Submitted:
19 March 2026
Posted:
20 March 2026
You are already at the latest version
Abstract
Background/Objectives: Drug-resistant tuberculosis (TB) remains a major challenge in high-burden settings, where timely identification of emerging resistance and effective governance responses are critical. While routine molecular diagnostics generate large volumes of resistance-associated mutation data, these outputs are typically used for individual patient management and remain underutilized for population-level surveillance and for the application of clinical governance approaches for improved TB care. Methods: We conducted a retrospective cross-sectional analysis of 1,386 molecular diagnostic records for Mycobacterium tuberculosis, collected between March 2021 and December 2024, from 30 health facilities in the K.S.D Local Municipality of O.R. Tambo District. Resistance-associated mutation proxies were identified for loci associated with isoniazid (katG, inhA), fluoroquinolone (gyrA), and second-line injectable agents (amikacin, kanamycin, and capreomycin) through mutations in the rrs locus. Mutation proxy prevalence was examined overall, by age group, over time, and across facilities. Persistence of resistance detection was assessed using consecutive-month analyses to characterize temporal continuity at the facility level. Results: At least one resistance-associated mutation proxy was detected in 72.7% of tests. Isoniazid-associated mutation proxies predominated, with katG detected in 52.2% and inhA in 20.2% of records, while fluoroquinolone- and injectable-associated proxies were less frequent. Resistance-associated mutation proxies were observed across all adult age groups, with the highest burden and greatest resistance diversity among individuals aged 25–44 years. Substantial temporal variation was evident, including declining annual prevalence for most mutation proxies between 2022 and 2024, alongside increasing inhA prevalence. Marked facility-level heterogeneity was observed, with high-volume referral sites contributing the largest absolute burden of resistant cases. Prolonged persistence of mutation detection, including uninterrupted runs of up to 15 months, was