Submitted:
18 March 2026
Posted:
19 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
- A.
- Datasets
- B.
- Data Cleaning and Preprocessing
- C.
- Machine Learning Algorithms
- D.
- Random Forest
- E.
- XGBoost
- F.
- Support Vector Machine
- G.
- Logistic Regression
- H.
- k-Nearest Neighbors
- I.
- Model Evaluation Metrics
- Accuracy (Acc) = (TP + TN) / (TP + TN + FP + FN)
- Precision (P) = TP / (TP + FP)
- Recall (R) = TP / (TP + FN)
- F1-score (F1) = 2 × (P × R) / (P + R)
- Area Under the Receiver Operating Characteristic Curve (ROC–AUC)
- J.
- Visualization and Feature Interpretation
- K.
- Phylogenetic Analysis
- L.
- Identification of Genomic Biomarkers
3. Results
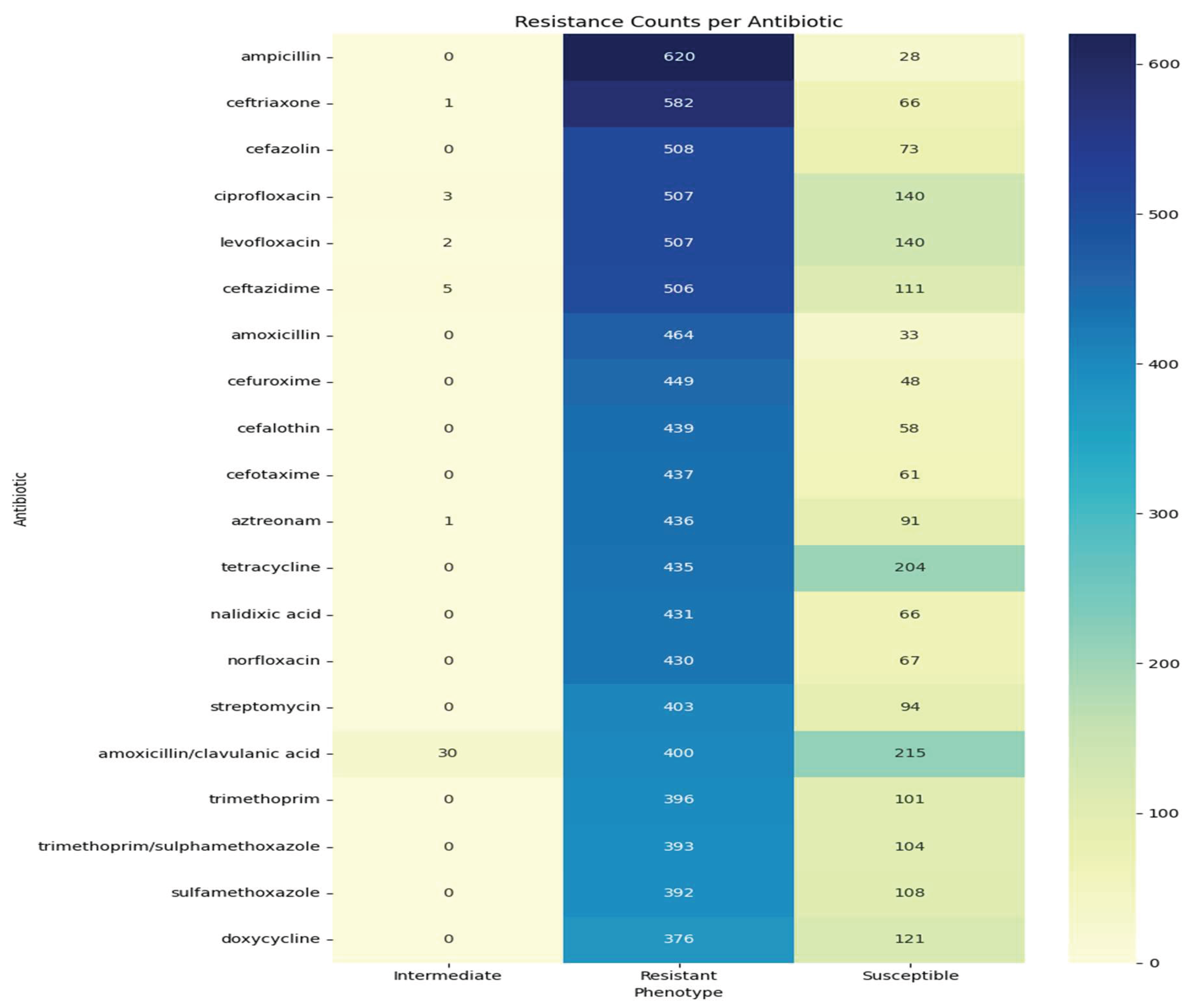
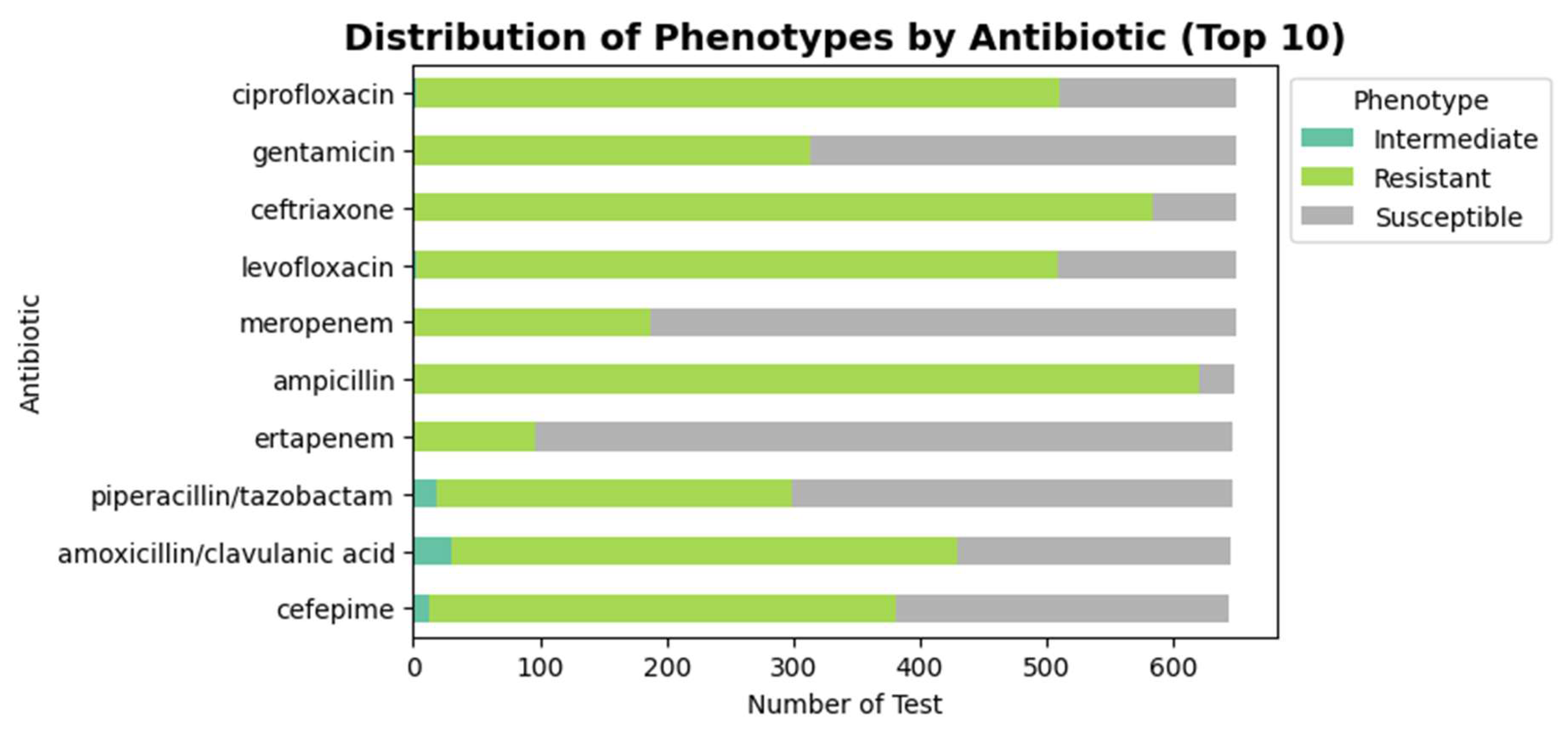
3.1. The Distribution of Antibiotics Testing and Resistance Patterns
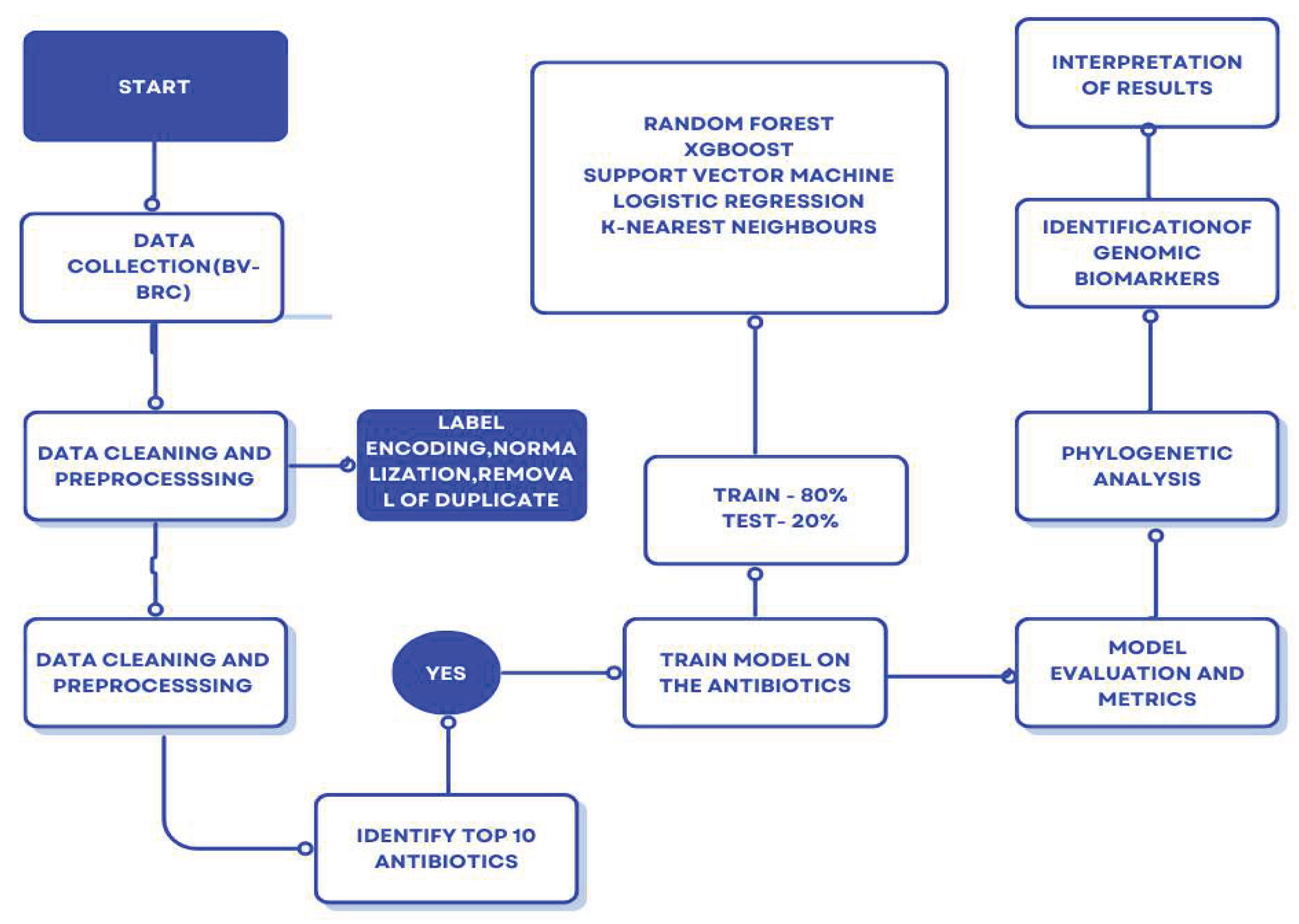
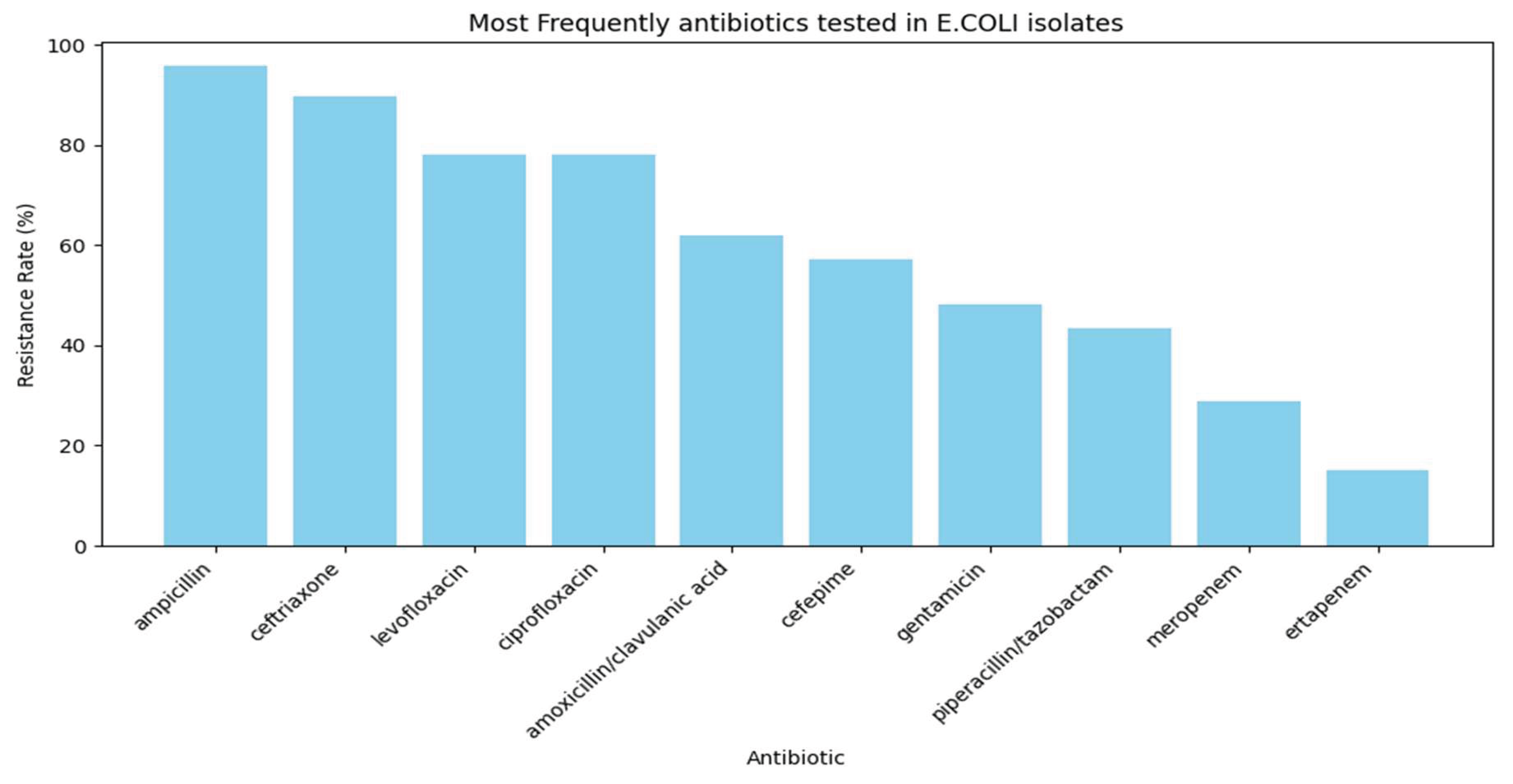
3.2. Machine Learning Model Performance
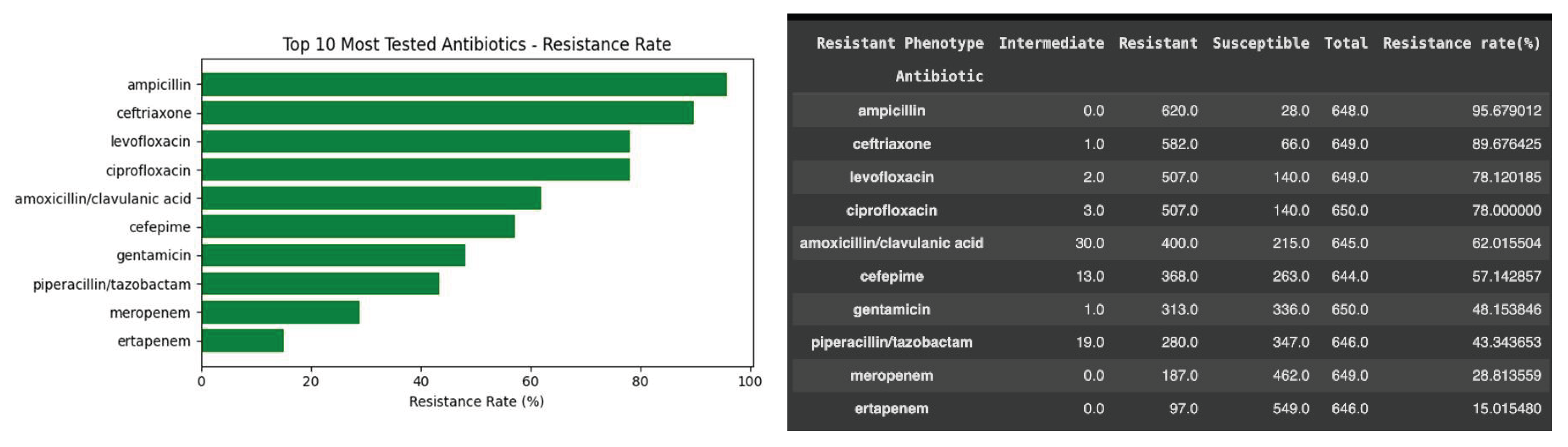
3.3. The Resistance Distribution Across Antibiotics
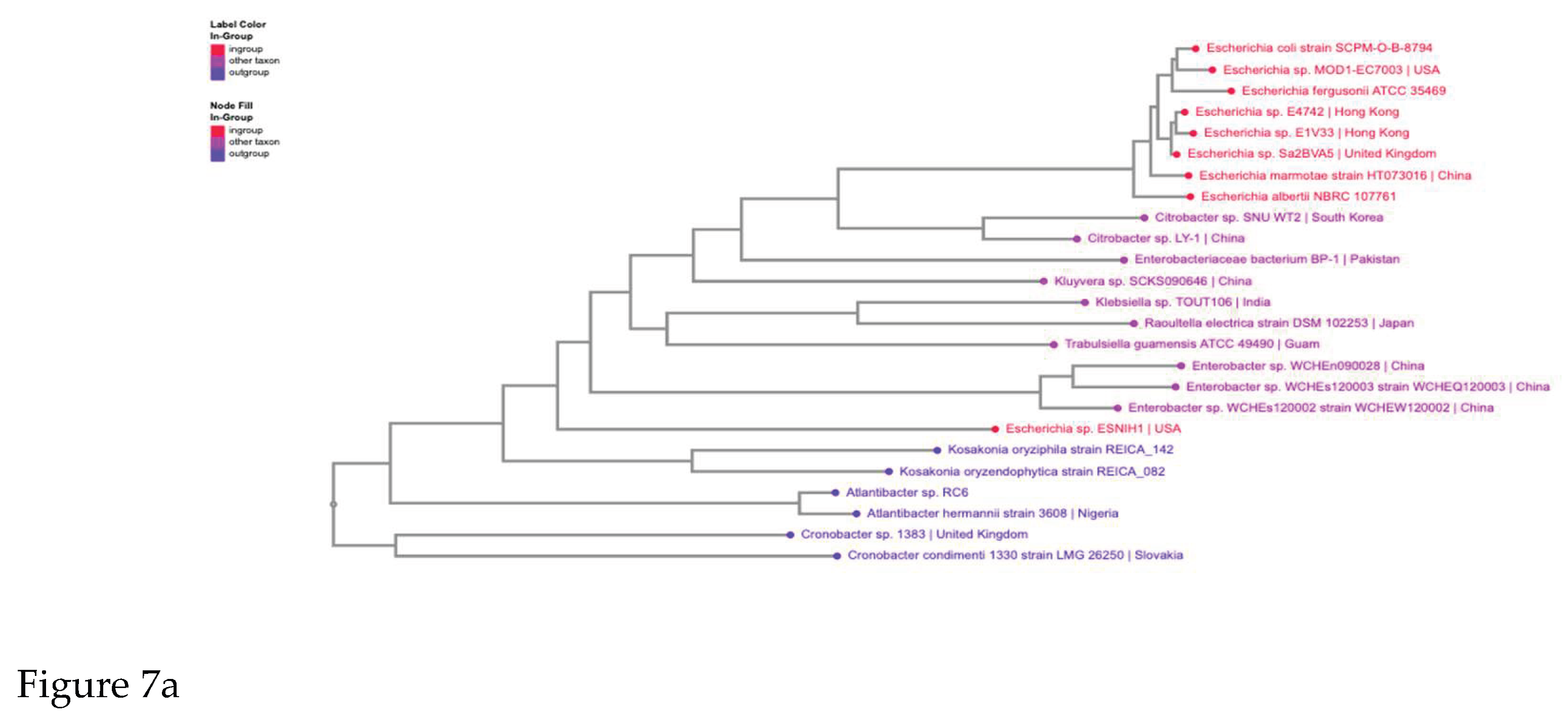
3.4. Phylogenetic Analysis

