Submitted:
24 March 2026
Posted:
25 March 2026
You are already at the latest version
Abstract
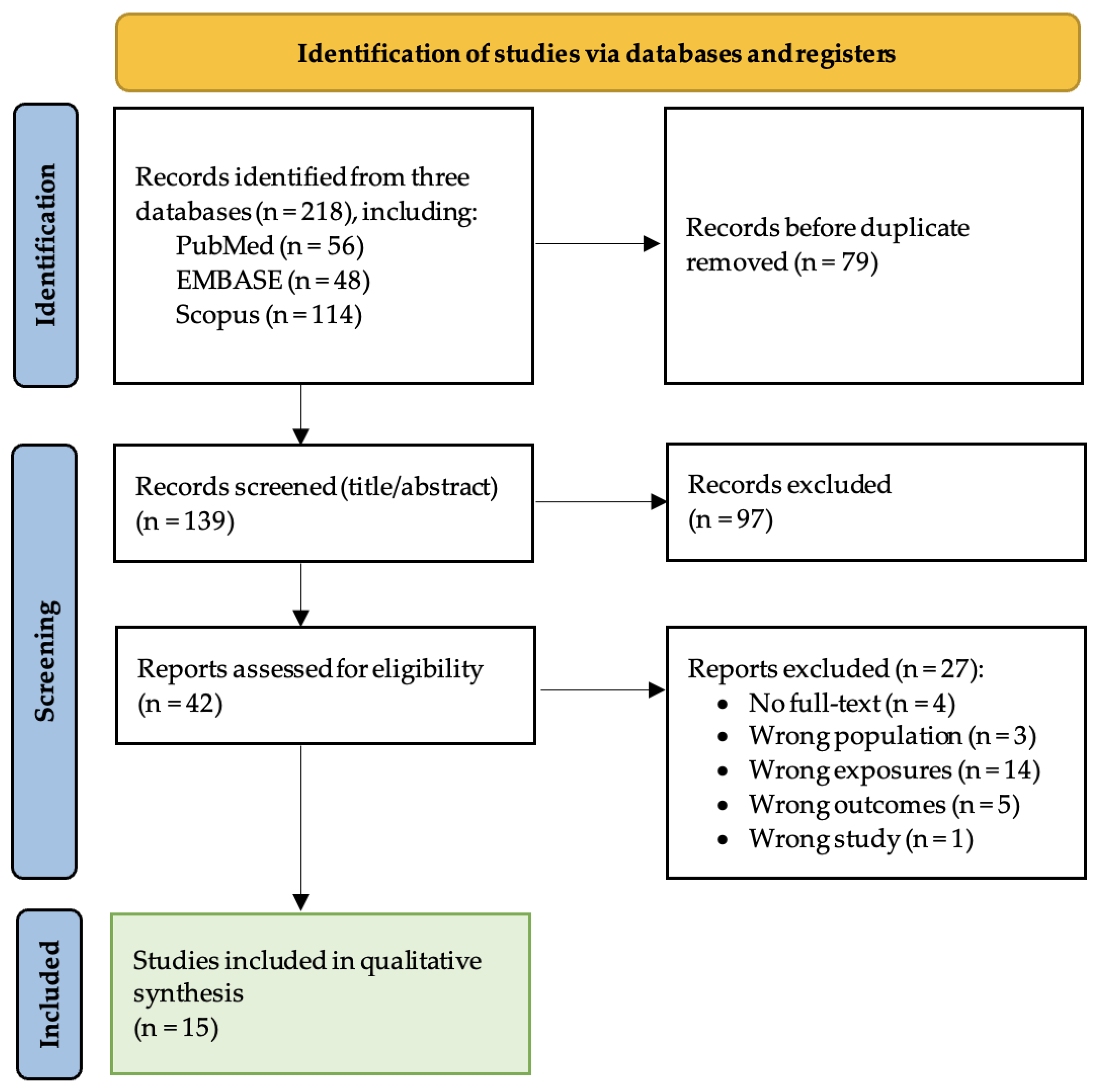
Background: Tuberculosis (TB) remains a major global cause of morbidity and mortality. Current tools for monitoring treatment response rely on sputum-based microscopy and culture, which may be insensitive, time-consuming, and impractical in extrapulmonary or pediatric TB and in individuals unable to produce sputum. Metabolomics has emerged as a promising approach to identify host-derived biomarkers reflecting treatment-associated immunometabolic changes, but evidence remains heterogeneous and incompletely synthesized. Methods: We conducted a comprehensive literature review of metabolomic biomarkers associated with TB treatment response. PubMed, Scopus, and Web of Science were searched for human studies evaluating targeted or untargeted metabolomics (NMR, LC-MS, GC-MS, CE-MS) in relation to treatment response or outcomes. Two reviewers independently screened studies, extracted data, and assessed risk of bias using QUIPS and PROBAST. Findings were synthesized using a structured framework across treatment stages and outcomes. Results: Of 218 records identified, 139 titles/abstracts were screened and 42 full texts assessed; 15 studies met inclusion criteria. Recurrent signals involved amino acid metabolism, particularly the tryptophan–kynurenine pathway, and vitamin/cofactor metabolites (pyridoxate, nicotinamide, trigonelline). Plasma studies frequently reported lipid remodeling and bile acid perturbations, while urine studies highlighted polyamine metabolism (e.g., N¹,N¹²-diacetylspermine) and fatty acid β-oxidation markers. Common limitations included inadequate adjustment for confounders and, in prediction models, small sample sizes and limited