identified at selected facilities. Conclusions: Routine molecular diagnostic data revealed a substantial and heterogeneous burden of drug-resistant Mycobacterium tuberculosis in K.S.D. Local Municipality, characterized by age-specific patterns, temporal shifts, and sustained facility-level persistence. Beyond descriptive epidemiology, routinely generated mutation proxy data can serve as early-warning indicators of clinical governance stress, signaling emerging pressures on TB care systems when resistance patterns persist or worsen. Interpreting these trends can support more anticipatory clinical governance, strengthen resistance surveillance, and guide prioritized interventions in high-burden, resource-constrained settings.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Data Source
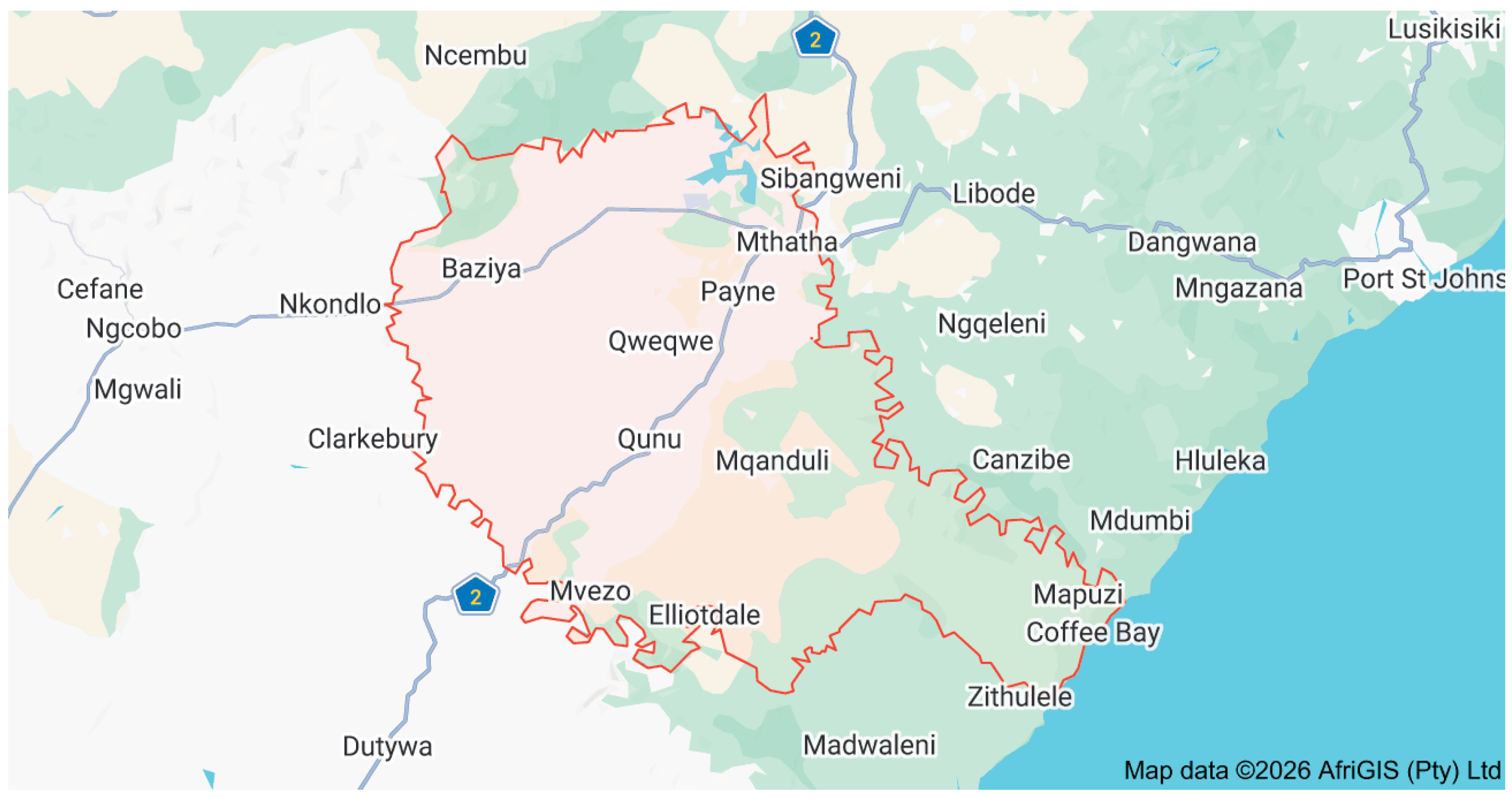
2.2. Study Setting and Population
2.3. Definition of Resistance-Associated Mutation Proxies
2.4. Outcomes and Stratification
2.5. Temporal and Facility-Level Analyses
2.6. Statistical Analysis
2.7. Ethical Considerations
3. Results
3.1. Diagnostic Volume and Dataset Characteristics
3.2. Overall Prevalence of Drug-Resistance Mutation Proxies
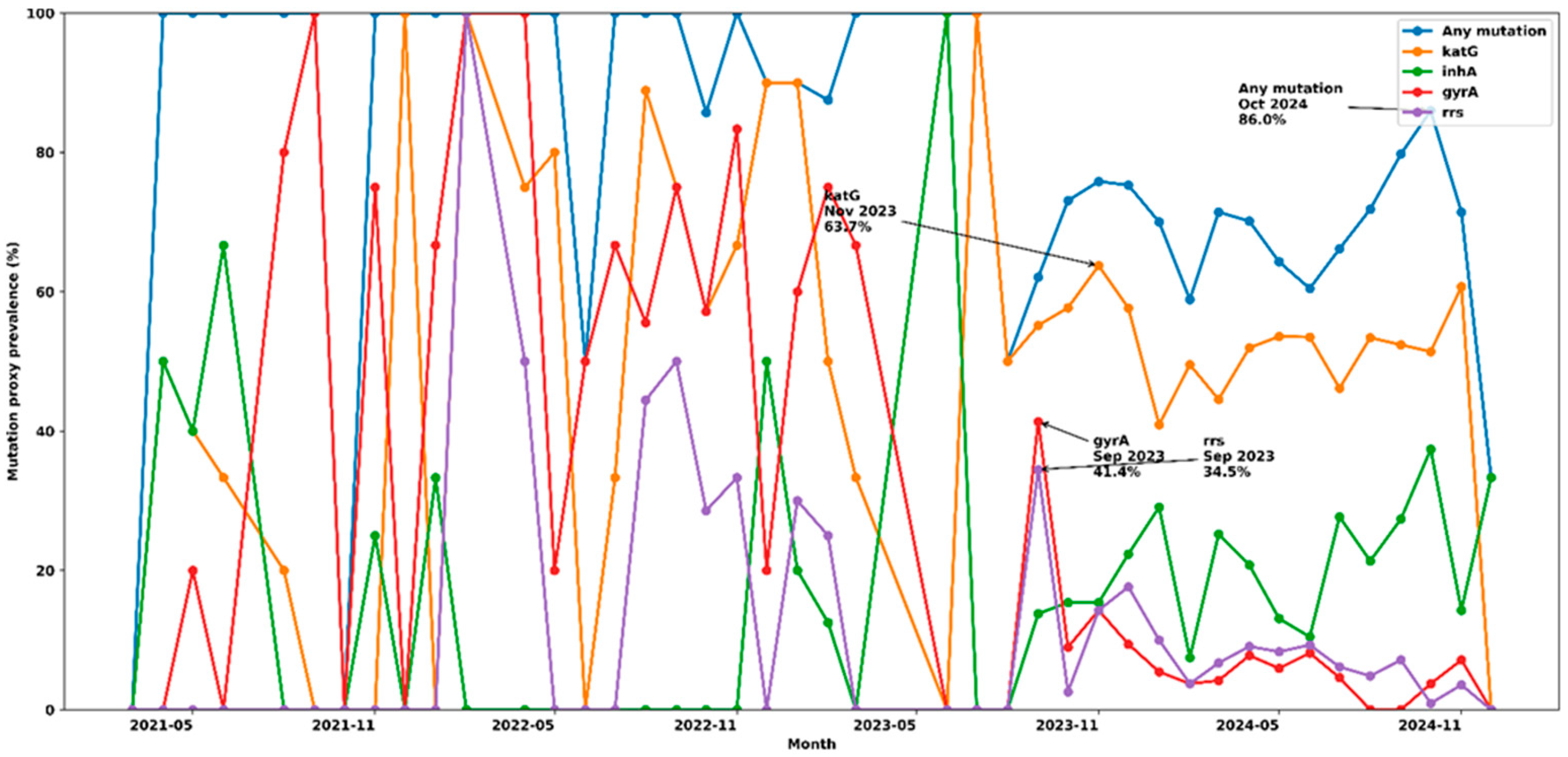
3.3. Temporal Patterns in Drug-Resistance Mutation Proxies
3.3.1. Monthly Variation
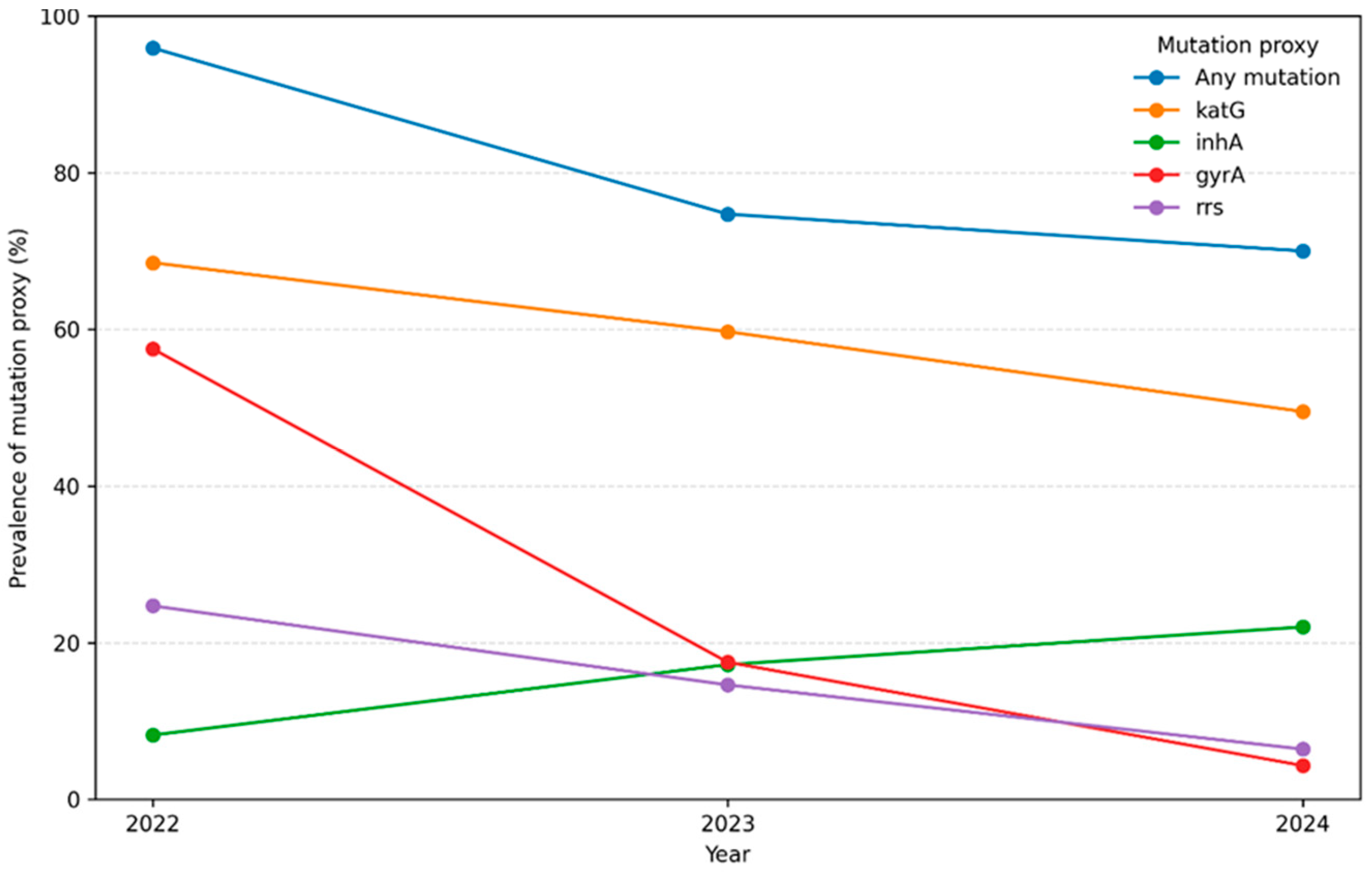
3.3.2. Annual Variation
3.4. Age-Stratified Distribution of Drug-Resistance Mutation Proxies
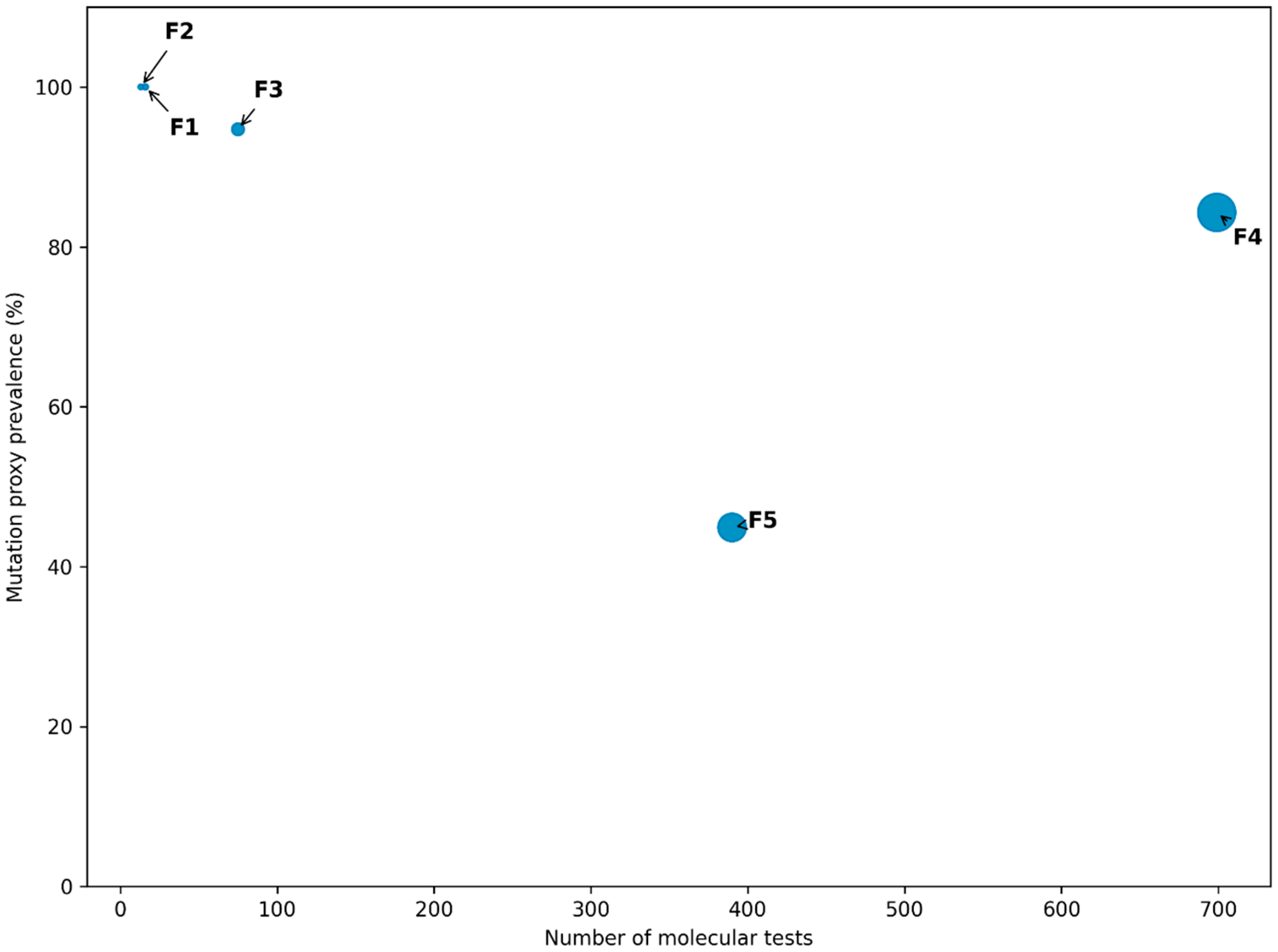
3.5. Facility-Level Distribution of Drug-Resistance Mutation Proxies
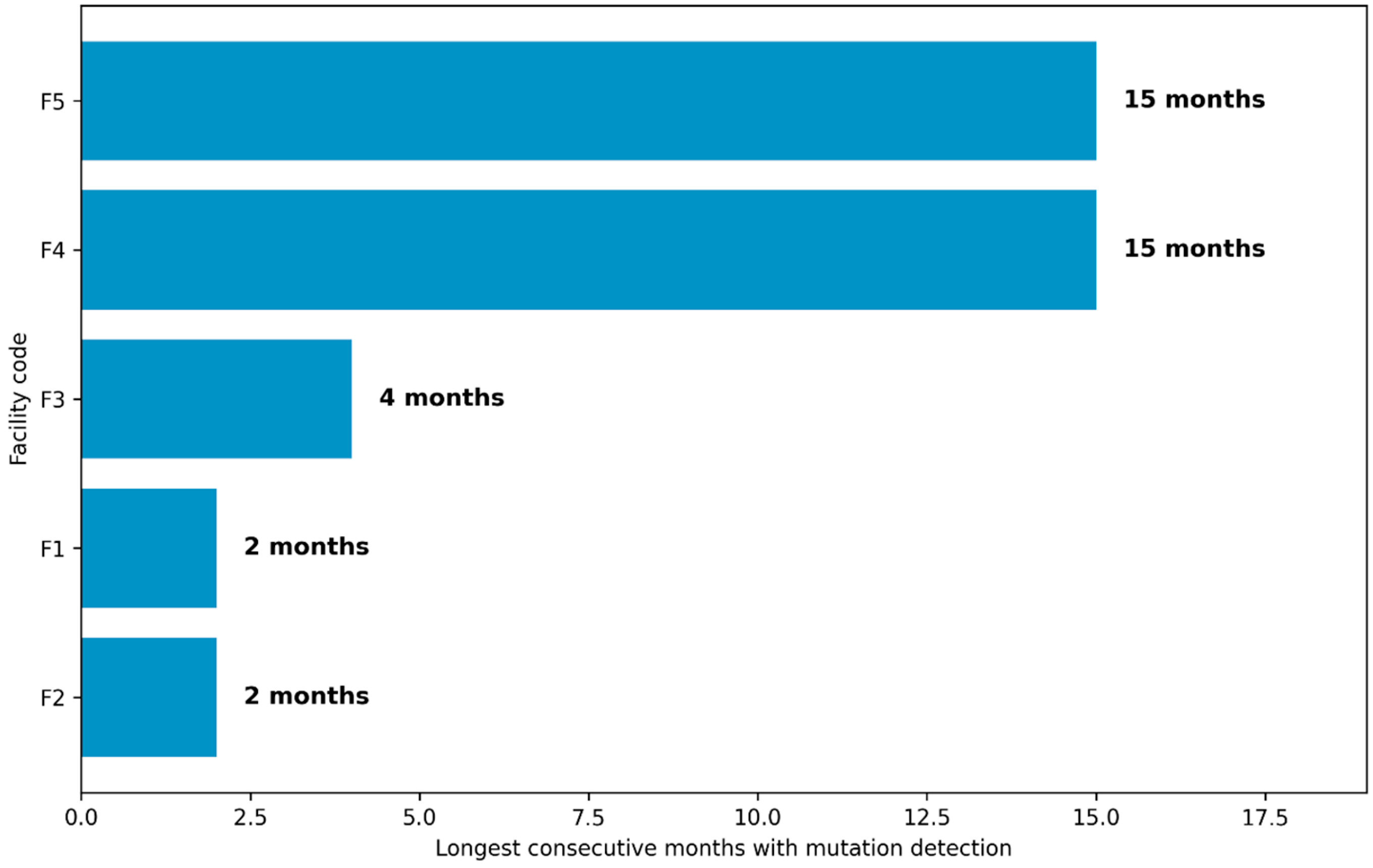
3.6. Persistence of Drug-Resistance Detection Over Time
4. Discussion
4.1. Diagnostic Volume and Dataset Maturity
4.2. Burden of Drug-Resistance Mutation Proxies
4.3. Temporal Patterns in Drug-Resistance Prevalence
4.4. Age-Stratified Distribution of Drug-Resistance Mutation Proxies
4.5. Facility-Level Heterogeneity in Drug-Resistance Burden
4.6. Persistence of Resistance Detection over Time
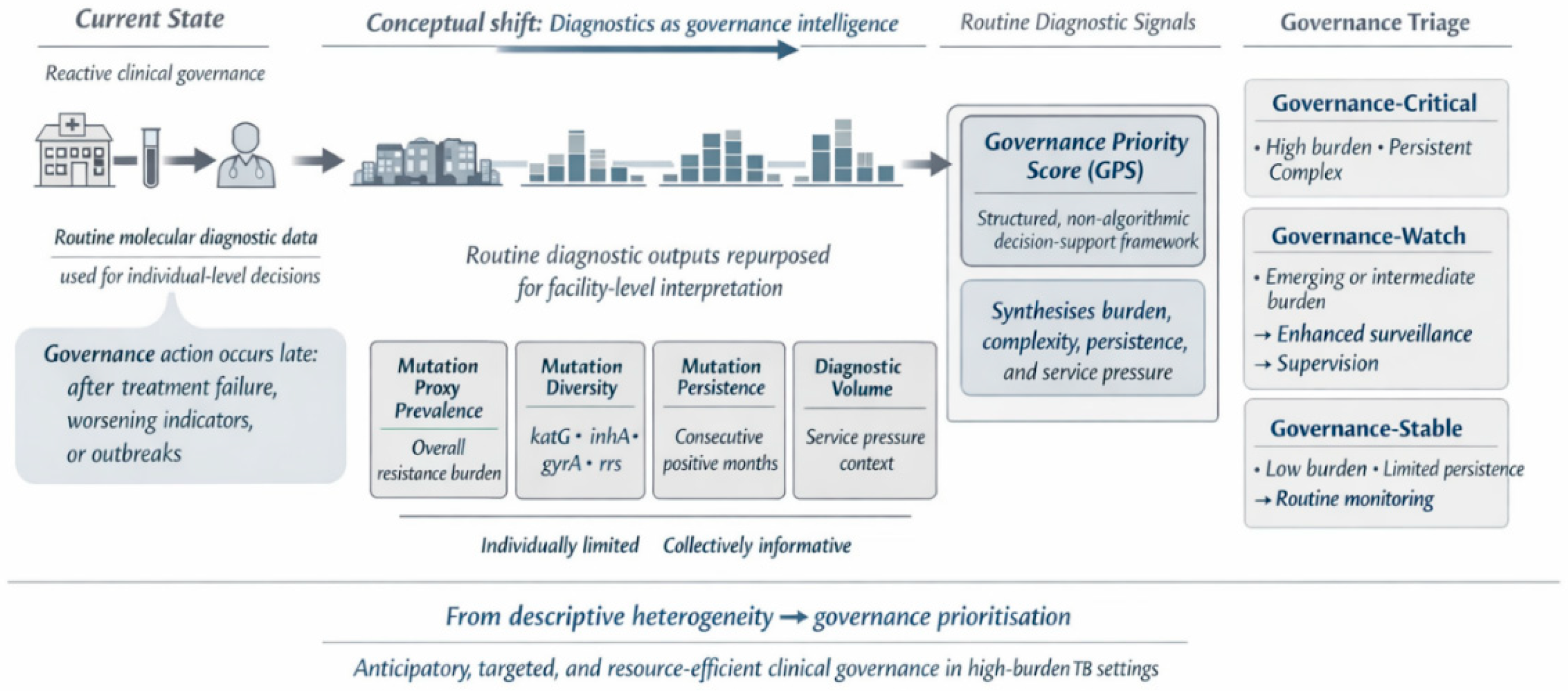
4.7. Mutation Persistence as a Proxy Indicator of Clinical Governance Stress
4.8. Facility-Level Governance Triage Using a Governance Priority Score
