Submitted:
19 March 2026
Posted:
19 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Study Population
Sample Collection and Processing
ELISA Measurements
Additional Analyses
Statistical Analyses
3. Results
- Patient Characteristics
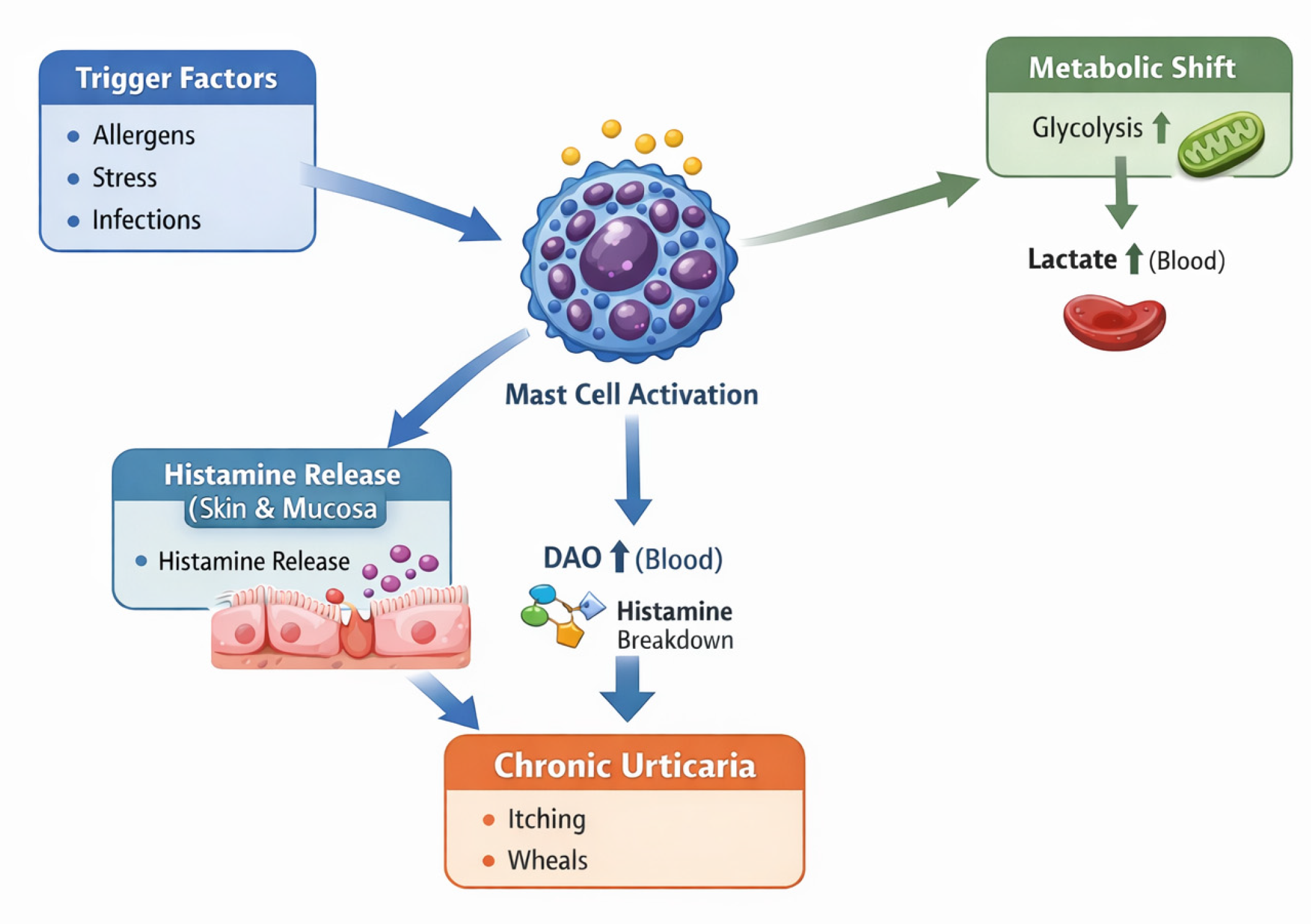
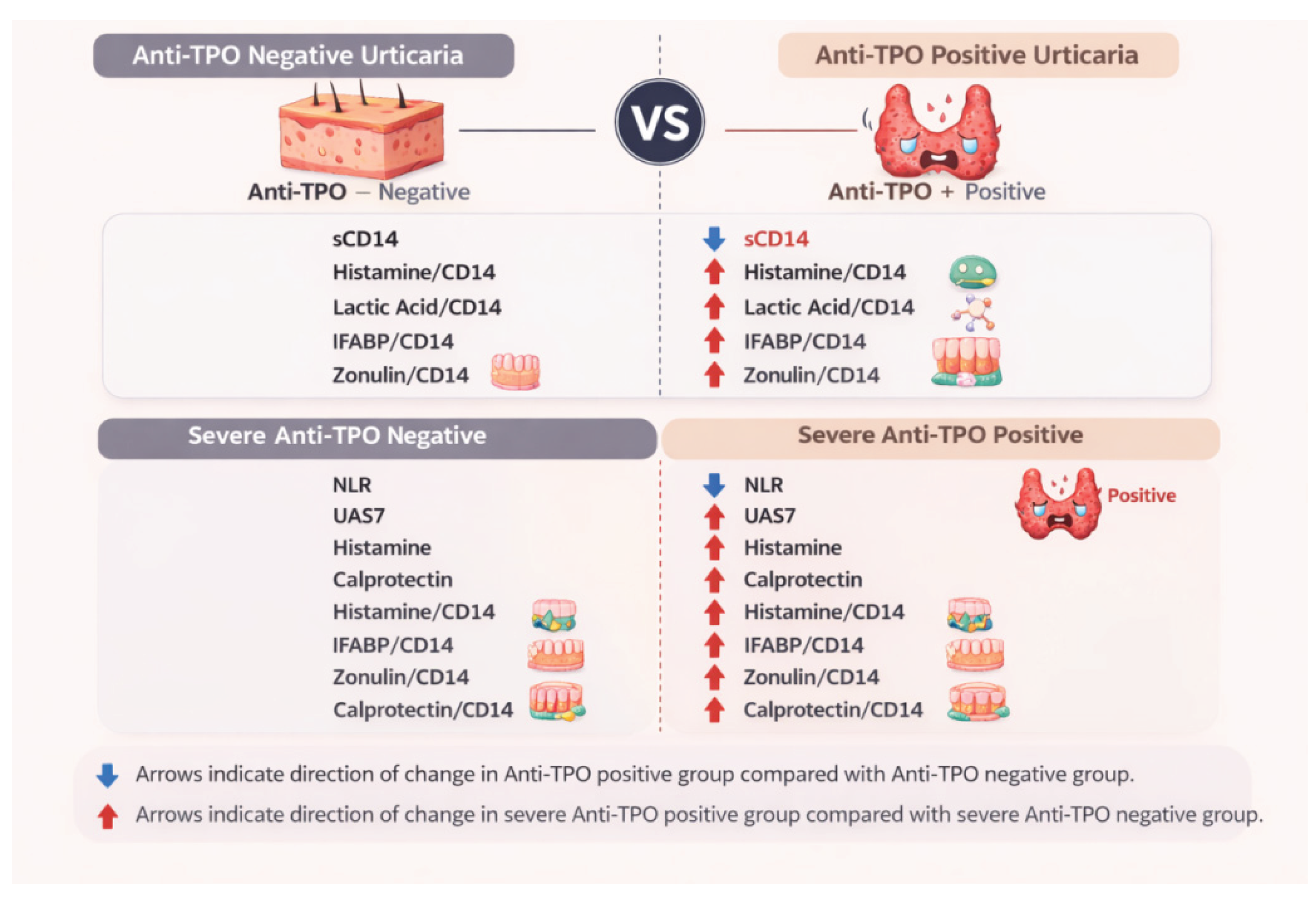
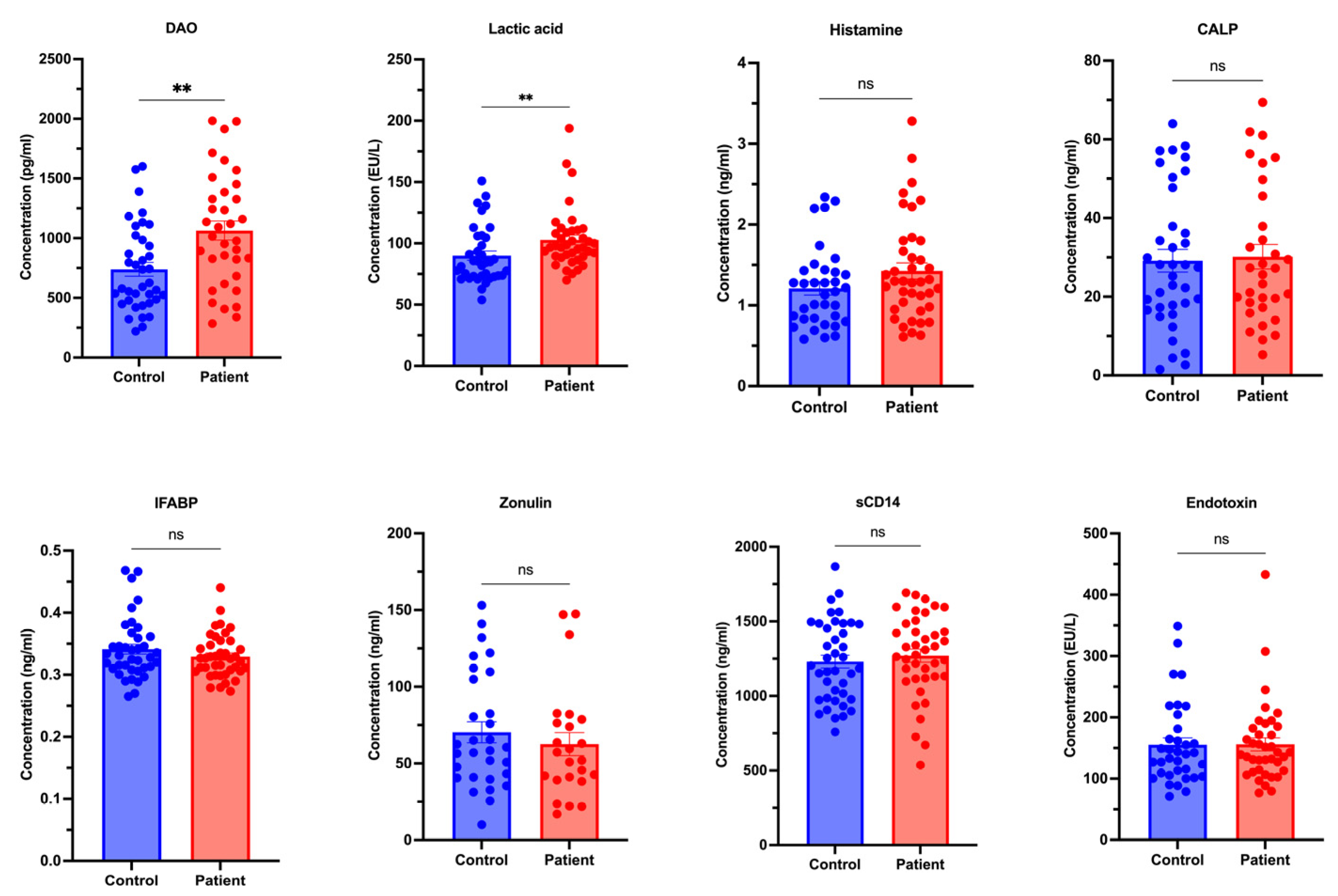
- Biomarker Profiles
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| DAO | Diamine oxidase |
| IFABP | Intestinal fatty acid-binding protein |
| sCD14 | Soluble CD14 |
| UAS7 | Urticaria Activity Score 7 |
| UCT | Urticaria Control Test |
| CRP | C-reactive protein |
| NLR | Neutrophil-to-lymphocyte ratio |
| ELR | Eosinophil-to-lymphocyte ratio |
| GI | Gastrointestinal |
| ELISA | Enzyme-linked immunosorbent assay |
| IQR | Interquartile range |
| SD | Standard deviation |
References
- Beck, LA; Bernstein, JA; Maurer, M. A Review of International Recommendations for the Diagnosis and Management of Chronic Urticaria. Acta Derm Venereol 2017, 97(2), 149–158. [Google Scholar] [CrossRef] [PubMed]
- Zuberbier, T; Bernstein, JA; Maurer, M. Chronic spontaneous urticaria guidelines: What is new? J Allergy Clin Immunol. Erratum in: J Allergy Clin Immunol. 2023 Feb;151(2):580. PMID: 36481045. 2022, 150(6), 1249–1255. