Submitted:
23 March 2026
Posted:
24 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- 1.
- A FHIR-faithful RDF graph that follows the HL7 FHIR-RDF specifications [8] and therefore preserves native nesting, references, and round-trip fidelity to JSON and XML representations; and
- 2.
- An OWL-compliant knowledge graph obtained via ontologization, optimized for concise data exploration and semantic integration.
- 1.
- Structural Ontologization: We present a transformative architecture that intentionally relaxes FHIR’s strict round-trip constraints to prioritize analytic accessibility. By collapsing the multi-hop blank-node traversals of standard FHIR.ttl into single-step property assertions, we deliver an OWL-compliant knowledge graph optimized for complex, cohort-level exploration without sacrificing clinical fidelity.
- 2.
- Terminology-Grounded Querying: We integrate ICD-9-CM and ICD-10-CM terminologies directly into the graph model, elevating clinical codes from flat strings to explicit, interoperable resource instances. This enables reproducible, terminology-driven validation and cross-system consistency.
- 3.
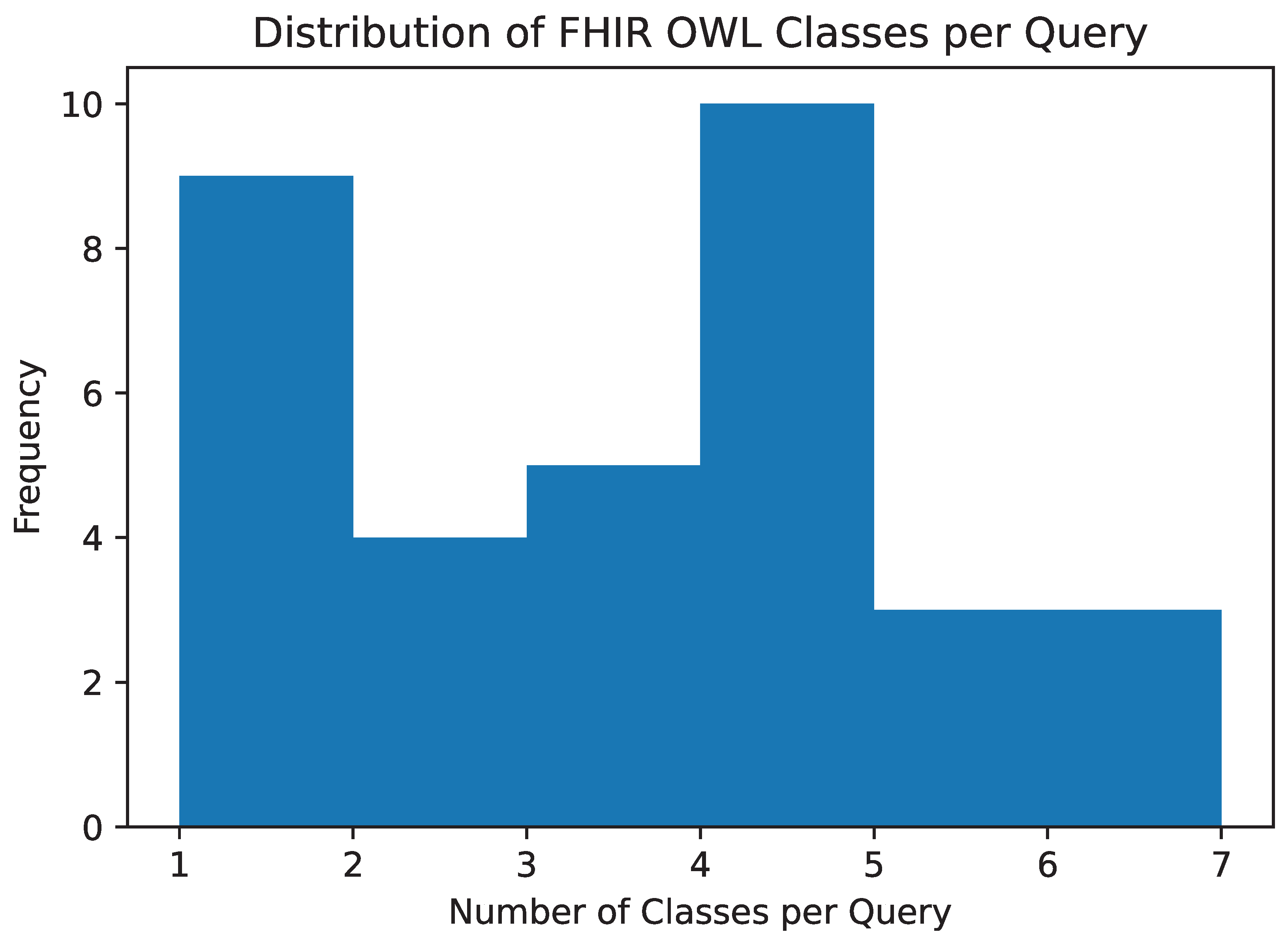
- Democratized Analytic Access: We introduce a highly accurate (mean accuracy >95%) NL2SPARQL interface that effectively bridges the gap between foundation large language models and formal graph query languages. Guided directly by the OWL schema, this tool allows domain experts to execute robust exploratory analyses using natural language, significantly lowering the barrier to entry for clinical research.