4.9. Facility-Level Governance Triage Model Based on Routine Molecular Diagnostic Data
4.10. Temporal Lag Between Resistance Detection and Clinical Governance Response
4.11. Strengths and Limitations
Strengths
Limitations
4.12. Recommendations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| F | Facility |
| GPS | Governance Priority Score |
| KSD | King Sabatha Dalindyebo |
| KSDLM | King Sabatha Dalindyebo Local Municipality |
| MDR | Multidrug resistant |
| O.R | Oliver Reginald |
| TB | Tuberculosis |
| XDR | Extensively drug-resistant |
References
- Farhat, M.; Cox, H.; Ghanem, M.; Denkinger, C.M.; Rodrigues, C.; Abd El Aziz, M.S.; Enkh-Amgalan, H.; Vambe, D.; Ugarte-Gil, C.; Furin, J.; Pai, M. Drug-resistant tuberculosis: a persistent global health concern. Nat Rev Microbiol. 2024, 22(10), 617–635. [Google Scholar] [CrossRef]
- Mancuso, G.; Midiri, A.; De Gaetano, S.; Ponzo, E.; Biondo, C. Tackling drug-resistant tuberculosis: new challenges from the old pathogen Mycobacterium tuberculosis. Microorganisms 2023, 11(9), 2277. [Google Scholar] [CrossRef]
- Dookie, N.; Ngema, S.L.; Perumal, R.; Naicker, N.; Padayatchi, N.; Naidoo, K. The changing paradigm of drug-resistant tuberculosis treatment: successes, pitfalls, and future perspectives. Clin Microbiol Rev. 2022, 35(4), e00180-19. [Google Scholar] [CrossRef]
- Liebenberg, D.; Gordhan, B.G.; Kana, B.D. Drug-resistant tuberculosis: implications for transmission, diagnosis, and disease management. Front Cell Infect Microbiol. 2022, 12, 943545. [Google Scholar] [CrossRef]
- Xiong, X.S.; Zhang, X.D.; Yan, J.W.; Huang, T.T.; Liu, Z.Z.; Li, Z.K.; Wang, L.; Li, F. Identification of Mycobacterium tuberculosis resistance to common antibiotics: an overview of current methods and techniques. Infect Drug Resist. 2024, 17, 1491–1506. [Google Scholar] [CrossRef]
- Bartolomeu-Gonçalves, G.; Souza, J.M.; Fernandes, B.T.; Spoladori, L.F.; Correia, G.F.; Castro, I.M.; Borges, P.H.; Silva-Rodrigues, G.; Tavares, E.R.; Yamauchi, L.M.; Pelisson, M. Tuberculosis diagnosis: current, ongoing, and future approaches. Diseases 2024, 12(9), 202. [Google Scholar] [CrossRef]