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L; Jian, X; Zhou, B; Liu, R; Muñoz, M; Sun, W; Xie, L; Chen, X; Peng, C; Maurer, M; Li, J. Gut microbiota facilitate chronic spontaneous urticaria. Nat Commun. 2024, 15(1), 112. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zuberbier, T; Abdul Latiff, AH; Abuzakouk, M; Aquilina, S; Asero, R; Baker, D; Ballmer-Weber, B; et al. The international EAACI/GA²LEN/EuroGuiDerm/APAAACI guideline for the definition, classification, diagnosis, and management of urticaria. Allergy 2022, 77(3), 734–766. [Google Scholar] [CrossRef] [PubMed]
- Di Tommaso, N; Gasbarrini, A; Ponziani, FR. Intestinal Barrier in Human Health and Disease. Int J Environ Res Public Health 2021, 18(23), 12836. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Malgesini, A; Marsiglia, MD; Borghi, E; Marzano, AV; Nazzaro, G. The Emerging Role of Gut Microbiota in Inflammatory Skin Diseases: A Systematic Review. Exp Dermatol 2026, 35(3), e70234. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Guo, S.; Gillingham, T.; Guo, Y.; Meng, D.; Zhu, W.; Walker, W.A.; Ganguli, K. Secretions of Bifidobacterium infantis and Lactobacillus acidophilus Protect Intestinal Epithelial Barrier Function. J. Pediatr. Gastroenterol. Nutr. 2017, 64, 404–412. [Google Scholar] [CrossRef]