external validation. Conclusions: Metabolomic reveals reproducible but heterogeneous immunometabolic changes during TB therapy. Key pathways include tryptophan-kynurenine metabolism, vitamin/cofactor metabolism, lipid remodeling, and urine polyamine pathways. Standardization and prospective multicenter validation are needed for clinical translation.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Literature Search Strategy and Study Selection
2.2. Eligibility Criteria (PECOS Framework)
2.2.1. Population
2.2.2. Exposure/Intervention
2.2.3. Comparators
2.2.4. Outcomes
2.2.5. Study Design
2.3. Information Sources and Search Strategy
2.4. Study Selection
2.5. Data Extraction
2.6. Risk of Bias Assessment
2.7. Data Synthesis and Analysis
3. Results
3.1. Study Selection
3.2. Study Characteristics
3.3. Biospecimens and Metabolomic Platforms
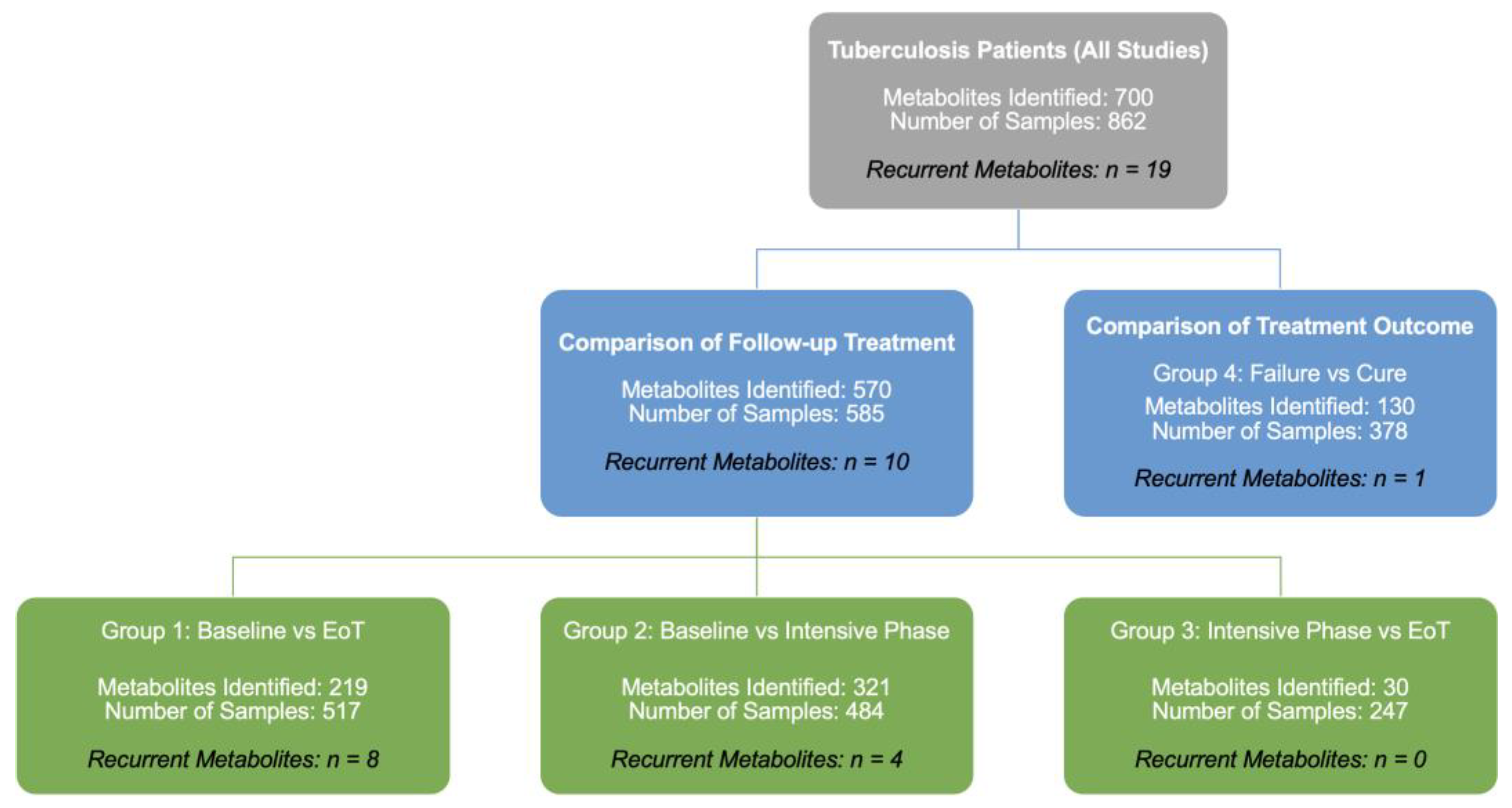
3.4. Treatment Response Comparison Group
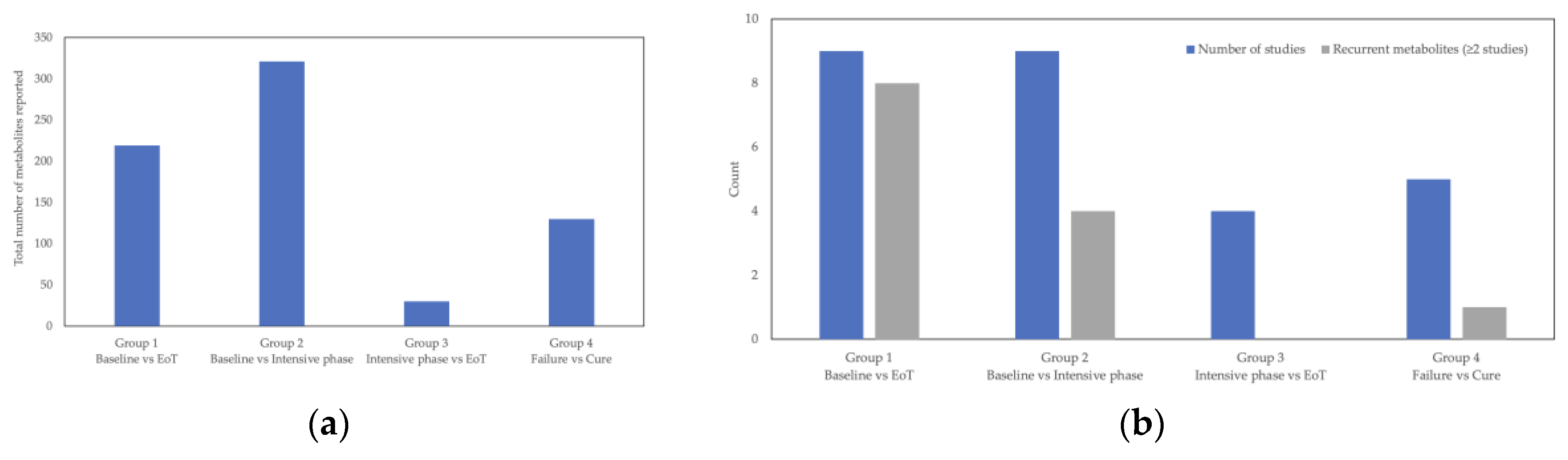
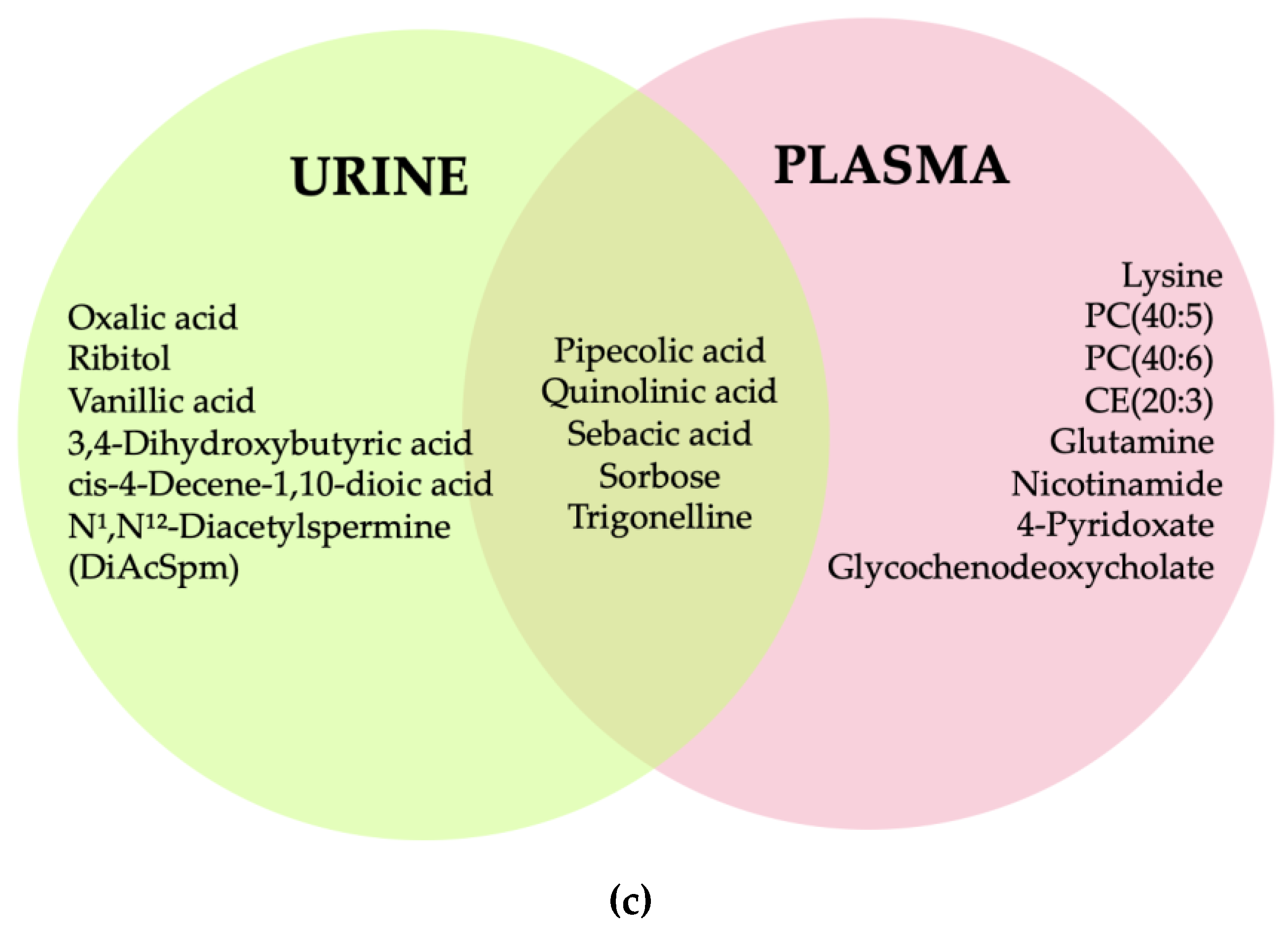
3.5. Summary of Metabolites by Comparison Group
3.6. Risk of Bias Assessment
| Table 3. A. Prediction model studies (PROBAST). | ||||||||
| Study (Year) [Ref] | Study classification | Participants | Predictors | Outcome | Analysis | Overall RoB | Overall applicability | Key concern(s) |
| Collins et al., 2025 [17] | Prediction model (external evaluation) | Low | Low | Low | Low | Low | Low | Externally evaluated; clear modeling |
| Nguyen et al., 2023 [23] | Prediction model / ML classifier | Unclear | Low | Low | High | High | Unclear | Small N vs predictors; limited validation |
| Mahapatra et al., 2014 [16] | Prediction model / ML classifier | High | High | Unclear | High | High | Unclear | Feature-level reporting; unclear model handling |
