, Dan Jurafsky, Peter Szolovits, David W Bates, Raja-Elie E Abdulnour, and et al. 2024. Assessing the potential of gpt-4 to perpetuate racial and gender biases in health care: a model evaluation study. The Lancet Digital Health 6, 1: e12–e22. [Google Scholar] [CrossRef]
- Zech, John R, Marcus A Badgeley, Manway Liu, Anthony B Costa, Joseph J Titano, and Eric Karl Oermann. 2018. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: a cross-sectional study. PLoS medicine 15, 11: e1002683. [Google Scholar] [CrossRef]
| 1 |
Zack et al. (2024) find that GPT-4 differentially diagnoses patients by stereotyping races, ethnicities, and genders. Assessments and plans made by the model were correlated to demographic attributes, and demographics could be used to predict the cost of procedures recommended. |
| Condition | Top-1 accuracy | Top-3 accuracy | Flip rate |
|---|---|---|---|
| Baseline (clean) | 62.0% [48.2–74.1] | 72.0% [58.3–82.5] | — |
| swap | 50.0% [36.6–63.4] | 70.0% [56.2–80.9] | 34.0% [22.4–47.8] |
| add | 60.0% [46.2–72.4] | 72.0% [58.3–82.5] | 8.0% [3.2–18.8] |
| drop chief | 56.0% [42.3–68.8] | 70.0% [56.2–80.9] | 34.0% [22.4–47.8] |
| drop last | 54.0% [40.4–67.0] | 74.0% [60.4–84.1] | 16.0% [8.3–28.5] |
| combined | 38.0% [25.9–51.8] | 56.0% [42.3–68.8] | 38.0% [25.9–51.8] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