- Fan, Y.; Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 2021, 19, 55–71. [Google Scholar] [CrossRef]
- Choi, W.; Yeruva, S.; Turner, J.R. Contributions of intestinal epithelial barriers to health and disease. Exp. Cell Res. 2017, 358, 71–77. [Google Scholar] [CrossRef]
- Paray, B.A.; Albeshr, M.F.; Jan, A.T.; Rather, I.A. Leaky Gut and Autoimmunity: An Intricate Balance in Individuals Health and the Diseased State. Int. J. Mol. Sci. 2020, 21, 9770. [Google Scholar] [CrossRef]
- Foster, J.A.; Neufeld, K.-A.M. Gut–brain axis: How the microbiome influences anxiety and depression. Trends Neurosci. 2013, 36, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Mayer, E.A.; Tillisch, K.; Gupta, A. Gut/brain axis and the microbiota. J. Clin. Investig. 2015, 125, 926–938. [Google Scholar] [CrossRef] [PubMed]
- Kesika, P.; Suganthy, N.; Sivamaruthi, B.S.; Chaiyasut, C. Role of gut-brain axis, gut microbial composition, and probiotic intervention in Alzheimer’s disease. Life Sci. 2021, 264, 118627. [Google Scholar] [CrossRef] [PubMed]
- Long, C; Zhou, X; Xia, F; Zhou, B. Intestinal Barrier Dysfunction and Gut Microbiota in Non-Alcoholic Fatty Liver Disease: Assessment, Mechanisms, and Therapeutic Considerations. Biology (Basel) 2024, 13(4), 243. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Selvakumar, B; Eladham, MW; Hafezi, S; Ramakrishnan, R; Hachim, IY; Bayram, OS; Sharif-Askari, NS; Sharif-Askari, FS; Ibrahim, SM; Halwani, R. Allergic Airway Inflammation Emerges from Gut Inflammation and Leakage in Mouse Model of Asthma. Adv Biol (Weinh) Epub. 2024, 8(1), e2300350. [Google Scholar] [CrossRef] [PubMed]
- Varela Trinidad, Gael; Domínguez Díaz, Carolina; Solórzano Castanedo, Karla; Iñiguez Gutiérrez, Liliana; Hernández, Teresita; Fafutis-Morris, Mary. Probiotics: Protecting Our Health from the Gut. Microorganisms 2022. [Google Scholar] [CrossRef]
- Chu, CY; Zuberbier, T. Urticaria and the gut. Curr Opin Allergy Clin Immunol 2020, 20(4), 381–385. [Google Scholar] [CrossRef] [PubMed]
- Zheleznov, S; Urzhumtseva, G; Petrova, N; et al. Gastritis can cause and trigger chronic spontaneous urticaria independent of the presence of Helicobacter pylori. Int Arch Allergy Immunol 2018, 175, 246–251. [Google Scholar] [CrossRef]
- Bakos, N; Fekete, B; Prohaszka, Z; et al. High prevalence of IgG and IgA antibodies to 19-kDa Helicobacter pylori-associated lipoprotein in chronic urticaria. Allergy 2003, 58, 663–667. [Google Scholar] [CrossRef]
- Liutu, M; Kalimo, K; Uksila, J; Savolainen, J. Extraction of IgEbinding components.