| Luies et al., 2017 [30] | Prediction model / ML classifier | High | Low | Low | High | High | Unclear | Small N; overfitting risk |
| Dutta et al., 2020 [28] | Prediction model / ML classifier | Unclear | Unclear | Low | High | High | Unclear | Complex integrated analysis; limited validation |
| Shivakoti et al., 2022 [27] | Prediction model / ML classifier | Unclear | Unclear | Unclear | High | High | Unclear | Confounding; limited model reporting |
| Tornheim et al., 2022 [18] | Prediction model / ML classifier | Unclear | Unclear | Unclear | High | High | Unclear | Small cohort; limited performance reporting |
| Study (Year) [Ref] | Study classification | Participation | Factor measurement | Outcome measurement | Confounding | Analysis/reporting | Overall RoB |
|---|---|---|---|---|---|---|---|
| Luies et al., 2017a [29] | Association / longitudinal metabolite study | Moderate | Moderate | Low | Moderate | Moderate | Moderate |
| Nguyen et al., 2024 [24] | Association / longitudinal metabolite study | Moderate | Low | Low | Moderate | Moderate | Moderate |
| Fitzgerald et al., 2019 [26] | Association / longitudinal metabolite study | Moderate | Low | Low | High | High | High |
| Combrink et al., 2019 [20] | Association / longitudinal metabolite study | High | Low | Moderate | Moderate | Moderate | High |
| Xia et al., 2020 [21] | Association / longitudinal metabolite study | Low | Low | Low | Low | Low | Low |
| Gatechompol et al., 2024 [19] | Association / longitudinal metabolite study | Moderate | Low | Moderate | Moderate | Moderate | Moderate |
| Yang et al., 2024 [22] | Association / longitudinal metabolite study | Low | Low | Low | High | High | High |
| Arriaga et al., 2022 [25] | Association / longitudinal metabolite study | Low | Low | Low | Low | Low | Low |
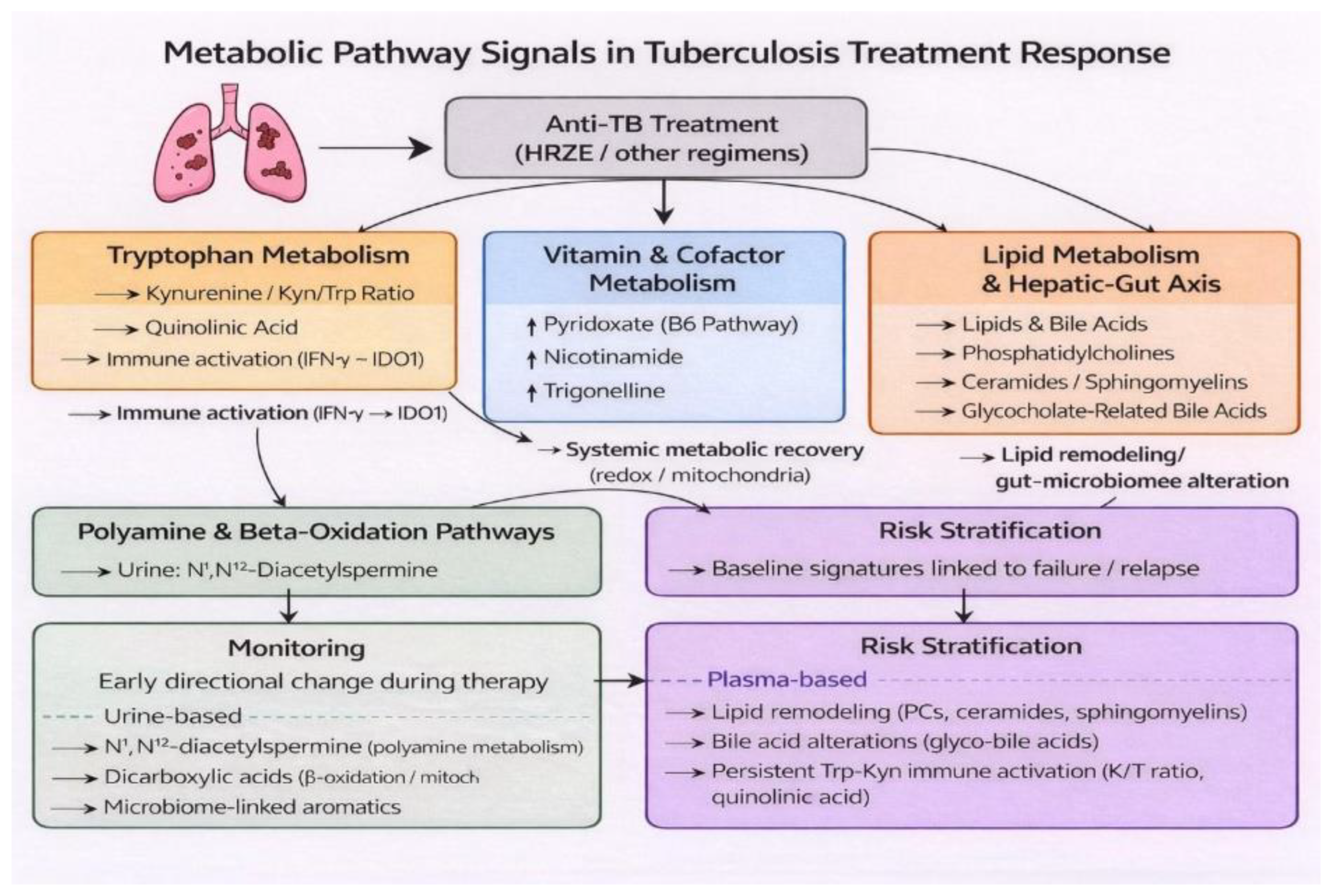