3.5. Genomic Biomarkers Identifications
4. Discussion
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMR | Antimicrobial Resistance |
| E. coli | Escherichia coli |
| BV-BRC | Bacterial and Viral Bioinformatics Resource Center |
| CARD | Comprehensive Antibiotic Resistance Database |
References
- Murray, C. J.; Ikuta, K. S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Naghavi, M. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The Lancet 2022, 399(10325), 629–655. [Google Scholar] [CrossRef]
- Shrestha, P; Cooper, BS; Coast, J; Oppong, R; Do Thi Thuy, N; Phodha, T; Celhay, O; Guerin, PJ; Wertheim, H; Lubell, Y. Enumerating the economic cost of antimicrobial resistance per antibiotic consumed to inform the evaluation of interventions affecting their use. Antimicrob Resist Infect Control 2018, 7, 98. [Google Scholar] [CrossRef] [PubMed]
- Kollef, M.H.; Shorr, A.F.; Bassetti, M.; Timsit, J.F.; Micek, S.T.; Michelson, A.P.; Garnacho-Montero, J. Timing of antibiotic therapy in the ICU. Crit. Care 2021, 25, 360. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Blanco; Ruiz-Blanco, YB; Agüero-Chapin, G; Romero-Molina, S; Antunes, A; Olari, LR; Spellerberg, B; Münch, J; Sanchez-Garcia, E.; et al. ABP-Finder: a tool to identify antibacterial peptides and the gram-staining type of targeted bacteria. Antibiotics 2022, 11(12), 1708. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Executive summary: the selection and use of essential medicines 2019. report of the 22nd WHO Expert Committee on the selection and use of essential medicines, WHO Headquarters, Geneva, 1-5 April 2019; 2019. Available online: https://apps.who.int/iris/handle/10665/325773.