- Rosendal, E.; Isidro, J.; Carneiro, S.; Gomes, J.P.; Macedo, R. Rapid drug resistance prediction in positive Mycobacterium tuberculosis clinical samples using an extensive targeted next-generation sequencing panel. Emerging Microbes Infect 2026, 15(1), 2627072. [Google Scholar] [CrossRef]
- King Sabatha Dalindyebo Municipality Draft Annual Report 2024/2025. Available online: https://ksd.gov.za/download/documents/reports/annual_reports/KSDM-Draft-Annual-Report-2024-2025.pdf (accessed on 9 March 2026).
- World Health Organization. Guidance for the surveillance of drug resistance in tuberculosis; World Health Organization: Geneva, 2020. [Google Scholar]
- Mehl, G.; Tunçalp, Ö.; Ratanaprayul, N.; Tamrat, T.; Barreix, M.; Lowrance, D.; Bartolomeos, K.; Say, L.; Kostanjsek, N.; Jakob, R.; Grove, J. WHO SMART guidelines: optimising country-level use of guideline recommendations in the digital age. Lancet Digit Health 2021, 3(4), e213–e216. [Google Scholar] [CrossRef]
- Zimmer, A.J.; Klinton, J.S.; Oga-Omenka, C.; Heitkamp, P.; Nyirenda, C.N.; Furin, J.; Pai, M. Tuberculosis in times of COVID-19. J Epidemiol Commun Health 2022, 76(3), 310–316. [Google Scholar] [CrossRef]
- Valafar, S.J. Systematic review of mutations associated with isoniazid resistance points to continuing evolution and subsequent evasion of molecular detection and potential for emergence of multidrug resistance in clinical strains of Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2021, 65(3), e01280-20. [Google Scholar] [CrossRef]
- Goig, G.A.; Menardo, F.; Salaam-Dreyer, Z.; Dippenaar, A.; Streicher, E.M.; Daniels, J.; Reuter, A.; Borrell, S.; Reinhard, M.; Doetsch, A.; Beisel, C. Effect of compensatory evolution in the emergence and transmission of rifampicin-resistant Mycobacterium tuberculosis in Cape Town, South Africa: a genomic epidemiology study. Lancet Microbe 2023, 4(7), e506–e515. [Google Scholar] [CrossRef]
- Nono, V.N.; Nantia, E.A.; Mutshembele, A.; Teagho, S.N.; Simo, Y.W.; Takong, B.S.; Djieugoue, Y.J.; Assolo, Y.P.; Ongboulal, S.M.; Awungafac, S.N.; Eyangoh, S. Prevalence of katG and inhA mutations associated with isoniazid resistance in Mycobacterium tuberculosis clinical isolates in Cameroon. BMC Microbiol. 2025, 25(1), 127. [Google Scholar] [CrossRef]