3.7. Metabolic Pathway Synthesis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| TB | Tuberculosis |
| LC | Liquid chromatography |
| LC-MS | Liquid chromatography-mass spectrometry |
| MS | Mass spectrometry |
| MS/MS | Tandem mass spectrometry |
| GC | Gas chromatography |
| GC-MS | Gas chromatography-mass spectrometry |
| GC×GC–MS | Two-dimensional gas chromatography time-of-flight |
| CE | Capillary electrophoresis |
| CE–MS | Capillary electrophoresis–mass spectrometry |
| NMR | Nuclear magnetic resonance |
| PRISMA | Preferred Reporting Items for Systematic Reviews and Meta-Analyses |
| PROSPERO | International Prospective Register of Systematic Reviews |
| RCT | Randomized controlled trial |
| GenAI | Generative artificial intelligence |
| PROBAST | Prediction Model Risk Of Bias Assessment Tool |
| QUIPS | Quality In Prognostic Studies |
| ELISA | Enzyme-linked immunosorbent assay |
| EoT | End of treatment |
| ML | Machine learning |
| MRM | Multiple Reaction Monitoring |
| TOF | Time of flight |
| GC×GC–TOFMS | Two-dimensional gas chromatography time-of-flight mass spectrometry |
| UPLC | Ultra performance liquid chromatography |
| K/T ratio | Kynurenine/tryptophan ratio |
| Trp–Kyn | Tryptophan-kynurenine |
References
- Team, W.; Global Programme on Tuberculosis and Lung Health (GTB) Global Tuberculosis Report 2025; 2025;
- Goletti, D.; Meintjes, G.; Andrade, B.B.; Zumla, A.; Shan Lee, S. Insights from the 2024 WHO Global Tuberculosis Report – More Comprehensive Action, Innovation, and Investments Required for Achieving WHO End TB Goals. Int. J. Infect. Dis. 2025, 150, 107325. [CrossRef]
- Pai, M.; Behr, M.A.; Dowdy, D.; Dheda, K.; Divangahi, M.; Boehme, C.C.; Ginsberg, A.; Swaminathan, S.; Spigelman, M.; Getahun, H.; et al. Tuberculosis. Nat. Rev. Dis. Prim. 2016.