- of Helicobacter pylori by immunoblotting analysis in chronic urticaria patients. Int Arch Allergy Immunol 2001, 126, 213–217. [CrossRef] [PubMed]
- Lu, T.; Chen, Y.; Guo, Y.; Sun, J.; Shen, W.; Yuan, M.; Zhang, S.; He, P.; Jiao, X.; Liu, R.; Peng, C.; Jing, D.; Xiao, Y.; Zhu, W.; Zhao, S.; Zhang, J.; Chen, X.; Li, J.; Altered gut microbiota diversity and composition in chronic urticaria. Biomarkers of Gut Microbiota in Chronic Spontaneous Urticaria and Symptomatic Dermographism. Dis. Markers;Front. Cell. Infect. Microbiol. 2019, 2019 11, 6417471. 22 703126. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Guo, S.; He, H.; Gong, L.; Cui, H. Gut microbiome and serum metabolome analyses identify unsaturated fatty acids and butanoate metabolism induced by gut microbiota in patients with chronic spontaneous urticaria. Front. Cell. Infect. Microbiol. 2020, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yi, W.; He, L.; Luo, S.; Wang, J.; Jiang, L.; Long, H.; Zhao, M.; Lu, Q. Abnormalities in Gut Microbiota and Metabolism in Patients with Chronic Spontaneous Urticaria. Front. Immunol. 2021, 12, 691304. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, J.; Chu, Z.; Shi, L.; Geng, S.; Guo, K. Gut Microbiome Alterations and Functional Prediction in Chronic Spontaneous Urticaria Patients. J. Microbiol. Biotechnol. 2021, 31, 747–755. [Google Scholar] [CrossRef]
- Krišto, M.; Lugović-Mihić, L.; Muñoz, M.; Rupnik, M.; Mahnic, A.; Ozretić, P.; Jaganjac, M.; Ćesić, D.; Kuna, M. Gut Microbiome Composition in Patients with Chronic Urticaria: A Review of Current Evidence and Data. Life 2023, 13, 152. [Google Scholar] [CrossRef]
- Chu, CY; Zuberbier, T. Urticaria and the gut. Curr Opin Allergy Clin Immunol 2020, 20(4), 381–385. [Google Scholar] [CrossRef] [PubMed]
- Pucino, V.; Certo, M.; Bulusu, V.; Cucchi, D.; Goldmann, K.; Pontarini, E.; Haas, R.; Smith, J.; Headland, S.; Blighe, K.; Ruscica, M.; Humby, F.; Lewis, M.; Kamphorst, J.; Bombardieri, M.; Pitzalis, C.; Mauro, C. Lactate Buildup at the Site of Chronic Inflammation Promotes Disease by Inducing CD4+ T Cell Metabolic Rewiring. Cell Metabolism 2019, 30, 1055–1074.e8. [Google Scholar] [CrossRef]
- Cucca, V.; Ramirez, G.; Pignatti, P.; Asperti, C.; Russo, M.; Della-Torre, E.; Breda, D.; Burastero, S.; Dagna, L.; Yacoub, M. Basal Serum Diamine Oxidase Levels as a Biomarker of Histamine Intolerance: A Retrospective Cohort Study. Nutrients 2022, 14. [Google Scholar] [CrossRef]
- Arih, K.; Đorđević, N.; Košnik, M.; Rijavec, M. Evaluation of Serum Diamine Oxidase as a Diagnostic Test for Histamine Intolerance. Nutrients 2023, 15. [Google Scholar] [CrossRef]