1.1. Other Approaches
2. Methods
-
FHIR-Faithful RDF Graph.As an initial transformation step, we converted the FHIR NDJSON resources into an RDF graph that preserves the native FHIR nesting structure, repeating elements, choice-type fields, and inter-resource references. This representation maintains the structural fidelity of the original FHIR specification and functions primarily as an intermediate waystation and baseline artifact for transformation validation and structural inspection, rather than as a standalone analytic model.
-
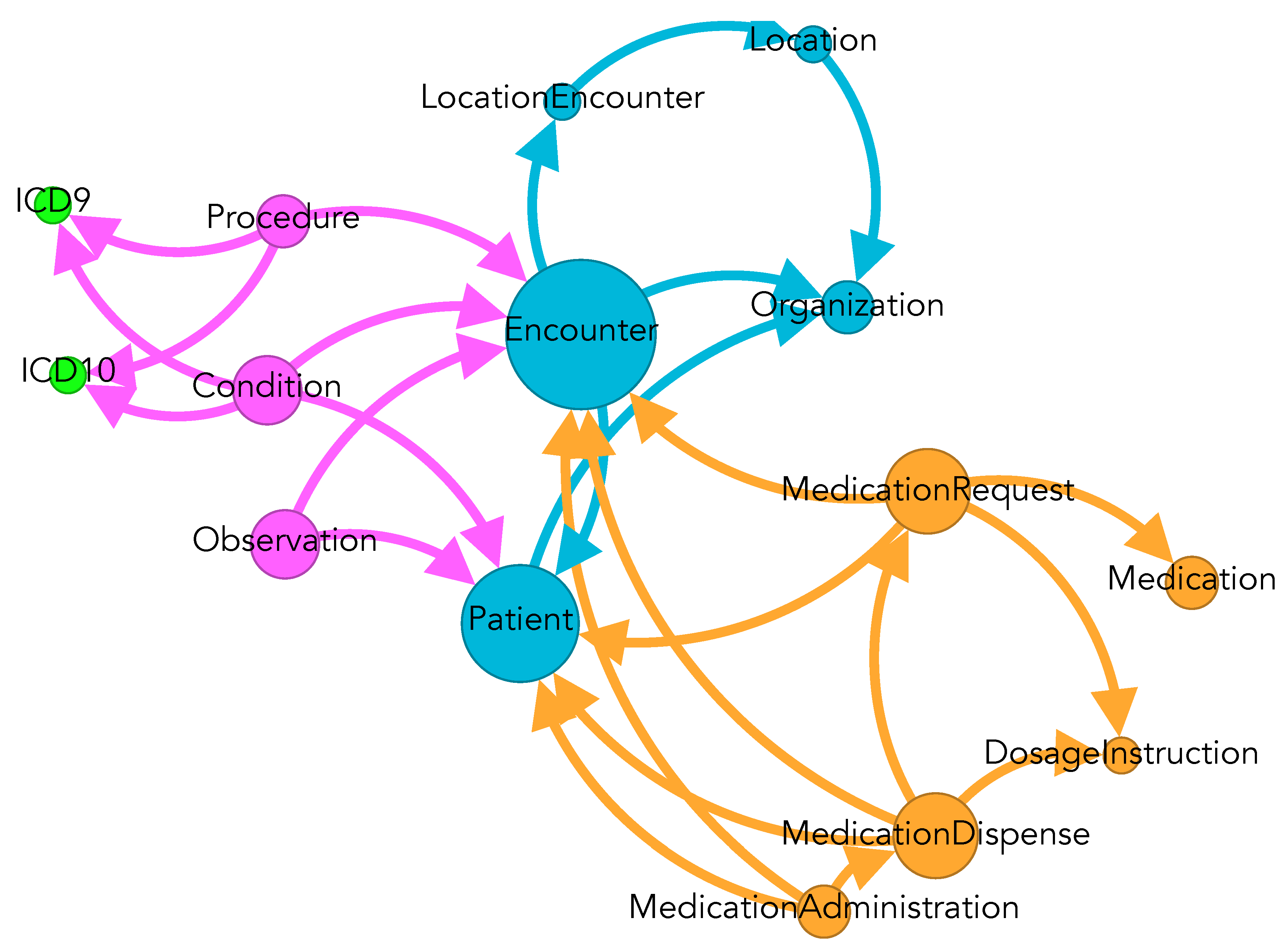
Ontologized OWL Graph.From the FHIR-faithful RDF graph, we then derived an OWL-compliant knowledge graph following the principles of ontology construction [10]. This transformation resolves deep nesting, variable-width arrays, and choice-type fields, enforces explicit distinctions between classes and individuals, and defines object properties and datatype properties. Selected FHIR resources were integrated with ICD-9-CM and ICD-10-CM terminologies using UMLS-derived mappings. For selected analyses, including evaluation of the NL2SPARQL interface and ontology inspection in Protégé [21], we derived task-specific projections of the ontologized OWL graph. These projections exclude high-volume resource types not required for a given task (e.g., Observation), and apply minor schema adjustments to improve tooling compatibility. They correspond to external schema views and do not constitute alternative ontologies or independent data models, but rather query-driven views of the same ontologized OWL graph [6].
2.1. Data Source and Conversion
2.2. Extract–Transform–Load Pipeline
@prefix se: <http://mimic-fhir.local/ontology#> .
@prefix fhir: <http://hl7.org/fhir/> .
@prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix owl: <http://www.w3.org/2002/07/owl#> .
@prefix xsd: <http://www.w3.org/2001/XMLSchema#> .
valueQuantity → flat:value_quantity
valueString → flat:value_string
valueCodeableConcept → flattened code/display properties
- 1.
- Code–label validation. Linking codes to explicit ICD resources enables internal consistency checks between stored codes and associated human-readable labels.
- 2.
- Terminology-grounded querying. Queries may reference standardized code identifiers directly, rather than relying on string matching of literal fields. This improves query robustness and reproducibility across datasets that share the same coding systems.
2.3. Schema-Grounded NL2SPARQL Query Interface
2.4. Validation of the Ontologized Graph
3. Results
How many unique patients have at least one diagnosis of essential hypertension (ICD-10 I10)?
4. Discussion
4.1. Limitations and Future Work
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mangalampalli, B.M.; Kolla, T. FHIR-Based Interoperability Frameworks For Real-Time Healthcare Data Exchange: Architecture Patterns And Performance Optimization. International Journal Of Advances in Signal and Image Sciences 2026, 1514–1536. [Google Scholar] [CrossRef]