- MacLean, E.L.H.; Zimmer, A.J.; den Boon, S.; Gupta-Wright, A.; Cirillo, D.M.; Cobelens, F.; Gillespie, S.H.; Nahid, P.; Phillips, P.P.; Ruhwald, M.; et al. Tuberculosis Treatment Monitoring Tests during Routine Practice: Study Design Guidance. Clin. Microbiol. Infect. 2024, 30, 481–488. [CrossRef]
- Zimmer, A.J.; Lainati, F.; Vasquez, N.A.; Chedid, C.; McGrath, S.; Benedetti, A.; MacLean, E.; Ruhwald, M.; Denkinger, C.M.; Kohli, M. Biomarkers That Correlate with Active Pulmonary Tuberculosis Treatment Response: A Systematic Review and Meta-Analysis. J. Clin. Microbiol. 2022, 60. [CrossRef]
- WHO 2022 Update; 2022; ISBN 9789240065093.
- Organization, W.H. Companion Handbook; 2014; ISBN 9789241548441.
- Wallis, R.S.; Wang, C.; Doherty, T.M.; Onyebujoh, P.; Vahedi, M.; Laang, H.; Olesen, O.; Parida, S.; Zumla, A. Biomarkers for Tuberculosis Disease Activity, Cure, and Relapse. Lancet Infect. Dis. 2010, 10, 68–69. [CrossRef]
- Babu, S. Biomarkers for Treatment Monitoring in Tuberculosis: A New Hope. EBioMedicine 2017, 26, 13–14. [CrossRef]
- Tounta, V.; Liu, Y.; Cheyne, A.; Larrouy-Maumus, G. Metabolomics in Infectious Diseases and Drug Discovery. Mol. Omi. 2021, 17, 376–393. [CrossRef]
- Yu, Y.; Jiang, X.X.; Li, J.C. Biomarker Discovery for Tuberculosis Using Metabolomics. Front. Mol. Biosci. 2023, 10, 1–23. [CrossRef]
- Dalamaga, M. Clinical Metabolomics: Useful Insights, Perspectives and Challenges. Metab. Open 2024, 22, 100290. [CrossRef]
- Abuawad, A.; Romero, M.; Jarquin, S.M.; Ghaemmaghami, A.M.; Kim, D.-H. Metabolomic Profiling of Host–Pathogen Interactions: Differential Effects of Gram-Positive and Gram-Negative Bacterial Secretomes on THP-1 Macrophage Metabolism. RSC Adv. 2025, 15, 40607–40618. [CrossRef]
- Ahmadi, S.; Sedaghat, F.R.; Memar, M.Y.; Yekani, M. Metabolomics in the Diagnosis of Bacterial Infections. Clin. Chim. Acta 2025, 565, 120020. [CrossRef]
- Ogger, P.P.; Murray, P.J. Dissecting Inflammation in the Immunemetabolomic Era. Cell. Mol. Life Sci. 2025, 82. [CrossRef]
- Mahapatra, S.; Hess, A.M.; Johnson, J.L.; Eisenach, K.D.; DeGroote, M.A.; Gitta, P.; Joloba, M.L.; Kaplan, G.; Walzl, G.; Boom, W.H.; et al. A Metabolic Biosignature of Early Response to Anti-Tuberculosis Treatment. BMC Infect. Dis. 2014, 14, 1–11. [CrossRef]
- Collins, J.M.; Bobosha, K.; Narayanan, N.; Gandhi, N.R.; Day, C.L.; Rengarajan, J.; Kempker, R.R.; Lau, M.S.Y.; Nellis, M.; Tukvadze, N.; et al. A Plasma Metabolic Signature to Diagnose Pulmonary Tuberculosis and Monitor Treatment Response. J. Infect. Dis. 2025, 232, 578–587. [CrossRef]
- Tornheim, J.A.; Paradkar, M.; Zhao, H.; Kulkarni, V.; Pradhan, N.; Kinikar, A.; Kagal, A.; Gupte, N.; Mave, V.; Gupta, A.; et al. The Kynurenine/Tryptophan Ratio Is a Sensitive Biomarker for the Diagnosis of Pediatric Tuberculosis Among Indian Children. Front. Immunol. 2022, 12, 1–6. [CrossRef]
- Gatechompol, S.; Lutter, R.; Vaz, F.M.; Ubolyam, S.; Avihingsanon, A.; Kerr, S.J.; van Leth, F.; Cobelens, F. The Plasma Kynurenine-to-Tryptophan Ratio as a Biomarker of Tuberculosis Disease in People Living with HIV on Antiretroviral Therapy: An Exploratory Nested Case–Control Study. BMC Infect. Dis. 2024, 24, 1–11. [CrossRef]
- Combrink, M.; Du Preez, I.; Ronacher, K.; Walzl, G.; Loots, D.T. Time-Dependent Changes in Urinary Metabolome before and after Intensive Phase Tuberculosis Therapy: A Pharmacometabolomics Study. Omi. A J. Integr. Biol. 2019, 23, 560–572. [CrossRef]
- Xia, Q.; Lee, M.H.; Walsh, K.F.; McAulay, K.; Bean, J.M.; Fitzgerald, D.W.; Dupnik, K.M.; Johnson, W.D.; Pape, J.W.; Rhee, K.Y.; et al. Urinary Biomarkers of Mycobacterial Load and Treatment Response in Pulmonary Tuberculosis. JCI Insight 2020, 5, 1–16. [CrossRef]