- Klopper, M.; van der Merwe, C.J.; van der Heijden, Y.F.; Folkerts, M.; Loubser, J.; Streicher, E.M.; Mekler, K.; Hayes, C.; Engelthaler, D.M.; Metcalfe, J.Z.; Warren, R.M. The hidden epidemic of isoniazid-resistant tuberculosis in South Africa. Annals Amer Thoracic Soc 2024, 21(10), 1391–1397. [Google Scholar] [CrossRef]
- Dagne, B.; Desta, K.; Fekade, R.; Amare, M.; Tadesse, M.; Diriba, G.; Zerihun, B.; Getu, M.; Sinshaw, W.; Seid, G.; Gamtesa, D.F. The epidemiology of first- and second-line drug resistance in common Mycobacterium tuberculosis complex species: evidence from selected tuberculosis treatment-initiating centers in Ethiopia. PLoS One 2021, 16(1), e0245687. [Google Scholar] [CrossRef]
- Gupta, V.K.; Tiwari, S.K.; Sharma, S.; Nagar, L. Mathematical model of tuberculosis with drug resistance to the first and second line of treatment. J New Theory 2018, (21), 94–106. [Google Scholar]
- Mohiuddin, S.G.; Kavousi, P.; Figueroa, D.; Ghosh, S.; Orman, M.A. The diverse phenotypic and mutational landscape induced by fluoroquinolone treatment. mSystems 2025, 10(8), e00713-25. [Google Scholar] [CrossRef]
- Dheda, K.; Mirzayev, F.; Cirillo, D.M.; Udwadia, Z.; Dooley, K.E.; Chang, K.C.; Omar, S.V.; Reuter, A.; Perumal, T.; Horsburgh, C.R.; Murray, M. Multidrug-resistant tuberculosis. Nat Rev Dis Primers 2024, 10(1), 22. [Google Scholar] [CrossRef]
- Naidoo, K.; Perumal, R.; Cox, H.; Mathema, B.; Loveday, M.; Ismail, N.; Omar, S.V.; Georghiou, S.B.; Daftary, A.; O'Donnell, M.; Ndjeka, N. The epidemiology, transmission, diagnosis, and management of drug-resistant tuberculosis—lessons from the South African experience. Lancet Infect Dis. 2024, 24(9), e559–e575. [Google Scholar] [CrossRef]
- Napier, G.; Campino, S.; Phelan, J.E.; Clark, T.G. Large-scale genomic analysis of Mycobacterium tuberculosis reveals extent of target and compensatory mutations linked to multidrug-resistant tuberculosis. Sci Rep. 2023, 13(1), 623. [Google Scholar] [CrossRef]
- Shivekar, S.S.; Kaliaperumal, V.; Brammacharry, U.; Sakkaravarthy, A.; Raj, C.V.; Alagappan, C.; Muthaiah, M. Prevalence and factors associated with multidrug-resistant tuberculosis in South India. Sci Rep. 2020, 10(1), 17552. [Google Scholar] [CrossRef]