- Gruendner, J.; Gulden, C.; Kampf, M.; Mate, S.; Prokosch, H.U.; Zierk, J. A framework for criteria-based selection and processing of fast healthcare interoperability resources (FHIR) data for statistical analysis: design and implementation study. JMIR Medical Informatics 2021, 9, e25645. [Google Scholar] [CrossRef] [PubMed]
- Gruendner, J.; Deppenwiese, N.; Folz, M.; Köhler, T.; Kroll, B.; Prokosch, H.U.; Rosenau, L.; Rühle, M.; Scheidl, M.A.; Schüttler, C.; et al. The architecture of a feasibility query portal for distributed COVID-19 Fast Healthcare Interoperability Resources (FHIR) patient data repositories: design and implementation study. JMIR Medical Informatics 2022, 10, e36709. [Google Scholar] [CrossRef] [PubMed]
- Vorisek, C.N.; Lehne, M.; Klopfenstein, S.A.I.; Mayer, P.J.; Bartschke, A.; Haese, T.; Thun, S. Fast healthcare interoperability resources (FHIR) for interoperability in health research: systematic review. JMIR Medical Informatics 2022, 10, e35724. [Google Scholar] [CrossRef] [PubMed]
- Löbe, M.; Draeger, C.; Strübing, A.; Palm, J.; Meineke, F.A.; Winter, A. Pitfalls in Analyzing FHIR Data from Different University Hospitals. In Proceedings of the GMDS; 2023; pp. 146–151. [Google Scholar] [CrossRef]
- Smith, B. Beyond concepts: ontology as reality representation. In Proceedings of the Third International Conference on Formal Ontology in Information Systems (FOIS 2004), 2004; IOS Press: Amsterdam; pp. 73–84. [Google Scholar]
- Smith, B.; Ashburner, M.; Rosse, C.; Bard, J.; Bug, W.; Ceusters, W.; Goldberg, L.J.; Eilbeck, K.; Ireland, A.; Mungall, C.J.; et al. The OBO Foundry: coordinated evolution of ontologies to support biomedical data integration. Nature Biotechnology 2007, 25, 1251–1255. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.; Matentzoglu, N.; Overton, J.A.; Vita, R.; Balhoff, J.P.; Buttigieg, P.L.; Carbon, S.; Courtot, M.; Diehl, A.D.; Dooley, D.M.; et al. OBO Foundry in 2021: operationalizing open data principles to evaluate ontologies. Database 2021, 2021, baab069. [Google Scholar] [CrossRef] [PubMed]
- Prud’hommeaux, E.; Collins, J.; Booth, D.; Peterson, K.J.; Solbrig, H.R.; Jiang, G. Development of a FHIR RDF data transformation and validation framework and its evaluation. Journal of Biomedical Informatics 2021, 117, 103755. [Google Scholar] [CrossRef]
- Noy, N.F.; McGuinness, D.L.; et al. Ontology development 101: A guide to creating your first ontology. In Stanford University; 2001. [Google Scholar]
- Kendall, E.F.; McGuinness, D.L. Ontology Engineering; Morgan & Claypool Publishers, 2019. [Google Scholar]