- Yang, M.; Wang, W.; Zhang, P.; Chen, X.; Liu, G.; Lu, H.; He, M.; Deng, G. Variations in Quinolinic Acid Levels in Tuberculosis Patients with Diabetes Comorbidity: A Pilot Prospective Cohort Study. Infect. Drug Resist. 2024, 17, 2975–2985. [CrossRef]
- Anh, N.K.; Phat, N.K.; Yen, N.T.H.; Jayanti, R.P.; Thu, V.T.A.; Park, Y.J.; Cho, Y.S.; Shin, J.G.; Kim, D.H.; Oh, J.Y.; et al. Comprehensive Lipid Profiles Investigation Reveals Host Metabolic and Immune Alterations during Anti-Tuberculosis Treatment: Implications for Therapeutic Monitoring. Biomed. Pharmacother. 2023, 158, 114187. [CrossRef]
- Anh, N.K.; Yen, N.T.H.; Tien, N.T.N.; Phat, N.K.; Park, Y.J.; Kim, H.S.; Vu, D.H.; Oh, J.Y.; Kim, D.H.; Long, N.P. Metabolic Phenotyping and Global Functional Analysis Facilitate Metabolic Signature Discovery for Tuberculosis Treatment Monitoring. Biochim. Biophys. Acta - Mol. Basis Dis. 2024, 1870, 167064. [CrossRef]
- Arriaga, M.B.; Karim, F.; Queiroz, A.T.L.; Araújo-Pereira, M.; Barreto-Duarte, B.; Sales, C.; Moosa, M.Y.S.; Mazibuko, M.; Milne, G.L.; Maruri, F.; et al. Effect of Dysglycemia on Urinary Lipid Mediator Profiles in Persons With Pulmonary Tuberculosis. Front. Immunol. 2022, 13, 1–12. [CrossRef]
- Fitzgerald, B.L.; Islam, M.N.; Graham, B.; Mahapatra, S.; Webb, K.; Boom, W.H.; Malherbe, S.T.; Joloba, M.L.; Johnson, J.L.; Winter, J.; et al. Elucidation of a Human Urine Metabolite as a Seryl-Leucine Glycopeptide and as a Biomarker of Effective Anti-Tuberculosis Therapy. ACS Infect. Dis. 2019, 5, 353–364. [CrossRef]
- Shivakoti, R.; Newman, J.W.; Hanna, L.E.; Queiroz, A.T.L.; Borkowski, K.; Gupte, A.N.; Paradkar, M.; Satyamurthi, P.; Kulkarni, V.; Selva, M.; et al. Host Lipidome and Tuberculosis Treatment Failure. Eur. Respir. J. 2022, 59, 1–13. [CrossRef]
- Dutta, N.K.; Tornheim, J.A.; Fukutani, K.F.; Paradkar, M.; Tiburcio, R.T.; Kinikar, A.; Valvi, C.; Kulkarni, V.; Pradhan, N.; Shivakumar, S.V.B.Y.; et al. Integration of Metabolomics and Transcriptomics Reveals Novel Biomarkers in the Blood for Tuberculosis Diagnosis in Children. Sci. Rep. 2020, 10, 1–11. [CrossRef]
- Luies, L.; Reenen, M. Van; Ronacher, K.; Walzl, G.; Loots, D.T. Predicting Tuberculosis Treatment Outcome Using Metabolomics. Biomark. Med. 2017, 11, 1057–1067. [CrossRef]
- Luies, L.; Mienie, J.; Motshwane, C.; Ronacher, K.; Walzl, G.; Loots, D.T. Urinary Metabolite Markers Characterizing Tuberculosis Treatment Failure. Metabolomics 2017, 13, 0. [CrossRef]
- Kirwan, J.A. Translating Metabolomics into Clinical Practice. Nat. Rev. Bioeng. 2023, 1, 228–229.
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. Bmj 2021, 372. [CrossRef]
- Scalbert, A.; Brennan, L.; Fiehn, O.; Hankemeier, T.; Kristal, B.S.; van Ommen, B.; Pujos-Guillot, E.; Verheij, E.; Wishart, D.; Wopereis, S. Mass-Spectrometry-Based Metabolomics: Limitations and Recommendations for Future Progress with Particular Focus on Nutrition Research. Metabolomics 2009, 5, 435–458. [CrossRef]
- Long, N.P.; Heo, D.Y.; Park, S.; Yen, N.T.H.; Cho, Y.S.; Shin, J.G.; Oh, J.Y.; Kim, D.H. Molecular Perturbations in Pulmonary Tuberculosis Patients Identified by Pathway-Level Analysis of Plasma Metabolic Features. PLoS One 2022, 17, 1–13. [CrossRef]
- Mellor, A. Indoleamine 2,3 Dioxygenase and Regulation of T Cell Immunity. Biochem. Biophys. Res. Commun. 2005, 338, 20–24. [CrossRef]
- Suzuki, Y.; Miwa, S.; Akamatsu, T.; Suzuki, M.; Fujie, M.; Nakamura, Y.; Inui, N.; Hayakawa, H.; Chida, K.;; Suda, T.; Indoleamine 2,3-Dioxygenase in the Pathogenesis of Tuberculous Pleurisy. Int. J. Tuberc. Lung Dis. 2013, 17, 1501–1506, doi:DOI: 10.5588/ijtld.13.0082.