- Chowers, Michal; Zehavi, Tamir; Sheva Gottesman, Bat; Baraz, Avi; Nevo, Daniel; Obolski, Uri. Estimating the impact of cefuroxime versus cefazolin and amoxicillin/clavulanate use on future collateral resistance: a retrospective comparison. Journal of Antimicrobial Chemotherapy 2022, Volume 77(Issue 7), Pages 1992–1995. [Google Scholar] [CrossRef]
- Tribble, Alison C; Lee, Brian R; Flett, Kelly B; Handy, Lori K; Gerber, Jeffrey S; Hersh, Adam L; Kronman, Matthew P; Terrill, Cindy M; Sharland, Mike; Newland, Jason G. for the Sharing Antimicrobial Reports for Pediatric Stewardship (SHARPS) Collaborative, Appropriateness of Antibiotic Prescribing in United States Children’s Hospitals: A National Point Prevalence Survey. Clinical Infectious Diseases 2020, Volume 71(Issue 8), Pages e226–e234. [Google Scholar] [CrossRef] [PubMed]
- eEML - Electronic Essential Medicines List. Available online: https://list.essentialmeds.org/.
- Loscalzo, J.; et al. Harrison’s Principles of Internal Medicine; McGraw Hill Professional, 2022; Vol. 1 & Vol. 2. [Google Scholar]
- Sharma, P. C.; Jain, A.; Jain, S.; Pahwa, R.; Yar, M. S. Ciprofloxacin: review on developments in synthetic, analytical, and medicinal aspects. Journal of Enzyme Inhibition and Medicinal Chemistry 2010, 25(4), 577–589. [Google Scholar] [CrossRef]
- Thomson, CJ. The global epidemiology of resistance to ciprofloxacin and the changing nature of antibiotic resistance: a 10 year perspective. J Antimicrob Chemother 1999, 43 Suppl A, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Dalhoff, A. Global fluoroquinolone resistance epidemiology and implictions for clinical use. Interdiscip. Perspect. Infect. Dis. 2012, 2012, 976273. [Google Scholar] [CrossRef]
- Low, M; et al. Association between urinary community-acquired fluoroquinolone-resistant Escherichia coli and neighbourhood antibiotic consumption: a population-based case-control study. Lancet Infect. Dis. 2019, 19, 419–428. [Google Scholar] [CrossRef]
- Eliopoulos, GM; Cosgrove, SE; Carmeli, Y. The impact of antimicrobial resistance on health and economic outcomes. Clin. Infect. Dis. 2003, 36, 1433–1437. [Google Scholar] [CrossRef]
- Gottesman, BS; Carmeli, Y; Shitrit, P; Chowers, M. Impact of quinolone restriction on resistance patterns of Escherichia coli isolated from urine by culture in a community setting. Clin. Infect. Dis. 2009, 49, 869–875. [Google Scholar] [CrossRef]
- Mintz, I; Chowers, M; Obolski, U. Prediction of ciprofloxacin resistance in hospitalized patients using machine learning. Commun Med (Lond) 2023, 3(1), 43. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Breiman, L.; Friedman, J.; Olshen, R.A.; Stone, C.J. Classification and Regression Trees. In Chapman and Hall/CRC, 1st ed.; 1984. [Google Scholar] [CrossRef]