- Grimes, J.; Szul, P.; Metke-Jimenez, A.; Lawley, M.; Loi, K. Pathling: analytics on FHIR. Journal of Biomedical Semantics 2022, 13, 23. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.; Bach, E.; Yang, E.; Pollard, T.; Johnson, A.; Choi, E.; Jia, Y.; Lee, J.H. FHIR-AgentBench: Benchmarking LLM Agents for Realistic Interoperable EHR Question Answering. Proceedings of Machine Learning Research Machine Learning for Health (ML4H) 2025. 2025, 297, 1–19. [Google Scholar]
- Idrissi-Yaghir, A.; Arzideh, K.; Schäfer, H.; Eryilmaz, B.; Bahn, M.; Wen, Y.; Borys, K.; Hartmann, E.; Schmidt, C.; Pelka, O.; et al. Using a Diverse Test Suite to Assess Large Language Models on Fast Health Care Interoperability Resources Knowledge: Comparative Analysis. Journal of Medical Internet Research 2025, 27, e73540. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Camero, I.V.; Garcia-Lopez, E.; Garcia-Cabot, A.; Caro-Alvaro, S. Context-aware few-shot learning SPARQL query generation from natural language on an aviation knowledge graph. Machine Learning and Knowledge Extraction 2025, 7, 52. [Google Scholar] [CrossRef]
- Rosenau, L.; Gruendner, J.; Behrend, P.; Triefenbach, L.; Kurscheidt, M.; Majeed, R.; Prokosch, H.U.; Ingenerf, J. From Feasibility to Insight: Piloting Feature Extraction from FHIR Cohorts to Advance Clinical Research. Research Square 2024. [Google Scholar] [CrossRef]
- Grimes, J.; Brush, R.; Rhyzhikov, N.; Szul, P.; Mandel, J.; Gottlieb, D.; Grieve, G.; Sadjad, B.; Sanyal, A. SQL on FHIR-Tabular views of FHIR data using FHIRPath. npj Digital Medicine 2025, 8, 342. [Google Scholar] [CrossRef] [PubMed]
- Hosch, R.; Baldini, G.; Parmar, V.; Borys, K.; Koitka, S.; Engelke, M.; Arzideh, K.; Ulrich, M.; Nensa, F. FHIR-PYrate: a data science friendly Python package to query FHIR servers. BMC Health Services Research 2023, 23, 734. [Google Scholar] [CrossRef] [PubMed]
- Palm, J.; Meineke, F.A.; Przybilla, J.; Peschel, T. “fhircrackr”: an R package unlocking fast Healthcare Interoperability resources for statistical analysis. Applied Clinical Informatics 2023, 14, 054–064. [Google Scholar] [CrossRef] [PubMed]
- Tomaszuk, D.; Smajevic, A.; Sagi, T.; Hose, K. FHIR Lens: A Graph-Based Approach to Semantic EHR Exploration. 2025 IEEE 38th International Symposium on Computer-Based Medical Systems (CBMS) 2025, 1–6. [Google Scholar] [CrossRef]
- Tudorache, T.; Noy, N.F.; Tu, S.; Musen, M.A. Supporting collaborative ontology development in Protégé. In Proceedings of the International Semantic Web Conference, 2008; Springer; pp. 17–32. [Google Scholar]
- Moody, G.B.; Mark, R.G.; Goldberger, A.L. PhysioNet: Physiologic signals, time series and related open source software for basic, clinical, and applied research. In Proceedings of the 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society. IEEE, 2011; pp. 8327–8330. [Google Scholar] [CrossRef]