- Badawy, A.A.B. Kynurenine Pathway of Tryptophan Metabolism: Regulatory and Functional Aspects. Int. J. Tryptophan Res. 2017, 10. [CrossRef]
- Adu-Gyamfi, C.G.; Savulescu, D.; George, J.A.; Suchard, M.S. Indoleamine 2, 3-Dioxygenase-Mediated Tryptophan Catabolism: A Leading Star or Supporting Act in the Tuberculosis and HIV Pas-de-Deux? Front. Cell. Infect. Microbiol. 2019, 9, 1–12. [CrossRef]
- Cervenka, I.; Agudelo, L.Z.; Ruas, J.L. Kynurenines: Tryptophan’s Metabolites in Exercise, Inflammation, and Mental Health. Science (80-. ). 2017, doi:DOI: 10.1126/science.aaf9794.
- Seo, S.K.; Kwon, B. Immune Regulation through Tryptophan Metabolism. Exp. Mol. Med. 2023, 55, 1371–1379. [CrossRef]
- Russell, D.G.; Cardona, P.-J.; Kim, M.-J.; Allain, S.; Altare, F. Foamy Macrophages and the Progression of the Human Tuberculosis Granuloma. Nat. Immunol. 2009, 10, 943–948.
- Lovewell, R.R.; Sassetti, C.M.; VanderVen, B.C. Chewing the Fat: Lipid Metabolism and Homeostasis during M. Tuberculosis Infection. Curr. Opin. Microbiol. 2016, 29, 30–36. [CrossRef]
- Fleishman, J.S.; Kumar, S. Bile Acid Metabolism and Signaling in Health and Disease: Molecular Mechanisms and Therapeutic Targets. Signal Transduct. Target. Ther. 2024, 9. [CrossRef]
- Tamber, S.S.; Bansal, P.; Sharma, S.; Singh, R.B.; Sharma, R. Biomarkers of Liver Diseases. Mol. Biol. Rep. 2023, 50, 7815–7823. [CrossRef]
- Qi, L.; Chen, Y. Circulating Bile Acids as Biomarkers for Disease Diagnosis and Prevention. J. Clin. Endocrinol. Metab. 2023, 108, 251–270. [CrossRef]
- Zou, P.; Wang, L. Dietary Pattern and Hepatic Lipid Metabolism. Liver Res. 2023, 7, 275–284. [CrossRef]
- Kang, S.; Jeong, D.Y.; Seo, J.; Daily, J.W.; Park, S. Microbiota-Mediated Bile Acid Metabolism as a Mechanistic Framework for Precision Nutrition in Gastrointestinal and Metabolic Diseases. Cells 2025, 15, 1–30. [CrossRef]
- González-Domínguez, R.; González-Domínguez, Á.; Sayago, A.; Fernández-Recamales, Á. Recommendations and Best Practices for Standardizing the Pre-Analytical Processing of Blood and Urine Samples in Metabolomics. Metabolites 2020, 10, 1–18. [CrossRef]
- Ali, A.M.; Monaghan, C.; Muggeridge, D.J.; Easton, C.; Watson, D.G. LC/MS-Based Discrimination between Plasma and Urine Metabolomic Changes Following Exposure to Ultraviolet Radiation by Using Data Modelling. Metabolomics 2023, 19, 1–15. [CrossRef]
- Qiu, S.; Cai, Y.; Yao, H.; Lin, C.; Xie, Y.; Tang, S.; Zhang, A. Small Molecule Metabolites: Discovery of Biomarkers and Therapeutic Targets. Signal Transduct. Target. Ther. 2023, 8, 1–37. [CrossRef]
- Wolff, R.F.; Moons, K.G.M.; Riley, R.D.; Whiting, P.F.; Westwood, M.; Collins, G.S.; Reitsma, J.B.; Kleijnen, J.; Mallett;, S.; Group, P. PROBAST: A Tool to Assess the Risk of Bias and Applicability of Prediction Model Studies. Ann. Intern. Med. 2019, 170.
- Hayden, J.A.; Windt, D.A. van der; Cartwright, J.L.; Côté, P.; Bombardier, C. Assessing Bias in Studies of Prognostic Factors. Ann. Intern. Med. 2013, 158.
| Author, Publication Year [Ref] | Setting and population | Study Design | Biospeciment | Platform and approach | Sample size | Sampling timpoints | Metabolite focus | Outcome comparison | |
|---|---|---|---|---|---|---|---|---|---|
| TB | Non-TB | ||||||||
| Luies et all., 2017 [30] | Adults with pulmonary TB; South Africa | Prospective longitudinal cohort | Urine | GC–MS/GC×GC–MS, Untargeted | 41 | 0 | Baseline | Global metabolites | Group 4 |
| Nguyen Ky Anh et al., 2024 [24] | Adults with pulmonary TB; diabetes (17.1%); Korea | Prospective observational longitudinal cohort | Plasma | LC–MS, Untargeted & targeted | 41 | 0 | Baseline; intensive (between week 6 & 11); EoT (between week 18 & 26) | Polar and bile acids/ lipids | Group 1 and Group 2 |
| Nguyen Ky Anh et al., 2023 [23] | Adults with pulmonary TB; diabetes (17.1%); Korea | Longitudinal cohort study | Plasma | LC–MS, Targeted lipidomics | 41 | 0 | Baseline; intensive (between week 6 & 11); EoT (between week 18 & 26) | Lipids | Group 1 and Group 2 |
| Fitzgerald BL et al., 2019 [26] | Adults with pulmonary TB; HIV (-); Multi-center (Uganda & South Africa) | Uganda cohort (KCHS): Cohort subset / case-contact study South Africa cohort (Catalysis Study): Longitudinal outcome cohort |
Urine | LC–MS, Untargeted | 45 | 39 | Baseline; week 1/2/4/8, EoT | Global metabolites | Group 2 andGroup 4 |
| Dutta NK et al., 2020 [28] | Children with pulmonary and extrapulmonary TB; India | Longitudinal nested case–control study (within the CTRIUMPH cohort) | Plasma | LC–MS/MS, Untargeted | 16 | 16 | Baseline; month 1; month 6 (EoT) | Polar metabolites | Group 1 andGroup 2 and Group 3 |
| Combrink M et al., 2019 [20] | Adults with pulmonary TB; South Africa | Prospective longitudinal pharmacometabolomics study | Urine | GC×GC–TOFMS, Untargeted | 23 | 0 | Baseline; week 1/2/4 | Global metabolites | Group 2 |