- Diriba, G; Alemu, A; Yenew, B; Ayano, BZ; Hailu, M; Buta, B; Wondimu, A; Tefera, Z; Meaza, A; Seid, G; Getahun, M; Dagne, B; Mollalign, H; Abebaw, Y; Getu, M; Tadesse, M; Belhu, T; Alemu, E; Demissie, M; Erresso, A; Tadesse, G. Second-line drug resistance among multidrug-resistant tuberculosis patients in Ethiopia: a laboratory-based surveillance. J Glob Antimicrob Resist. 2025, 42, 167–174. [Google Scholar] [CrossRef]
- Charan, A.S.; Gupta, N.; Dixit, R. Pattern of InhA and KatG mutations in isoniazid monoresistant Mycobacterium tuberculosis isolates. Lung India 2020, 37(3), 227–231. [Google Scholar] [CrossRef]
- Naik, J.H.; Bhuvaneshwari, G.; Krishnamurthy, V.; Kumar, N. Age-independent molecular resistance patterns in tuberculosis patients: a cross-sectional analytical study. J Pioneering Med Sci. 2026, 15(2), 96–104. [Google Scholar] [CrossRef]
- Adisa, R.; Ayandokun, T.T.; Ige, O.M. Knowledge about tuberculosis, treatment adherence and outcome among ambulatory patients with drug-sensitive tuberculosis in two directly-observed treatment centres in Southwest Nigeria. BMC Public Health 2021, 21(1), 677. [Google Scholar] [CrossRef]
- Mphande-Nyasulu, F.A.; Puengpipattrakul, P.; Praipruksaphan, M.; Keeree, A.; Ruanngean, K. Prevalence of tuberculosis, including multidrug-resistant and extensively drug-resistant tuberculosis, and association with occupation in adults at Sirindhorn Hospital, Bangkok. IJID Reg. 2022, 2, 141–148. [Google Scholar] [CrossRef]
- Faye, L.M.; Hosu, M.C.; Apalata, T. Drug-resistant tuberculosis hotspots in Oliver Reginald Tambo District Municipality, Eastern Cape, South Africa. Infect Dis Rep 2024, 16(6), 1197–1213. [Google Scholar] [CrossRef]
- Daneshi, S.; Mehni, E.B.; Kamali, M.; Barfar, E.; Barahouei, F.B.; Hushmandi, K.; Raesi, R. Prevalence and contributing factors of drug-resistant tuberculosis in Iran: a systematic review. BMC Infect Dis. 2025, 25(1), 1004. [Google Scholar] [CrossRef]
- Wei, W.; Xia, L.; Wu, J.; Zhou, Z.; Zhang, W.; Luan, R. The environmental and socioeconomic effects and prediction of patients with tuberculosis in different age groups in Southwest China: a population-based study. JMIR Public Health Surveill. 2023, 9, e40659. [Google Scholar] [CrossRef]