- Bennett, A.M.; Ulrich, H.; Van Damme, P.; Wiedekopf, J.; Johnson, A.E. MIMIC-IV on FHIR: converting a decade of in-patient data into an exchangeable, interoperable format. Journal of the American Medical Informatics Association 2023, 30, 718–725. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.; Brochhausen, M. Putting biomedical ontologies to work. Methods of Information in Medicine 2010, 49, 135–140. [Google Scholar] [CrossRef] [PubMed]
- Overton, J.A.; Jackson, R.C.; Matentzoglu, N.; Duncan, W.D.; Vita, R.; Harris, N.L.; Mungall, C.J.; Peters, B. COB: A Core Ontology for Biology and Biomedicine. In Proceedings of the ICBO, 2022; p. 1. [Google Scholar]
- Das, S.; Hussey, P. HL7-FHIR-based ContSys formal ontology for enabling continuity of care data interoperability. Journal of Personalized Medicine 2023, 13, 1024. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef] [PubMed]
- Bennett, A.; Ulrich, H.; Wiedekopf, J.; Szul, P.; Grimes, J.; Johnson, A. MIMIC-IV Clinical Database Demo on FHIR. Version 2.1.0; PhysioNet. 2025. [Google Scholar]



| Structural feature | Implication |
|---|---|
| Deep nesting | Hierarchical data structures extend across multiple levels. |
| Variable-width Arrays | Unbounded repeating elements require explicit modeling. |
| Choice Fields | Multiple value type fields that introduce heterogeneity. |
| Class | Resources | At Least One Outgoing Reference |
|---|---|---|
| Organization | 1 | 0 |
| Location | 39 | 39 |
| Patient | 100 | 100 |
| Encounter | 401 | 401 |
| LocationEncounter (auxiliary) | 972 | 972 |
| Procedure | 2,105 | 2,105 |
| Condition | 4,181 | 4,181 |
| MedicationMix (auxiliary) | 6,461 | 6,461 |
| Specimen | 11,817 | 11,817 |
| MedicationDispense | 13,455 | 13,455 |
| MedicationRequest | 16,366 | 16,366 |
| Medication | 20,075 | 0 |
| DosageInstruction (auxiliary) | 27,768 | 0 |
| MedicationAdministration | 49,635 | 49,635 |
| Observation | 762,633 | 762,633 |
| Type | Max. Array Len. | Max. Nesting Depth | Choice Fields |
|---|---|---|---|
| Observation | 26 | 6 | 1 (4) |
| MedicationAdministration | 1 | 6 | 1 (2) |
| MedicationRequest | 1 | 8 | 1 (2) |
| MedicationDispense | 1 | 8 | 0 (0) |
| Specimen | 1 | 5 | 0 (0) |
| Condition | 1 | 6 | 0 (0) |
| Procedure | 1 | 6 | 1 (2) |
| Medication | 4 | 6 | 0 (0) |
| Encounter | 9 | 6 | 0 (0) |
| Patient | 2 | 7 | 1 (3) |
| Location | 1 | 5 | 0 (0) |
| Organization | 1 | 6 | 0 (0) |
| Locations per encounter | Count | % |
|---|---|---|
| 1 | 153 | 36.9 |
| 2 | 106 | 25.5 |
| 3 | 65 | 15.7 |
| 4 | 45 | 10.8 |
| 5 | 23 | 5.5 |
| 6 | 14 | 3.4 |
| 7 | 4 | 1.0 |
| 9 | 5 | 1.2 |
| Total | 415 | 100.0 |
| Category | FHIR-faithful RDF | Ontologized OWL |
|---|---|---|
| Structure | Nested | Flattened/ontologized |
| Nodes | 20,782,214 | 4,252,079 |
| Edges | 33,133,045 | 15,582,681 |
| Classes (declared) | 12 | 15 |
| OWL Datatype Properties | 0 | 103 |
| OWL Object Properties | 0 | 16 |
| FHIR-faithful representation | Yes | Transformed |
| OWL-compliant | No | Yes |
| RDF-compliant | Yes | Yes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