| Collins JM et al., 2025 [17] | Adults with pulmonary TB; Ethiopia | Case-control + longitudinal follow-up | Plasma | LC–MS (panel) & ML, Targeted & ML | 82 | 104 | Baseline; Month 2/6/12 after treatment | 153-metabolite panel | Group 1 and Group 2 and Group 3 |
| Xia Q et al., 2020 [21] | Adults with pulmonary TB; HIV (Africa 11.8% & Haiti 0%); Multi-center (Africa & Haiti) | Prospective longitudinal cohort analysis (2 cohorts) | Urine | LC–MS & ELISA, Targeted | 69 | 0 | Baseline; weeks 2/4/8/17/26; week 52 post-treatment) | DiAcSpm | Group 1 and Group 4 |
| Shivakoti R et al., 2022 [27] | Adults with pulmonary TB; diabetes (32%), HIV (2%); India | Case-control study, nested within a prospective cohort | Plasma | LC–MS, Untargeted | 192 | 0 | Baseline | Global metabolites | Group 4 |
| Mahapatra S et al., 2014 [16] | Adults with pulmonary TB; HIV (-); Multi-center (Uganda & South Africa) | Prospective observational cohort of TB patients with longitudinal treatment sampling | Urine | LC–MS, Untargeted | 87 | 0 | Baseline; month 1/2/6 | Global metabolites | Group 1 and Group 2 |
| Gatechompol S et al., 2024 [19] | Adults with pulmonary TB; HIV (+); Thailand | Nested case-control (within prospective HIV cohort on ART) | Plasma | LC–MS/MS, Targeted | 13 | 13 | Pre-TB (6 months before TB diagnosis); diagnosis (baseline); EoT (6 months after TB treatment) | Tryptophan–Kynurenine pathway | Group 1 and Group 4 |
| Yang et al., 2024 [22] | Adults with pulmonary TB; type 2 diabetes mellitus (50%), HIV (-); China | Prospective cohort (targeted metabolite quantification) | Plasma | UPLC–MRM, Targeted | 32 | 32 | Baseline; month 6 of post-treatment | Quinolinic acid panel | Group 1 |
| Luies et al., 2017 [29] | Adults with pulmonary TB; South Africa | Prospective observational cohort study | Urine | GC×GC–TOFMS, Untargeted | 31 | 0 | Baseline | Global metabolites | Group 4 |
| Tornheim JA et al., 2022 [18] | Children with pulmonary and extrapulmonary TB; HIV (-); India | Targeted diagnostic accuracy analysis (secondary analysis from cohort biorepository) | Plasma | LC–MS/MS, Targeted | 16 | 32 | Baseline; month 1; EoT | Tryptophan–Kynurenine pathway |
Group 1 and Group 2 |
| Arriaga MB et al., 2022 [25] | Adults with pulmonary TB; dysglycemia (31.1%), HIV (22.3%); Multi-center (Brazil & South Africa) | Prospective longitudinal cohort | Urine | UPLC–MS/MS, Targeted | 133 | 60 | Baseline; month 2; month 6 (EoT) | Eicosanoids | Group 1 and Group 2 and Group 3 |
| Comp. group | Definition | Biospecimen | No. studies | Total metabolite | No. recurrent | Recurrent metabolites (≥2 studies)1 | Predominant direction with successful therapy | Supporting studies [Ref] |
|---|---|---|---|---|---|---|---|---|
| Group 1 | Baseline vs EoT | Plasma | 6 | 157 | 7 | 4-Pyridoxate; Glutamine; Glycochenodeoxycholate; Lysine; Nicotinamide; Quinolinic acid; Trigonelline (N’-methylnicotinate) | Mostly ↓ from baseline to EoT (normalization) | [17,18,22,24,28] |
| Urine | 3 | 62 | 1 | N¹,N¹²-Diacetylspermine (DiAcSpm) | Mostly ↓ from baseline to EoT (normalization) | [16,21,25] | ||
| Group 2 | Baseline vs intensive phase | Plasma | 5 | 138 | 4 | 4-Pyridoxate; Glycochenodeoxycholate; Nicotinamide; Trigonelline (N’-methylnicotinate) | Mostly ↓ early during intensive phase | [17,18,23,24,28] |
| Urine | 4 | 183 | 0 | NR | — | [16,20,25,26] | ||
| Group 3 | Intensive phase vs EoT | Plasma | 3 | 18 | 0 | NR | — | [17,18,28] |
| Urine | 1 | 12 | 0 | NR | — | [25] | ||
| Group 4 | Treatment failure vs cure | Plasma | 1 | 62 | 0 | NR | — | [27] |
| Urine | 4 | 68 | 1 | cis-4-Decene-1,10-dioic acid | Higher in failure/non-response (unfavorable outcome) | [21,26,29,30] |
| Comparison group | Plasma (representative metabolites) | Urine (representative metabolites) | Key pathway theme(s) | Key supporting studies [Ref] | Clinical relevance |
|---|---|---|---|---|---|
| Group 1 (baseline vs EoT) | K/T ratio; quinolinic acid; nicotinamide; glutamine; glycochenodeoxycholate | N¹,N¹²-diacetylspermine | Trp–Kyn; vitamin/cofactor; bile acids | [16,17,18,21,22,24,25,28] | Treatment monitoring |
| Group 2 (baseline vs intensive) | K/T ratio; 4-Pyridoxate; nicotinamide; trigonelline; bile acids | NR; multiple unique features | Trp–Kyn; Early immunometabolic shift; polyamines; β-oxidation | [16,17,18,20,23,24,25,26,28] | Treatment monitoring |
| Group 3 (intensive vs EoT) | K/T ratio | NR | Trp–Kyn; Late-phase metabolic normalization (heterogeneous) | [17,18,25,28] | Treatment monitoring |
| Group 4 (failure vs cure) | NR | cis-4-Decene-1,10-dioic acid; aromatic metaboloties | Lipid remodeling; β-oxidation; microbiome-related aromatics | [21,26,27,29,30] | Risk stratification |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