- Collins, M.; Schapire, R.E.; Singer, Y. Logistic Regression, AdaBoost and Bregman Distances. Mach. Learn. 2002, 48, 253–285. Available online: https://api.semanticscholar.org/CorpusID:207651918. [CrossRef]
- Burges, C.J.C. A Tutorial on Support Vector Machines for Pattern Recognition. Data Min. Knowl. Discov. 1998, 2, 121–167. [Google Scholar] [CrossRef]
- Mangasarian, O. L.; Musicant, David R. Lagrangian support vector machines. J. Mach. Learn. Res. 2001, 1(9/1/2001), 161–177. [Google Scholar] [CrossRef]
- Arana-Daniel, N.; Gallegos, A.A.; López-Franco, C.; Alanís, A.Y.; Morales, J.; López-Franco, A. Support Vector Machines Trained with Evolutionary Algorithms Employing. [CrossRef]
- Kernel Adatron for Large Scale Classification of Protein Structures. Evol. Bioinform. 2016, 12, EBO.S40912–302. [CrossRef]
- Hosmer, D.W., Jr.; Lemeshow, S.; Sturdivant, R.X. Applied Logistic Regression. In John Wiley & Sons; Hoboken, NJ, USA, 2013. [Google Scholar] [CrossRef]
- Lewis, CM; Knight, J. Introduction to genetic association studies. Cold Spring Harb Protoc 2012, 2012(3), 297–306. [Google Scholar] [CrossRef] [PubMed]
- Lowrie, EG; Lew, NL. Death risk in hemodialysis patients: the predictive value of commonly measured variables and an evaluation of death rate differences between facilities. Am J Kidney Dis. 1990, 15(5), 458–82. [Google Scholar] [CrossRef] [PubMed]
- Langer, DL; van der Kwast, TH; Evans, AJ; Trachtenberg, J; Wilson, BC; Haider, MA. Prostate cancer detection with multi-parametric MRI: logistic regression analysis of quantitative T2, diffusion-weighted imaging, and dynamic contrast-enhanced MRI. J Magn Reson Imaging 2009, 30(2), 327–34. [Google Scholar] [CrossRef] [PubMed]
- Redgrave, L. S.; Sutton, S. B.; Webber, M. A.; Piddock, L. J. V. Fluoroquinolone resistance: Mechanisms, impact on bacteria, and role in evolutionary success. Trends in Microbiology 2014, 22(8), 438–445. [Google Scholar] [CrossRef]
- Shaik, S; Ranjan, A; Tiwari, SK; Hussain, A; Nandanwar, N; Kumar, N; Jadhav, S; Semmler, T; Baddam, R; Islam, MA; Alam, M; Wieler, LH; Watanabe, H; Ahmed, N. Comparative Genomic Analysis of Globally Dominant ST131 Clone with Other Epidemiologically Successful Extraintestinal Pathogenic Escherichia coli (ExPEC) Lineages. mBio 2017, 8(5), e01596-17. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Huang, L; Wu, C; Gao, H; Xu, C; Dai, M; Huang, L; Hao, H; Wang, X; Cheng, G. Bacterial Multidrug Efflux Pumps at the Frontline of Antimicrobial Resistance: An Overview. Antibiotics (Basel) 2022, 11(4), 520. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Thänert, R; Choi, J; Reske, KA; et al. Persisting uropathogenic Escherichia coli lineages show signatures of niche-specific within-host adaptation mediated by mobile genetic elements. Cell Host & Microbe 2022, 30(7), 1034–1047.e6. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