- Sy, K.T.; Leavitt, S.V.; de Vos, M.; Dolby, T.; Bor, J.; Horsburgh, C.R.; Warren, R.M.; Streicher, E.M.; Jenkins, H.E.; Jacobson, K.R. Spatial heterogeneity of extensively drug-resistant tuberculosis in Western Cape Province, South Africa. Sci Rep. 2022, 12(1), 10844. [Google Scholar] [CrossRef]
- Faye, L.M.; Hosu, M.C.; Vasaikar, S.; Dippenaar, A.; Oostvogels, S.; Warren, R.M.; Apalata, T. Spatial distribution of drug-resistant Mycobacterium tuberculosis infections in rural Eastern Cape Province of South Africa. Pathogens 2023, 12(3), 475. [Google Scholar] [CrossRef]
- Omoteso, O.A.; Fadaka, A.O.; Walker, R.B.; Khamanga, S.M. Innovative strategies for combating multidrug-resistant tuberculosis: advances in drug delivery systems and treatment. Microorganisms 2025, 13(4), 722. [Google Scholar] [CrossRef]
- Shah, H.D.; Nazli Khatib, M.; Syed, Z.Q.; Gaidhane, A.M.; Yasobant, S.; Narkhede, K.; Bhavsar, P.; Patel, J.; Sinha, A.; Puwar, T. Gaps and interventions across the diagnostic care cascade of TB patients at the level of patient, community, and health system: a qualitative review of the literature. Trop Med Infect Dis. 2022, 7(7), 136. [Google Scholar] [CrossRef]
- Saktiawati, AMI; Vasiliu, A; Saluzzo, F; Akkerman, OW. Strategies to enhance diagnostic capabilities for the new drug-resistant tuberculosis drugs. Pathogens 2024, 13(12), 1045. [Google Scholar] [CrossRef]
- Jassat, W.; Moshabela, M.; Nicol, M.P.; Dickson, L.; Cox, H.; Mlisana, K.; Black, J.; Loveday, M.; Grant, A.D.; Kielmann, K.; Schneider, H. Decentralising drug-resistant tuberculosis care: the trade-off between quality of care and service coverage in the early phase of implementation. Public Health Action 2025, 15(3), 97–102. [Google Scholar] [CrossRef]
- King Sabata Dalindyebo Municipality. Draft Annual Report 2024/2025; O.R. Tambo District, Eastern Cape, South Africa, 2025. Available online: https://ksd.gov.za/download/documents/reports/annual_reports/KSDM-Draft-Annual-Report-2024-2025.pdf (accessed on 19 March 2026).
| Resistance-associated locus | Mutation proxy detected, n (%) | Total tests |
| Any mutation proxy | 1,007 (72.7) | 1,386 |