- Mikuls, T.; LeVan, T.; Sayles, H.; Yu, F.; Caplan, L.; Cannon, G.; Kerr, G.; Reimold, A.; Johnson, D.; Thiele, G. Soluble CD14 and CD14 Polymorphisms in Rheumatoid Arthritis. The Journal of Rheumatology 2011, 38, 2509–2516. [Google Scholar] [CrossRef]
| Parameter | Values |
|---|---|
| Age (mean ± SD) | 40,45 ± 15,4 |
| Sex (F/M) | 29/11 |
| Chronic urticaria subtype, n (%) | |
| Spontaneous | 32 (80) |
| Inducible | 5 (12,5) |
| Spontaneous + inducible | 3 (7,5) |
| Presence of angioedema, n (%) | 15 (35,7) |
| Antihistamine resistance, n (%) | 22 (55) |
| Anti-TPO positivity, n (%) | 12 (30,8) |
| Elevated IgE, n (%) | 23 (59) |
| Elevated CRP, n (%) | 11 (27,5) |
| UAS7 (mean ± SD) | 31,25 ± 12,25 |
| UCT score (mean ± SD) | 6,3 ± 4,24 |
| Parameter | Control | Chronic urticaria | p |
|---|---|---|---|
| Histamine (ng/mL), median (IQR) | 1,17 (0,6) | 1,30 (0,8) | 0,122 |
| DAO (pg/mL), median (IQR) | 795,7 (618,3) | 964,6 (673,7) | 0,003 |
| sCD14 (ng/mL)*, mean ± SD | 1230,74 ± 271,74 | 1269,97 ± 277,43 | 0,525 |
| Lactic acid (EU/L), median (IQR) | 85,5 (31,2) | 96,7 (14,6) | 0,004 |
| Endotoxin (EU/L), median (IQR) | 143,9 (52,5) | 147,5 (53,6) | 0,711 |
| IFABP (ng/mL), median (IQR) | 0,34 (0,06) | 0,32 (0,05) | 0,346 |
| Zonulin (ng/mL), median (IQR) | 56,5 (30,6) | 51,4 (38,9) | 0,497 |
| Calprotectin (ng/mL), median (IQR) | 21,2 (19,4) | 25,3 (40,5) | 0,861 |
| Histamine/DAO, median (IQR) | 0,0015 (0,0014) | 0,0011 (0,0011) | ,3 |
| Lactic acid/DAO, median (IQR) | 0,1 (0,08) | 0,10 (0,08) | ,052 |
| Other parameters, median (IQR) | ↔ | ↔ | NS |
| Parameter | Mild-moderate CU | Severe CU | P value |
|---|---|---|---|
| Histamine (ng/mL), mean ± SD | 1,16 ± 0,69 | 1,50± 0,60 | 0,156 |
| DAO (pg/mL), mean ± SD | 1314,03 ± 513,74 | 976,66 ± 434,39 | 0,064 |
| sCD14 (ng/mL), mean ± SD | 1065,73 ± 319,67 | 1329,26 ± 238,03 | 0,010 |
| Lactic acid (EU/L), mean ± SD | 101,35± 7,64 | 103,16 ± 27,35 | 0,856 |
| Endotoxin (EU/L), median (IQR) | 146,28 ± 47,52 | 158,50 ± 70,58 | 0,648 |
| IFABP (ng/mL), median (IQR) | 0,33 ± 0,023 | 0,33± 0,04 | 0,828 |
| Zonulin (ng/mL), median (IQR) | 69,28 ± 55,21 | 59,85 ± 27,29 | 0,679 |
| Calprotectin (ng/mL), median (IQR) | 28,40 ± 19,86 | 30,85± 17,05 | 0,30 |
| Histamine/DAO* | 0,0008 (0,0005) | 0,0016 (0,0021) | 0,006 |
| Lactic acid/DAO*, median (IQR) | 0,08 (0,05) | 0,11 (0,14) | 0,042 |
| Lactic acid/Histamine* | 118, 14 (67,7) | 72,97 (46,95) | ,012 |
| IFABP/sCD14* | 0,00029 (0,00024) | 0,00024 (0,00008) | ,034 |
| Other parameters | ↔ | ↔ | NS |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