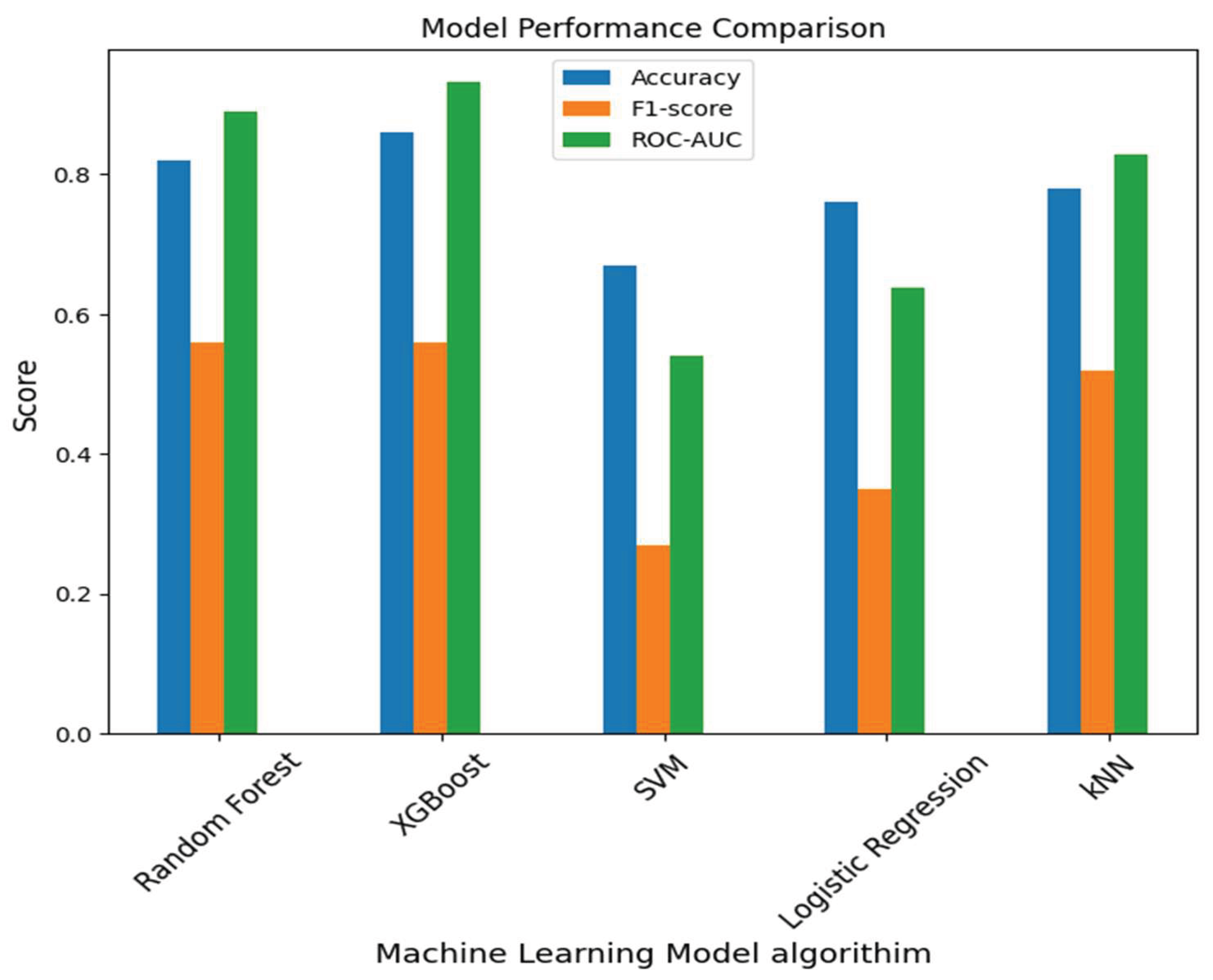

| MODEL | ACCURACY | ROC - AUC | Precision | Recall | F1 |
|---|---|---|---|---|---|
| Random Forest | 0.82 | 0.890 | 0.56 | 0.55 | 0.56 |
| XGBoost | 0.86 | 0.932 | 0.57 | 0.55 | 0.56 |
| SVM | 0.67 | 0.540 | 0.56 | 0.34 | 0.27 |
| Logistic Regression | 0.76 | 0.637 | 0.40 | 0.37 | 0.35 |
| KNN | 0.78 | 0.828 | 0.52 | 0.51 | 0.52 |
| Gene/Mutation | Antibiotics class | Phenotypic Resistance/Associated Drugs | Resistance Mechanism | Confidence/Source |
|---|---|---|---|---|
| gyrA (S83L, D87N) | Fluoroquinolones | Ciprofloxacin, Levofloxacin | Target alteration (DNA gyrase mutation) | Perfect, CARD |
| parC (S80I) | Fluoroquinolones | Ciprofloxacin | Target alteration (Topoisomerase IV mutation) | Perfect, CARD |
| aac(3)-IIa | Aminoglycosides | Gentamicin, Tobramycin | N(3)-acetyltransferase enzyme | 28636609, ResFinder |
| aac(6’)-Ib-cr | Aminoglycosides / Fluoroquinolones | Tobramycin, Amikacin, Ciprofloxacin | N(6’)-acetyltransferase (fluoroquinolone acetylation) | DQ303918, ResFinder |
| blaCTX-M-15 | β-lactams / Cephalosporins | Amoxicillin, Cefotaxime, Cefepime, Ceftriaxone | Extended-spectrum β-lactamase (Class A) | 11470367, ResFinder |
| blaOXA-1 | β-lactams | Ampicillin, Amoxicillin-clavulanate, Piperacillin | Class D OXA-type β-lactamase | 10898672, ResFinder |
| blaTEM, ampC | β-lactams / Cephalosporins | Ampicillin, Cefazolin | Antibiotic inactivation (β-lactamase) | Strict, CARD |
| acrA, acrB, acrE, acrF | Multidrug / Quinolones | Multiple drug substrates | Efflux pump complex (RND family) | Strict, CARD |
| mdtK, mdtH, mdtM, mdtG, mdtN | Multidrug / Macrolides | Erythromycin, Azithromycin | Efflux transporters and regulators | Strict, CARD |
| emrB, emrR, emrY | Macrolides / Phenicols | Chloramphenicol, Erythromycin | Multidrug efflux and regulatory proteins | Strict, CARD |
| catB3 | Phenicols | Chloramphenicol | O-acetyltransferase enzyme (drug inactivation) | 1662753 / 7793874, ResFinder |
| tet(A), tetR | Tetracyclines | Tetracycline, Doxycycline | MFS efflux pump system | 12654659, ResFinder |
| msbA, tolC | Disinfectant / Multidrug | Broad substrate range | Membrane transport and antibiotic efflux | Loose, CARD |
| vanG, vanD | Glycopeptides | Vancomycin | Target alteration (cell wall modification) | Loose, CARD |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