| katG | 724 (52.2) | 1,386 |
| inhA | 280 (20.2) | 1,386 |
| gyrA (any region) | 149 (10.8) | 1,386 |
| rrs | 125 (9.0) | 1,386 |
| eis | 0 (0.0) | 1,386 |
| Age group (years) | Tests, n | Any mutation n (%) | katG n (%) | inhA n (%) | gyrA n (%) | rrs n (%) |
| <15 | 4 | 0 (0.0) | 0 | 0 | 0 | 0 |
| 15–24 | 157 | 153 (97.5) | 129 (82.2) | 24 (15.3) | 1 (0.6) | 0 (0.0) |
| 25–34 | 322 | 231 (71.7) | 128 (39.8) | 60 (18.6) | 89 (27.6) | 34 (10.6) |
| 35–44 | 389 | 265 (68.1) | 155 (39.8) | 134 (34.4) | 17 (4.4) | 46 (11.8) |
| 45–54 | 244 | 149 (61.1) | 103 (42.2) | 46 (18.9) | 0 (0.0) | 3 (1.2) |
| ≥55 | 270 | 209 (77.4) | 209 (77.4) | 16 (5.9) | 42 (15.6) | 42 (15.6) |
| Facility code | Diagnostic volume (n) | Any mutation proxy n (%) | Key resistance features | Persistence/volume context | Governance triage category | Rationale for classification |
| F1 | 16 | 16 (100%) | Predominantly first-line resistance; no gyrA or rrs detected | Very low volume; no documented persistence | Governance-Watch | Extremely high proportional prevalence based on small numbers; requires monitoring rather than immediate intensive intervention. |
| F2 | 13 | 13 (100%) | High second-line resistance burden (gyrA 69.2%, rrs 46.2%) | Low volume but high resistance complexity | Governance-Critical | Extensive fluoroquinolone and injectable resistance signals indicate elevated governance risk. |
| F3 | 75 | 71 (94.7%) | Resistance is largely limited to first-line drugs | Moderate volume; referral-level case mix | Governance-Watch | High prevalence expected in a tertiary referral setting; patterns reflect case complexity |
| F4 | 699 | 589 (84.3%) | Significant second-line resistance (gyrA 17.5%, rrs 11.7%) | Very high volume; prolonged persistence (up to 15 months) | Governance-Critical | Convergence of high absolute burden, resistance diversity, and persistence indicates sustained governance stress. |
| F5 | 13 | 11 (84.6%) | Prominent fluoroquinolone resistance (gyrA 46.2%) | Low–moderate volume; no RRs detected | Governance-Watch | High proportional resistance to the second-line signal warrants enhanced surveillance. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




